Submitted:
24 April 2026
Posted:
27 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
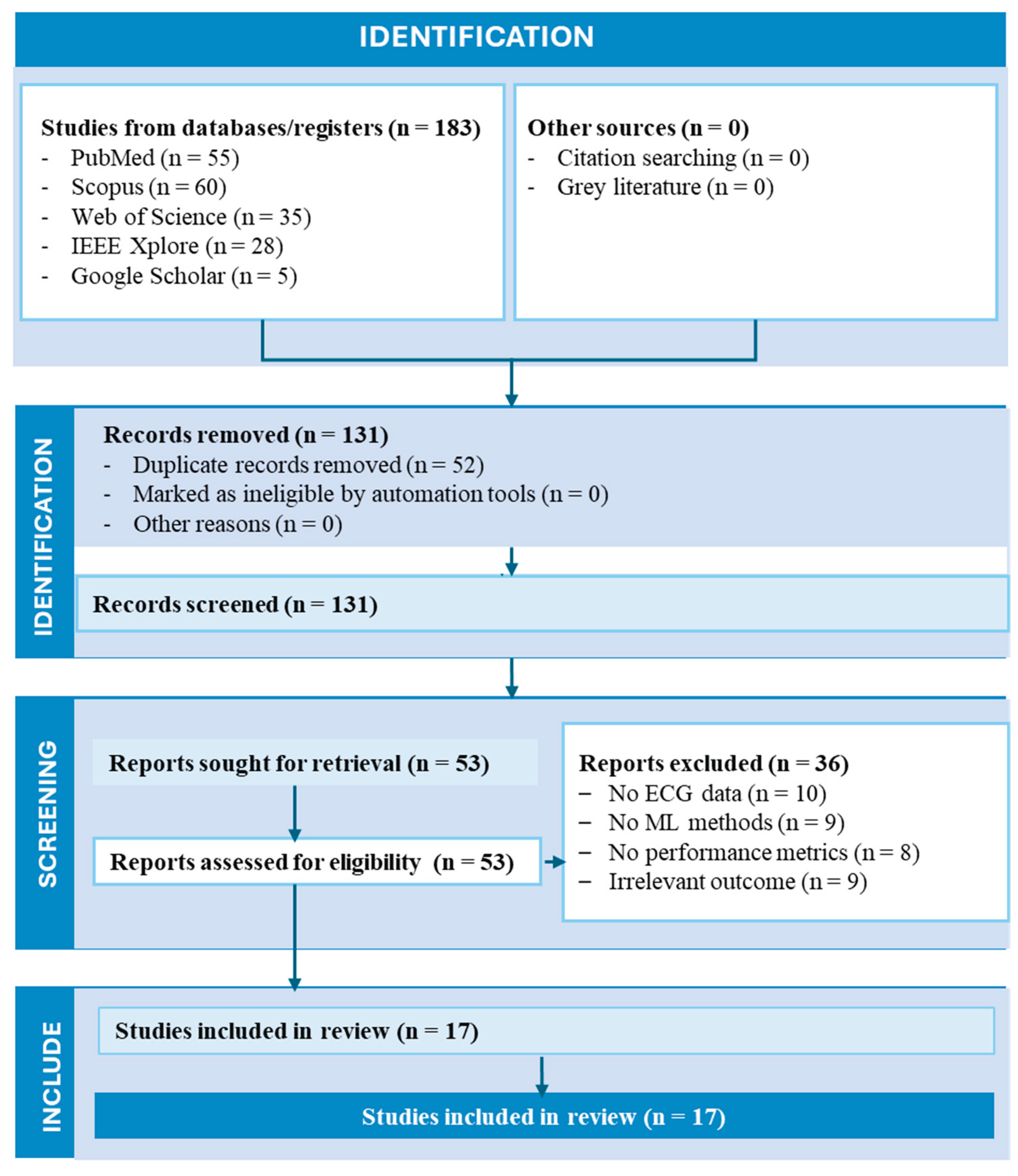
2. Literature Search Methodology
2.1. Search Strategy
2.2. Eligibility Criteria
2.3. Study Selection and Data Extraction
2.4. Risk of Bias Assessment
2.5. Data Synthesis
3. Results
3.1. General Characteristics and Aims
| Study | Year | Signal | Dataset | ECG | N | Population | Marker | Design | Model | Performance | Limitations | Maturity |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| [29] | 2021 | ECG | Outpatient cohort (hospital-based) | 12-lead, 500 Hz, 10 s; intervals (HR, PR, QRS, QT, QTc), axes (P, QRS, T) | 4,832 | Non-DM, prediabetes, T2DM; mean duration ~4.7 y | HbA1c | Retrospective cohort (validated) | CNN-based DL (ResNet + SE + attention) | AUC 0.826; Sens 71.9%; Spec 77.7% | Moderate accuracy; reduced performance in severe DM; single-center | Level 3 (retrospective clinical validation) |
| [30] | 2023 | ECG | Ethnic cohort (Sindhi, India; high-risk families) | 12-lead, 10 s, 1000 Hz | 1,262 (10,461 beats) | Mean age ~48 y; 61% female; high cardiometabolic burden | HbA1c, FPG, RBG | Observational; train/val/test split | XGBoost (best); compared with RF, MLP, LSTM, CNN, Transformer | Acc 96.8%; Prec 97.1%; Rec 96.2%; F1 96.6% | Selection bias; no external validation; beat-level analysis; limited generalizability | Level 2 (model development, internal validation) |
| [31] | 2022 | ECG | Private dataset (non-public) | Single-lead; 256 Hz; resting | 86 (24,630 segments) | 35 T2DM / 51 healthy; age 20–70 y | Glucose (≥160 mg/dL) | Supervised classification | Decision Tree (DTC); compared with FT, MT, CT | Acc 86.9%; Sens 81.9%; Spec 90.6%; F1 82.8% | Small sample; no external validation; private dataset; limited generalizability | Level 2–3 (prototype; limited clinical validation) |
| [32] | 2021 | ECG | Private dataset (Taiwan; ECG + glucose) | Single-lead; 1000 Hz; 60 s | 1,119 | Age 38–80 y; mixed glycemic status | Blood glucose (≥100 mg/dL) | Retrospective; binary classification; 80/20 sp lit + CV | Deep NN (10-layer); compared with LR, SVM | AUC 0.945; Sens 87.6%; Spec 85.0% | Private dataset; no external validation; sensitive to signal quality | Level 3 (advanced ML validation) |
| [33] | 2020 | ECG | Self-collected dataset | 3-electrode setup (wrist + ankles) | 24 (~1,500 samples) | 10 diabetic / 14 healthy | Clinical status (no HbA1c/glucose) | Experimental; 5-fold CV | SVM (cubic); compared with DT, LDA, NB, KNN | Acc 96.8% | Very small sample; no objective biomarkers; no external validation; high overfitting risk | Level 1–2 (proof-of-concept) |
| [34] | 2021 | ECG | Hospital-based dataset (wearable ECG) | Single-lead; 60 s segments | 370 (~317k segments) | T2DM only; mean age ~43.5 y | HbA1c | Retrospective; 5-fold CV | CNN-MFVW; compared with CNN, CNN-LSTM | Acc 90.2%; AUC 0.990; F1 0.901 | No control group; small cohort; no external validation; sensitive to preprocessing | Level 2–3 (model development; limited clinical validation) |
| [35] | 2021 | ECG | Self-collected experimental dataset | Single-lead; 1000 Hz | 21 (~22k segments) | Young adults; mixed glycemic status | Blood glucose (OGTT) | Prospective; 3-class classification | DBSCAN + CNN | Acc 81.7%; Sens 98.5%; Spec 76.8% | Very small sample; controlled setting; selection bias; no external validation | Level 2 (early-stage experimental study) |
| [36] | 2025 | ECG + clinical (multimodal) | Population-based cohort (Qatar Biobank) | 12-lead (clinical) | 2,043 + 395 (test) | Middle Eastern; mean age ~46 y | HbA1c, FPG | Cross-sectional + longitudinal (5-year follow-up) | DNN (ECG-DiaNet; ECG + CRFs) | AUC 0.845 (multimodal); 0.822 (CRF); 0.675 (ECG) | No external validation; single-region cohort; small longitudinal test set | Level 3 (advanced clinical ML; longitudinal validation) |
| [37] | 2025 | HRV (ECG-derived) | Retrospective cohort (AFT lab, India) | Lead II; 1000 Hz; 5-min segments | 519 (261 T2DM / 258 controls) | Age 18–55 y; no major comorbidities | FBG, PPBG, HbA1c | Retrospective; binary classification; 80/20 split | CatBoost (best); compared with LR, KNN, RF, GBM | Acc 91.3%; AUC 0.91; Sens 90.6%; Spec 91.9% | No external validation; controlled setting; HRV-only features; limited generalizability | Level 2–3 (validated ML model) |
| [38] | 2025 | ECG (engineered features) | Population-based cohort (Japan; external validation) | 12-lead; 10 s; 500 Hz | 16,766 + 2,456 (external) | General population; higher risk in older subjects | FPG, HbA1c | Retrospective; internal + external validation | LightGBM (best); compared with LR, RF, XGBoost, DNN | AUC 0.851 (internal); 0.785 (external) | Feature-based (no raw ECG DL); moderate specificity; class imbalance | Level 4 (advanced clinical ML with external validation) |
| [39] | 2022 | ECG + demographics (multimodal) | EHR cohort (NYU Langone) | 12-lead; 10 s; 250–500 Hz | 25,951 (test); large training cohort | Outpatients; new-onset diabetes subgroup | HbA1c ≥ 6.5% | Retrospective; prediction; external validation | DL (ResNet); ECG + demographics | AUC 0.80 (model); 0.68 (risk score) | Selection bias; multimodal dependence; no real-world validation; data not public | Level 4 (advanced clinical ML; near-translational) |
| [40] | 2022 | ECG (image-based) | Hospital cohort (China; 3 centers) | 12-lead ECG images; 5 s | ~2,914 | Middle-aged/elderly; high-risk | FPG, OGTT | Retrospective; binary classification; CV + test set | CNN (JGRNet); compared with AlexNet, GoogleNet, SVM | Acc 0.781; AUC 0.777 | Image-based ECG (information loss); no external validation; moderate performance | Level 2–3 (early DL with internal validation) |
| [41] | 2023 | Multimodal (ECG + glucose + ACC + respiration) | DINAMO wearable dataset (free-living) | Wearable ECG; 250 Hz; continuous (~4 days) | 29 (20 healthy / 9 diabetic) | Mixed cohort; continuous monitoring | Continuous glucose | Experimental; supervised classification | XGBoost (best); compared with LR, DT, RF, SVM | Acc 98.2% (multimodal); ~87.5% (ECG only) | Very small sample; uses glucose input; no external validation; high overfitting risk | Level 1–2 (exploratory multimodal study) |
| [42] | 2024 | ECG (high-density) | Private dataset (self-collected) | HD-ECG (up to 98 leads) | 50 | Healthy volunteers | Not specified | Experimental; supervised classification | CNN (HD-MVCNN) | Acc 99.0%; F1 94.5% | No glycemic ground truth; unclear labels; small sample; unrealistic setup (98 leads); no validation | Level 1 (concept study) |
| [43] | 2023 | ECG | MIMIC-III (ICU subset) | Single-lead; 125 Hz; 1 s windows | 50 | ICU patients; median age 64 y | Blood glucose | Retrospective; personalized classification | One-class SVM | AUC 0.92 (beat); 0.97 (10 s) | ICU-only cohort; small sample; personalized model; no external validation | Level 3 (advanced ML validation) |
| [44] | 2017 | HRV (RR-interval) | Public dataset (PhysioNet) | RR intervals (QRS-based) | 50 (33 normal / 17 diabetic) | Not specified | Not reported | Supervised classification | SVM | Acc ~95% | Very small sample; no glycemic markers; unclear labels; no external validation | Level 2 (early-stage study) |
| [45] | 2024 | HRV (ECG-derived) | Hospital cohort (Korea; prospective) | Wearable ECG; 250 Hz | 83 → 21 (final) | T2DM only; elderly (mean ~69 y) | Continuous glucose | Observational; temporal prediction | 1D CNN (ResNet-like; HRV input) | Acc 90.5%; Sens 87.5%; Spec 92.7% | Very small final cohort; no control group; HRV-only; no external validation | Level 3 (clinical ML validation) |
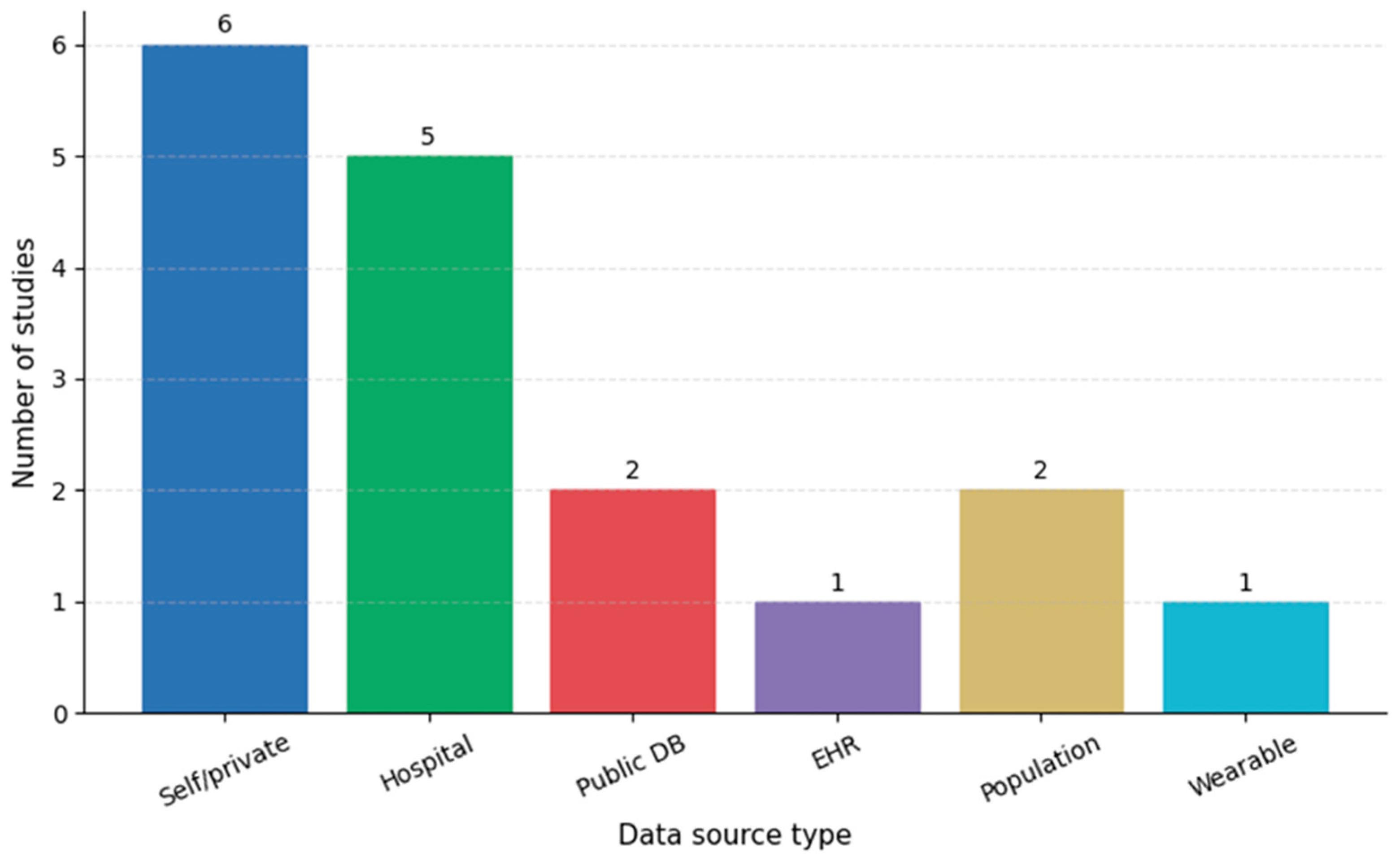
3.2. Data Sources and Study Populations
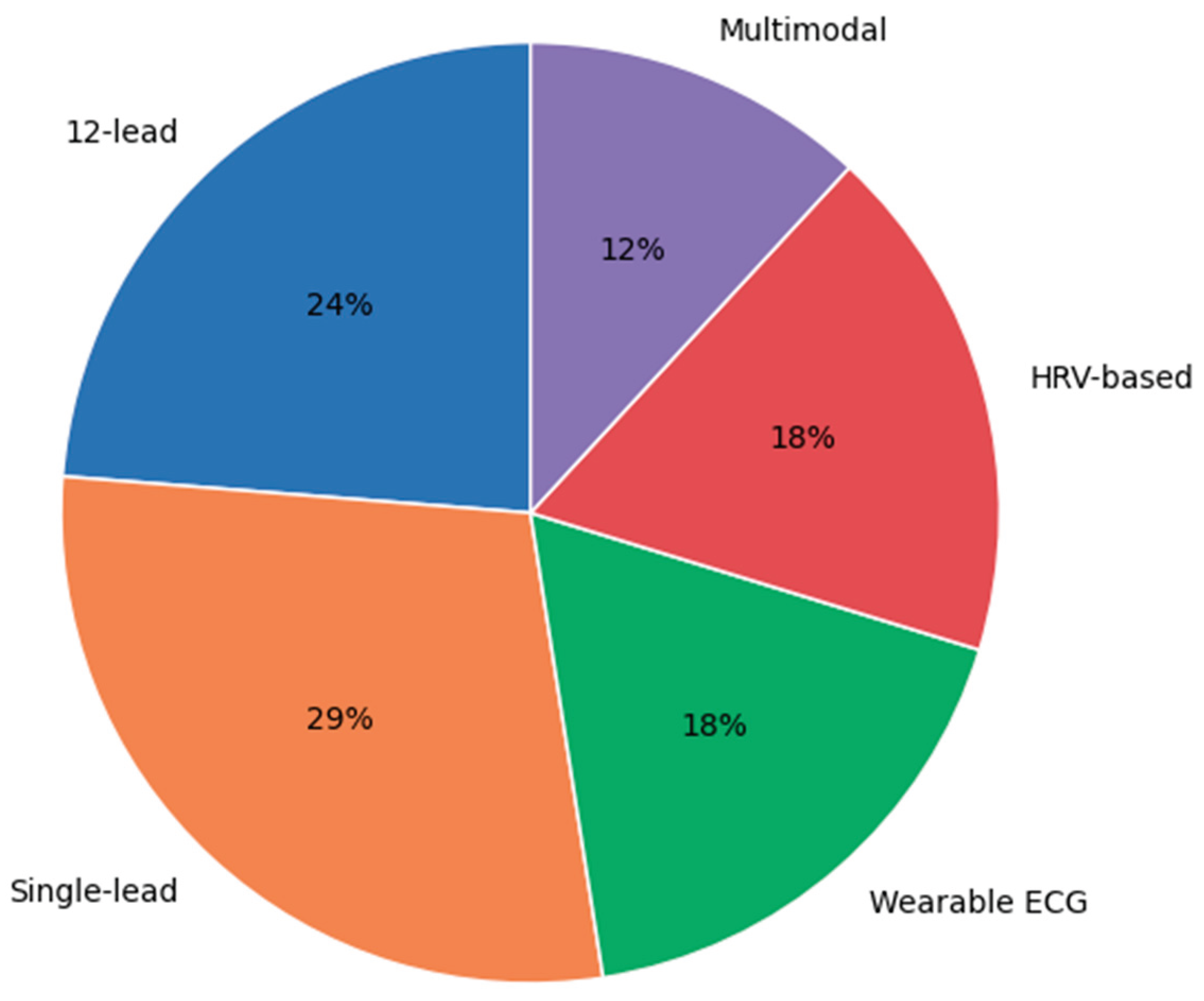
3.3. ECG Acquisition and Signal Configuration
3.4. Feature Representation and ECG-Derived Biomarkers
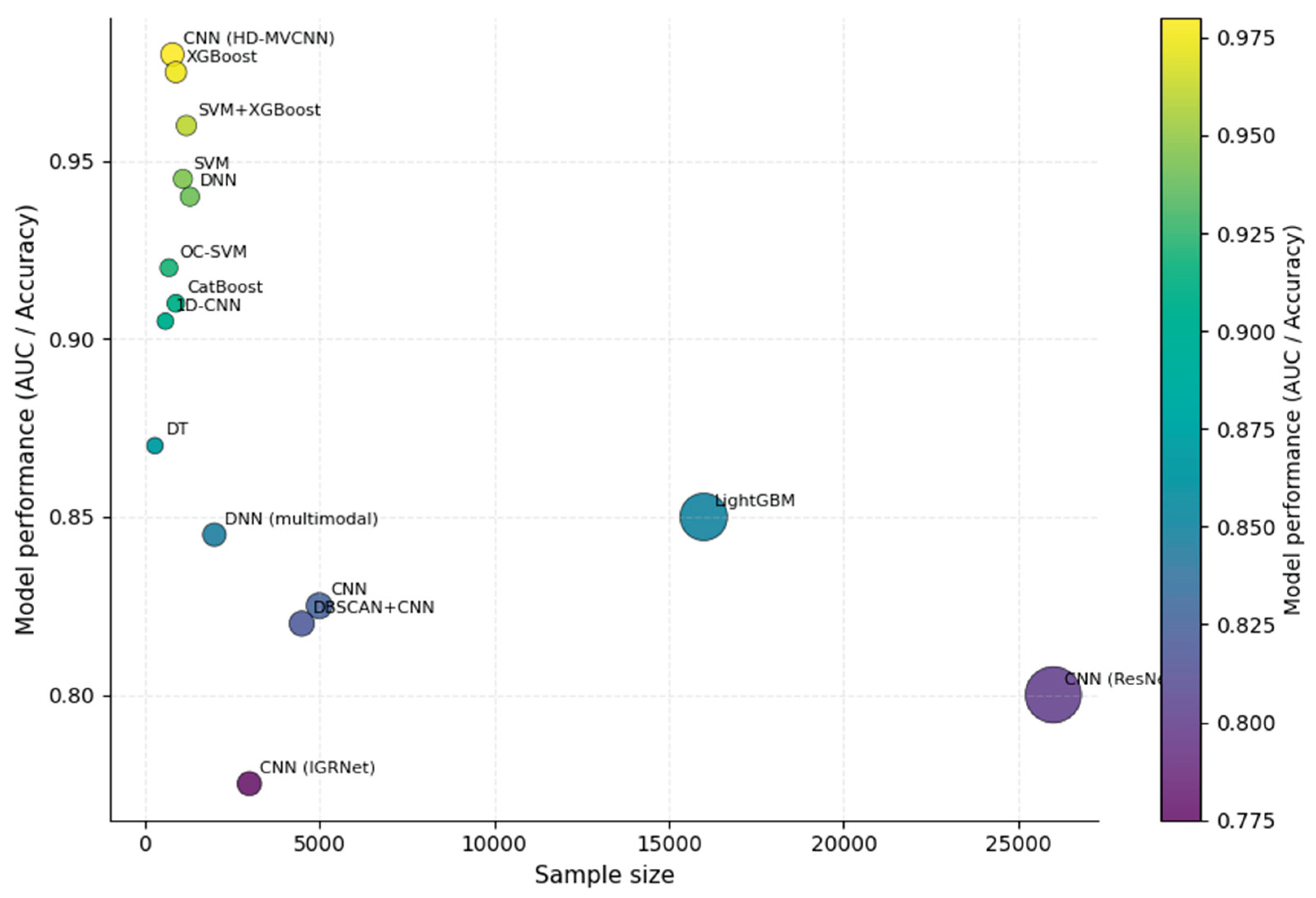
3.5. Machine Learning Models
3.6. Model Performance and Validation
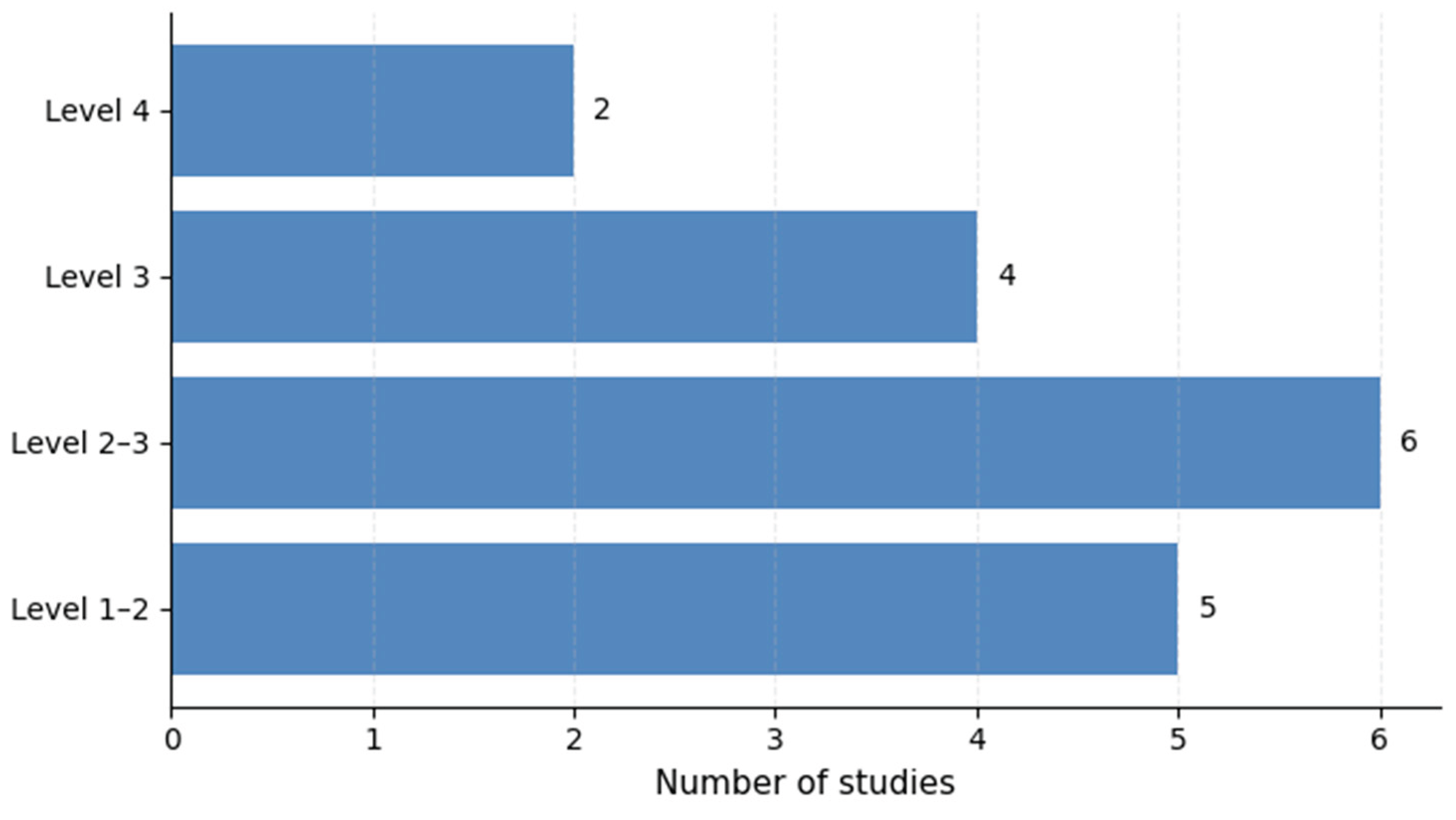
3.7. Model Maturity and Translational Readiness
4. Discussion
| Domain | Key finding | Strengths | Limitations | Required improvement |
|---|---|---|---|---|
| Data characteristics | ECG-based dysglycemia detection is feasible across multiple datasets | Large-scale studies demonstrate predictive potential | Most studies rely on small, single-center datasets; limited diversity | Large, multi-center, population-level datasets |
| ECG acquisition | Signal characteristics strongly influence model performance | Multilead ECG provides richer physiological information | Heterogeneous acquisition protocols; frequent use of single-lead ECG | Standardized, wearable multilead ECG systems |
| Feature representation | Both engineered and deep features capture relevant information | HRV and repolarization features show physiological relevance | Lack of standardization; inconsistent preprocessing | Hybrid feature frameworks with standardized pipelines |
| Machine learning models | ML and DL models achieve high performance under controlled conditions | CNN, boosting models show strong results | Performance depends on dataset rather than model; limited interpretability | Robust, interpretable models validated across datasets |
| Model performance | High reported accuracy in many studies | Strong results in experimental settings | Overfitting, optimistic bias, poor comparability | Standardized evaluation metrics and protocols |
| Validation strategy | Validation is a key bottleneck | Some studies include external validation | Most rely on internal validation; data leakage risk | External and prospective validation |
| Model maturity | Majority of studies at early/intermediate levels | Emerging Level 3–4 studies | Limited translational readiness | Maturity-driven development frameworks |
| Clinical applicability | ECG has potential for non-invasive screening | Scalable and low-cost modality | No real-world deployment; lack of screening studies | Integration into clinical workflows and screening programs |
| System integration | End-to-end systems are required | Advances in wearable ECG devices | Fragmented pipelines; lack of standardization | Integrated acquisition–ML–validation systems |
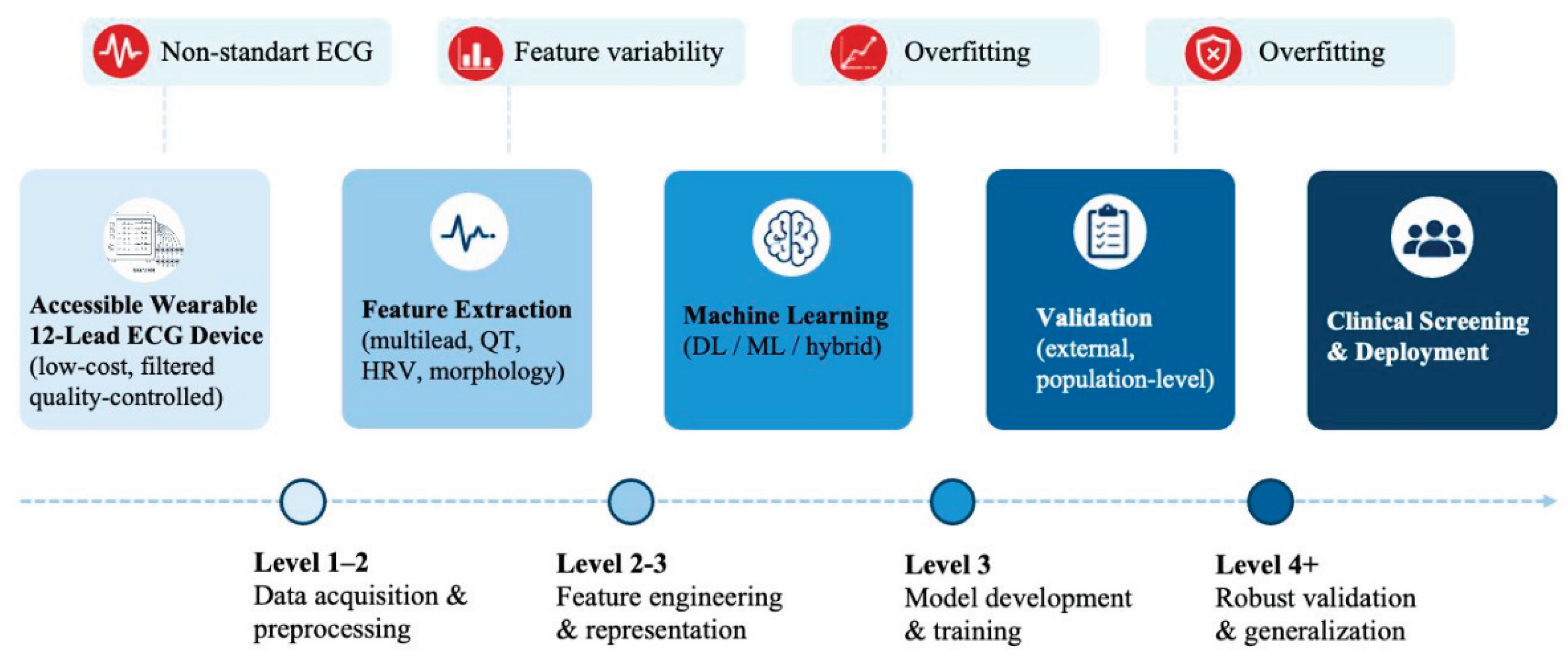
4.1. Requirements for Clinical Translation
4.2. Toward Practical ECG-Based Screening Systems
5. Limitations of This Review
6. Conclusion
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| ANOVA | Analysis of Variance |
| AUC | Area Under the Curve |
| BMI | Body Mass Index |
| CAN | Cardiac Autonomic Neuropathy |
| CAD | Coronary Artery Disease |
| CEEMDAN | Complete Ensemble Empirical Mode Decomposition with Adaptive Noise |
| CGM | Continuous Glucose Monitoring |
| CMR | Cardiac Magnetic Resonance |
| CMD | Coronary Microvascular Dysfunction |
| CNN | Convolutional Neural Network |
| CV | Cross-Validation |
| DCM | Diabetic Cardiomyopathy |
| DL | Deep Learning |
| DLM | Deep Learning Model |
| ECG | Electrocardiogram |
| EHR | Electronic Health Record |
| EMD | Empirical Mode Decomposition |
| F1-score | Harmonic Mean of Precision and Recall |
| FPG | Fasting Plasma Glucose |
| GBM | Gradient Boosting Machine |
| GRI | Glycaemia Risk Index |
| Grad-CAM | Gradient-weighted Class Activation Mapping |
| HbA1c | Glycated Hemoglobin |
| HR | Heart Rate |
| HRV | Heart Rate Variability |
| IFG | Impaired Fasting Glucose |
| KNN | k-Nearest Neighbors |
| LR | Logistic Regression |
| LSTM | Long Short-Term Memory |
| ML | Machine Learning |
| NB | Naïve Bayes |
| OGTT | Oral Glucose Tolerance Test |
| PPBG | Postprandial Blood Glucose |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses |
| RF | Random Forest |
| ROC | Receiver Operating Characteristic |
| SE | Squeeze-and-Excitation |
| SVM | Support Vector Machine |
| T2DM | Type 2 Diabetes Mellitus |
References
- Nanda, M., Sharma, R., Mubarik, S. et al. Type-2 Diabetes Mellitus (T2DM): Spatial-temporal Patterns of Incidence, Mortality and Attributable Risk Factors from 1990 to 2019 among 21 World Regions. Endocrine 77, 444–454 (2022). [CrossRef]
- Atageldiyeva, K., Syssoyev, D., Mussina, K. et al. All-cause hospital admissions and incidence of type 2 diabetes among adolescents in Kazakhstan. Sci Rep 15, 20746 (2025). [CrossRef]
- Ziegler D, Herder C, Papanas N. Neuropathy in prediabetes. Diabetes Metab Res Rev. 2023;39(8):e3693. [CrossRef]
- Neil H. White, Qing Pan, William C. Knowler, Emily B. Schroeder, Dana Dabelea, Emily Y. Chew, Barbara Blodi, Ronald B. Goldberg, Xavier Pi-Sunyer, Christine Darwin, Mathias Schlögl, David M. Nathan, for the Diabetes Prevention Program Outcome Study (DPPOS) Research Group; Risk Factors for the Development of Retinopathy in Prediabetes and Type 2 Diabetes: The Diabetes Prevention Program Experience. Diabetes Care 1 November 2022; 45 (11): 2653–2661. [CrossRef]
- Ahmad, A., Lim, LL., Morieri, M.L. et al. Precision prognostics for cardiovascular disease in Type 2 diabetes: a systematic review and meta-analysis. Commun Med 4, 11 (2024). [CrossRef]
- Genuth SM, Palmer JP, Nathan DM. Classification and Diagnosis of Diabetes. In: Diabetes in America. 3rd ed. National Institute of Diabetes and Digestive and Kidney Diseases (US), Bethesda (MD); 2018. PMID: 33651569.
- Thomas, A., Shenoy, M. T., Shenoy, K., & George, N. . (2021). Glucometers for Patients with Type 2 Diabetes Mellitus: Are they helpful?. International Journal of Medical Students, 9(2), 140–144. [CrossRef]
- Samuel Seidu, Setor K. Kunutsor, Ramzi A. Ajjan, Pratik Choudhary; Efficacy and Safety of Continuous Glucose Monitoring and Intermittently Scanned Continuous Glucose Monitoring in Patients With Type 2 Diabetes: A Systematic Review and Meta-analysis of Interventional Evidence. Diabetes Care 2 January 2024; 47 (1): 169–179. [CrossRef]
- Swapna, G., Soman, K.P., Vinayakumar, R. (2020). Diabetes Detection Using ECG Signals: An Overview. In: Dash, S., Acharya, B., Mittal, M., Abraham, A., Kelemen, A. (eds) Deep Learning Techniques for Biomedical and Health Informatics. Studies in Big Data, vol 68. Springer, Cham. [CrossRef]
- Balcıoğlu AS, Müderrisoğlu H. Diabetes and cardiac autonomic neuropathy: Clinical manifestations, cardiovascular consequences, diagnosis and treatment. World J Diabetes. 2015 Feb 15;6(1):80-91. doi: 10.4239/wjd.v6.i1.80. PMID: 25685280; PMCID: PMC4317320. [CrossRef]
- Chirag H. Mandavia, Annayya R. Aroor, Vincent G. DeMarco, James R. Sowers, Molecular and metabolic mechanisms of cardiac dysfunction in diabetes, Life Sciences, Volume 92, Issue 11, 2013, Pages 601-608, ISSN 0024-3205. [CrossRef]
- Adeghate, E., Singh, J. Structural changes in the myocardium during diabetes-induced cardiomyopathy. Heart Fail Rev 19, 15–23 (2014). [CrossRef]
- Jonas L. Isaksen, Christian B. Sivertsen, Christian Zinck Jensen, Claus Graff, Dominik Linz, Christina Ellervik, Magnus T. Jensen, Peter G. Jørgensen, Jørgen K. Kanters, Electrocardiographic markers in patients with type 2 diabetes and the role of diabetes duration, Journal of Electrocardiology, Volume 84, 2024, Pages 129-136, ISSN 0022-0736. [CrossRef]
- Kuehl, M., Stevens, M. Cardiovascular autonomic neuropathies as complications of diabetes mellitus. Nat Rev Endocrinol 8, 405–416 (2012). [CrossRef]
- Filipović N, Marinović Guić M, Košta V, Vukojević K. Cardiac innervations in diabetes mellitus-Anatomical evidence of neuropathy. Anat Rec (Hoboken). 2023 Sep;306(9):2345-2365. doi: 10.1002/ar.25090. Epub 2022 Oct 17. PMID: 36251628. [CrossRef]
- Sudo, S.Z.; Montagnoli, T.L.; Rocha, B.d.S.; Santos, A.D.; de Sá, M.P.L.; Zapata-Sudo, G. Diabetes-Induced Cardiac Autonomic Neuropathy: Impact on Heart Function and Prognosis. Biomedicines 2022, 10, 3258. [CrossRef]
- Balcıoğlu AS, Müderrisoğlu H. Diabetes and cardiac autonomic neuropathy: Clinical manifestations, cardiovascular consequences, diagnosis and treatment. World J Diabetes. 2015 Feb 15;6(1):80-91. doi: 10.4239/wjd.v6.i1.80. PMID: 25685280; PMCID: PMC4317320. [CrossRef]
- Evans, A.J.; Li, Y.-L. Remodeling of the Intracardiac Ganglia During the Development of Cardiovascular Autonomic Dysfunction in Type 2 Diabetes: Molecular Mechanisms and Therapeutics. Int. J. Mol. Sci. 2024, 25, 12464. [CrossRef]
- Tarvainen MP, Laitinen TP, Lipponen JA, Cornforth DJ and Jelinek HF (2014) Cardiac Autonomic Dysfunction in Type 2 Diabetes – Effect of Hyperglycemia and Disease Duration. Front. Endocrinol. 5:130. [CrossRef]
- Qian LL, Liu XY, Li XY, Yang F, Wang RX. Effects of Electrical Remodeling on Atrial Fibrillation in Diabetes Mellitus. Rev Cardiovasc Med. 2023 Jan 3;24(1):3. doi: 10.31083/j.rcm2401003. PMID: 39076858; PMCID: PMC11270397. [CrossRef]
- Charlotte Coopmans, Tan Lai Zhou, Ronald M.A. Henry, Jordi Heijman, Nicolaas C. Schaper, Annemarie Koster, Miranda T. Schram, Carla J.H. van der Kallen, Anke Wesselius, Robert J.A. den Engelsman, Harry J.G.M. Crijns, Coen D.A. Stehouwer; Both Prediabetes and Type 2 Diabetes Are Associated With Lower Heart Rate Variability: The Maastricht Study. Diabetes Care 1 May 2020; 43 (5): 1126–1133. [CrossRef]
- Alam, Krishna Chaitanya; Dasari, Dhanunjaya; Modampuri, Akhil Koundinya. A clinical study of corrected QT interval in type 2 diabetes mellitus patients. MRIMS Journal of Health Sciences 12(4):p 268-273, Oct–Dec 2024. [CrossRef]
- Chávez-González E, Calero YME, Harrichand S, Mensah EB. QRS and QT Interval Modifications in Patients with Type 2 Diabetes Mellitus. Curr Health Sci J. 2022 Jul-Sep;48(3):270-276. doi: 10.12865/CHSJ.48.03.04. Epub 2022 Sep 30. PMID: 36815079; PMCID: PMC9940933. [CrossRef]
- Singh, R.M., Waqar, T., Howarth, F.C. et al. Hyperglycemia-induced cardiac contractile dysfunction in the diabetic heart. Heart Fail Rev 23, 37–54 (2018). [CrossRef]
- Bakkar, N.-M.Z.; Dwaib, H.S.; Fares, S.; Eid, A.H.; Al-Dhaheri, Y.; El-Yazbi, A.F. Cardiac Autonomic Neuropathy: A Progressive Consequence of Chronic Low-Grade Inflammation in Type 2 Diabetes and Related Metabolic Disorders. Int. J. Mol. Sci. 2020, 21, 9005. [CrossRef]
- Kiruthika Balakrishnan, Durgadevi Velusamy, Karthikeyan Ramasamy, Hana E. Hinkle, Holly J. Hudson, Ram Bilas Pachori, Hikmat Khan, Artificial intelligence approaches for non-invasive diabetes prediction using ECG signals: A systematic review, Computer Methods and Programs in Biomedicine, Volume 278, 2026, 109264, ISSN 0169-2607. [CrossRef]
- Alimbayev, C.; Alimbayeva, Z.; Ozhikenov, K.; Karibayev, K.; Orynbay, Z.; Igembay, Y.; Daniyalov, M.; Nurdanali, A. Electrocardiographic Signatures of Dysglycaemia: Mechanistic Foundations, Digital Biomarkers, and Artificial Intelligence for Non-Invasive Diabetes Risk Stratification. Appl. Sci. 2026, 16, 2902. [CrossRef]
- Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA, Stewart LA, Thomas J, Tricco AC, Welch VA, Whiting P, Moher D. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. Int J Surg. 2021 Apr;88:105906. doi: 10.1016/j.ijsu.2021.105906. Epub 2021 Mar 29. PMID: 33789826. [CrossRef]
- Lin, C.-S.; Lee, Y.-T.; Fang, W.-H.; Lou, Y.-S.; Kuo, F.-C.; Lee, C.-C.; Lin, C. Deep Learning Algorithm for Management of Diabetes Mellitus via Electrocardiogram-Based Glycated Hemoglobin (ECG-HbA1c): A Retrospective Cohort Study. J. Pers. Med. 2021, 11, 725. [CrossRef]
- Kulkarni AR, Patel AA, Pipal KV, et alMachine-learning algorithm to non-invasively detect diabetes and pre-diabetes from electrocardiogramBMJ Innovations 2023;9:32-42. [CrossRef]
- K. Gupta and V. Bajaj, "A Robust Framework for Automated Screening of Diabetic Patient Using ECG Signals," in IEEE Sensors Journal, vol. 22, no. 24, pp. 24222-24229, 15 Dec.15, 2022. [CrossRef]
- Cordeiro, R.; Karimian, N.; Park, Y. Hyperglycemia Identification Using ECG in Deep Learning Era. Sensors 2021, 21, 6263. [CrossRef]
- S. Z. H. Naqvi, S. Aziz, M. U. Khan, M. Abbas, A. Haider and H. A. Hashmi, "Electrocardiography based System for Characterization of Diabetes," 2020 International Conference on Electrical, Communication, and Computer Engineering (ICECCE), Istanbul, Turkey, 2020, pp. 1-6. [CrossRef]
- Jingzhen Li, Jingyi Lu, Igbe Tobore, Yuhang Liu, Abhishek Kandwal, Lei Wang, Jian Zhou, Zedong Nie, Towards noninvasive and fast detection of Glycated hemoglobin levels based on ECG using convolutional neural networks with multisegments fusion and Varied-weight, Expert Systems with Applications, Volume 186, 2021, 115846, ISSN 0957-4174. [CrossRef]
- J. Li, I. Tobore, Y. Liu, A. Kandwal, L. Wang and Z. Nie, "Non-invasive Monitoring of Three Glucose Ranges Based On ECG By Using DBSCAN-CNN," in IEEE Journal of Biomedical and Health Informatics, vol. 25, no. 9, pp. 3340-3350, Sept. 2021. [CrossRef]
- Mohsen, F., Safa, A. & Shah, Z. ECG features improve multimodal deep learning prediction of incident T2DM in a Middle Eastern cohort. Sci Rep 15, 27164 (2025). [CrossRef]
- Fengade VS, Swati H, Chandak M, Rattan R, Singhal A, Kamble P, Phatak M, John N. Development of Enhanced Machine Learning Models for Predicting Type 2 Diabetes Mellitus Using Heart Rate Variability: A Retrospective Study. Cureus. 2025 Mar 21;17(3):e80933. doi: 10.7759/cureus.80933. PMID: 40255847; PMCID: PMC12009493. [CrossRef]
- Koga, D., Kaneda, R., Komiya, C. et al. Artificial intelligence identifies individuals with prediabetes using single-lead electrocardiograms. Cardiovasc Diabetol 24, 415 (2025). [CrossRef]
- Jethani, Neil & Manas Puli, Aahlad & Zhang, Hao & Garber, Leonid & Jankelson, Lior & Aphinyanaphongs, Yindalon & Ranganath, Rajesh. (2022). New-Onset Diabetes Assessment Using Artificial Intelligence-Enhanced Electrocardiography. [CrossRef]
- Wang, L.; Mu, Y.; Zhao, J.; Wang, X.; Che, H. IGRNet: A Deep Learning Model for Non-Invasive, Real-Time Diagnosis of Prediabetes through Electrocardiograms. Sensors 2020, 20, 2556. [CrossRef]
- A. Site, J. Nurmi and E. S. Lohan, "Machine-Learning-Based Diabetes Prediction Using Multisensor Data," in IEEE Sensors Journal, vol. 23, no. 22, pp. 28370-28377, 15 Nov.15, 2023. [CrossRef]
- D. Santhakumar, K. Dhana Shree, M. Buvanesvari, A. Saran Kumar, Ayodeji Olalekan Salau, HD-MVCNN: High-density ECG signal based diabetic prediction and classification using multi-view convolutional neural network, Egyptian Informatics Journal, Volume 28, 2024, 100573, ISSN 1110-8665. [CrossRef]
- Chiu, I.-M.; Cheng, C.-Y.; Chang, P.-K.; Li, C.-J.; Cheng, F.-J.; Lin, C.-H.R. Utilization of Personalized Machine-Learning to Screen for Dysglycemia from Ambulatory ECG, toward Noninvasive Blood Glucose Monitoring. Biosensors 2023, 13, 23. [CrossRef]
- R. Musale and A. N. Paithane, "Design and develop an algorithm for a diabetic detection using ECG signal," 2017 International Conference on Computing Methodologies and Communication (ICCMC), Erode, India, 2017, pp. 961-966. [CrossRef]
- Song, H.-J.; Han, J.-H.; Cho, S.-P.; Im, S.-I.; Kim, Y.-S.; Park, J.-U. Predicting Dysglycemia in Patients with Diabetes Using Electrocardiogram. Diagnostics 2024, 14, 2489. [CrossRef]
- Xiaowei Zhang, Changning Liu, Yang Sun, Liangzhen You, Xiaoyu Zhang, Hongcai Shang, Clinical research on artificial intelligence medical diagnostic devices: A scoping review, EngMedicine, Volume 3, Issue 1, 2026, 100120, ISSN 2950-4899. [CrossRef]
- Fahim YA, Hasani IW, Kabba S, Ragab WM. Artificial intelligence in healthcare and medicine: clinical applications, therapeutic advances, and future perspectives. Eur J Med Res. 2025 Sep 23;30(1):848. doi: 10.1186/s40001-025-03196-w. PMID: 40988064; PMCID: PMC12455834. [CrossRef]
- Bartusik-Aebisher, D.; Justin Raj, D.R.; Aebisher, D. Artificial Intelligence in Medical Diagnostics: Foundations, Clinical Applications, and Future Directions. Appl. Sci. 2026, 16, 728. [CrossRef]
- Kelly, C.J., Karthikesalingam, A., Suleyman, M. et al. Key challenges for delivering clinical impact with artificial intelligence. BMC Med 17, 195 (2019). [CrossRef]
- American Diabetes Association Professional Practice Committee; 17. Diabetes Advocacy: Standards of Care in Diabetes—2024. Diabetes Care 1 January 2024; 47 (Supplement_1): S307–S308. [CrossRef]
- Hannun AY, Rajpurkar P, Haghpanahi M, Tison GH, Bourn C, Turakhia MP, Ng AY. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nat Med. 2019 Jan;25(1):65-69. doi: 10.1038/s41591-018-0268-3. Epub 2019 Jan 7. Erratum in: Nat Med. 2019 Mar;25(3):530. doi: 10.1038/s41591-019-0359-9. PMID: 30617320; PMCID: PMC6784839. [CrossRef]
- Neri, L.; Oberdier, M.T.; van Abeelen, K.C.J.; Menghini, L.; Tumarkin, E.; Tripathi, H.; Jaipalli, S.; Orro, A.; Paolocci, N.; Gallelli, I.; et al. Electrocardiogram Monitoring Wearable Devices and Artificial-Intelligence-Enabled Diagnostic Capabilities: A Review. Sensors 2023, 23, 4805. [CrossRef]
- Alimbayev, Chingiz and Alimbayeva, Zhadyra and Ozhikenov, Kassymbek and Bodin, Oleg and Mukazhanov, Yerkat, Development of Measuring System for Determining Life-Threatening Cardiac Arrhythmias in a Patient’s Free Activity (February 29, 2020). Eastern-European Journal of Enterprise Technologies, 1(9 (103)), 12-22, 2020, doi: 10.15587/1729-4061.2020.197079, Available at SSRN: https://ssrn.com/abstract=3703319. [CrossRef]
- Rudin C. Stop Explaining Black Box Machine Learning Models for High Stakes Decisions and Use Interpretable Models Instead. Nat Mach Intell. 2019 May;1(5):206-215. doi: 10.1038/s42256-019-0048-x. Epub 2019 May 13. PMID: 35603010; PMCID: PMC9122117. [CrossRef]
- Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med. 2019 Jan;25(1):44-56. doi: 10.1038/s41591-018-0300-7. Epub 2019 Jan 7. PMID: 30617339. [CrossRef]
- Shaffer F and Ginsberg JP (2017) An Overview of Heart Rate Variability Metrics and Norms. Front. Public Health 5:258. [CrossRef]
- Rieke, N., Hancox, J., Li, W. et al. The future of digital health with federated learning. npj Digit. Med. 3, 119 (2020). [CrossRef]
- Tuija Leinonen, David Wong, Antti Vasankari, Ali Wahab, Ramesh Nadarajah, Matti Kaisti, Antti Airola, Empirical investigation of multi-source cross-validation in clinical ECG classification, Computers in Biology and Medicine, Volume 183, 2024, 109271, ISSN 0010-4825. [CrossRef]
- Nasef, D.; Nasef, D.; Basco, K.J.; Singh, A.; Hartnett, C.; Ruane, M.; Tagliarino, J.; Nizich, M.; Toma, M. Clinical Applicability of Machine Learning Models for Binary and Multi-Class Electrocardiogram Classification. AI 2025, 6, 59. [CrossRef]
- Attia IZ, Tseng AS, Benavente ED, Medina-Inojosa JR, Clark TG, Malyutina S, Kapa S, Schirmer H, Kudryavtsev AV, Noseworthy PA, Carter RE, Ryabikov A, Perel P, Friedman PA, Leon DA, Lopez-Jimenez F. External validation of a deep learning electrocardiogram algorithm to detect ventricular dysfunction. Int J Cardiol. 2021 Apr 15;329:130-135. doi: 10.1016/j.ijcard.2020.12.065. Epub 2021 Jan 2. PMID: 33400971; PMCID: PMC7955278. [CrossRef]
- Kalmady, S.V., Salimi, A., Sun, W. et al. Development and validation of machine learning algorithms based on electrocardiograms for cardiovascular diagnoses at the population level. npj Digit. Med. 7, 133 (2024). [CrossRef]
- Ong Ly C, Unnikrishnan B, Tadic T, Patel T, Duhamel J, Kandel S, Moayedi Y, Brudno M, Hope A, Ross H, McIntosh C. Shortcut learning in medical AI hinders generalization: method for estimating AI model generalization without external data. NPJ Digit Med. 2024 May 14;7(1):124. doi: 10.1038/s41746-024-01118-4. PMID: 38744921; PMCID: PMC11094145. [CrossRef]
- Quer G, Arnaout R, Henne M, Arnaout R. Machine Learning and the Future of Cardiovascular Care: JACC State-of-the-Art Review. J Am Coll Cardiol. 2021 Jan 26;77(3):300-313. doi: 10.1016/j.jacc.2020.11.030. PMID: 33478654; PMCID: PMC7839163. [CrossRef]
- Steinhubl SR, Muse ED, Topol EJ. The emerging field of mobile health. Sci Transl Med. 2015 Apr 15;7(283):283rv3. doi: 10.1126/scitranslmed.aaa3487. PMID: 25877894; PMCID: PMC4748838. [CrossRef]
- Alimbayev, C.; Alimbayeva, Z.; Ozhikenov, K.; Karibayev, K.; Orynbay, Z.; Igembay, Y.; Daniyalov, M.; Nurdanali, A. Development and Pilot Evaluation of a Wearable 12-Lead ECG System for Multilead Feature Analysis in Individuals with Different Glycemic Status. Sensors 2026, 26, 1598. [CrossRef]
- Yang Y, Gao W. Wearable and flexible electronics for continuous molecular monitoring. Chem Soc Rev. 2019 Mar 18;48(6):1465-1491. doi: 10.1039/c7cs00730b. PMID: 29611861. [CrossRef]
- Lin CS, Liu WT, Chen YH, Lin SH, Lin C. Artificial intelligence-enabled electrocardiography from scientific research to clinical application. EMBO Mol Med. 2026 Jan;18(1):22-40. doi: 10.1038/s44321-025-00351-y. Epub 2025 Dec 1. PMID: 41326714; PMCID: PMC12808761. [CrossRef]
- Liu X, Cruz Rivera S, Moher D, Calvert MJ, Denniston AK; SPIRIT-AI and CONSORT-AI Working Group. Reporting guidelines for clinical trial reports for interventions involving artificial intelligence: the CONSORT-AI extension. Lancet Digit Health. 2020 Oct;2(10):e537-e548. doi: 10.1016/S2589-7500(20)30218-1. Epub 2020 Sep 9. PMID: 33328048; PMCID: PMC8183333. [CrossRef]
- Alimbayeva, Z.; Alimbayev, C.; Ozhikenov, K.; Bayanbay, N.; Ozhikenova, A. Wearable ECG Device and Machine Learning for Heart Monitoring. Sensors 2024, 24, 4201. [CrossRef]
| Maturity level | General characteristics | Validation status | Dataset requirements | Translational meaning |
|---|---|---|---|---|
| Level 1 | Proof-of-concept / exploratory study | Internal only or absent | Small, highly selective cohorts | Technical feasibility only |
| Level 2 | Initial model development | Cross-validation / train–test split | Single-center datasets | Early methodological evidence |
| Level 3 | Clinical ML validation | Independent test set, structured retrospective evaluation | Larger clinical datasets | Moderate translational potential |
| Level 4 | Advanced clinical validation | External validation across cohorts | Multi-center / population-based datasets | Near-translational readiness |
| Level 5 | Real-world deployment | Prospective and implementation evaluation | Representative screening populations | Clinically deployable screening system |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




