2. Materials and Methods
2.1. Study Design and Public Datasets
This study integrated four core publicly available transcriptomic and epigenomic datasets representing complementary biological layers of psoriatic disease (Supplementary Table S1). GSE200376, a peripheral blood DNA methylation cohort containing healthy controls (HC), psoriasis without arthritis (PsO), and psoriatic arthritis (PsA), was used as the primary systemic discovery dataset to define the disease coordinates [
14,
15]. GSE194315, a subject-level peripheral blood mononuclear cell (PBMC) single-cell RNA sequencing dataset with surface epitope profiling, was used as an independent systemic transcriptomic comparison cohort for subject-level positioning, intermediate-state analysis in PSX, early-mover compartment analysis, and pathway-state remodeling [
5,
16]. GSE186063, a skin RNA sequencing dataset with lesional and non-lesional biopsies plus healthy-appearing skin from ankylosing spondylitis comparators, was used as the cutaneous anchoring cohort to determine whether the psoriasis-versus-PsA distinction could be explained by skin inflammatory burden [
6,
17]. GSE236694, a small exploratory purified circulating CD4
+ T-cell DNA methylation cohort, was included only to examine whether disease-axis structure might also be detectable in a more cell-enriched methylation context [
7,
18]. Two additional public skin cohorts, GSE205748 and GSE202011, were used later as external contextual checks for the skin-layer interpretation [
19,
20,
21,
22].
All analyses were performed on publicly available, de-identified data. No new human participants were recruited for this study. Additional notes on how DIR and CRS were operationalized and interpreted across modalities are summarized in Appendix Table A1.
2.2. Metadata Harmonization and Cohort Definitions
Sample- and subject-level metadata were harmonized across datasets using the integrated project metadata tables and dataset-specific design tables generated during preprocessing. Disease groups were standardized to HC, PsO, PsA, and, where available, PSX. In GSE194315, the source study defined PSX as the subgroup of psoriasis subjects reporting joint pain but not meeting CASPAR classification criteria for PsA [
5,
23]. We therefore treated PSX here as a dataset-defined arthralgia subgroup rather than as a newly assigned clinical label. Because the public source records did not adjudicate alternative causes of joint pain, the PSX label in the current study should not be interpreted as proof of prodromal PsA. For GSE186063, skin state was retained separately as healthy-appearing ankylosing spondylitis comparator skin (AS-H), non-lesional (N), or lesional (L), and the analysis unit was defined as AS-H, PsO-N, PsO-L, PsA-N, or PsA-L. Accordingly, AS-H in this dataset denotes a within-dataset non-psoriatic skin comparator rather than skin from unaffected healthy volunteers. For GSE194315, subject rather than individual cell was treated as the primary statistical unit. Subject-level metadata were linked to cell-level metadata by subject identifier, and treated subjects were excluded from the primary analysis set. A prespecified threshold of 100 retained cells per subject was used as a basic quality-control safeguard to avoid unstable subject-level cell-type proportion estimates and pseudobulk analyses in very low-coverage subjects. No subject in the final primary cohort fell below this threshold. A disease-stage factor was prespecified in the fixed order HC → PsO → PSX → PsA. For ordered trend analyses, this stage order was encoded as an ordinal numeric variable (HC = 0, PsO = 1, PSX = 2, and PsA = 3) to test monotonic associations of cell-type proportions and signature scores with disease-stage rank. This coding was used only to represent rank order and was not intended to imply equal biological spacing or observed longitudinal progression between groups.
Across the four core datasets, group standardization and subject-level alignment were applied during metadata harmonization, and harmonized age- and sex-adjusted score-level consistency checks were performed. Dataset characteristics are summarized in Supplementary Table S1, and harmonized age- and sex-adjusted score-level consistency checks are summarized in Supplementary Table S2.
2.3. Preprocessing of DNA Methylation Datasets
Raw Illumina methylation IDAT files were processed in R using
minfi (v1.56.0) [
24]. For both GSE200376 and GSE236694, raw Illumina methylation IDAT basenames were matched to dataset-specific target tables keyed to GEO sample identifiers (GSM) to define the analysis cohorts. Detection
p values were computed with
minfi::detectionP, and per-sample quality-control metrics, including call rate and median methylated and unmethylated signal intensities, were recorded. Background correction and normalization were performed using
preprocessNoob [
25]. Beta values and M values were extracted with
getBeta and
getM, respectively.
For the primary GSE200376 differential methylation analysis, probes were retained only if detection
in at least 95% of samples, after which non-CpG probes, sex-chromosome probes, and published cross-reactive and SNP-overlapping probes were excluded using the EPIC blacklist files corresponding to Supplementary Tables S1, S4, S5, and S6 of the original resource [
26].
For GSE236694, the primary differential methylation analysis likewise retained probes with detection in at least 95% of samples and then restricted the analysis to autosomal probes. Sample-level quality-control metrics were reviewed for both methylation cohorts, and this review did not identify any sample requiring additional exclusion in the primary differential methylation analyses. Filtered beta matrices, M-value matrices, detection p matrices, and per-sample quality-control tables were retained as downstream analysis inputs.
Because GSE200376 served as the primary whole-blood discovery methylation cohort, a stringent blacklist-based probe filter was applied, whereas GSE236694 was treated as a small exploratory purified CD4+ T-cell cohort and therefore used a minimal probe filter limited to detection-passing autosomal probes in order to preserve analyzable probe coverage.
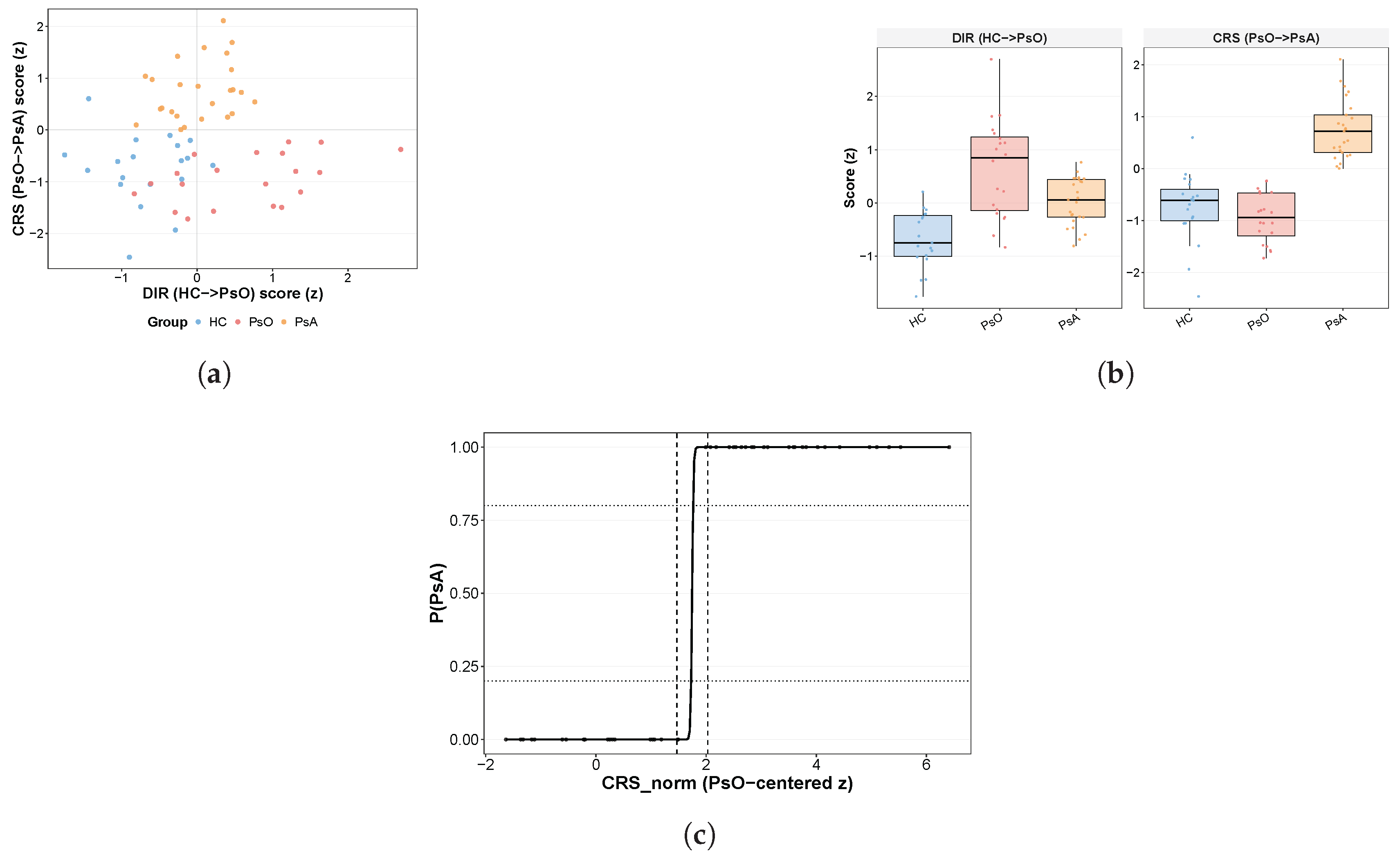
2.4. Construction of the GSE200376 Discovery Coordinate System
GSE200376 was used to construct the primary discovery coordinate system. Two axes were defined. The inflammatory disease axis (DIR) was anchored to the PsO-versus-HC contrast and was intended to capture the shared inflammatory component of psoriatic disease. The contrast-resolved systemic-state coordinate (CRS) was anchored to the PsA-versus-PsO contrast and was intended to capture the additional systemic state associated with PsA relative to PsO.
Differentially methylated probes (DMPs) were identified from M values using
limma (v3.66.0), and differentially methylated regions (DMRs) were identified using
DMRcate (v3.6.0) [
27,
28]. For pairwise comparisons, the primary total-signal model was specified as
where
group denotes the harmonized disease-group factor (HC, PsO, or PsA) and
sentrix_id denotes the Illumina BeadChip/Sentrix slide identifier derived from the IDAT basename. A cell-adjusted sensitivity model additionally included the estimated whole-blood cell fractions CD8T, CD4T, NK, Bcell, and Mono. Ordered trend analyses across HC, PsO, and PsA used a numeric trend term coded as HC
, PsO
, and PsA
. This coding was used only to represent ordered disease-stage rank for trend testing and was not intended to imply equal biological spacing or observed longitudinal progression. The total-signal trend model was specified as
and a cell-adjusted sensitivity trend model additionally included the same whole-blood cell-fraction covariates. Statistical testing was performed on M values, and strict DMPs were defined as probes with Benjamini–Hochberg false discovery rate (FDR)
and
. DMRs were called using a strict seed threshold of FDR
in the reported primary run.
Whole-blood cell fractions were estimated with the FlowSorted.Blood.EPIC reference-based deconvolution workflow using
estimateCellCounts2 with the
preprocessNoob processing method and IDOL probe selection [
29]. Sample identifiers were harmonized to GSM IDs before merging estimated cell fractions with the design table, and cell-fraction row sums were checked and renormalized when minor numerical drift from 1.0 was detected.
Final GSE200376 DIR and CRS scores were computed from CpG signatures derived from the internal five-fold resampling procedure used for score construction. Within each training fold, CpGs were selected de novo from the relevant
limma contrast, and probes selected in at least three valid folds, defined here as folds in which the training contrast was estimable and a fold-specific score model could be fit, were retained for final scoring; this three-fold rule was used as a heuristic stability filter rather than as a formal optimality criterion. This yielded 25 CpGs for DIR and 45 CpGs for CRS in the final scoring set, reflecting axis-specific fold stability rather than a matched target feature count. For each sample
j, the raw score was computed as
where
is the
limma moderated
t statistic for CpG
i from the relevant discovery contrast, and
denotes CpG-wise standardization of the sample M value using the mean and standard deviation of that CpG within the relevant reference set. For each axis, the M value of each retained CpG was first standardized using the mean and standard deviation of that CpG within the relevant reference groups (DIR: HC+PsO; CRS: PsO+PsA). These standardized CpG values were then combined as a weighted sum to obtain the raw score. The raw score was subsequently centered and scaled using the mean and standard deviation of the raw-score distribution within the same reference set, and the resulting reference-centered standardized scores were saved as the final DIR and CRS values used for downstream positioning and visualization. These axis-specific reference sets were chosen to preserve contrast-specific interpretability rather than to place DIR and CRS on a single shared assay-level scale.
2.5. Internal Stability and Calibration Assessment in GSE200376
The GSE200376 discovery framework was internally evaluated using leakage-aware nested resampling, but this procedure was used only to characterize the stability of the coordinate system and the discrimination and calibration of the derived score-based prediction models; it was not treated as external validation. Five-fold outer cross-validation was performed. Fold assignment was grouped by sentrix_id whenever feasible to reduce batch leakage; when grouped splitting was infeasible, a stratified fallback preserving class balance was used.
Within each training fold, feature selection was performed de novo using
limma, and the weighted score was reconstructed using training data only. Fold-specific score standardization was based exclusively on the training mean and standard deviation. For each axis, prediction models were then fitted for the corresponding discovery contrast (DIR: HC versus PsO; CRS: PsO versus PsA) using ridge logistic regression (
glmnet, v4.1-10), with a generalized linear model fallback implemented as a safeguard when penalized fitting was infeasible in a given fold [
30]. The ridge penalty parameter was selected within each training fold by inner cross-validation using the 1-SE rule, defined as the largest penalty yielding cross-validated error within one standard error of the minimum [
31]. The full prediction model included the fold-specific standardized score, age, sex, and the first two principal components of the cell-fraction matrix.
The area under the receiver operating characteristic curve (AUC), Brier score, calibration intercept, and calibration slope were recorded together with five-bin calibration tables and fold-level diagnostics.
2.6. Subject-Level and Cell-Type-Level DIR/CRS Scoring in GSE194315
GSE194315 was analyzed at the subject level. Subject-level cell-type proportions were first calculated from the filtered cell metadata. Pseudobulk RNA count matrices were then generated for each subject and annotated cell type by summing pre-normalization single-cell counts within each subject–cell type combination. Subject–cell type combinations represented by fewer than 30 retained cells were excluded from pseudobulk generation. Per-cell-type pseudobulk matrices and matching subject-level metadata tables were then constructed for downstream scoring and pathway analysis.
Cell-type-specific DIR and CRS signatures were derived independently within GSE194315 from pseudobulk contrasts using
edgeR (v4.8.2), and signature scores were subsequently computed by single-sample gene set enrichment analysis using
GSVA (v2.4.4) [
32,
33]. For each cell type, a signed ranking statistic was defined as
where
F is the quasi-likelihood test statistic from
glmQLFTest [
34]. This quantity was used only to rank genes for signature construction and was not interpreted as a formal inferential effect-size estimate.
DIR signatures were learned using only HC and PsO subjects, whereas CRS signatures were learned using only PsO and PsA subjects; PSX subjects were explicitly excluded from signature learning and were scored only after the signatures had been fixed. For each contrast and cell type, signature learning required at least five subjects in each contrast group. The top 100 genes with the most positive signed statistic and the top 100 genes with the most negative signed statistic were retained as the UP and DOWN components of the signature, respectively. Per-cell-type scoring was then performed only when at least five genes remained in both the UP and DOWN components after matching to the expression matrix.
Per-cell-type scores were computed by ssGSEA using
GSVA, with
kcdf = "Gaussian",
absRanking = TRUE, and
ssgsea.norm = TRUE. The final score was defined as
Subject-level overall DIR and CRS scores were then computed as weighted means across cell types, using weights proportional to when subject–cell type cell counts were available and equal weighting otherwise. This weighting scheme was used as a heuristic compromise between equal weighting and direct cell-count weighting.
2.7. Definition of the CRS Reference Band and Early-Mover Compartments
To formalize a reference interval along the systemic CRS continuum, CRS scores in GSE200376 were first normalized relative to the PsO distribution:
A logistic model for was then fitted using the PsO and PsA samples only. The initial lower and upper band limits were defined as the normalized CRS values corresponding to predicted probabilities of 0.20 and 0.80, respectively. To summarize uncertainty and obtain the final applied band, class-balanced bootstrap resampling (1000 iterations) was performed, and the bootstrap medians of the lower and upper bounds were retained as the final CRS reference band. These probability cut points were used as a heuristic reference interval for the central reference zone rather than as optimized diagnostic thresholds.
Band stability in the discovery cohort was further assessed by repeated stratified five-fold cross-validation (50 repeats), in which CRS normalization and logistic fitting were repeated within each training split, and the resulting band limits were summarized across repeats. Concise summaries of the bootstrap, repeated-CV, and cutoff-sensitivity analyses are reported in Supplementary Figure S1.
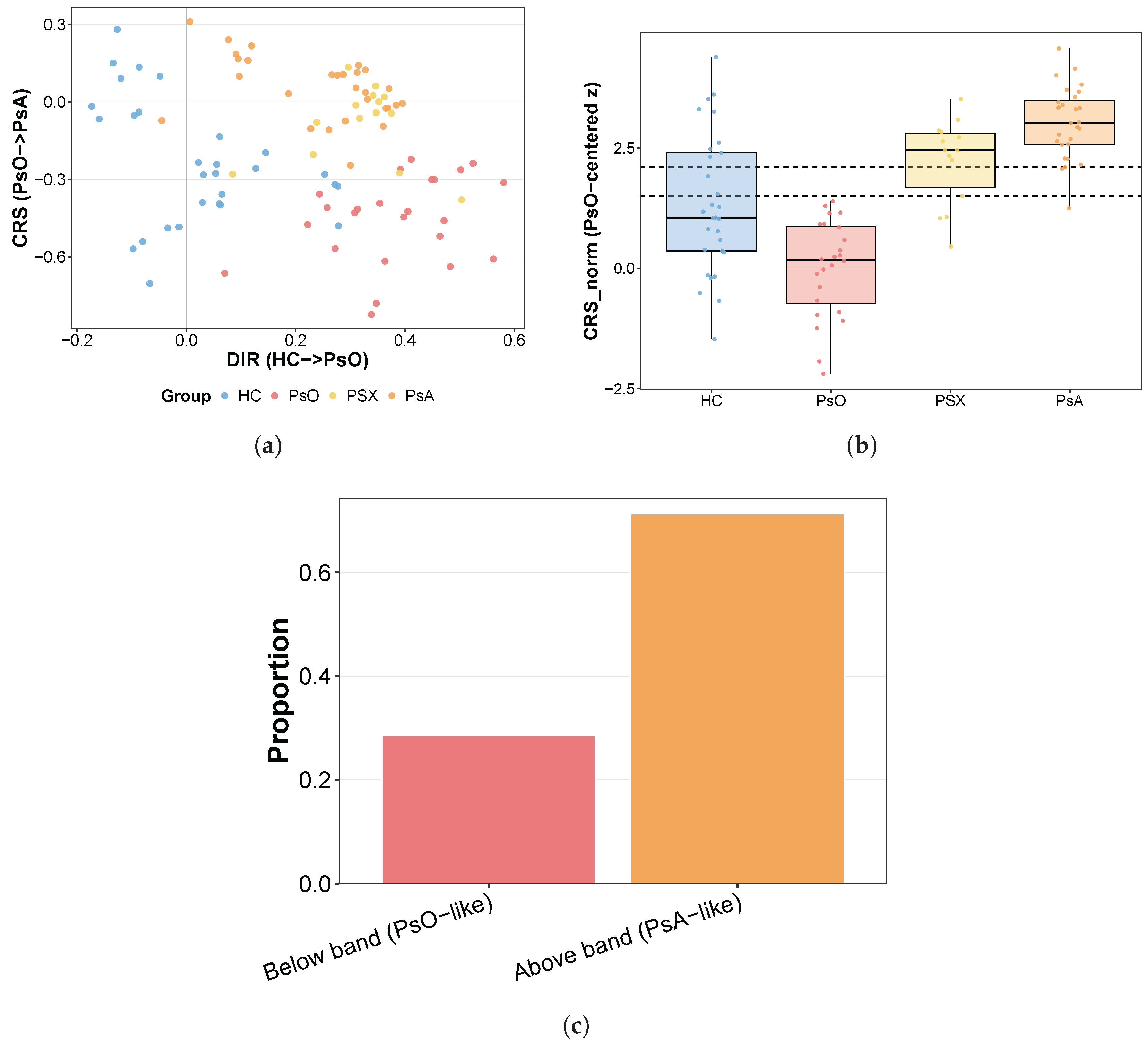
The resulting GSE200376-derived band was transferred to GSE194315 after recalculating the PsO mean and standard deviation within the scRNA-seq cohort: for subject-level overall scores, normalization used the overall PsO distribution, whereas for subject–cell type scores, normalization was performed separately within each cell type using the corresponding PsO distribution. This transfer was based on contrast-level rather than feature-level equivalence: in both datasets, CRS was defined independently from the same PsO-versus-PsA biological contrast within the assay-specific data layer. PsO-centered normalization therefore aligned each cohort to its own internal psoriasis reference state, so that transferred band labels reflected relative displacement away from PsO rather than equality of methylation-derived and transcript-derived measurement scales. The transfer was interpreted only as a test of whether a discovery-derived positional interval preserved PsO-like versus PsA-like ordering in an independent transcriptomic cohort. Subject-level and subject–cell type observations were then classified as below-band (PsO-like), in-band (central reference zone), or above-band (PsA-like).
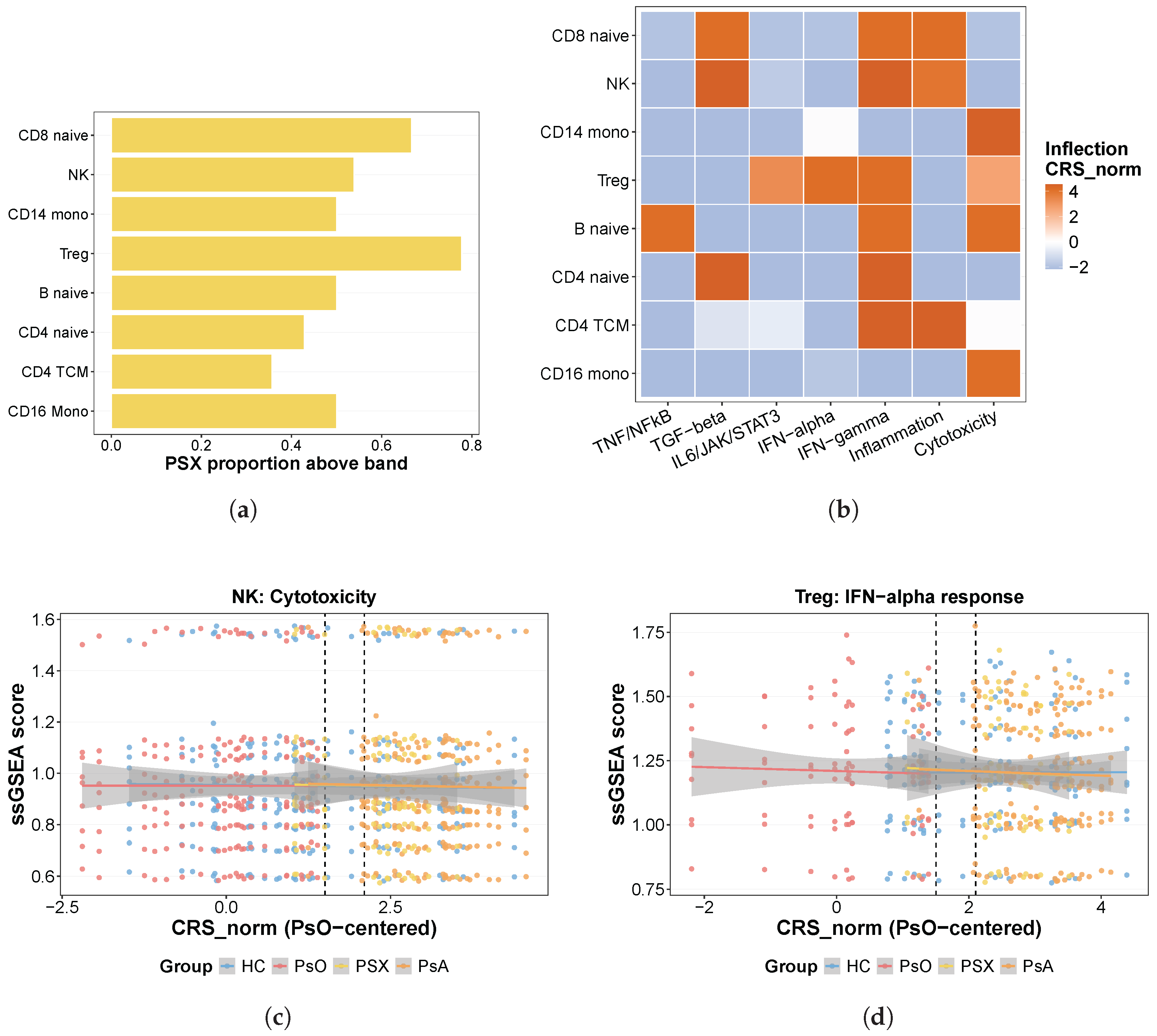
Early-mover compartments were defined as cell types in which PSX subjects were enriched in the above-band state relative to PsO subjects. For each cell type, this was tested only when at least six PsO and six PSX subjects were available, using one-sided Fisher’s exact tests for enrichment of PSX in the above-band state relative to PsO and permutation testing based on 5000 shuffles of the PSX/PsO labels, followed by Benjamini–Hochberg correction across cell types.
2.8. Cell-Type Pathway Dynamics along the CRS Continuum
To characterize within-cell-type pathway-state remodeling associated with CRS, a focused Hallmark pathway panel was evaluated in GSE194315 pseudobulk data. The panel included interferon-alpha response, interferon-gamma response, TNF/NFB signaling, IL6/JAK/STAT3 signaling, inflammatory response, complement, IL2/STAT5 signaling, and TGF- signaling, together with a custom cytotoxicity gene set defined as NKG7, PRF1, GZMB, GZMH, GNLY, KLRD1, FCGR3A, TRAC, TRBC1, and TRBC2. Gene symbols were harmonized across pseudobulk matrices, and Ensembl identifiers were mapped to symbols using the org.Hs.eg.db annotation package when necessary.
Pathway analysis was restricted to a focused eight-cell-type panel selected for focused modeling on sufficiently represented immune compartments: CD8.Naive, NK, CD14.Mono, Treg, B.naive, CD4.Naive, CD4.TCM, and CD16.Mono. This analysis was intended as a focused rather than exhaustive survey of all annotated cell types. For each selected cell type, pseudobulk counts were normalized using edgeR::calcNormFactors and transformed to log-CPM values using cpm(log = TRUE, prior.count = 1), and ssGSEA pathway scores were then computed per subject using GSVA (v2.4.4). Cell types with fewer than eight subjects in common with the subject-level CRS table were excluded from the primary pathway analysis. A broader exploratory survey across all eligible annotated pseudobulk cell types, defined here as cell types with at least eight subjects in common with the subject-level CRS table, is reported in Supplementary Figure S7; this broader screen was not used to redefine the primary eight-cell-type panel.
Within each selected cell type and pathway, pathway activity was modeled across the cross-sectional CRS continuum spanning PsO, PSX, and PsA subjects as a smooth function of
using generalized additive models implemented in the
mgcv package (v1.9-4) [
35] of the form
For each pathway–cell type pair, the pathway inflection point was operationally defined as the
value at which the absolute first derivative of the fitted smooth function was maximal. This procedure was used as a descriptive summary of pathway-specific positional dynamics across the cross-sectional CRS continuum.
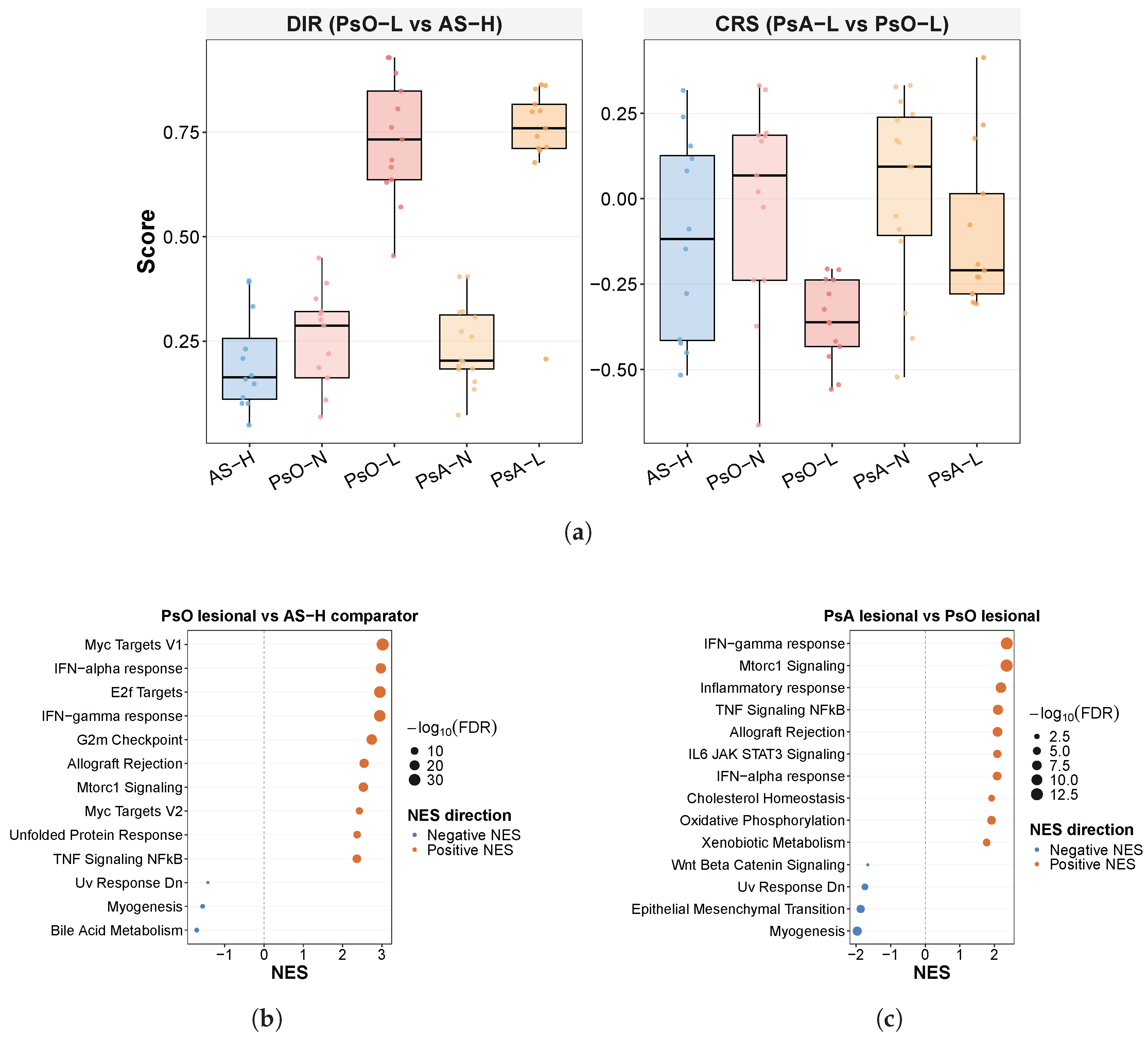
2.9. Skin Transcriptomic Analysis in GSE186063
GSE186063 skin RNA-seq data were analyzed from a preprocessed dataset containing an edgeR DGEList, a logCPM expression matrix, and parsed GEO series-matrix metadata. Metadata rows were aligned to the expression matrix primarily by sample identifier, with fallback matching by sample title and GSM when necessary. Disease group, tissue state, and detailed unit-group labels were standardized to AS-H, PsO-N, PsO-L, PsA-N, and PsA-L, where AS-H denotes healthy-appearing ankylosing spondylitis comparator skin rather than skin from unaffected healthy volunteers. Subject identifiers were parsed from the aligned sample identifiers to allow evaluation of possible paired lesional/non-lesional analyses.
Differential expression analysis was performed using the
edgeR quasi-likelihood framework with robust dispersion estimation. The tested contrasts were PsA-L versus AS-H, PsA-N versus AS-H, PsA-L versus PsA-N, PsA-L versus PsO-L, PsA-N versus PsO-N, PsO-L versus AS-H, PsO-N versus AS-H, and PsO-L versus PsO-N. For unpaired contrasts, the design was specified as
For within-disease lesional-versus-non-lesional comparisons, paired models of the form
were prespecified when at least five paired subjects were available, where
group denotes the harmonized analysis-group factor (AS-H, PsO-N, PsO-L, PsA-N, or PsA-L) and
subject_block denotes the subject-specific blocking factor for paired lesional/non-lesional samples; otherwise an unpaired design was used. In the final analysis-ready dataset, both within-disease contrasts were analyzed using the unpaired design. Differential expression results were exported in full, and FDR-significant gene counts were summarized across contrasts.
Pathway analysis was performed from the differential expression results using pre-ranked GSEA with
fgsea (v1.36.2) [
36]. Ranking was based on signed
statistics derived from the edgeR quasi-likelihood test, defined as
, where
P denotes the edgeR quasi-likelihood test
p value. Hallmark, GO biological process, KEGG legacy, and Reactome collections were analyzed by GSEA. In addition, Hallmark over-representation analyses for upregulated and downregulated gene sets were performed using
clusterProfiler::enricher (v4.18.4) [
37] when at least 10 FDR-significant mapped genes were available in the corresponding direction, with all mapped tested genes in the corresponding contrast used as the background universe. For the primary skin-axis scores, DIR
skin was anchored to the PsO-L versus AS-H contrast and CRS
skin> to the PsA-L versus PsO-L contrast. Scores were computed by ssGSEA as UP minus DOWN using the same GSVA-based ssGSEA settings described in
Section 2.6. Core signatures from the prespecified strict and relaxed GMT signature sets defined in the core-signature pipeline were used when available, with fallback first to the relaxed GMT set and then to ranked-signature TOP-100 sets when core signatures were unavailable or too small. This ranked-signature fallback was required for the primary CRS
skin score because the corresponding strict and relaxed core signatures were empty.
2.10. Exploratory Purified CD4+ T-Cell Methylation Analysis in GSE236694
GSE236694 was analyzed as an exploratory purified-cell methylation cohort. Upstream preprocessing was performed from raw IDAT files using the same
minfi Noob-based workflow described above, and the present analysis used the resulting analysis-ready beta-value, M-value, and detection-
p matrices. Differential methylation analysis was performed on M values using
limma with robust empirical Bayes moderation [
27]. The design matrix included disease group and, when available and non-degenerate within the analyzed subset, age, sex,
batch_slide, and
batch_array. Three contrasts were evaluated: PsO versus HC (DIR), PsA versus PsO (CRS), and PsA versus HC (disease spectrum).
DMRs were identified using
DMRcate, with array type inferred automatically from probe identifiers. Primary DMR calling used CpG-level annotation with a strict seed threshold of FDR
and region calling under the default strict
DMRcate setting (without relaxed
pcutoff override). When no reportable regions were returned,
pcutoff was relaxed sequentially to 0.05, 0.10, and then 0.20 while other
DMRcate settings were held constant. In the reported run, all three contrasts yielded reportable regions at the first
pcutoff-relaxation step (
pcutoff = 0.05), and no further relaxation was required [
28]. Gene-set enrichment was performed with
missMethyl::gometh (v1.44.0) for GO and KEGG categories [
38]. Because no contrast yielded FDR-significant CpG sets of sufficient size for enrichment, the reported
gometh analyses used the fallback set of probes with nominal
.
Exploratory DIR and CRS methylation scores were constructed separately from the GSE236694 DMP tables for the DIR (PsO versus HC) and CRS (PsA versus PsO) contrasts using a tiered feature-selection strategy, because the prespecified strict thresholds yielded too few FDR-significant CpGs to support stable score construction in this small cohort. The selection tiers were applied in the following order: (i) FDR and ; (ii) FDR and ; (iii) FDR and ; (iv) fallback to top probes irrespective of FDR; and (v) fallback to top probes. The top 30 CpGs passing the best available tier were used for each axis. For each probe, beta values were standardized relative to the appropriate baseline group (HC for DIR and PsO for CRS), multiplied by the sign of (or the sign of the t statistic if needed), averaged across probes, and then re-standardized within the baseline group. Because these scores were built and evaluated within the same small cohort, the resulting DIR and CRS scores from GSE236694 were interpreted only as exploratory within-cohort findings.
2.11. Additional Robustness Analyses
Several additional robustness analyses were performed using previously generated subject-level, cell-type, and pathway summary outputs in order to quantify the stability of the main subject-level and pathway-level conclusions. In GSE194315, subject-level DIR and CRS scores were recalculated using alternative cell-type aggregation weights proportional to 1, , , and , and the resulting subject-level scores were compared with the primary implementation by Pearson and Spearman correlation, preservation of the broad HC/PsO/PSX/PsA group ordering, and group-wise above-band occupancy.
In parallel, the discovery-derived CRS reference interval was transferred to GSE194315 using three positional strategies: the primary PsO-centered z-score transfer, pooled quantile transfer, and a within-cohort PsO-versus-PsA refit.
To refine interpretation of the early-mover Fisher analyses when PsO above-band counts were zero, Haldane–Anscombe continuity-corrected odds ratios and bounded Newcombe–Wilson risk-difference confidence intervals were also computed as descriptive sensitivity summaries; these quantities did not replace the primary Fisher/permutation inference.
For the main-text pathway-dynamic analyses, evaluated across the 72 pathway–cell type pairs in the primary pathway-localization panel, the generalized additive model basis dimension was varied from to , and bootstrap resampling (100 resamples per pathway–cell type pair) was used to summarize the directional stability of the inferred pathway-localization patterns. These robustness summaries are reported in Supplementary Tables S3 and S4.
2.12. External Skin Cohort Checks in GSE205748 and GSE202011
Two additional public skin datasets were analyzed to provide external context for the limited additional PsA-related separation observed in GSE186063.
GSE205748 is a small independent bulk skin RNA-seq cohort containing healthy control skin, uninvolved PsA skin, and PsA lesional skin (9 samples per group) [
19]. Gene-level count data were normalized with
edgeR::calcNormFactors and transformed to logCPM values using
cpm(log = TRUE, prior.count = 2) [
32]. LogCPM expression values were then scored by ssGSEA against a focused Hallmark panel comprising TNF/NF
B signaling, interferon-alpha response, interferon-gamma response, IL6/JAK/STAT3 signaling, inflammatory response, and TGF-
signaling, using the same GSVA-based ssGSEA settings applied elsewhere in the manuscript [
33]. The tested contrasts were PsA lesional versus healthy control skin and PsA lesional versus uninvolved PsA skin, evaluated by exact rank-sum permutation testing with Benjamini–Hochberg correction. Because the cohort did not include psoriasis without arthritis, it was used only for pathway-level PsA skin corroboration rather than for direct PsO-versus-PsA testing.
GSE202011 is an external spatial transcriptomic skin cohort containing healthy control, PsO, and PsA samples, including both lesional and non-lesional skin states [
21]. Because the aim of this analysis was sample-level external projection of the GSE186063-derived bulk-like skin signatures rather than inference on spatial microenvironments, raw spot-level matrices were collapsed to sample-level pseudobulk counts by summing transcript counts across spots within each Visium sample. These sample-level pseudobulk counts were normalized with
edgeR::calcNormFactors and transformed to logCPM values using
cpm(log = TRUE, prior.count = 2) [
32]. The official GSE186063-derived DIR
skin and CRS
skin signatures were then projected to these pseudobulk matrices using the same ssGSEA UP-minus-DOWN scoring logic and GSVA-based settings described above [
33]. External score contrasts were evaluated by exact rank-sum permutation testing with Benjamini–Hochberg correction for the lesional comparisons PsO-L versus healthy control skin, PsA-L versus healthy control skin, and PsA-L versus PsO-L.
2.13. Covariate-Adjusted Consistency Checks
To verify that the principal directional conclusions were not explained solely by age- or sex-imbalance, we performed harmonized score-level consistency checks across the four core datasets. For selected key disease-axis findings emphasized in the main figures, the relevant DIR/CRS score was fitted in a pairwise linear model including disease group, age, and sex:
These checks were performed on the already generated dataset-specific DIR/CRS score tables rather than by repeating each full modality-specific pipeline, and they were interpreted as harmonized sensitivity summaries rather than as replacements for the primary analyses. The resulting summary is provided in Supplementary Table S2.
2.14. Statistical Analysis and Software
Correlations between DIR and CRS were assessed using Pearson and Spearman coefficients. Multiple-testing correction was performed using the Benjamini–Hochberg method where relevant. All tests were two-sided unless a directional alternative was explicitly specified (e.g., in the early-mover enrichment analysis).
Figure 3(b)–(d) should be interpreted as descriptive summaries of pathway-dynamic localization and representative fitted trajectories rather than as per-tile or per-curve hypothesis tests.
All analyses were performed in R version 4.5.1, and principal software packages and version numbers are reported at first use in the relevant subsections. During preparation of this study and manuscript, OpenAI GPT-5.4 Pro was used to assist with code review and debugging, as well as English-language editing and wording refinement. All analytical decisions, code execution, result verification, and final interpretation were performed and verified directly by the authors.