Submitted:
24 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
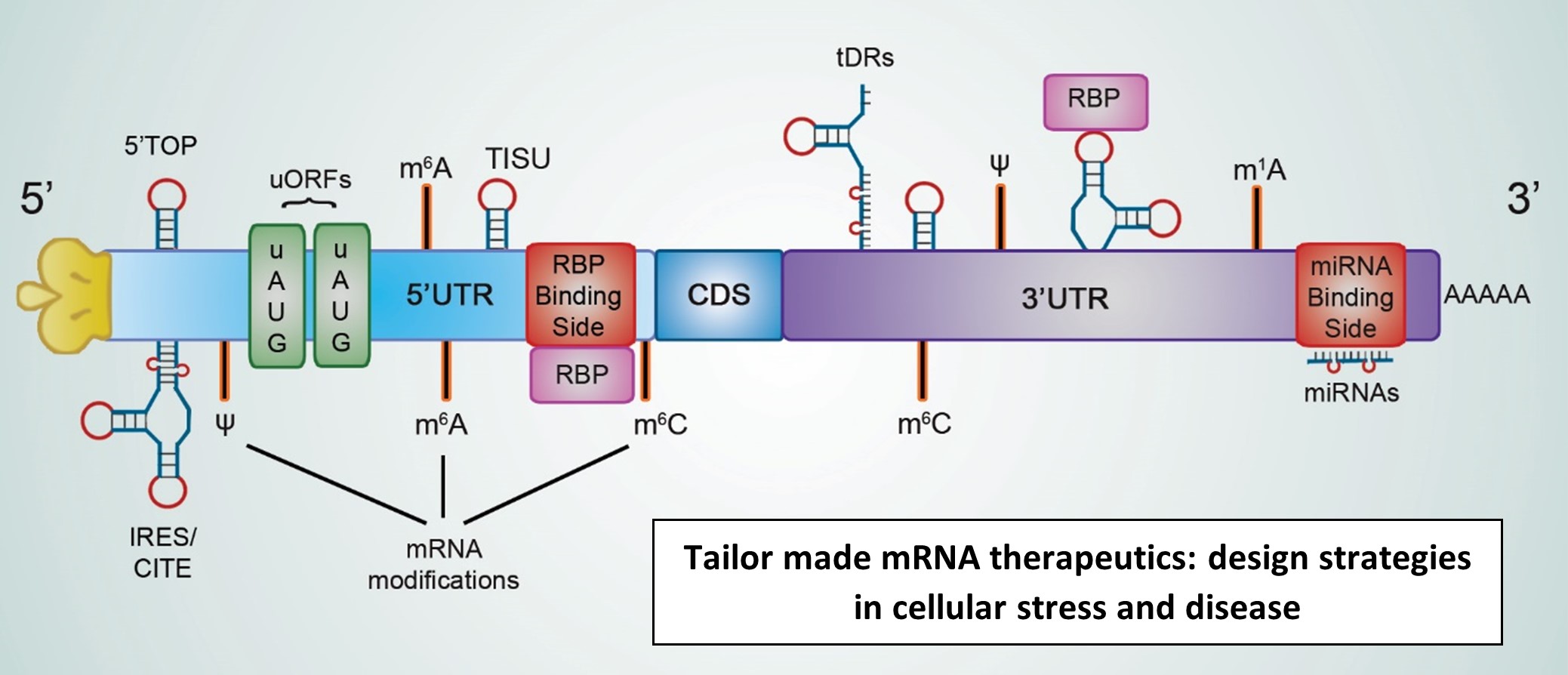
Keywords:
1. Introduction
2. Canonical Mode of mRNA Translation
3. mRNA Translation Initiation in Cellular Stress and Disease
4. Cellular Pathways Involved in Stress Responses
4.1. Stress-Induced eIF2a Phosphorylation and Disease Development
4.2. Regulation of eIF4E Activity Under Cellular Stress
4.2.1. The mTOR Pathway
5. mRNA Specific Regulation
5.1. Trans-Acting Factors Regulate mRNA Fate in Cellular Stress Response
5.1.1. RNA-Binding Proteins (RBPs)
5.1.2. MicroRNAs
5.1.3. tRFs and tiRNAs
5.2. Specific cis-Acting Features of mRNA Control the Rate and Mode of Translation During Stress
5.2.1. Stress-Induced mRNA Modification
5.2.2. Upstream Open Reading Frames (uORFs)
5.2.3. 5’Terminal OligoPyrimidines (5’TOP) Motifs
5.2.4. Translation Initiator of Short 5′UTR (TISU)
5.2.5. Internal Ribosome Entry Site (IRES)
5.2.6. Cap-independent translation enhancers (CITE)
5.2.7. Alternative Cap-Dependent Mechanism in Stress Response
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| mRNA | Messenger RNA |
| 5’UTR | 5’ Untranslated Region |
| 3’UTR | 3’ Untranslated Region |
| tRNA | Transfer RNA |
| 43S PIC | 43S pre-initiation complex |
| eIF | Eukaryotic Translation Initiation Factor |
| GEF | Guanine Nucleotide Exchange Factor |
| eIF2–GDP | Eukaryotic Initiation Factor 2-Guanosine Diphosphate |
| eIF2–GTP | Eukaryotic Initiation Factor 2-Guanosine Triphosphate |
| eIF4 | Eukaryotic Translation Initiation Factor 4 |
| eEF2K | Eukaryotic Elongation Factor 2 Kinase |
| TC | Ternary Complex |
| GTP | Guanosine triphosphate |
| ISR | Integrated Stress Response |
| GCN2 | General Control Non-Derepressible-2 |
| PERK | PKR-like ER Kinase |
| PKR | Protein Kinase RNA |
| HRI | Heme-Regulated Inhibitor Kinase |
| uORFs | Upstream Open Reading Frames |
| ATF4 | Activating Transcription Factor 4 |
| IRESs | Internal Ribosome Entry Sites |
| BiP | Binding Immunoglobulin Protein |
| APAF-1 | Apoptotic Protease Activating Factor-1 |
| RBPs | RNA-Binding Protein |
| HuR | Human Antigen R |
| TIA-1 | T-cell intracellular antigen-1 |
| mTOR | Mammalian Target of Rapamycin |
| 4E-BPs | eIF4E-Binding Proteins |
| PI3K | Class I Phosphoinositide 3-Kinase |
| AMPK | 5′-Adenosine Monophosphate-Activated Protein Kinase |
| 5′TOP | 5′Terminal Oligopyrimidine |
| CCND1 | Cyclin D1 |
| VEGFA | Vascular Endothelial Growth Factor A |
| ODC1 | Ornithine Decarboxylase |
| MNKs | Mitogen-Activated Protein Kinase (MAPK)-Interacting Kinases |
| TSC | hamartin |
| MDM2 | Mouse Double Minute 2 |
| BAX | Bcl-2-Associated X Protein |
| AGO2 | Argonaute 2 |
| FXR1 | Fragile X-Related Protein 1 |
| tRFs | tRNA-Derived RNA Fragments |
| tiRNAs | tRNA-Derived Stress-Induced RNAs |
| ANG | Angiogenin |
| YTHDF1 | YTH N6-Methyladenosine RNA Binding Protein 1 |
| CHOP | DNA Damage-Inducible Transcript 3 |
| ATG5 | Autophagy Related 5 |
| 5’TOP | 5’Terminal OligoPyrimidines |
| PABP | Poly(A)-Binding Protein |
| TISU | Translation Initiator of Short 5′ UTR |
| TSS | Transcription Start Site |
| IRES | Internal Ribosome Entry Site |
| VEGF | Vascular Endothelial Growth |
| HIF-1α | Hypoxia-Inducible Factor 1-alpha |
| CITE | Cap-Independent Translation Enhancers |
References
- Rohner, E.; Yang, R.; Foo, K.S.; Goedel, A.; Chien, K.R. Unlocking the Promise of MRNA Therapeutics. Nat. Biotechnol. 2022, 40. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, N.; Weissman, D.; Whitehead, K.A. MRNA Vaccines for Infectious Diseases: Principles, Delivery and Clinical Translation. Nat. Rev. Drug. Discov. 2021, 20. [Google Scholar] [CrossRef]
- Mauger, D.M.; Joseph Cabral, B.; Presnyak, V.; Su, S. V.; Reid, D.W.; Goodman, B.; Link, K.; Khatwani, N.; Reynders, J.; Moore, M.J.; et al. MRNA Structure Regulates Protein Expression through Changes in Functional Half-Life. Proc. Natl. Acad. Sci. U. S. A. 2019, 116. [Google Scholar] [CrossRef]
- Lee, J.; Woodruff, M.C.; Kim, E.H.; Nam, J.H. Knife’s Edge: Balancing Immunogenicity and Reactogenicity in MRNA Vaccines. Exp. Mol. Med. 2023, 55. [Google Scholar] [CrossRef]
- Berger, S.; Lächelt, U.; Wagner, E. Dynamic Carriers for Therapeutic RNA Delivery. Proc. Natl. Acad. Sci. U. S. A. 2024, 121. [Google Scholar] [CrossRef]
- Rosa, S.S.; Prazeres, D.M.F.; Azevedo, A.M.; Marques, M.P.C. MRNA Vaccines Manufacturing: Challenges and Bottlenecks. Vaccine 2021, 39. [Google Scholar] [CrossRef]
- Potužník, J.F.; Cahova, H. If the 5’ Cap Fits (Wear It)–Non-Canonical RNA Capping. RNA Biol. 2024, 21, 1–13. [Google Scholar] [CrossRef]
- Chu, Y.; Yu, D.; Li, Y.; Huang, K.; Shen, Y.; Cong, L.; Zhang, J.; Wang, M. A 5′ UTR Language Model for Decoding Untranslated Regions of MRNA and Function Predictions. Nat. Mach. Intell. 2024, 6. [Google Scholar] [CrossRef]
- Li, X.; Kazan, H.; Lipshitz, H.D.; Morris, Q.D. Finding the Target Sites of RNA-Binding Proteins. Wiley Interdiscip. Rev. RNA 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Ryczek, N.; Łyś, A.; Makałowska, I. The Functional Meaning of 5′UTR in Protein-Coding Genes. Int. J. Mol. Sci. 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Mayr, C. What Are 3′ Utrs Doing? Cold Spring Harb. Perspect. Biol. 2019, 11. [Google Scholar] [CrossRef]
- Hong, D.; Jeong, S. 3’UTR Diversity: Expanding Repertoire of RNA Alterations in Human MRNAs. Mol. Cells 2023, 46. [Google Scholar] [CrossRef] [PubMed]
- Passmore, L.A.; Coller, J. Roles of MRNA Poly(A) Tails in Regulation of Eukaryotic Gene Expression. Nat. Rev. Mol. Cell. Biol. 2022, 23. [Google Scholar] [CrossRef]
- Harnisch, C.; Moritz, B.; Rammelt, C.; Temme, C.; Wahle, E. Chapter Nine—Activity and Function of Deadenylases. In The Enzymes; Chanfreau, G.F., Tamanoi, F., Eds.; Academic Press, 2012; Vol. 31, pp. 181–211. ISSN ISBN 1874-6047. [Google Scholar]
- Vavilis, T.; Stamoula, E.; Ainatzoglou, A.; Sachinidis, A.; Lamprinou, M.; Dardalas, I.; Vizirianakis, I.S. MRNA in the Context of Protein Replacement Therapy. Pharmaceutics 2023, 15. [Google Scholar] [CrossRef]
- Bicknell, A.A.; Reid, D.W.; Licata, M.C.; Jones, A.K.; Cheng, Y.M.; Li, M.; Hsiao, C.J.; Pepin, C.S.; Metkar, M.; Levdansky, Y.; et al. Attenuating Ribosome Load Improves Protein Output from MRNA by Limiting Translation-Dependent MRNA Decay. Cell. Rep. 2024, 43. [Google Scholar] [CrossRef]
- Reinhart, A.G.; Osterwald, A.; Ringler, P.; Leiser, Y.; Lauer, M.E.; Martin, R.E.; Ullmer, C.; Schumacher, F.; Korn, C.; Keller, M. Investigations into MRNA Lipid Nanoparticles Shelf-Life Stability under Nonfrozen Conditions. Mol. Pharm. 2023, 20. [Google Scholar] [CrossRef] [PubMed]
- Leppek, K.; Byeon, G.W.; Kladwang, W.; Wayment-Steele, H.K.; Kerr, C.H.; Xu, A.F.; Kim, D.S.; Topkar, V. V.; Choe, C.; Rothschild, D.; et al. Combinatorial Optimization of MRNA Structure, Stability, and Translation for RNA-Based Therapeutics. Nat. Commun. 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Hanson, G.; Alhusaini, N.; Morris, N.; Sweet, T.; Coller, J. Translation Elongation and MRNA Stability Are Coupled through the Ribosomal A-Site. RNA 2018, 24. [Google Scholar] [CrossRef]
- Courel, M.; Clément, Y.; Bossevain, C.; Foretek, D.; Cruchez, O.V.; Yi, Z.; Bénard, M.; Benassy, M.N.; Kress, M.; Vindry, C.; et al. Gc Content Shapes MRNA Storage and Decay in Human Cells. Elife 2019, 8. [Google Scholar] [CrossRef]
- Wieder, N.; D’Souza, E.N.; Martin-Geary, A.C.; Lassen, F.H.; Talbot-Martin, J.; Fernandes, M.; Chothani, S.P.; Rackham, O.J.L.; Schafer, S.; Aspden, J.L.; et al. Differences in 5’untranslated Regions Highlight the Importance of Translational Regulation of Dosage Sensitive Genes. Genome Biol. 2024, 25. [Google Scholar] [CrossRef]
- Dave, P.; Roth, G.; Griesbach, E.; Mateju, D.; Hochstoeger, T.; Chao, J.A. Single-Molecule Imaging Reveals Translation-Dependent Destabilization of MRNAs. Mol. Cell. 2023, 83. [Google Scholar] [CrossRef]
- Lindqvist, L.; Imataka, H.; Pelletier, J. Cap-Dependent Eukaryotic Initiation Factor-MRNA Interactions Probed by Cross-Linking. RNA 2008, 14. [Google Scholar] [CrossRef]
- Jia, X.; He, X.; Huang, C.; Li, J.; Dong, Z.; Liu, K. Protein Translation: Biological Processes and Therapeutic Strategies for Human Diseases. Signal Transduct. Target. Ther. 2024, 9. [Google Scholar] [CrossRef]
- Advani, V.M.; Ivanov, P. Translational Control under Stress: Reshaping the Translatome. BioEssays 2019, 41. [Google Scholar] [CrossRef]
- Sonenberg, N.; Hinnebusch, A.G. Regulation of Translation Initiation in Eukaryotes: Mechanisms and Biological Targets. Cell 2009, 136. [Google Scholar] [CrossRef]
- Sharma, A.K.; Sormanni, P.; Ahmed, N.; Ciryam, P.; Friedrich, U.A.; Kramer, G.; O’Brien, E.P. A Chemical Kinetic Basis for Measuring Translation Initiation and Elongation Rates from Ribosome Profiling Data. PLoS Comput. Biol. 2019, 15. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G.; Lorsch, J.R. The Mechanism of Eukaryotic Translation Initiation: New Insights and Challenges. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef] [PubMed]
- Kozak, M. Pushing the Limits of the Scanning Mechanism for Initiation of Translation. Gene 2002, 299. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, S.; Niimura, Y.; Gojobori, T.; Tanaka, H.; Miura, K. ichiro Diversity of Preferred Nucleotide Sequences around the Translation Initiation Codon in Eukaryote Genomes. Nucleic Acids Res. 2008, 36. [Google Scholar] [CrossRef]
- Lee, J.H.; Pestovat, T. V.; Shin, B.S.; Cao, C.; Choi, S.K.; Dever, T.E. Initiation Factor EIF5B Catalyzes Second GTP-Dependent Step in Eukaryotic Translation Initiation. Proc. Natl. Acad. Sci. U. S. A. 2002, 99. [Google Scholar] [CrossRef]
- Huang, B.Y.; Fernández, I.S. Long-Range Interdomain Communications in EIF5B Regulate GTP Hydrolysis and Translation Initiation. Proc. Natl. Acad. Sci. U. S. A. 2020, 117. [Google Scholar] [CrossRef] [PubMed]
- Costa-Mattioli, M.; Walter, P. The Integrated Stress Response: From Mechanism to Disease. Science 2020, 368. [Google Scholar] [CrossRef]
- Guan, B.J.; van Hoef, V.; Jobava, R.; Elroy-Stein, O.; Valasek, L.S.; Cargnello, M.; Gao, X.H.; Krokowski, D.; Merrick, W.C.; Kimball, S.R.; et al. A Unique ISR Program Determines Cellular Responses to Chronic Stress. Mol. Cell. 2017, 68. [Google Scholar] [CrossRef]
- Chee, N.T.; Lohse, I.; Brothers, S.P. MRNA-to-Protein Translation in Hypoxia. Mol. Cancer 2019, 18. [Google Scholar] [CrossRef]
- Shu, X.E.; Swanda, R. V.; Qian, S.B. Nutrient Control of MRNA Translation. Annu. Rev. Nutr. 2020, 40. [Google Scholar] [CrossRef]
- Panniers, R. Translational Control during Heat Shock. Biochimie 1994, 76. [Google Scholar] [CrossRef] [PubMed]
- Stern-Ginossar, N.; Thompson, S.R.; Mathews, M.B.; Mohr, I. Translational Control in Virus-Infected Cells. Cold Spring Harb. Perspect. Biol. 2019, 11. [Google Scholar] [CrossRef]
- Ghosh, A.; Shcherbik, N. Effects of Oxidative Stress on Protein Translation: Implications for Cardiovascular Diseases. Int. J. Mol. Sci. 2020, 21. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Hu, Y.; Wang, J.L.; Chatterjee, M.; Shi, Y.; Kaufman, R.J. Ultraviolet Light Inhibits Translation through Activation of the Unfolded Protein Response Kinase PERK in the Lumen of the Endoplasmic Reticulum. J. Biol. Chem. 2002, 277. [Google Scholar] [CrossRef]
- Mazroui, R.; Di Marco, S.; Kaufman, R.J.; Gallouzi, I.E. Inhibition of the Ubiquitin-Proteasome System Induces Stress Granule Formation. Mol. Biol. Cell. 2007, 18. [Google Scholar] [CrossRef]
- Castilho, B.A.; Shanmugam, R.; Silva, R.C.; Ramesh, R.; Himme, B.M.; Sattlegger, E. Keeping the EIF2 Alpha Kinase Gcn2 in Check. Biochim. Biophys. Acta Mol. Cell. Res. 2014, 1843. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lv, Y.; Zhao, N.; Guan, G.; Wang, J. Protein Kinase R-like ER Kinase and Its Role in Endoplasmic Reticulum Stress-Decided Cell Fate. Cell. Death Dis. 2015, 6. [Google Scholar] [CrossRef]
- Williams, B.R.G. PKR; A Sentinel Kinase for Cellular Stress. Oncogene 1999, 18. [Google Scholar] [CrossRef]
- Girardin, S.E.; Cuziol, C.; Philpott, D.J.; Arnoult, D. The EIF2α Kinase HRI in Innate Immunity, Proteostasis, and Mitochondrial Stress. FEBS J. 2021, 288. [Google Scholar] [CrossRef]
- Komar, A.A.; Hatzoglou, M. Cellular IRES-Mediated Translation: The War of ITAFs in Pathophysiological States. Cell. Cycle 2011, 10. [Google Scholar] [CrossRef]
- Spriggs, K.A.; Bushell, M.; Willis, A.E. Translational Regulation of Gene Expression during Conditions of Cell Stress. Mol. Cell. 2010, 40. [Google Scholar] [CrossRef]
- Topisirovic, I.; Svitkin, Y. V.; Sonenberg, N.; Shatkin, A.J. Cap and Cap-Binding Proteins in the Control of Gene Expression. Wiley Interdiscip. Rev. RNA 2011, 2. [Google Scholar] [CrossRef]
- Igreja, C.; Peter, D.; Weiler, C.; Izaurralde, E. 4E-BPs Require Non-Canonical 4E-Binding Motifs and a Lateral Surface of EIF4E to Repress Translation. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Niedzwiecka, A.; Marcotrigiano, J.; Stepinski, J.; Jankowska-Anyszka, M.; Wyslouch-Cieszynska, A.; Dadlez, M.; Gingras, A.C.; Mak, P.; Darzynkiewicz, E.; Sonenberg, N.; et al. Biophysical Studies of EIF4E Cap-Binding Protein: Recognition of MRNA 5′ Cap Structure and Synthetic Fragments of EIF4G and 4E-BP1 Proteins. J. Mol. Biol. 2002, 319. [Google Scholar] [CrossRef] [PubMed]
- Volarević, S.; Thomas, G. Role of S6 Phosphorylation and S6 Kinase in Cell Growth. Prog. Nucleic Acid. Res. Mol. Biol. 2000, 65. [Google Scholar]
- Ballard, D.J.; Peng, H.Y.; Das, J.K.; Kumar, A.; Wang, L.; Ren, Y.; Xiong, X.; Ren, X.; Yang, J.M.; Song, J. Insights Into the Pathologic Roles and Regulation of Eukaryotic Elongation Factor-2 Kinase. Front. Mol. Biosci. 2021, 8. [Google Scholar] [CrossRef]
- Dibble, C.C.; Cantley, L.C. Regulation of MTORC1 by PI3K Signaling. Trends Cell. Biol. 2015, 25. [Google Scholar] [CrossRef]
- Ling, N.X.Y.; Kaczmarek, A.; Hoque, A.; Davie, E.; Ngoei, K.R.W.; Morrison, K.R.; Smiles, W.J.; Forte, G.M.; Wang, T.; Lie, S.; et al. MTORC1 Directly Inhibits AMPK to Promote Cell Proliferation under Nutrient Stress. Nat. Metab. 2020, 2. [Google Scholar] [CrossRef]
- Hsieh, A.C.; Liu, Y.; Edlind, M.P.; Ingolia, N.T.; Janes, M.R.; Sher, A.; Shi, E.Y.; Stumpf, C.R.; Christensen, C.; Bonham, M.J.; et al. The Translational Landscape of MTOR Signalling Steers Cancer Initiation and Metastasis. Nature 2012, 485, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Deng, X.; Chen, J. RNA-Binding Proteins in Regulating MRNA Stability and Translation: Roles and Mechanisms in Cancer. Semin. Cancer Biol. 2022, 86. [Google Scholar] [CrossRef]
- Vind, A.C.; Genzor, A.V.; Bekker-Jensen, S. Ribosomal Stress-Surveillance: Three Pathways Is a Magic Number. Nucleic Acids Res. 2020, 48. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.K.L.; Sharp, P.A. MicroRNA Functions in Stress Responses. Mol. Cell. 2010, 40. [Google Scholar] [CrossRef] [PubMed]
- Gebetsberger, J.; Wyss, L.; Mleczko, A.M.; Reuther, J.; Polacek, N. A TRNA-Derived Fragment Competes with MRNA for Ribosome Binding and Regulates Translation during Stress. RNA Biol. 2017, 14. [Google Scholar] [CrossRef]
- Harvey, R.F.; Smith, T.S.; Mulroney, T.; Queiroz, R.M.L.; Pizzinga, M.; Dezi, V.; Villenueva, E.; Ramakrishna, M.; Lilley, K.S.; Willis, A.E. Trans-Acting Translational Regulatory RNA Binding Proteins. Wiley Interdiscip. Rev. RNA 2018, 9. [Google Scholar] [CrossRef]
- Van Nostrand, E.L.; Freese, P.; Pratt, G.A.; Wang, X.; Wei, X.; Xiao, R.; Blue, S.M.; Chen, J.Y.; Cody, N.A.L.; Dominguez, D.; et al. A Large-Scale Binding and Functional Map of Human RNA-Binding Proteins. Nature 2020, 583. [Google Scholar] [CrossRef]
- Rowe, W.; Kershaw, C.J.; Castelli, L.M.; Costello, J.L.; Ashe, M.P.; Grant, C.M.; Sims, P.F.G.; Pavitt, G.D.; Hubbard, S.J. Puf3p Induces Translational Repression of Genes Linked to Oxidative Stress. Nucleic Acids Res. 2014, 42. [Google Scholar] [CrossRef]
- Masuda, K.; Abdelmohsen, K.; Kim, M.M.; Srikantan, S.; Lee, E.K.; Tominaga, K.; Selimyan, R.; Martindale, J.L.; Yang, X.; Lehrmann, E.; et al. Global Dissociation of HuR-MRNA Complexes Promotes Cell Survival after Ionizing Radiation. EMBO J. 2011, 30. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Liu, S.; Fan, B.; Jin, D.; Miao, L.; Liu, L.; Du, S.; Lin, J. Enhancing MRNA Translation Efficiency by Introducing Sequence Optimized AU-Rich Elements in 3’ UTR via HuR Anchorage. Mol. Ther. Nucleic Acids 2025, 36. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. . 2018, 9. [Google Scholar] [CrossRef]
- Varghese, J.; Lim, S.F.; Cohen, S.M. Drosophila MiR-14 Regulates Insulin Production and Metabolism through Its Target, Sugarbabe. Genes. Dev. 2010, 24. [Google Scholar] [CrossRef]
- Rupaimoole, R.; Wu, S.Y.; Pradeep, S.; Ivan, C.; Pecot, C. V.; Gharpure, K.M.; Nagaraja, A.S.; Armaiz-Pena, G.N.; McGuire, M.; Zand, B.; et al. Hypoxia-Mediated Downregulation of MiRNA Biogenesis Promotes Tumour Progression. Nat. Commun. 2014, 5, 5202. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Lopez-Berestein, G.; Sood, A. MicroRNA Therapeutics: Principles, Expectations, and Challenges. Chin. J. Cancer 2011, 30, 368–370. [Google Scholar] [CrossRef]
- Vasudevan, S.; Tong, Y.; Steitz, J. Switching from Repression to Activation: MicroRNAs Can Up-Regulate Translation. Science 2008, 318, 1931–1934. [Google Scholar] [CrossRef]
- Lee, Y.S.; Shibata, Y.; Malhotra, A.; Dutta, A. A Novel Class of Small RNAs: TRNA-Derived RNA Fragments (TRFs). Genes. Dev. 2009, 23. [Google Scholar] [CrossRef] [PubMed]
- Fu, H.; Feng, J.; Liu, Q.; Sun, F.; Tie, Y.; Zhu, J.; Xing, R.; Sun, Z.; Zheng, X. Stress Induces TRNA Cleavage by Angiogenin in Mammalian Cells. FEBS Lett. 2009, 583. [Google Scholar] [CrossRef]
- Yamasaki, S.; Ivanov, P.; Hu, G.F.; Anderson, P. Angiogenin Cleaves TRNA and Promotes Stress-Induced Translational Repression. J. Cell. Biol. 2009, 185. [Google Scholar] [CrossRef]
- Goodarzi, H.; Liu, X.; Nguyen, H.C.B.; Zhang, S.; Fish, L.; Tavazoie, S.F. Endogenous TRNA-Derived Fragments Suppress Breast Cancer Progression via YBX1 Displacement. Cell 2015, 161. [Google Scholar] [CrossRef] [PubMed]
- Medina-Muñoz, S.G.; Kushawah, G.; Castellano, L.A.; Diez, M.; DeVore, M.L.; Salazar, M.J.B.; Bazzini, A.A. Crosstalk between Codon Optimality and Cis-Regulatory Elements Dictates MRNA Stability. Genome Biol. 2021, 22. [Google Scholar] [CrossRef]
- Delaunay, S.; Helm, M.; Frye, M. RNA Modifications in Physiology and Disease: Towards Clinical Applications. Nat. Rev. Genet. 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Meyer, K.D.; Patil, D.P.; Zhou, J.; Zinoviev, A.; Skabkin, M.A.; Elemento, O.; Pestova, T.V.; Qian, S.-B.; Jaffrey, S.R. 5’ UTR M6A Promotes Cap-Independent Translation. Cell 2015, 163, 999–1010. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Yi, Y.; Gao, X.; Wang, X.; Zhao, D.; Wang, R.; Zhang, L.-S.; Gao, B.; Zhang, Y.; Zhang, L.; et al. 2-O-Methylation at Internal Sites on MRNA Promotes MRNA Stability. Mol. Cell. 2024, 84, 2320–2336.e6. [Google Scholar] [CrossRef]
- Calvo, S.E.; Pagliarini, D.J.; Mootha, V.K. Upstream Open Reading Frames Cause Widespread Reduction of Protein Expression and Are Polymorphic among Humans. Proc. Natl. Acad. Sci. U. S. A. 2009, 106. [Google Scholar] [CrossRef]
- Xiao, W.; Sun, Y.; Xu, J.; Zhang, N.; Dong, L. UORF-Mediated Translational Regulation of ATF4 Serves as an Evolutionarily Conserved Mechanism Contributing to Non-Small-Cell Lung Cancer (NSCLC) and Stress Response. J. Mol. Evol. 2022, 90. [Google Scholar] [CrossRef]
- Jousse, C.; Bruhat, A.; Carraro, V.; Urano, F.; Ferrara, M.; Ron, D.; Fafournoux, P. Inhibition of CHOP Translation by a Peptide Encoded by an Open Reading Frame Localized in the Chop 5′UTR. Nucleic Acids Res. 2001, 29. [Google Scholar] [CrossRef]
- Chen, H.H.; Tarn, W.Y. UORF-Mediated Translational Control: Recently Elucidated Mechanisms and Implications in Cancer. RNA Biol. 2019, 16. [Google Scholar] [CrossRef]
- Yang, Y.; Gatica, D.; Liu, X.; Wu, R.; Kang, R.; Tang, D.; Klionsky, D.J. Upstream Open Reading Frames Mediate Autophagy-Related Protein Translation. Autophagy 2023, 19. [Google Scholar] [CrossRef]
- Russo, G.L.; Stampone, E.; Cervellera, C.; Borriello, A. Regulation of P27kip1 and P57kip2 Functions by Natural Polyphenols. Biomolecules 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Liang, J. Activating Transcription Factor 4: A Regulator of Stress Response in Human Cancers. Front. Cell. Dev. Biol. 2024, 12, 1370012. [Google Scholar] [CrossRef]
- Somers, J.; Pöyry, T.; Willis, A.E. A Perspective on Mammalian Upstream Open Reading Frame Function. Int. J. Biochem. Cell. Biol. 2013, 45, 1690–1700. [Google Scholar] [CrossRef]
- Young-Baird, S.; Wek, R. Upstream Open Reading Frames Differentially Regulate Gene-Specific Translation in the Integrated Stress Response. J. Biol. Chem. 2016, 291, jbc.R116.733899. [Google Scholar] [CrossRef]
- Cockman, E.; Anderson, P.; Ivanov, P. Top Mrnps: Molecular Mechanisms and Principles of Regulation. Biomolecules 2020, 10. [Google Scholar] [CrossRef]
- Philippe, L.; van den Elzen, A.M.G.; Watson, M.J.; Thoreen, C.C. Global Analysis of LARP1 Translation Targets Reveals Tunable and Dynamic Features of 5′ TOP Motifs. Proc. Natl. Acad. Sci. U. S. A. 2020, 117. [Google Scholar] [CrossRef] [PubMed]
- Tuxworth, W.J.; Shiraishi, H.; Moschella, P.C.; Yamane, K.; McDermott, P.J.; Kuppuswamy, D. Translational Activation of 5′-TOP MRNA in Pressure Overload Myocardium. Basic Res. Cardiol. 2008, 103, 41–53. [Google Scholar] [CrossRef]
- Jefferies, H.B.J.; Fumagalli, S.; Dennis, P.B.; Reinhard, C.; Pearson, R.B.; Thomas, G. Rapamycin Suppresses 5′TOP MRNA Translation through Inhibition of P70s6k. EMBO J. 1997, 16, 3693–3704. [Google Scholar] [CrossRef]
- Elfakess, R.; Sinvani, H.; Haimov, O.; Svitkin, Y.; Sonenberg, N.; Dikstein, R. Unique Translation Initiation of MRNAs-Containing TISU Element. Nucleic Acids Res. 2011, 39. [Google Scholar] [CrossRef] [PubMed]
- Elfakess, R.; Dikstein, R. A Translation Initiation Element Specific to MRNAs with Very Short 5′UTR That Also Regulates Transcription. PLoS ONE 2008, 3. [Google Scholar] [CrossRef]
- Sinvani, H.; Haimov, O.; Svitkin, Y.; Sonenberg, N.; Tamarkin-Ben-Harush, A.; Viollet, B.; Dikstein, R. Translational Tolerance of Mitochondrial Genes to Metabolic Energy Stress Involves TISU and EIF1-EIF4GI Cooperation in Start Codon Selection. Cell. Metab. 2015, 21. [Google Scholar] [CrossRef]
- Terenin, I.M.; Smirnova, V. V.; Andreev, D.E.; Dmitriev, S.E.; Shatsky, I.N. A Researcher’s Guide to the Galaxy of IRESs. Cell. Mol. Life Sci. 2017, 74. [Google Scholar] [CrossRef]
- Mokrejš, M.; Mašek, T.; Vopálenský, V.; Hlubuček, P.; Delbos, P.; Pospíšek, M. IRESite A Tool for the Examination of Viral and Cellular Internal Ribosome Entry Sites. Nucleic Acids Res. 2009, 38. [Google Scholar] [CrossRef]
- Komar, A.A.; Hatzoglou, M. Internal Ribosome Entry Sites in Cellular MRNAs: Mystery of Their Existence. J. Biol. Chem. 2005, 280, 23425–23428. [Google Scholar] [CrossRef]
- Holcik, M.; Sonenberg, N. Translational Control in Stress and Apoptosis. Nat. Rev. Mol. Cell. Biol. 2005, 6. [Google Scholar] [CrossRef]
- Gao, X.; Wu, Z. IRES-Mediated Translation: Expanding the Toolkits of RNA Therapy. Int. J. Mol. Sci. 2025, 26, 10542. [Google Scholar] [CrossRef]
- Sorokin, I.I.; Vassilenko, K.S.; Terenin, I.M.; Kalinina, N.O.; Agol, V.I.; Dmitriev, S.E. Non-Canonical Translation Initiation Mechanisms Employed by Eukaryotic Viral MRNAs. Biochemistry 2021, 86. [Google Scholar] [CrossRef]
- Andreev, D.E.; Dmitriev, S.E.; Terenin, I.M.; Prassolov, V.S.; Merrick, W.C.; Shatsky, I.N. Differential Contribution of the M7G-Cap to the 5′ End-Dependent Translation Initiation of Mammalian MRNAs. Nucleic Acids Res. 2009, 37. [Google Scholar] [CrossRef]
- Shatsky, I.N.; Terenin, I.M.; Smirnova, V. V.; Andreev, D.E. Cap-Independent Translation: What’s in a Name? Trends Biochem. Sci. 2018, 43, 882–895. [Google Scholar] [CrossRef]
- Liu, Y.; Cui, J.; Hoffman, A.R.; Hu, J.-F. Eukaryotic Translation Initiation Factor EIF4G2 Opens Novel Paths for Protein Synthesis in Development, Apoptosis and Cell Differentiation. Cell. Prolif. 2023, 56, e13367. [Google Scholar] [CrossRef]
- Roiuk, M.; Neff, M.; Teleman, A.A. EIF4E-Independent Translation Is Largely EIF3d-Dependent. Nat. Commun. 2024, 15, 6692. [Google Scholar] [CrossRef]
- Lee, A.S.Y.; Kranzusch, P.J.; Cate, J.H.D. EIF3 Targets Cell-Proliferation Messenger RNAs for Translational Activation or Repression. Nature 2015, 522. [Google Scholar] [CrossRef]
- Hayek, H.; Gross, L.; Janvier, A.; Schaeffer, L.; Martin, F.; Eriani, G.; Allmang, C. EIF3 Interacts with Histone H4 Messenger RNA to Regulate Its Translation. J. Biol. Chem. 2021, 296. [Google Scholar] [CrossRef]
- Lamper, A.M.; Fleming, R.H.; Ladd, K.M.; Lee, A.S.Y. A Phosphorylation-Regulated EIF3d Translation Switch Mediates Cellular Adaptation to Metabolic Stress. Science (1979) . 2020, 370. [Google Scholar] [CrossRef]
- Castillo-Hair, S.; Fedak, S.; Wang, B.; Linder, J.; Havens, K.; Certo, M.; Seelig, G. Optimizing 5’UTRs for MRNA-Delivered Gene Editing Using Deep Learning. Nat. Commun. 2024, 15. [Google Scholar] [CrossRef]
- Castillo-Hair, S.M.; Seelig, G. Machine Learning for Designing Next-Generation MRNA Therapeutics. Acc. Chem. Res. 2022, 55. [Google Scholar] [CrossRef]
- Koletsou, E.; Huppertz, I. RNA-Binding Proteins as Versatile Metabolic Regulators. Npj Metab. Health Dis. 2025, 3, 1. [Google Scholar] [CrossRef]
- Singh, S.; Shyamal, S.; Das, A.; Panda, A.C. Global Identification of MRNA-Interacting Circular RNAs by CLiPPR-Seq. Nucleic Acids Res. 2024, 52. [Google Scholar] [CrossRef]
- Raza, F.; Waldron, J.A.; Le Quesne, J. Translational Dysregulation in Cancer: EIF4A Isoforms and Sequence Determinants of EIF4A Dependence. Biochem. Soc. Trans. 2015, 43. [Google Scholar] [CrossRef]
- Weber, R.; Ghoshdastider, U.; Spies, D.; Duré, C.; Valdivia-Francia, F.; Forny, M.; Ormiston, M.; Renz, P.F.; Taborsky, D.; Yigit, M.; et al. Monitoring the 5′UTR Landscape Reveals Isoform Switches to Drive Translational Efficiencies in Cancer. Oncogene 2023, 42. [Google Scholar] [CrossRef] [PubMed]
- Drazkowska, K.; Tomecki, R.; Warminski, M.; Baran, N.; Cysewski, D.; Depaix, A.; Kasprzyk, R.; Kowalska, J.; Jemielity, J.; Sikorski, P.J. 2′-O-Methylation of the Second Transcribed Nucleotide within the MRNA 5′ Cap Impacts the Protein Production Level in a Cell-Specific Manner and Contributes to RNA Immune Evasion. Nucleic Acids Res. 2022, 50. [Google Scholar] [CrossRef] [PubMed]
- Anderson, B.R.; Muramatsu, H.; Nallagatla, S.R.; Bevilacqua, P.C.; Sansing, L.H.; Weissman, D.; Karikó, K. Incorporation of Pseudouridine into MRNA Enhances Translation by Diminishing PKR Activation. Nucleic Acids Res. 2010, 38, 5884–5892. [Google Scholar] [CrossRef]
- Warminski, M.; Trepkowska, E.; Smietanski, M.; Sikorski, P.J.; Baranowski, M.R.; Bednarczyk, M.; Kedzierska, H.; Majewski, B.; Mamot, A.; Papiernik, D.; et al. Trinucleotide MRNA Cap Analogue N6-Benzylated at the Site of Posttranscriptional M6Am Mark Facilitates MRNA Purification and Confers Superior Translational Properties In Vitro and In Vivo. J. Am. Chem. Soc. 2024, 146. [Google Scholar] [CrossRef]
- Sultana, N.; Hadas, Y.; Sharkar, M.; Kaur, K.; Magadum, A.; Kurian, A.; Hossain, N.; Alburquerque, B.; Ahmed, S.; Chepurko, E.; et al. Optimization of 5’ Untranslated Region of Modified MRNA for Use in Cardiac or Hepatic Ischemic Injury. Mol. Ther. Methods Clin. Dev. 2020, 17. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



