Submitted:
21 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset and Data Preparation
2.2. Model Architecture
2.3. Training Procedure
2.4. Temporal Analysis of Bleeding
2.5. Evaluation Metrics
3. Results
3.1. Segmentation Performance
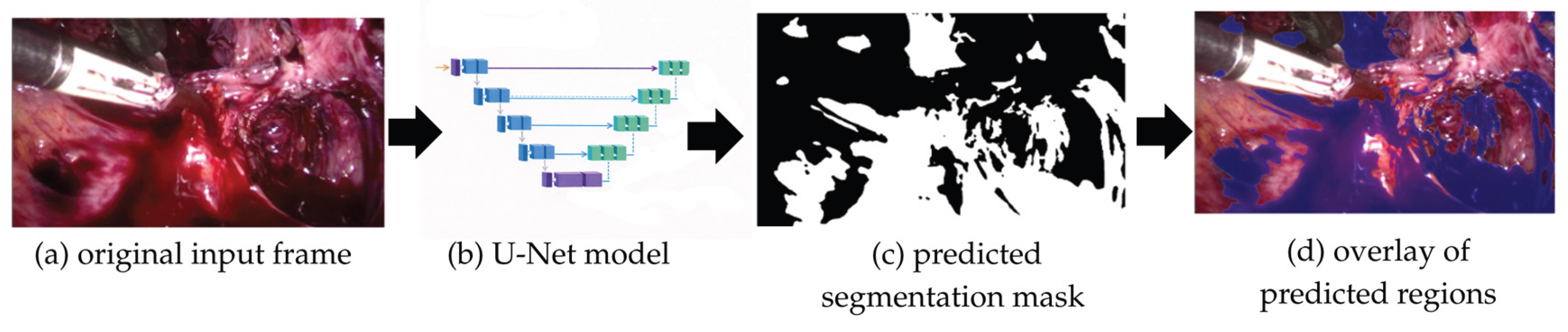
3.2. Temporal Analysis of Bleeding Dynamics
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ghezzi, T.L.; Corleta, O.C. 30 Years of Robotic Surgery. World J. Surg. 2016, 40, 2550–2557. [CrossRef]
- King, A.B.; Fowler, G.E.; Macefield, R.C.; Walker, H.; Thomas, C.; Markar, S.; Higgins, E.; Blazeby, J.M.; Blencowe, N.S. Use of artificial intelligence in the analysis of digital videos of invasive surgical procedures: Scoping review. BJS Open 2025, 9, zraf073. [CrossRef]
- Nawrat, Z. MIS AI – the artificial intelligence application in minimally invasive surgery. Mini-invasive Surg. 2020, 4, 28. [CrossRef]
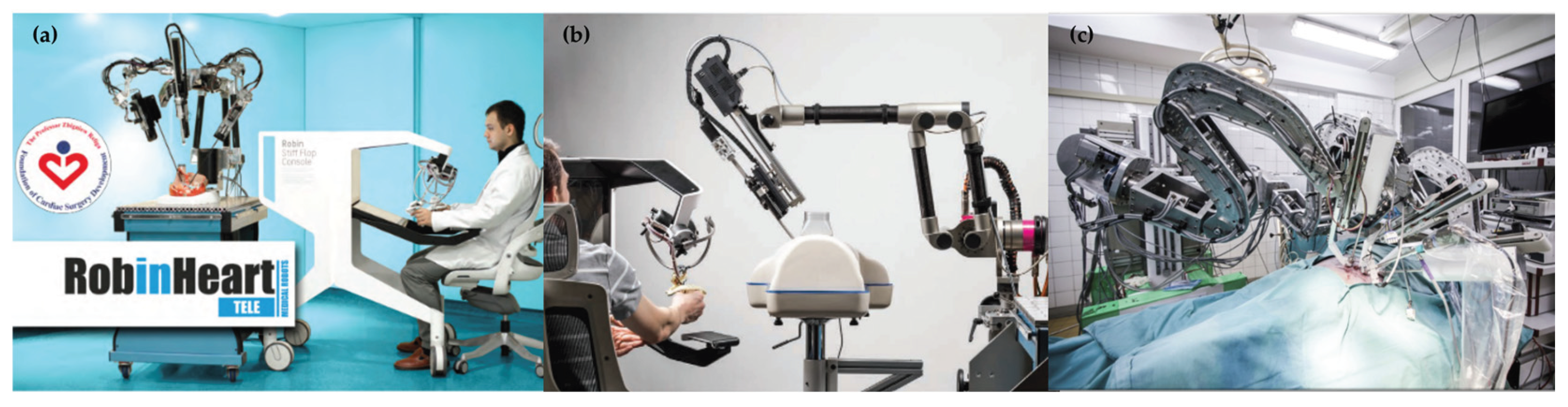
- Nawrat, Z.; Kostka, P.; Polański, A.; Rohr, K.; Sadowski, W.; Krzysztofik, K. Polish cardio-robot “Robin Heart”. System description and technical evaluation. Int. J. Med. Robot. Comput. Assist. Surg. 2006, 2, 36–44.
- Nawrat, Z.; Mucha, Ł.; Krawczyk, D.; Lis, K.; Lehrich, K.; Rohr, K.; Földesy, P.; Radó, J.; Dücső, C.; Sántha, H.; Szebényi, G.; Fürjes, P. Robin Heart INCITE surgical tele manipulator controlled by system equipped with 3D force sensor. Med. Robot. Rep. 2017, 6, 37-46.
- Nawrat, Z. Introduction to AI-driven surgical robots. Artif. Intell. Surg. 2023, 3, 90–97. [CrossRef]
- Arakaki, S.; Takenaka, S.; Sasaki, K.; Kitaguchi, D.; Hasegawa, H.; Takeshita, N.; Takatsuki, M.; Ito, M. Artificial Intelligence in Minimally Invasive Surgery: Current State and Future Challenges. JMA J. 2025, 8, 86–90. [CrossRef]
- Hua, S.; Gao, J.; Wang, Z.; Yeerkenbieke, P.; Li, J.; Wang, J.; He, G.; Jiang, J.; Lu, Y.; Yu, Q.; Han, X.; Liao, Q.; Wu, W. Automatic bleeding detection in laparoscopic surgery based on a faster region-based convolutional neural network. Ann. Transl. Med. 2022, 10, 546. [CrossRef]
- Sunakawa, T.; Kitaguchi, D.; Kobayashi, S.; Aoki, K.; Kujiraoka, M.; Sasaki, K.; Azuma, L.; Yamada, A.; Kudo, M.; Sugimoto, M.; Hasegawa, H.; Takeshita, N.; Gotohda, N.; Ito, M. Deep learning-based automatic bleeding recognition during liver resection in laparoscopic hepatectomy. Surg. Endosc. 2024, 38, 7656–7662. [CrossRef]
- Okamoto, T.; Ohnishi, T.; Kawahira, H.; Dergachyava, O.; Jannin, P.; Haneishi, H. Real-time identification of blood regions for hemostasis support in laparoscopic surgery. Signal Image Video Process. 2019, 13, 405–412. [CrossRef]
- Grammatikopoulou, M.; Sanchez-Matilla, R.; Bragman, F.; Owen, D.; Culshaw, L.; Kerr, K.; Stoyanov, D.; Luengo, I. A spatio-temporal network for video semantic segmentation in surgical videos. Int. J. Comput. Assist. Radiol. Surg. 2024, 19, 375–382. [CrossRef]
- Gao, Y.; Jiang, Y.; Peng, Y.; Yuan, F.; Zhang, X.; Wang, J. Medical Image Segmentation: A Comprehensive Review of Deep Learning-Based Methods. Tomography 2025, 11, 52. [CrossRef]
- Kamtam, D.N.; Shrager, J.B.; Malla, S.D.; Lin, N.; Cardona, J.J.; Kim, J.J.; Hu, C. Deep learning approaches to surgical video segmentation and object detection: A scoping review. Comput. Biol. Med. 2025, 194, 110482. [CrossRef]
- Sibilano, E.; Delprete, C.; Marvulli, P.M.; Brunetti, A.; Marino, F.; Lucarelli, G.; Battaglia, M.; Bevilacqua, V. Deep learning strategies for semantic segmentation in robot-assisted radical prostatectomy. Appl. Sci. 2025, 15, 10665. [CrossRef]
- Caballero, D.; Sánchez-Margallo, J.A.; Pérez-Salazar, M.J.; Sánchez-Margallo, F.M. Applications of Artificial Intelligence in Minimally Invasive Surgery Training: A Scoping Review. Surgeries 2025, 6, 7. [CrossRef]
- Nasirihaghighi, S.; Ghamsarian, N.; Peschek, L.; Munari, M.; Husslein, H.; Sznitman, R.; Schoeffmann, K. GynSurg: A Comprehensive Gynecology Laparoscopic Surgery Dataset. arXiv 2025, arXiv:2506.11356.
- Computer Vision Annotation Tool (CVAT). Available online: https://cvat.ai/ (accessed on 7 April 2026).
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015; Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F., Eds.; Springer: Cham, Switzerland, 2015; Volume 9351, pp. 234–241. [CrossRef]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. In Proceedings of the 32nd International Conference on Machine Learning, Lille, France, 6–11 July 2015; Bach, F.; Blei, D., Eds.; PMLR: Lille, France, 2015; Volume 37, pp. 448–456.
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. In Proceedings of the IEEE International Conference on Computer Vision (ICCV 2015), Santiago, Chile, 7–13 December 2015; IEEE: Santiago, Chile, 2015; pp. 1026–1034. [CrossRef]
- Milletari, F.; Navab, N.; Ahmadi, S.-A. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. In Proceedings of the Fourth International Conference on 3D Vision (3DV 2016), Stanford, CA, USA, 25–28 October 2016; IEEE: Stanford, CA, USA, 2016; pp. 565–571. [CrossRef]
- Sudre, C.H.; Li, W.; Vercauteren, T.; Ourselin, S.; Cardoso, M.J. Generalised Dice overlap as a deep learning loss function for highly unbalanced segmentations. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Wang, Q., Shi, Y., Suk, H.-I., Suzuki, K., Eds.; Springer: Cham, Switzerland, 2017; Volume 10553, pp. 240–248. [CrossRef]
- Loshchilov, I.; Hutter, F. Decoupled Weight Decay Regularization. arXiv 2017, arXiv:1711.05101. [CrossRef]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; Desmaison, A.; Köpf, A.; Yang, E.; DeVito, Z.; Raison, M.; Tejani, A.; Chilamkurthy, S.; Steiner, B.; Fang, L.; Bai, J.; Chintala, S. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Advances in Neural Information Processing Systems 32 (NeurIPS 2019); Wallach, H.; Larochelle, H.; Beygelzimer, A.; d’Alché-Buc, F.; Fox, E.; Garnett, R., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2019; pp. 8024–8035.
- Evangelidis, G.D.; Psarakis, E.Z. Parametric Image Alignment Using Enhanced Correlation Coefficient Maximization. IEEE Trans. Pattern Anal. Mach. Intell. 2008, 30, 1858–1865. [CrossRef]
- Farnebäck, G. Two-Frame Motion Estimation Based on Polynomial Expansion. In Image Analysis; Bigun, J.; Gustavsson, T., Eds.; Springer: Berlin, Heidelberg, Germany, 2003; pp. 363–370.
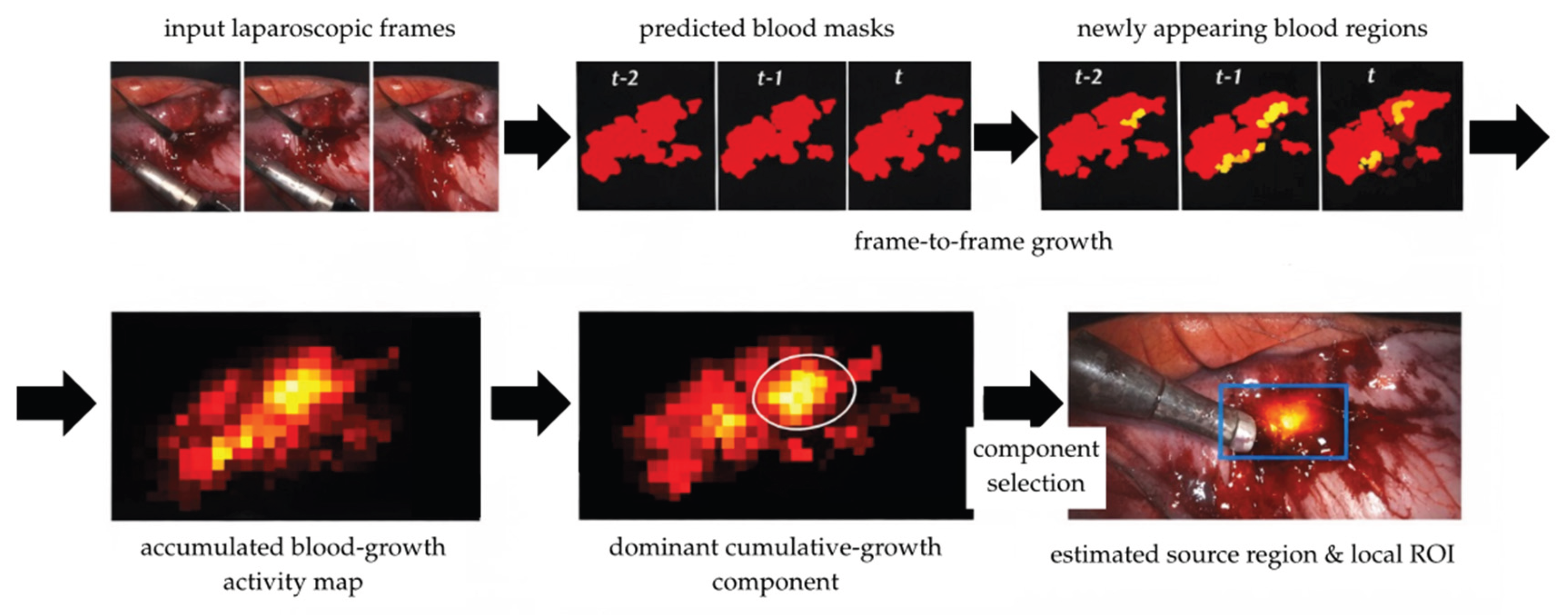
| Sequence | Source | Pattern | Duration [s] | Peak dA/dt (px/s) |
Mean temporal IoU |
Mean abs change ratio | Mean flow magnitude | P95 flow magnitude |
|---|---|---|---|---|---|---|---|---|
| int-1 | internal | burst-like coherent progression | 2.47 | 5.01106 | 0.909 | 0.036 | 1.978 | 20.590 |
| int-2 | internal | motion-active progression | 20.47 | 7.88106 | 0.849 | 0.068 | 2.916 | 27.537 |
| int-3 | internal | burst-like coherent progression | 20.00 | 1.62107 | 0.900 | 0.044 | 2.514 | 27.592 |
| int-4 | internal | more spatially coherent progression | 17.87 | 5.05106 | 0.882 | 0.055 | 2.526 | 23.957 |
| int-5 | internal | dynamic and unstable progression | 31.50 | 3.87106 | 0.789 | 0.258 | 3.583 | 26.291 |
| ext-static | external | static reference | 4.70 | 1.83106 | 0.894 | 0.065 | 0.006 | 0.046 |
| ext-1 | external | dynamic and unstable progression | 6.07 | 1.57106 | 0.690 | 0.179 | 7.213 | 31.167 |
| ext-2 | external | more spatially coherent progression | 4.83 | 5.23105 | 0.856 | 0.045 | 1.966 | 23.032 |
| ext-3 | external | motion-active despite relative stability | 6.97 | 3.46106 | 0.865 | 0.04 | 3.091 | 29.703 |
| ext-4 | external | localized burst | 1.53 | 4.01106 | 0.816 | 0.078 | 1.250 | 9.051 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




