Submitted:
21 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
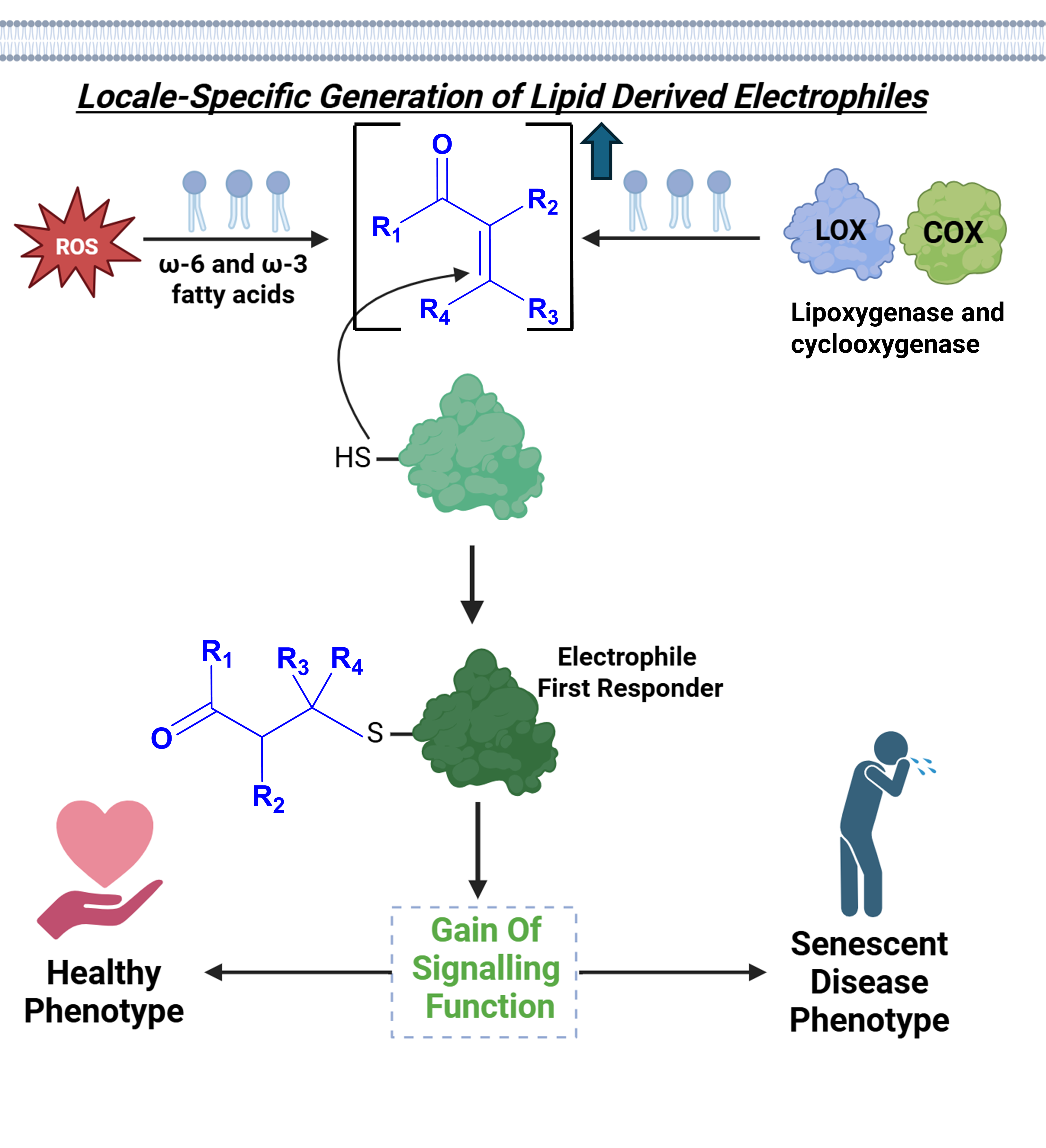
Keywords:
Introduction
Stress Upregulates a Host of REMs, Many of Which Can Covalently Modify Proteins
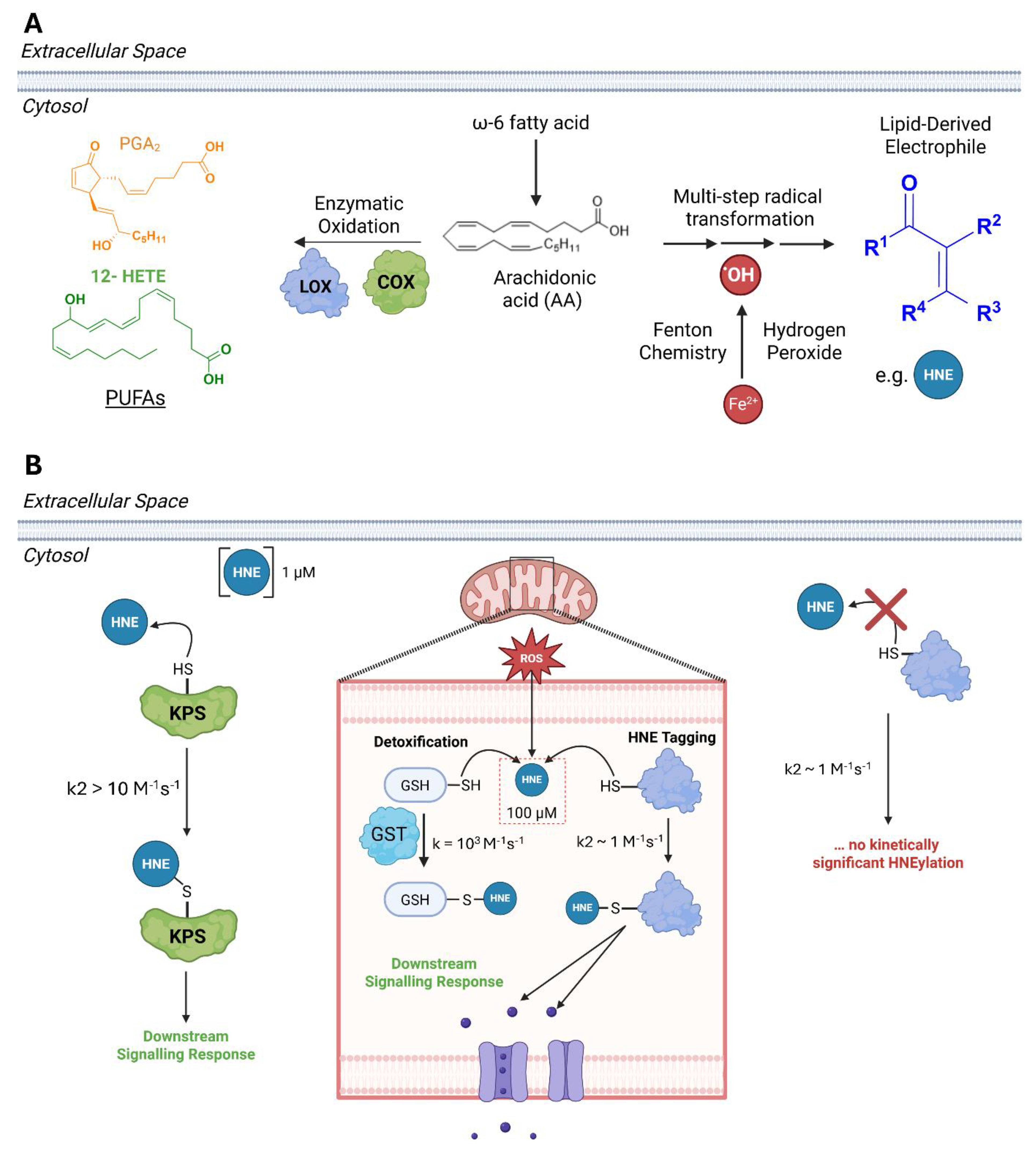
Reactive Carbon-Based REMs React with Proteins, but Not Sufficiently Enough to Achieve High Occupancy
Extent of HNEylation Can Be Increased in Cells
Kinetic Privilege: Context and Considerations
Signaling Modes
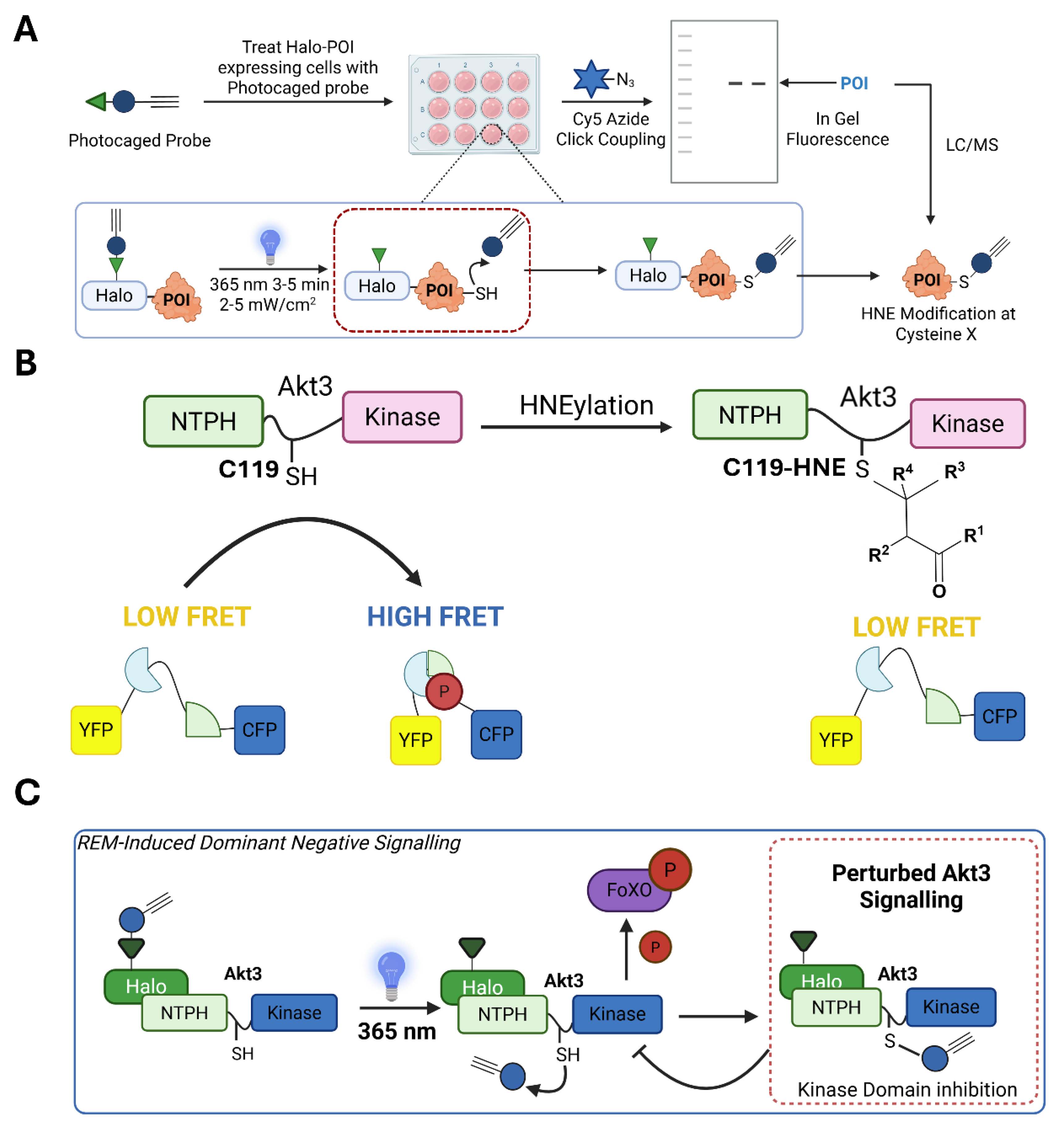
Turn on GOF Signaling
General Similarities of GOF Signaling
Factoring in Protein Stability on Labeling and Sensing
Protein Half-Lives Set Lower Limits on Reactivity Needed to Be Sensors
Basal Levels of Electrophiles Set Upper Limits on Reactivity
Two Different Paradigms of REM Sensors
Comparison with Current GOF Drugs
GOF Protein Sensors-Ideal for Drug Discovery
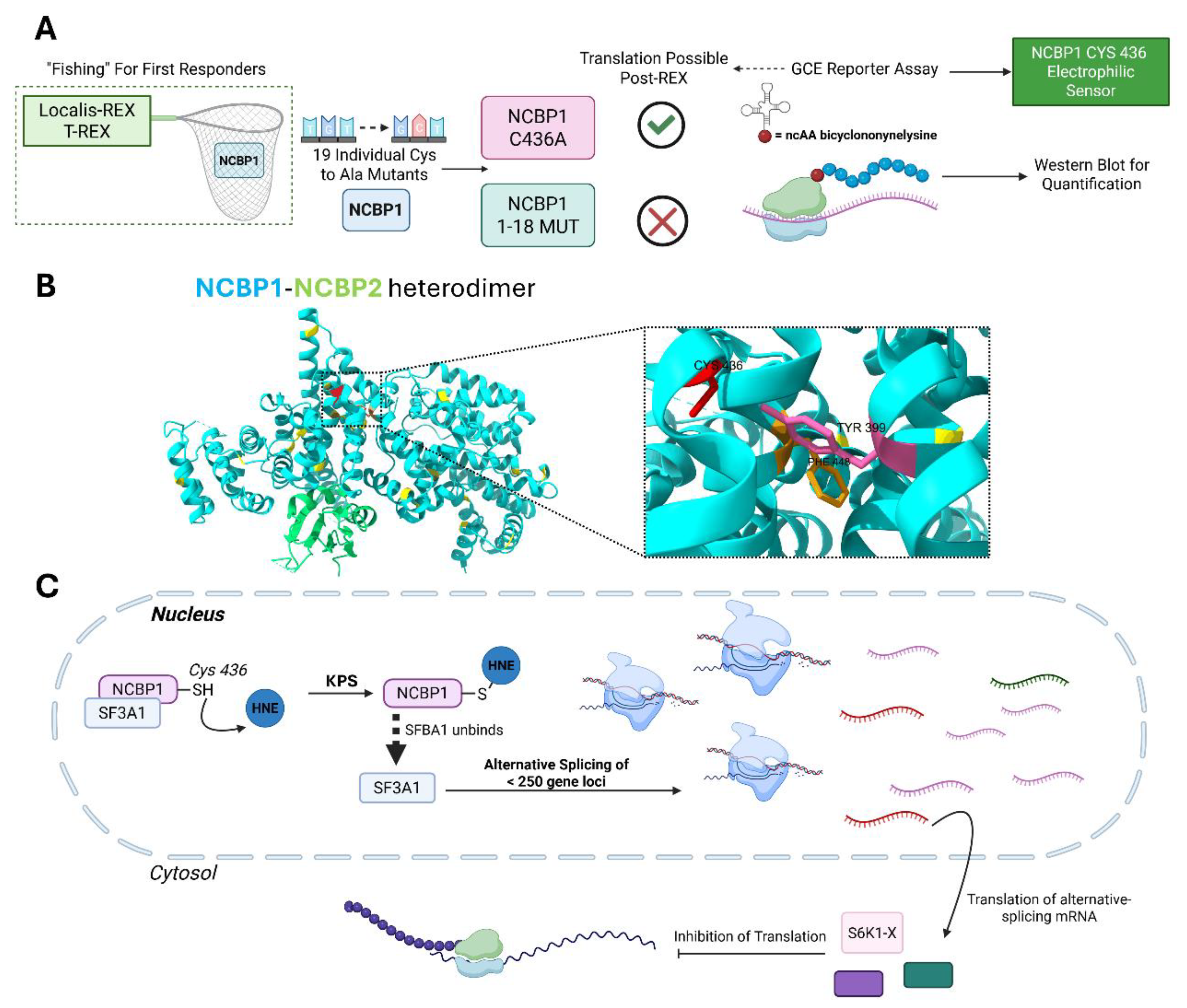
Identifying Sentinel Proteins in Unexpected Places
Author contributions
Acknowledgements
Conflict of interest
References
- Esterbauer, H.; Schaur, R. J.; Zollner, H. Chemistry and biochemistry of 4-hydroxynonenal, malonaldehyde and related aldehydes. Free Radic Biol Med. 1991, 11, 81–128. [Google Scholar] [CrossRef]
- Parvez, S.; Long, M. J. C.; Poganik, J. R.; Aye, Y. Redox Signaling by Reactive Electrophiles and Oxidants. Chem Rev. 2018, 118, 8798–8888. [Google Scholar] [CrossRef]
- Schopfer, F. J.; Cipollina, C.; Freeman, B. A. Formation and signaling actions of electrophilic lipids. Chem Rev. 2011, 111, 5997–6021. [Google Scholar] [CrossRef]
- Blair, I. A. DNA adducts with lipid peroxidation products. J Biol Chem. 2008, 283, 15545–9. [Google Scholar] [CrossRef] [PubMed]
- Sakanyan, V. Reactive Chemicals and Electrophilic Stress in Cancer: A Minireview. High Throughput 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Huang, K. T.; Aye, Y. Toward decoding spatiotemporal signaling activities of reactive immunometabolites with precision immuno-chemical biology tools. Commun Chem. 2024, 7, 195. [Google Scholar] [CrossRef] [PubMed]
- Poganik, J. R.; Huang, K. T.; Parvez, S.; Zhao, Y.; Raja, S.; Long, M. J. C.; Aye, Y. Wdr1 and cofilin are necessary mediators of immune-cell-specific apoptosis triggered by Tecfidera. Nat Commun. 2021, 12, 5736. [Google Scholar] [CrossRef]
- Hauer, L.; Sellner, J. Diroximel Fumarate as a Novel Oral Immunomodulating Therapy for Relapsing Forms of Multiple Sclerosis: A Review on the Emerging Data. Drug Des Devel Ther. 2022, 16, 3915–3927. [Google Scholar] [CrossRef]
- Montano, J. L.; Wang, B. J.; Volk, R. F.; Warrington, S. E.; Garda, V. G.; Hofmann, K. L.; Chen, L. C.; Zaro, B. W. Improved Electrophile Design for Exquisite Covalent Molecule Selectivity. ACS Chem Biol. 2022, 17, 1440–1449. [Google Scholar] [CrossRef]
- Gu, T. J.; Cai, J.; Auster, A.; Torres, E.; Zhang, D.; Khojasteh, S. C.; Wang, S. In vitro metabolism of targeted covalent inhibitors and their thiol conjugates by gut microbiome from rats, mice, and humans. Drug Metab Dispos. 2025, 53, 100027. [Google Scholar] [CrossRef]
- Baird, L.; Zhang, L.; Hidaka, T.; Xi, L.; Wang, K.; Tateno, K.; Iso, T.; Suzuki, T.; Kumada, K.; Katsuoka, F.; Kinoshita, K.; Yamamoto, M. Systemic activation of NRF2 contributes to the therapeutic efficacy of clinically-approved KRAS-G12C anti-cancer drugs. Br J Cancer 2025, 133, 1377–1390. [Google Scholar] [CrossRef]
- Long, M. J.; Parvez, S.; Zhao, Y.; Surya, S. L.; Wang, Y.; Zhang, S.; Aye, Y. Akt3 is a privileged first responder in isozyme-specific electrophile response. Nat Chem Biol. 2017, 13, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Long, M. J. C.; Hopkins, B. D.; Luo, C.; Wang, L.; Aye, Y. Precision Targeting of pten-Null Triple-Negative Breast Tumors Guided by Electrophilic Metabolite Sensing. ACS Cent Sci. 2020, 6, 892–902. [Google Scholar] [CrossRef] [PubMed]
- Alnasser, S. M. The role of glutathione S-transferases in human disease pathogenesis and their current inhibitors. Genes Dis. 2025, 12, 101482. [Google Scholar] [CrossRef]
- Liu, J.; Kulkarni, A.; Gao, Y. Q.; Urul, D. A.; Hamelin, R.; Novotny, B. A.; Long, M. J. C.; Aye, Y. Organ-specific electrophile responsivity mapping in live C. elegans. Cell. 2024, 187, 7450–7469 e29. [Google Scholar] [CrossRef] [PubMed]
- Recktenwald, C. V.; Kellner, R.; Lichtenfels, R.; Seliger, B. Altered detoxification status and increased resistance to oxidative stress by K-ras transformation. Cancer Res. 2008, 68, 10086–93. [Google Scholar] [CrossRef]
- Singh, R. R.; Reindl, K. M. Glutathione S-Transferases in Cancer. Antioxidants (Basel) 2021, 10. [Google Scholar] [CrossRef]
- Dalleau, S.; Baradat, M.; Gueraud, F.; Huc, L. Cell death and diseases related to oxidative stress: 4-hydroxynonenal (HNE) in the balance. Cell Death Differ. 2013, 20, 1615–30. [Google Scholar] [CrossRef]
- Huang, B. K.; Sikes, H. D. Quantifying intracellular hydrogen peroxide perturbations in terms of concentration. Redox Biol. 2014, 2, 955–62. [Google Scholar] [CrossRef]
- Gueraud, F.; Atalay, M.; Bresgen, N.; Cipak, A.; Eckl, P. M.; Huc, L.; Jouanin, I.; Siems, W.; Uchida, K. Chemistry and biochemistry of lipid peroxidation products. Free Radic Res. 2010, 44, 1098–124. [Google Scholar] [CrossRef]
- Lam, M.; Mast, N.; Pikuleva, I. A. Drugs and Scaffold That Inhibit Cytochrome P450 27A1 In Vitro and In Vivo. Mol Pharmacol. 2018, 93, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Normandin, K.; Coulombe-Huntington, J.; St-Denis, C.; Bernard, A.; Bourouh, M.; Bertomeu, T.; Tyers, M.; Archambault, V. Genetic enhancers of partial PLK1 inhibition reveal hypersensitivity to kinetochore perturbations. PLoS Genet. 2023, 19, e1010903. [Google Scholar] [CrossRef] [PubMed]
- Miseta, A.; Csutora, P. Relationship between the occurrence of cysteine in proteins and the complexity of organisms. Mol Biol Evol. 2000, 17, 1232–9. [Google Scholar] [CrossRef] [PubMed]
- Chang, D.; Assari, M.; Suwathep, C.; Sappakhaw, K.; Uttamapinant, C.; Long, M. J. C.; Aye, Y. NCBP1 stress signaling drives alternative S6K1 splicing inhibiting translation. Nat Chem Biol. 2026. [Google Scholar] [CrossRef]
- Parvez, S.; Fu, Y.; Li, J.; Long, M. J.; Lin, H. Y.; Lee, D. K.; Hu, G. S.; Aye, Y. Substoichiometric hydroxynonenylation of a single protein recapitulates whole-cell-stimulated antioxidant response. J Am Chem Soc. 2015, 137, 10–3. [Google Scholar] [CrossRef]
- Zhou, Y. F.; Zhang, L.; Niu, Z. L.; Wang, Z. A. Targeting the Reactive Proteome: Recent Advances in Activity-Based Protein Profiling and Probe Design. Biomolecules 2025, 15. [Google Scholar] [CrossRef]
- Wall, S. B.; Smith, M. R.; Ricart, K.; Zhou, F.; Vayalil, P. K.; Oh, J. Y.; Landar, A. Detection of electrophile-sensitive proteins. Biochim Biophys Acta 2014, 1840, 913–22. [Google Scholar] [CrossRef]
- Long, M. J. C.; Urul, D. A.; Aye, Y. REX technologies for profiling and decoding the electrophile signaling axes mediated by Rosetta Stone proteins. Methods Enzymol. 2020, 633, 203–230. [Google Scholar]
- Liu, X.; Long, M. J. C.; Aye, Y. Proteomics and Beyond: Cell Decision-Making Shaped by Reactive Electrophiles. Trends Biochem Sci. 2019, 44, 75–89. [Google Scholar] [CrossRef]
- Zhao, Y.; Miranda Herrera, P. A.; Chang, D.; Hamelin, R.; Long, M. J. C.; Aye, Y. Function-guided proximity mapping unveils electrophilic-metabolite sensing by proteins not present in their canonical locales. Proc Natl Acad Sci U S A. 2022, 119. [Google Scholar] [CrossRef]
- Guerau-de-Arellano, M.; Piedra-Quintero, Z. L.; Tsichlis, P. N. Akt isoforms in the immune system. Front Immunol. 2022, 13, 990874. [Google Scholar] [CrossRef]
- Kumar, B. H.; Kabekkodu, S. P.; Pai, K. S. R. Structural insights of AKT and its activation mechanism for drug development. Mol Divers. 2025, 29, 5443–5463. [Google Scholar] [CrossRef]
- Calabrese, E. J.; Kozumbo, W. J. The hormetic dose-response mechanism: Nrf2 activation. Pharmacol Res. 2021, 167, 105526. [Google Scholar] [CrossRef] [PubMed]
- Suda, T.; Takubo, K.; Semenza, G. L. Metabolic regulation of hematopoietic stem cells in the hypoxic niche. Cell Stem Cell. 2011, 9, 298–310. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K. K.; Santhoshkumar, P. Lens aging: effects of crystallins. Biochim Biophys Acta 2009, 1790, 1095–108. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Sheikh, M. S.; Huang, Y. Decision Making by p53: Life versus Death. Mol Cell Pharmacol. 2010, 2, 69–77. [Google Scholar]
- Gregory, M. A.; Hann, S. R. c-Myc proteolysis by the ubiquitin-proteasome pathway: stabilization of c-Myc in Burkitt’s lymphoma cells. Mol Cell Biol. 2000, 20, 2423–35. [Google Scholar] [CrossRef]
- Mailand, N.; Podtelejnikov, A. V.; Groth, A.; Mann, M.; Bartek, J.; Lukas, J. Regulation of G(2)/M events by Cdc25A through phosphorylation-dependent modulation of its stability. EMBO J. 2002, 21, 5911–20. [Google Scholar] [CrossRef]
- Lee, J. M.; Hammaren, H. M.; Savitski, M. M.; Baek, S. H. Control of protein stability by post-translational modifications. Nat Commun. 2023, 14, 201. [Google Scholar] [CrossRef]
- Basu, R.; Fletcher, S. Protein structural dynamics in covalent drug design: insights from irreversible and reversible covalent inhibitors. RSC Chem Biol. 2026, 7, 376–399. [Google Scholar] [CrossRef]
- Lai, A. C.; Crews, C. M. Induced protein degradation: an emerging drug discovery paradigm. Nat Rev Drug Discov. 2017, 16, 101–114. [Google Scholar] [CrossRef]
- Parvez, S.; Long, M. J.; Lin, H. Y.; Zhao, Y.; Haegele, J. A.; Pham, V. N.; Lee, D. K.; Aye, Y. T-REX on-demand redox targeting in live cells. Nat Protoc. 2016, 11, 2328–2356. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Long, M. J. C.; Wang, Y.; Zhang, S.; Aye, Y. Ube2V2 Is a Rosetta Stone Bridging Redox and Ubiquitin Codes, Coordinating DNA Damage Responses. ACS Central Science 2018, 4, 246–259. [Google Scholar] [CrossRef] [PubMed]
- Wakeling, A. E.; Dukes, M.; Bowler, J. A potent specific pure antiestrogen with clinical potential. Cancer Res. 1991, 51, 3867–73. [Google Scholar] [PubMed]
- Wang, G. Fulvestrant as a reference antiestrogen and estrogen receptor (ER) degrader in preclinical studies: treatment dosage, efficacy, and implications on development of new ER-targeting agents. Transl Cancer Res. 2020, 9, 4464–4468. [Google Scholar] [CrossRef]
- Ito, T.; Ando, H.; Suzuki, T.; Ogura, T.; Hotta, K.; Imamura, Y.; Yamaguchi, Y.; Handa, H. Identification of a primary target of thalidomide teratogenicity. Science 2010, 327, 1345–50. [Google Scholar] [CrossRef]
- Sakamoto, K. M.; Kim, K. B.; Kumagai, A.; Mercurio, F.; Crews, C. M.; Deshaies, R. J. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc Natl Acad Sci U S A 2001, 98, 8554–9. [Google Scholar] [CrossRef]
- Suo, Y.; Du, D.; Chen, C.; Zhu, H.; Wang, X.; Song, N.; Lu, D.; Yang, Y.; Li, J.; Wang, J.; Luo, Z.; Zhou, B.; Luo, C.; Zhou, H. Uncovering PROTAC Sensitivity and Efficacy by Multidimensional Proteome Profiling: A Case for STAT3. J Med Chem. 2024, 67, 4804–4818. [Google Scholar] [CrossRef]
- Pike, A.; Williamson, B.; Harlfinger, S.; Martin, S.; McGinnity, D. F. Optimising proteolysis-targeting chimeras (PROTACs) for oral drug delivery: a drug metabolism and pharmacokinetics perspective. Drug Discov Today 2020, 25, 1793–1800. [Google Scholar] [CrossRef]
- Tinworth, C. P.; Lithgow, H.; Dittus, L.; Bassi, Z. I.; Hughes, S. E.; Muelbaier, M.; Dai, H.; Smith, I. E. D.; Kerr, W. J.; Burley, G. A.; Bantscheff, M.; Harling, J. D. PROTAC-Mediated Degradation of Bruton’s Tyrosine Kinase Is Inhibited by Covalent Binding. ACS Chem Biol. 2019, 14, 342–347. [Google Scholar] [CrossRef]
- Gabizon, R.; Shraga, A.; Gehrtz, P.; Livnah, E.; Shorer, Y.; Gurwicz, N.; Avram, L.; Unger, T.; Aharoni, H.; Albeck, S.; Brandis, A.; Shulman, Z.; Katz, B. Z.; Herishanu, Y.; London, N. Efficient Targeted Degradation via Reversible and Irreversible Covalent PROTACs. J Am Chem Soc. 2020, 142, 11734–11742. [Google Scholar] [CrossRef]
- Zhang, X.; Crowley, V. M.; Wucherpfennig, T. G.; Dix, M. M.; Cravatt, B. F. Electrophilic PROTACs that degrade nuclear proteins by engaging DCAF16. Nat Chem Biol. 2019, 15, 737–746. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



