Submitted:
23 April 2026
Posted:
24 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset and Preprocessing
2.1.1. Children’s Hospital of Fudan University Dataset
2.1.2. MASS-SS3 Dataset
2.1.3. Preprocessing
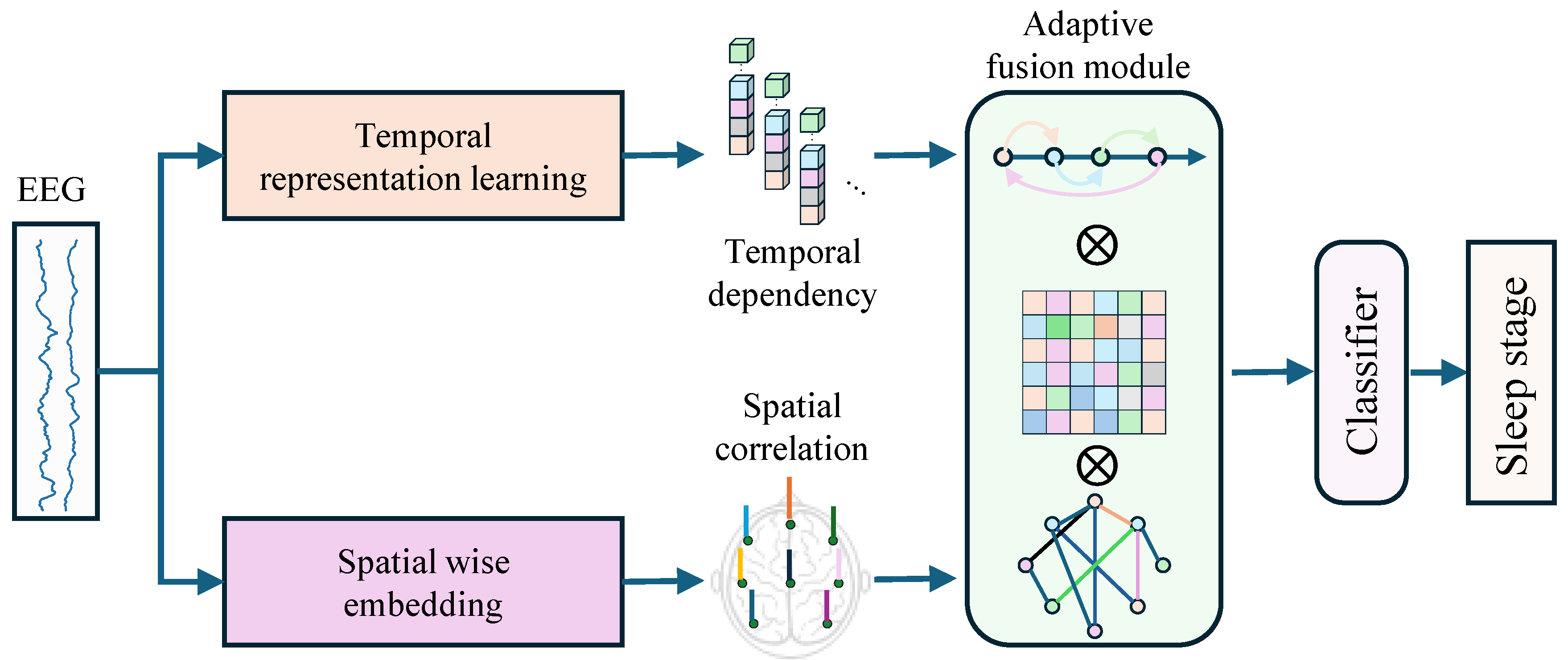
2.2. Temporal-spatial Feature Fusion Network
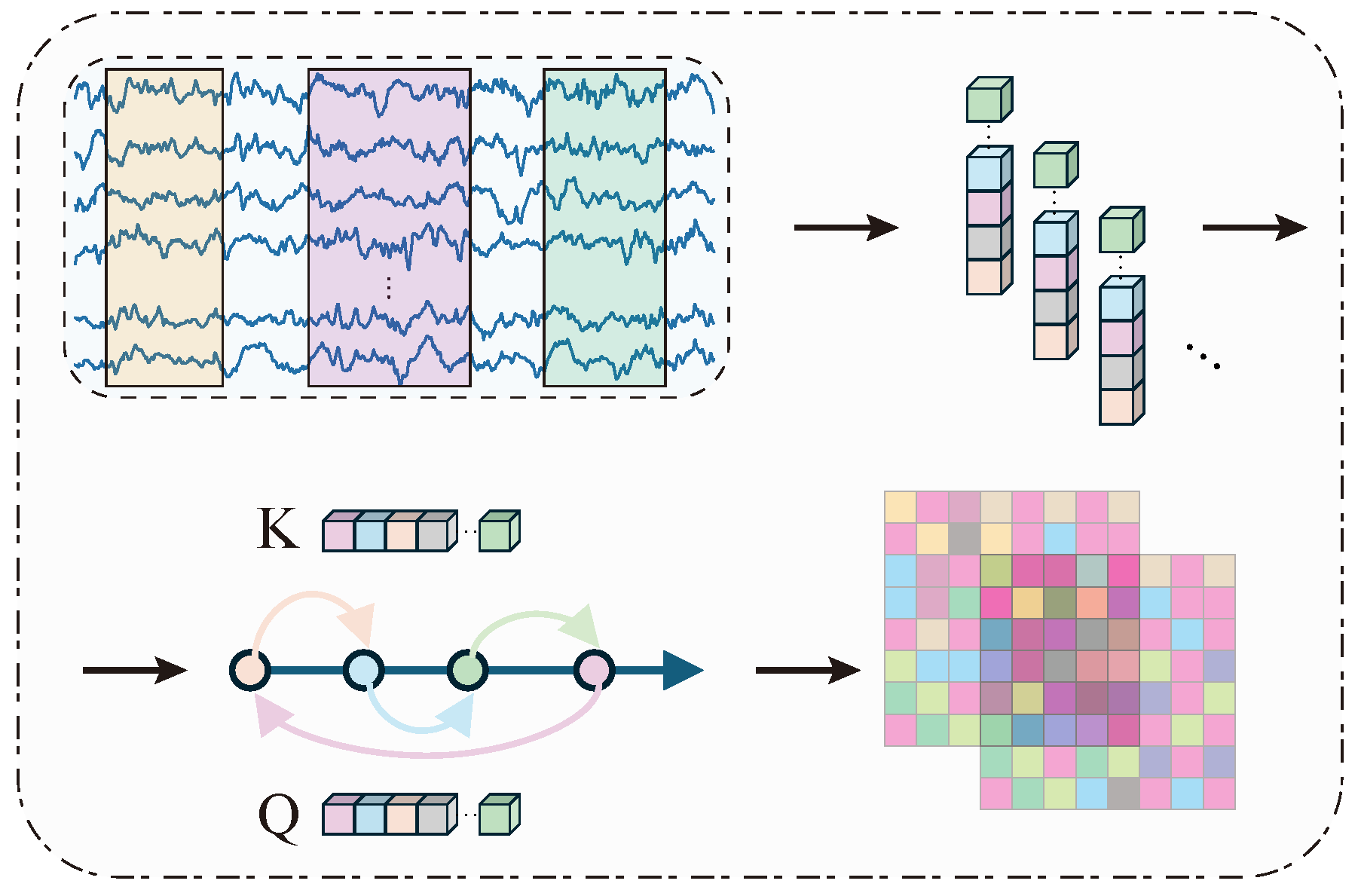
2.2.1. Temporal Representation Learning Branch
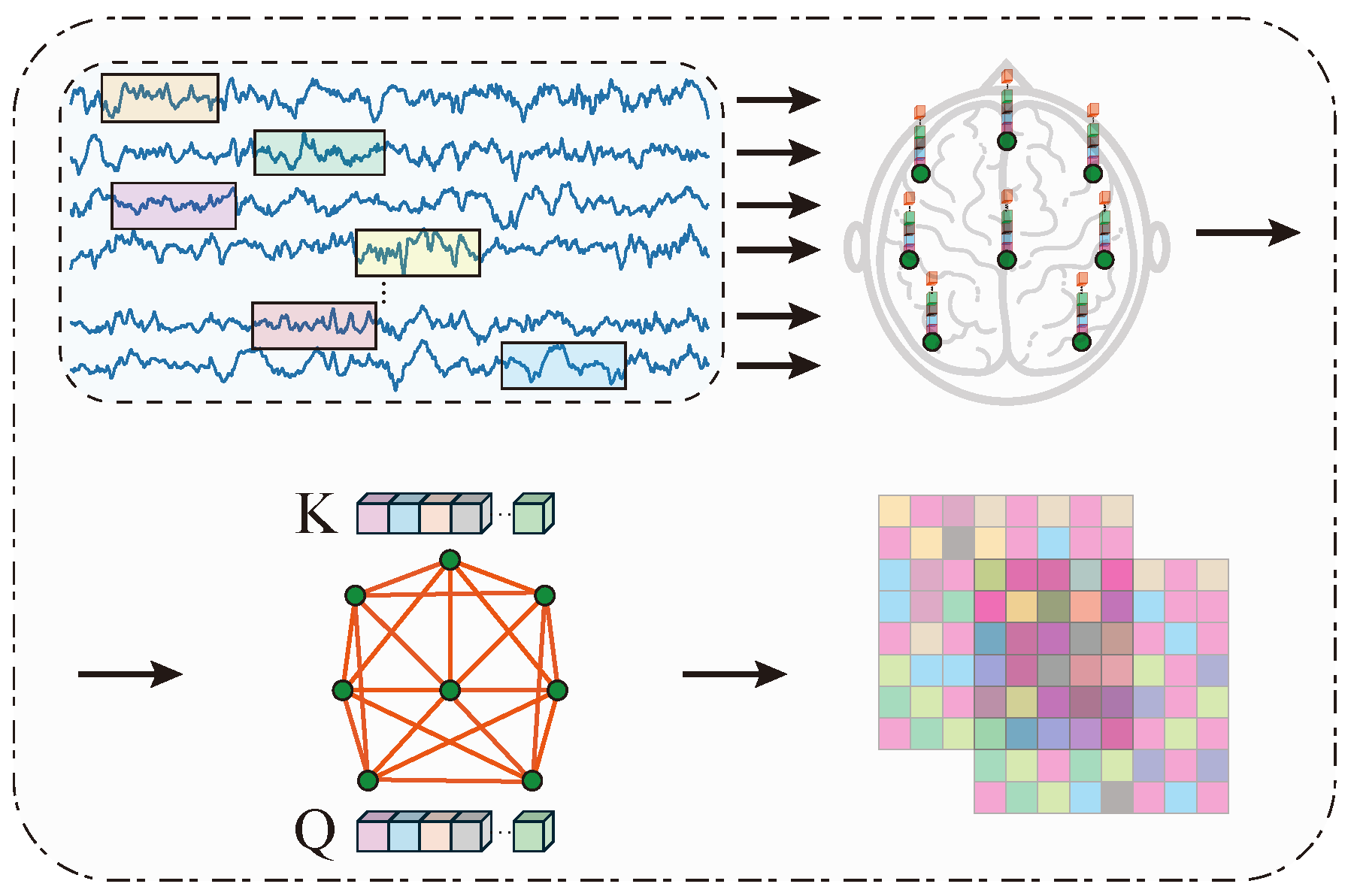
2.2.2. Spatial Correlation Learning Branch
2.2.3. Adaptive Fusion Module
2.3. Evaluation Metrics
3. Results
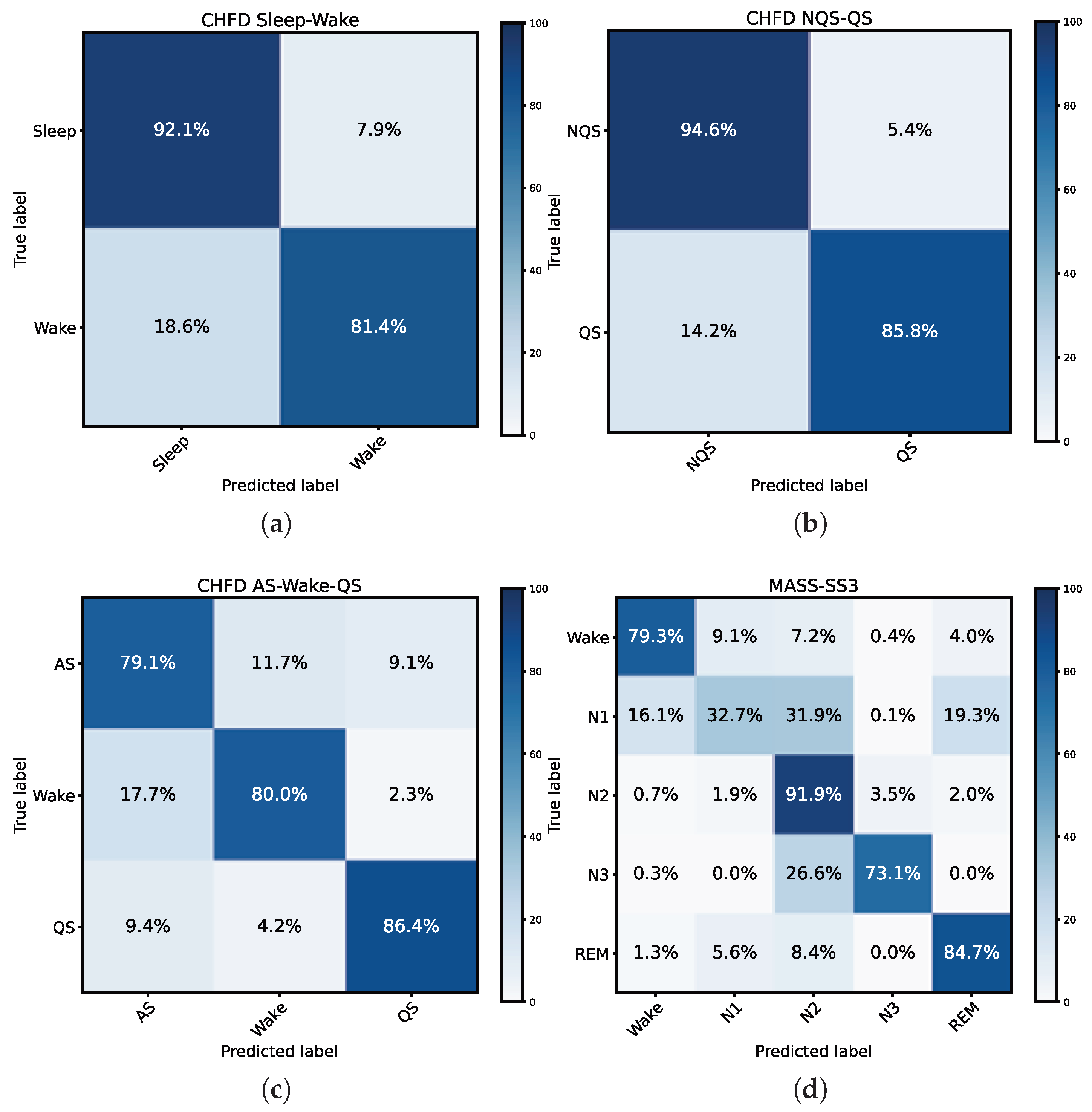
3.1. Performance on Sleep-Wake Task on CHFD Dataset
3.2. Performance on QS Detection Task on CHFD Dataset
3.3. Performance on AS-W-QS Task on CHFD Dataset
3.4. Performance on five-stage Classification Task on MASS-SS3 Dataset
3.5. Cross-Task Analysis
3.6. Comparison with Existing Methods
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PSG | Polysomnography |
| NICUs | Neonatal intensive care units |
| CHFD | Children’s hospital of fudan dataset |
| MASS | Montreal archive of sleep studies |
| SHHS | Sleep heart health study |
| EEG | Electroencephalograph |
| CNN | Convolutional neural network |
| RNN | Recurrent neural network |
| GNN | Graph neural network |
| LSTM | Long short-term memory |
| QS | Quiet sleep |
| AS | Active sleep |
References
- Tarullo, A.R.; Balsam, P.D.; Fifer, W.P. Sleep and infant learning. Infant and Child Development 2011, 20, 35–46. Available online: https://onlinelibrary.wiley.com/doi/pdf/10.1002/icd.685. [CrossRef] [PubMed]
- Ryan, M.A.J.; Mathieson, S.R.; Livingstone, V.; O’Sullivan, M.P.; Dempsey, E.M.; Boylan, G.B. Sleep state organisation of moderate to late preterm infants in the neonatal unit. Pediatric Research 2023, 93, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, S.F.; Abbas, A.; Ahmad, I.; Alshehri, M.S.; Almakdi, S.; Ghadi, Y.Y.; Ahmad, J. Automatic neonatal sleep stage classification: A comparative study. Heliyon 2023, 9, e22195. [Google Scholar] [CrossRef] [PubMed]
- Choo, B.P.; Mok, Y.; Oh, H.C.; Patanaik, A.; Kishan, K.; Awasthi, A.; Biju, S.; Bhattacharjee, S.; Poh, Y.; Wong, H.S. Benchmarking performance of an automatic polysomnography scoring system in a population with suspected sleep disorders. Frontiers in Neurology 2023, 14–2023. [Google Scholar] [CrossRef]
- Alhejaili, F. Comparing polysomnography auto scoring with the standard of care in sleep medicine. Annals of Thoracic Medicine 2025, 21, 21–28. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, X.; Huang, Q.; Lv, Y.; Chen, F. A review of automated sleep stage based on EEG signals. Biocybernetics and Biomedical Engineering 2024, 44, 651–673. [Google Scholar] [CrossRef]
- Alickovic, E.; Subasi, A. Ensemble SVM Method for Automatic Sleep Stage Classification. IEEE Transactions on Instrumentation and Measurement 2018, 67, 1258–1265. [Google Scholar] [CrossRef]
- Klok, A.B.; Edin, J.; Cesari, M.; Olesen, A.N.; Jennum, P.; Sorensen, H.B. A New Fully Automated Random-Forest Algorithm for Sleep Staging. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 2018; pp. 4920–4923. [Google Scholar] [CrossRef]
- Supratak, A.; Dong, H.; Wu, C.; Guo, Y. DeepSleepNet: A Model for Automatic Sleep Stage Scoring Based on Raw Single-Channel EEG. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2017, 25, 1998–2008. [Google Scholar] [CrossRef]
- Phan, H.; Andreotti, F.; Cooray, N.; Chén, O.Y.; De Vos, M. SeqSleepNet: End-to-End Hierarchical Recurrent Neural Network for Sequence-to-Sequence Automatic Sleep Staging. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2019, 27, 400–410. [Google Scholar] [CrossRef]
- Phan, H.; Mikkelsen, K.; Chén, O.Y.; Koch, P.; Mertins, A.; De Vos, M. SleepTransformer: Automatic Sleep Staging With Interpretability and Uncertainty Quantification. IEEE Transactions on Biomedical Engineering 2022, 69, 2456–2467. [Google Scholar] [CrossRef]
- Jia, Z.; Lin, Y.; Wang, J.; Zhou, R.; Ning, X.; He, Y.; Zhao, Y. GraphSleepNet: Adaptive Spatial-Temporal Graph Convolutional Networks for Sleep Stage Classification. In Proceedings of the Proceedings of the Twenty-Ninth International Joint Conference on Artificial Intelligence; IJCAI-20, Bessiere, C., Ed.; International Joint Conferences on Artificial Intelligence Organization; Main track, 7 2020; pp. 1324–1330. [Google Scholar] [CrossRef]
- Alix, J.J.; Ponnusamy, A.; Pilling, E.; Hart, A.R. An introduction to neonatal EEG. Paediatrics and Child Health 2017, 27, 135–142. [Google Scholar] [CrossRef]
- Vanhatalo, S.; Kaila, K. Development of neonatal EEG activity: From phenomenology to physiology. Seminars in Fetal and Neonatal Medicine;Assessing Brain Function in the Perinatal Period 2006, 11, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chan, G.S.H.; Tracy, M.B.; Lee, Q.Y.; Hinder, M.; Savkin, A.V.; Lovell, N.H. Spectral analysis of systemic and cerebral cardiovascular variabilities in preterm infants: relationship with clinical risk index for babies (CRIB). Physiological Measurement 2011, 32, 1913. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, T.N.; Wusthoff, C.J.; Shellhaas, R.A.; Abend, N.S.; Hahn, C.D.; Sullivan, J.E.; Nguyen, S.; Weinstein, S.; Scher, M.S.; Riviello, J.J.; et al. American clinical neurophysiology society standardized EEG terminology and categorization for the description of continuous EEG monitoring in neonates: report of the American Clinical Neurophysiology Society critical care monitoring committee. Journal of clinical neurophysiology 2013, 30, 161–173. [Google Scholar] [CrossRef]
- Bertelle, V.; Sevestre, A.; Laou-Hap, K.; Nagahapitiye, M.; Sizun, J. Sleep in the neonatal intensive care unit. The Journal of perinatal & neonatal nursing 2007, 21, 140–148. [Google Scholar]
- Dereymaeker, A.; Pillay, K.; Vervisch, J.; De Vos, M.; Van Huffel, S.; Jansen, K.; Naulaers, G. Review of sleep-EEG in preterm and term neonates. Early human development 2017, 113, 87–103. [Google Scholar] [CrossRef]
- Grigg-Damberger, M.M. The visual scoring of sleep in infants 0 to 2 months of age. Journal of clinical sleep medicine 2016, 12, 429–445. [Google Scholar] [CrossRef]
- O’Reilly, C.; Gosselin, N.; Carrier, J.; Nielsen, T. Montreal Archive of Sleep Studies: an open-access resource for instrument benchmarking and exploratory research. Journal of Sleep Research 2014, 23, 628–635. Available online: https://onlinelibrary.wiley.com/doi/pdf/10.1111/jsr.12169. [CrossRef]
- Kemp, B.; Zwinderman, A.; Tuk, B.; Kamphuisen, H.; Oberye, J. Analysis of a sleep-dependent neuronal feedback loop: the slow-wave microcontinuity of the EEG. IEEE Transactions on Biomedical Engineering 2000, 47, 1185–1194. [Google Scholar] [CrossRef]
- Zhang, G.Q.; Cui, L.; Mueller, R.; Tao, S.; Kim, M.; Rueschman, M.; Mariani, S.; Mobley, D.; Redline, S. The National Sleep Research Resource: towards a sleep data commons. Journal of the American Medical Informatics Association 2018, 25, 1351–1358. Available online: https://academic.oup.com/jamia/article-pdf/25/10/1351/34150622/ocy064.pdf. [CrossRef]
- Mazzotti, D.R.; Guindalini, C.; de Souza, A.A.L.; Sato, J.R.; Santos-Silva, R.; Bittencourt, L.R.A.; Tufik, S. Adenosine Deaminase Polymorphism Affects Sleep EEG Spectral Power in a Large Epidemiological Sample. PLOS ONE 2012, 7, 1–6. [Google Scholar] [CrossRef]
- Crespo-Garcia, M.; Atienza, M.; Cantero, J.L. Muscle Artifact Removal from Human Sleep EEG by Using Independent Component Analysis. Annals of Biomedical Engineering 2008, 36, 467–475. [Google Scholar] [CrossRef] [PubMed]
- Reis, P.; Hebenstreit, F.; Gabsteiger, F.; von Tscharner, V.; Lochmann, M. Methodological aspects of EEG and body dynamics measurements during motion. Frontiers in Human Neuroscience 2014, 8–2014. [Google Scholar] [CrossRef] [PubMed]
- Ansari, A.H.; Wel, O.D.; Lavanga, M.; Caicedo, A.; Dereymaeker, A.; Jansen, K.; Vervisch, J.; Vos, M.D.; Naulaers, G.; Huffel, S.V. Quiet sleep detection in preterm infants using deep convolutional neural networks. Journal of Neural Engineering 2018, 15, 066006. [Google Scholar] [CrossRef] [PubMed]
- Ansari, A.H.; Wel, O.D.; Pillay, K.; Dereymaeker, A.; Jansen, K.; Huffel, S.V.; Naulaers, G.; Vos, M.D. A convolutional neural network outperforming state-of-the-art sleep staging algorithms for both preterm and term infants. Journal of Neural Engineering 2020, 17, 016028. [Google Scholar] [CrossRef]
- Zhu, H.; Wang, L.; Shen, N.; Wu, Y.; Feng, S.; Xu, Y.; Chen, C.; Chen, W. MS-HNN: Multi-Scale Hierarchical Neural Network With Squeeze and Excitation Block for Neonatal Sleep Staging Using a Single-Channel EEG. IEEE Transactions on Neural Systems and Rehabilitation Engineering Conference Name: IEEE Transactions on Neural Systems and Rehabilitation Engineering 2023, 31, 2195–2204. [Google Scholar] [CrossRef]
- Siddiqa, H.A.; Tang, Z.; Xu, Y.; Wang, L.; Irfan, M.; Abbasi, S.F.; Nawaz, A.; Chen, C.; Chen, W. Single-Channel EEG Data Analysis Using a Multi-Branch CNN for Neonatal Sleep Staging. IEEE Access 2024, 12, 29910–29925. [Google Scholar] [CrossRef]
- Jia, Z.; Lin, Y.; Wang, J.; Ning, X.; He, Y.; Zhou, R.; Zhou, Y.; Lehman, L.w.H. Multi-View Spatial-Temporal Graph Convolutional Networks With Domain Generalization for Sleep Stage Classification. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2021, 29, 1977–1986. [Google Scholar] [CrossRef]
- Eldele, E.; Chen, Z.; Liu, C.; Wu, M.; Kwoh, C.K.; Li, X.; Guan, C. An Attention-Based Deep Learning Approach for Sleep Stage Classification With Single-Channel EEG. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2021, 29, 809–818. [Google Scholar] [CrossRef]
- Lyu, J.; Shi, W.; Zhang, C.; Yeh, C.H. A Novel Sleep Staging Method Based on EEG and ECG Multimodal Features Combination. IEEE Transactions on Neural Systems and Rehabilitation Engineering 2023, 31, 4073–4084. [Google Scholar] [CrossRef]
- Zhou, D.; Xu, Q.; Zhang, J.; Wu, L.; Xu, H.; Kettunen, L.; Chang, Z.; Zhang, Q.; Cong, F. Interpretable Sleep Stage Classification Based on Layer-Wise Relevance Propagation. IEEE Transactions on Instrumentation and Measurement 2024, 73, 1–10. [Google Scholar] [CrossRef]
| Terms | Details |
|---|---|
| Gender (b: g) | 32:32 |
| Gestational age (w + d) | 38.3 ± 1.8 |
| Postmenstrual age (w + d) | 40.5 ± 1.7 |
| Weight (kg) | 3.3 ± 0.6 |
| Number of wakefulness epochs | 5514 (32.8%) |
| Number of QS epochs | 5749 (34.2%) |
| Number of AS epochs | 5540 (33.0%) |
| EEG channel | F3, F4, C3, C4, T3, T4, P3, and P4 |
| Sampling rate | 500Hz |
| Terms | Details |
|---|---|
| Gender (m: f) | 28:34 |
| Scoring rules | AASM |
| Sampling rate | 256 Hz |
| Number of wakefulness epochs | 6442 |
| Number of N1 epochs | 4839 |
| Number of N2 epochs | 29802 |
| Number of N3 epochs | 7653 |
| Number of REM epochs | 10581 |
| Selected EEG channel | F3, F4, C3, C4, T3, T4, P3, and P4 |
| Dataset | Task | Accuracy | MF1 | Kappa | Macro-sensitivity | Macro-specificity |
|---|---|---|---|---|---|---|
| CHFD | Sleep-Wake | 0.886 | 0.870 | 0.740 | 0.868 | 0.868 |
| CHFD | QS Detection | 0.916 | 0.906 | 0.811 | 0.902 | 0.902 |
| CHFD | AS-W-QS | 0.819 | 0.819 | 0.729 | 0.818 | 0.910 |
| MASS-SS3 | W-N1-N2-N3-REM | 0.820 | 0.739 | 0.729 | 0.723 | 0.944 |
| Method | Accuracy | MF1 | Kappa | M-sens | M-spec | Parameters |
|---|---|---|---|---|---|---|
| MB-CNN [29] | 0.728 | 0.682 | 0.561 | 0.671 | 0.850 | <0.01M |
| Conv-2d [26] | 0.535 | 0.531 | 0.489 | 0.768 | 0.536 | <0.01M |
| Conv-2d [27] | 0.523 | 0.519 | 0.411 | 0.761 | 0.523 | <0.01M |
| DeepSleepNet [9] | 0.689 | 0.682 | 0.535 | 0.845 | 0.692 | 24.75M |
| AttnSleep [31] | 0.680 | 0.646 | 0.659 | 0.839 | 0.650 | 5.20M |
| MS-HNN [28] | 0.754 | 0.758 | 0.728 | 0.876 | 0.755 | 25.63M |
| GraphSleepNet [12] | 0.689 | 0.682 | 0.535 | 0.845 | 0.692 | <0.05M |
| MVST-GCN [30] | 0.697 | 0.696 | 0.547 | 0.849 | 0.699 | <0.05M |
| Proposed | 0.819 | 0.819 | 0.729 | 0.818 | 0.910 | 0.81M |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




