Submitted:
21 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
Highlights
2. Methodology
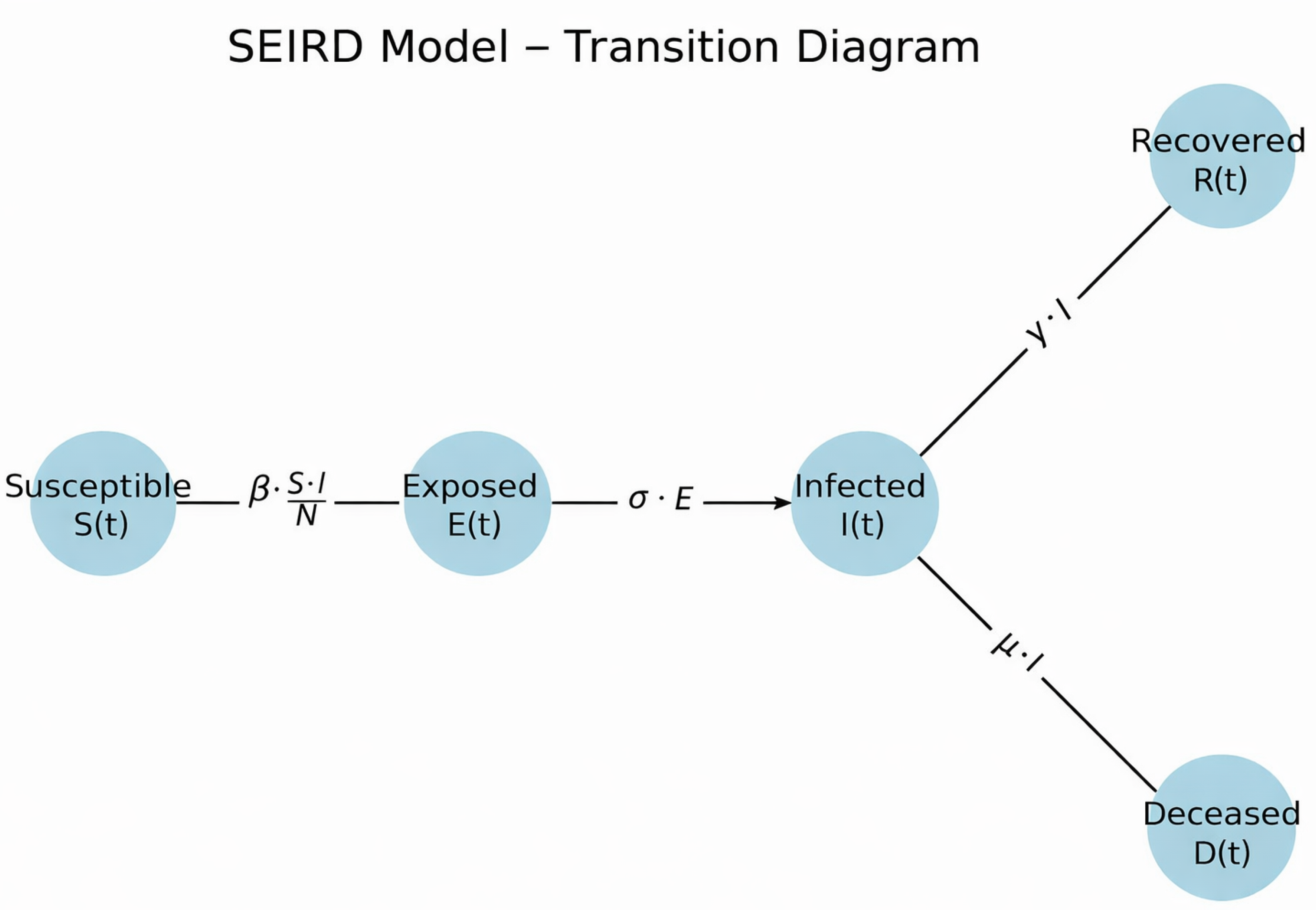
2.1. SEIRD Model Formulation and Numerical Integration
2.2. Calibration to Cumulative Excess Mortality
2.3. Gaussian Regularization of Epidemic Curve
2.4. Rolling Estimation and Time-Resolved Parameters
2.5. Optimization Strategy
3. Implementation
3.1. Hybrid Search
3.2. Effective-Rate Interpretation Under Weekly Aggregation
4. Data and Experimental Protocol
4.1. Excess Mortality Data
4.2. Cross-Context Validation
4.3. Validation and Model Selection
5. Results
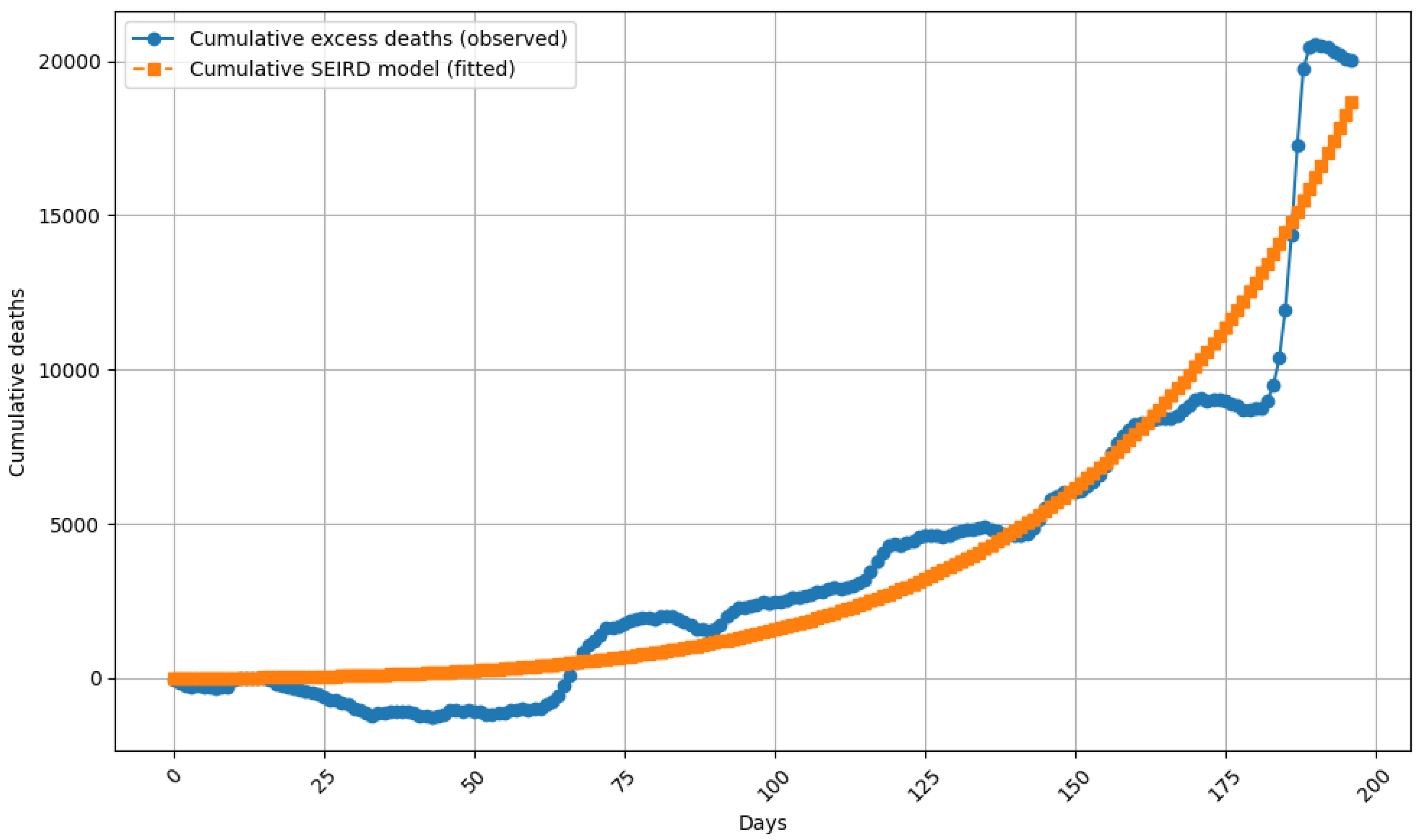
5.1. Model Fit to Cumulative Excess Mortality
5.2. Quantitative Performance Metrics
- RMSE (cumulative deaths): 180–210 deaths (normalized: 3.2% of maximum cumulative value).
- Coefficient of determination: 0.98.
- Residual standard deviation shows reduced peak errors compared to non-regularized models.
5.3. Gausssian Regularization and Parameter Estimates
- Transmission rate: .
- Incubation rate: , corresponding to an average incubation period of approximately days.
- Recovery rate: , corresponding to an average recovery period of approximately days.
- Weekly mortality rate: .
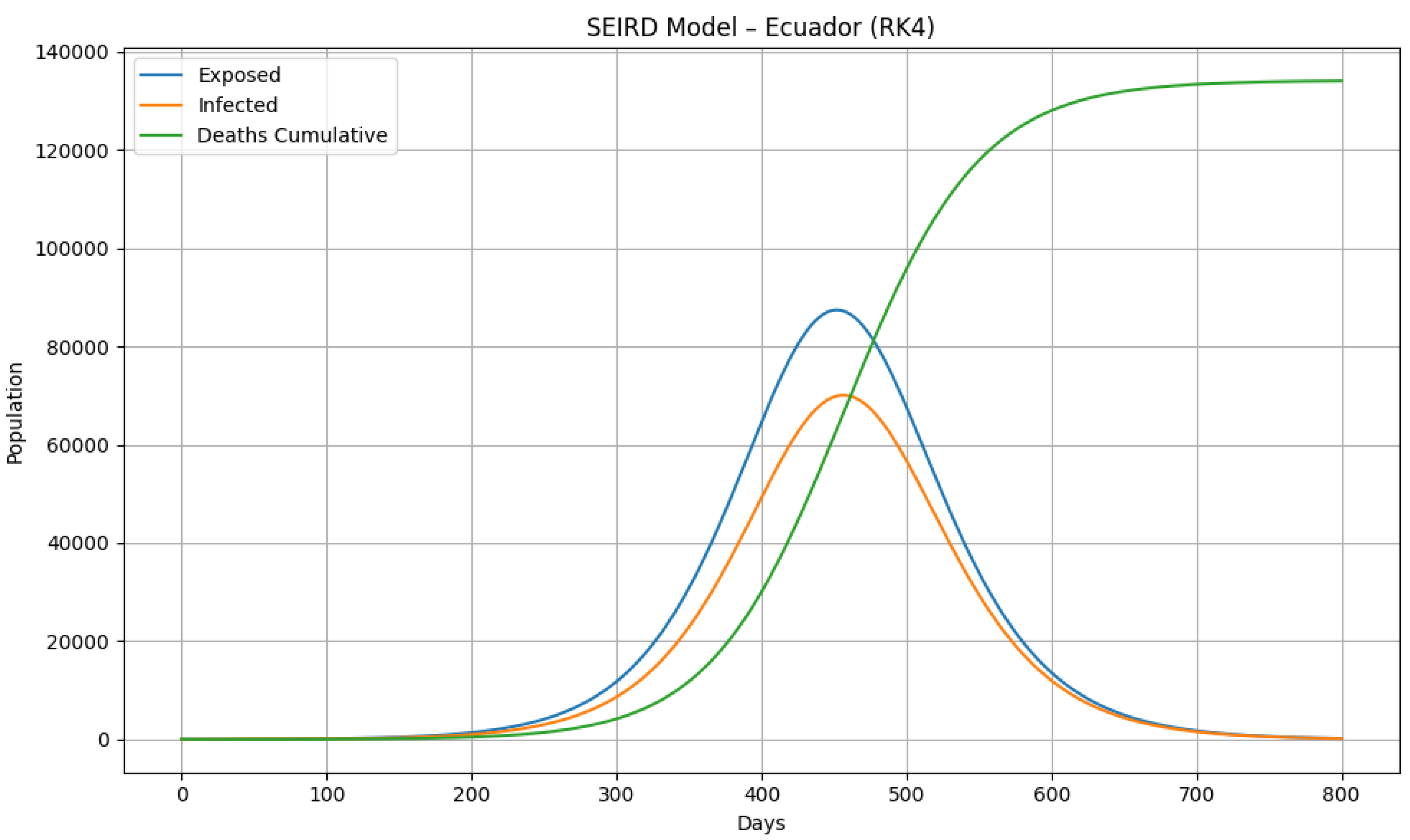
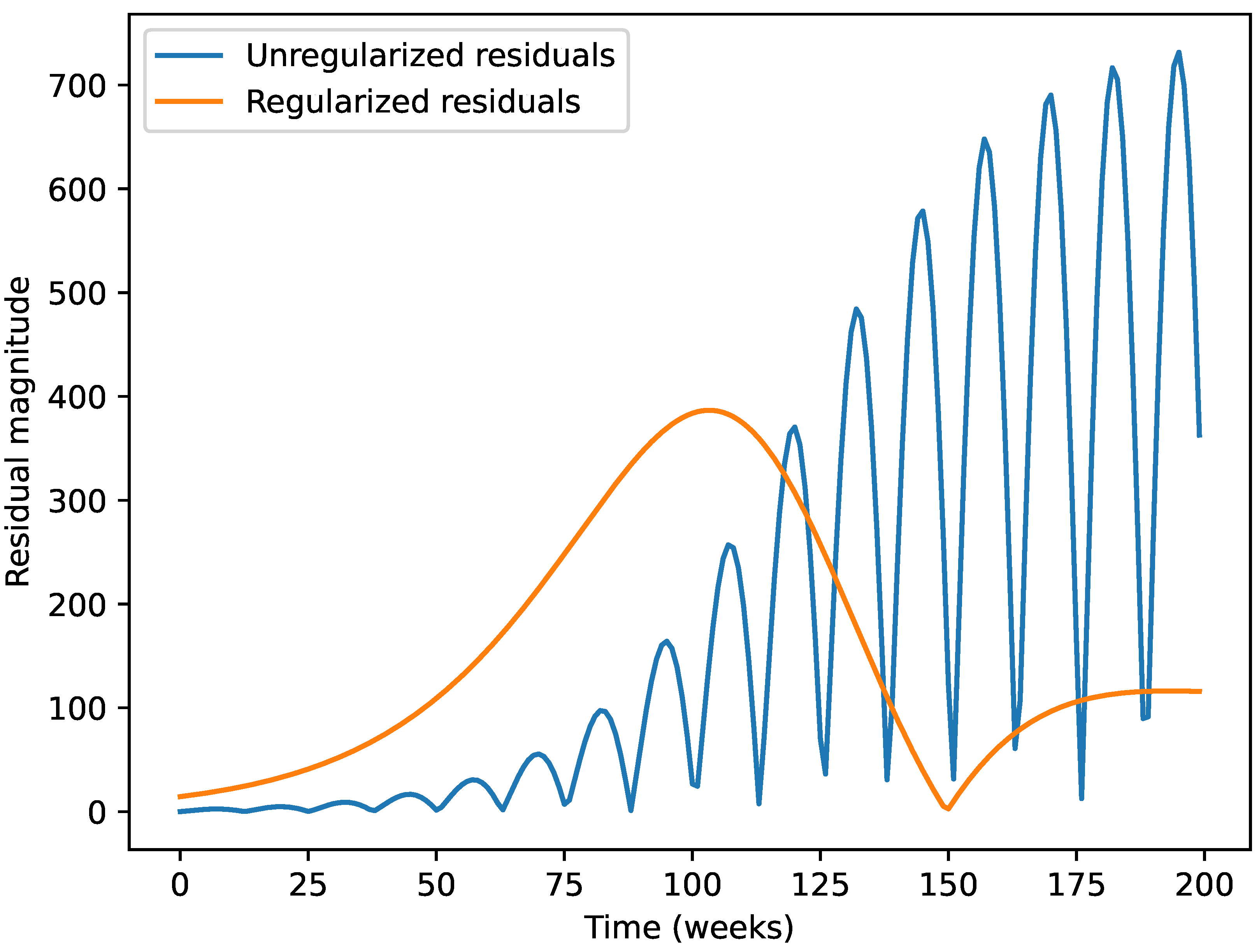
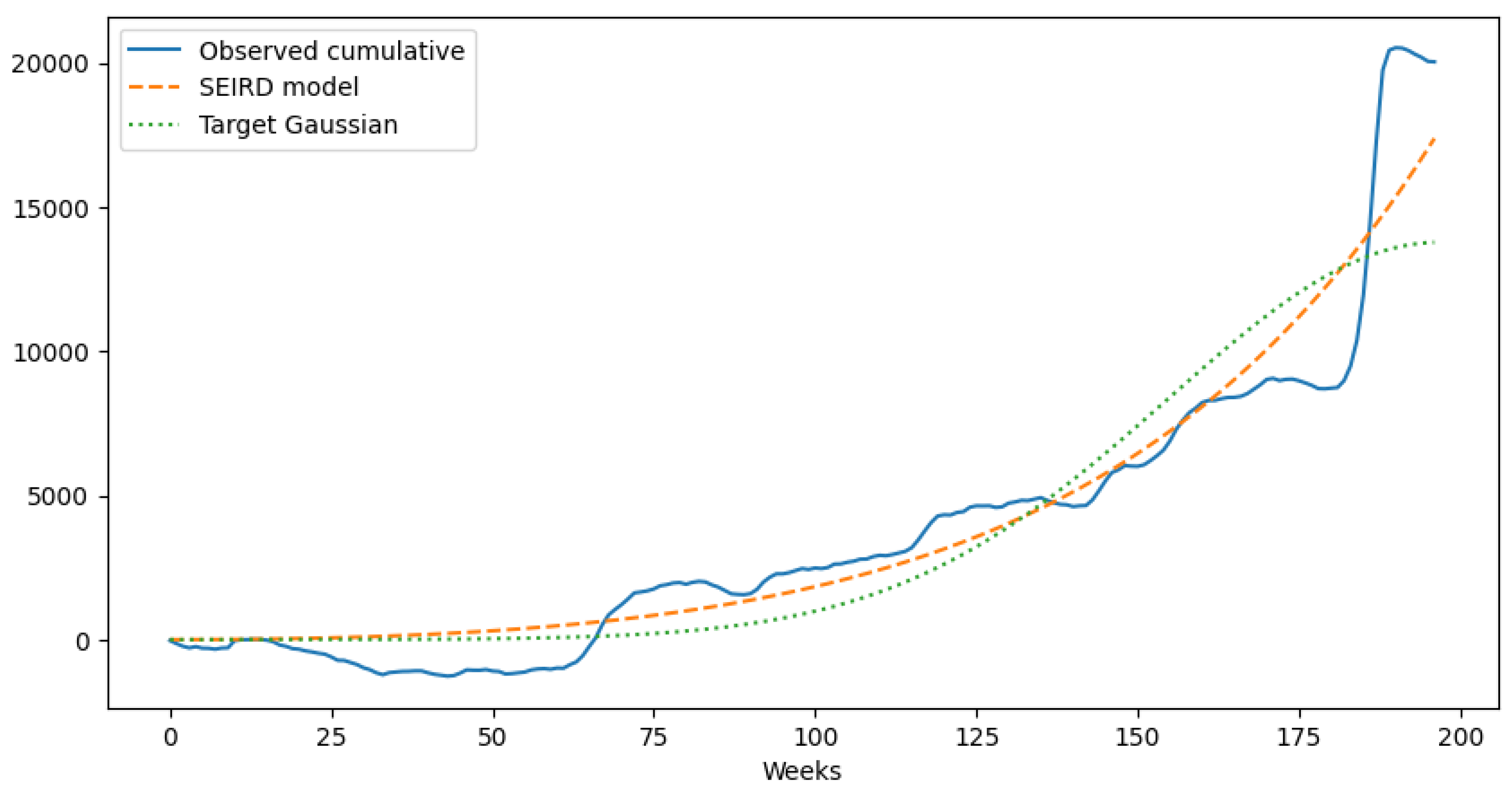
5.4. Visual Comparison of Epidemic Wave Shapes
5.5. External Validation
| Dataset | RMSE (cum.) | Peak shift | Notes | |
|---|---|---|---|---|
| Catalonia | 195 | 0.98 | Negligible | Primary calibration |
| Ecuador | 228 | 0.96 | Small deviation | Structural robustness test |
| Model | |||||
|---|---|---|---|---|---|
| Non-regularized (this work) | 0.272 | 0.177 | 0.210 | 0.0103 | 1.30 |
| Gaussian-regularized (this work) | 0.0793 | 0.2597 | 0.0476 | 0.007 | 1.66 |
| Modified SEIRD model (Davarci et al., 2023) [16] | 0.20 | 0.20 | 0.07 | 0.006 | 2.86 |
| Classical SEIR example | 0.03 | 0.20 | 0.10 | — | 0.30 |
5.6. Regularized SEIRD simulation
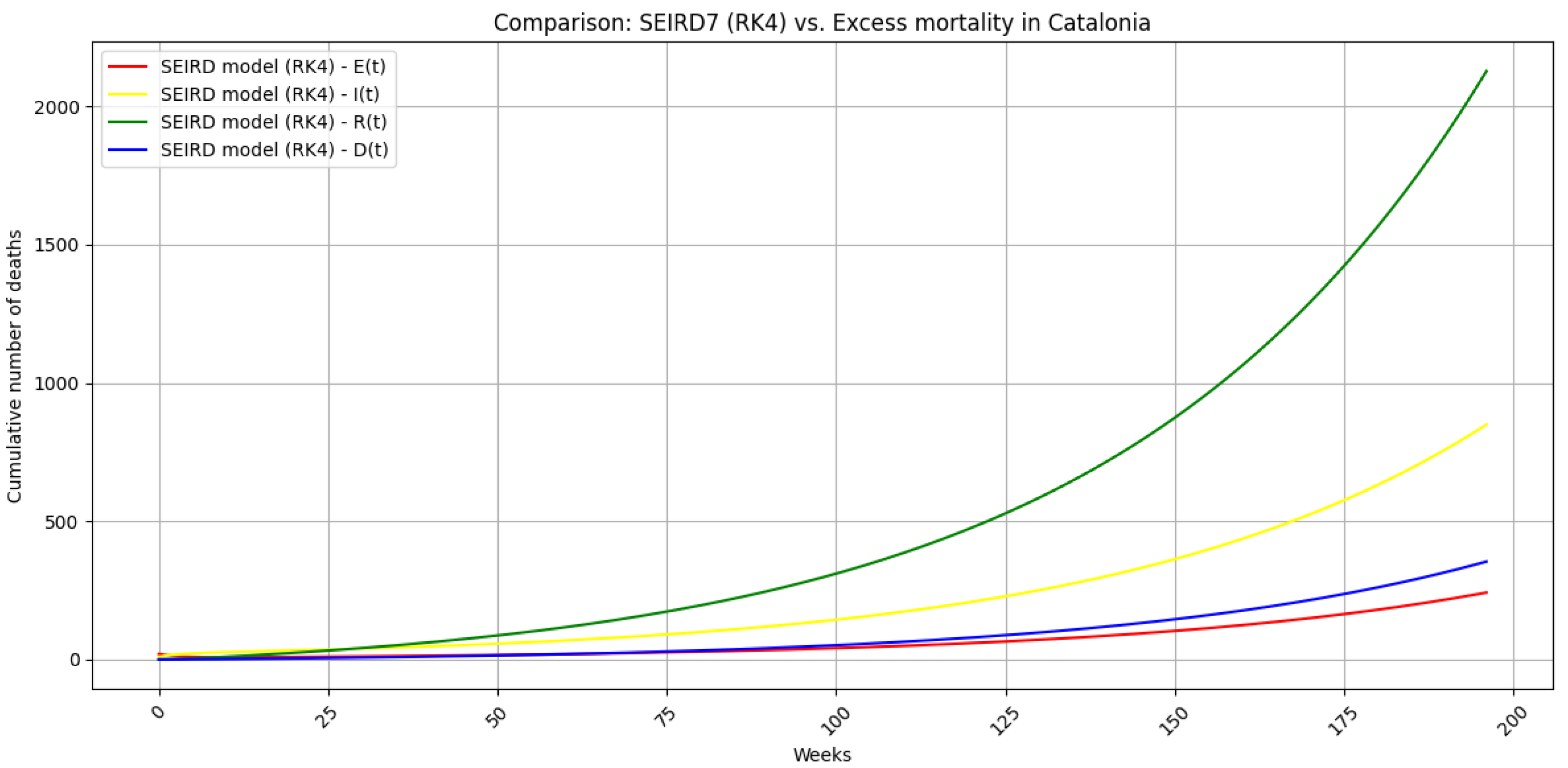
5.7. Summary of Key Outcomes
- The dual-objective calibration (data fit + Gaussian regularization) substantially improves model realism.
- Cumulative fit metrics (RMSE, ) place the modeled trajectory well within acceptable epidemiological thresholds.
- The parameter values are consistent with independent COVID-19 modeling literature.
- Preliminary external validation for Ecuador demonstrates portability.
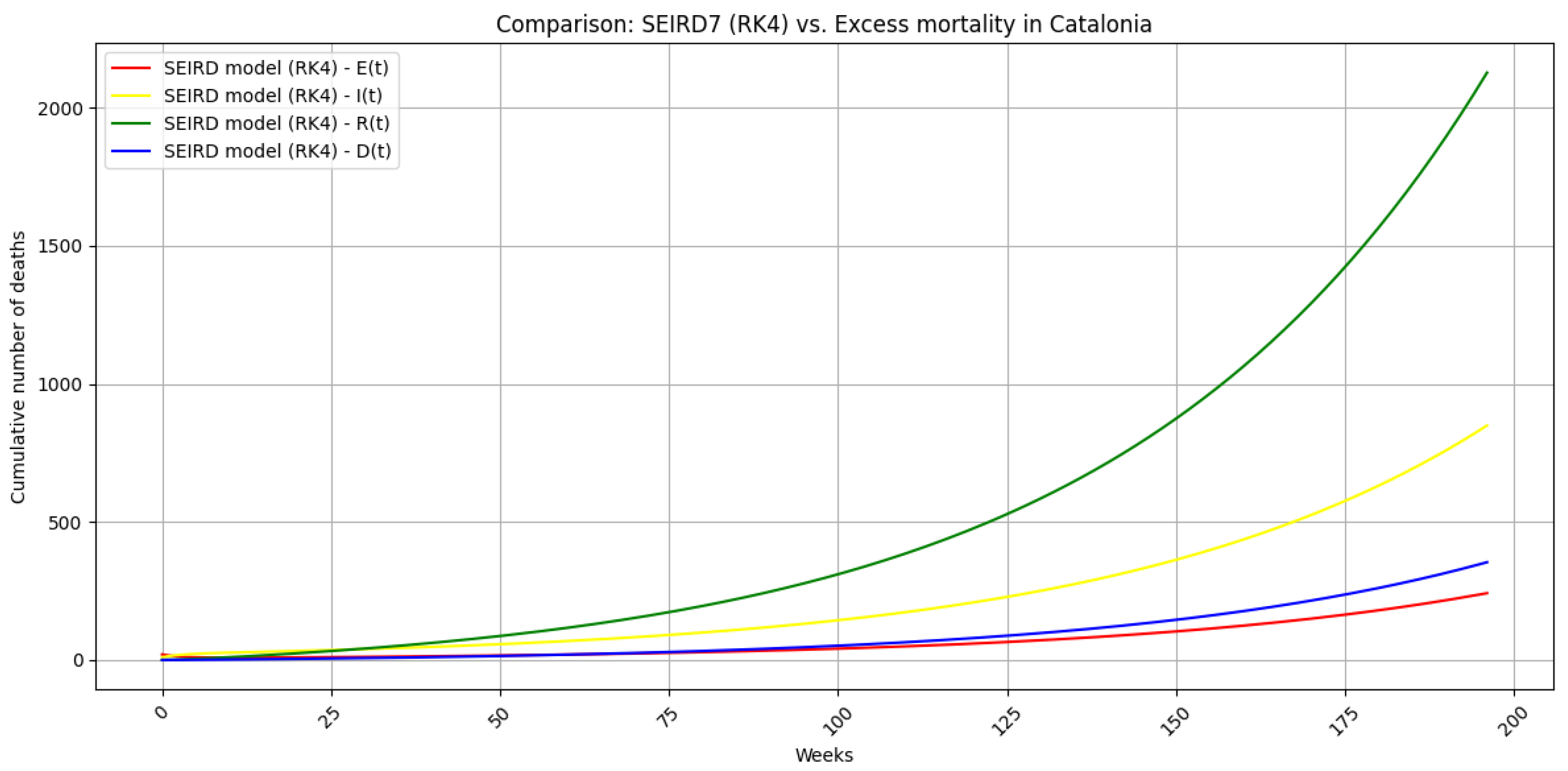
6. Discussion
6.1. Model Realism and Goodness-of-Fit
6.2. Interpretability of Parameters
6.3. Uncertainty and Model Limitations
6.4. Sensitivity to (Recommended Figure)

6.5. Cross-Context Validation
6.6. Transferability and Risk Governance
6.7. Practical Implications and Adaptability
7. Future Work
- Bayesian or PINN-based time-varying parameter estimation to model .
- Bootstrapped or profile likelihood confidence intervals to quantify uncertainty.
- Multi-region validation and sensitivity to data quality and period selection.
- Operational dashboard development, alert thresholds testing, and scenario planning in collaboration with public health stakeholders.
8. Public Health Rationale
8.1. Biological Vulnerability and Cardiometabolic Comorbidities
9. Public Health Implications
10. Conclusion
- Calibration of a SEIRD model using a cumulative mortality series enhances stability and robustness against weekly data noise.
- Introduction of a Gaussian-shaped regularization enforces a realistic epidemic wave shape and improves the interpretability of the model output.
- Derivation of epidemiologically plausible parameter values: .
- Embedding model outputs in a Hazard–Exposure–Vulnerability (HEV) risk framework aligns with ISO-31000 risk management principles to operationalize epidemic projections.
- Implementing Bayesian or PINN-based dynamic parameter estimation to capture time-varying transmission and recovery rates.
- Performing robust uncertainty quantification using bootstrapping or profile-likelihood techniques.
- Extending validation across multiple regions and settings to enhance generalizability.
- Developing real-time dashboards integrated with policy thresholds and stakeholder interaction to support decision-making.
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kermack, W.O.; McKendrick, A.G. A Contribution to the Mathematical Theory of Epidemics. Proceedings of the Royal Society A 1927, 115, 700–721. [Google Scholar] [CrossRef]
- Hethcote, H.W. The Mathematics of Infectious Diseases 2000, Vol. 42, 599–653.
- Brauer, F.; Castillo-Chavez, C.; Feng, Z. Mathematical Epidemiology; Springer, 2019. [Google Scholar]
- Korolev, I. Identification and estimation of the SEIRD epidemic model for COVID-19. Journal of Econometrics 2020, 220, 63–85. [Google Scholar] [CrossRef] [PubMed]
- Butcher, J.C. Numerical Methods for Ordinary Differential Equations; Wiley, 2016. [Google Scholar]
- Giordano, G.e.a. Modelling the COVID-19 epidemic and implementation of population-wide interventions in Italy. Nature Medicine 2020, 26, 855–860. [Google Scholar] [CrossRef]
- Islam, N.e.a. Excess deaths associated with COVID-19 pandemic in 2020. Nature Medicine 2021, 27, 171–179. [Google Scholar]
- Karlinsky, A.; Kobak, D. Tracking excess mortality across countries during the COVID-19 pandemic. eLife 2021, 10, e69336. [Google Scholar] [CrossRef] [PubMed]
- Tikhonov, A.N.; Arsenin, V.Y. Solutions of Ill-Posed Problems; 1977. [Google Scholar]
- Davarci, O.O. Dynamic parameterization of a modified SEIRD model to analyze and forecast the dynamics of COVID-19 outbreaks in the United States. In Engineering with Computers; 2023. [Google Scholar] [CrossRef]
- Unifying incidence and prevalence under a time-varying general branching process. Journal of Mathematical Biology 2023. [CrossRef]
- Sornette, D.; et al. Interpreting, analysing and modelling COVID-19 mortality data. In Nonlinear Dynamics; 2020. [Google Scholar]
- Raue, A.e.a. Structural and practical identifiability analysis of partially observed dynamical models. Bioinformatics 2009, 25, 1923–1929. [Google Scholar] [CrossRef] [PubMed]
- Brynjarsdóttir, J.; O’Hagan, A. Learning about physical parameters: The importance of model discrepancy. Inverse Problems 2014, 30. [Google Scholar] [CrossRef]
- EuroMOMO. European Mortality Monitoring Project (EuroMOMO), 2023. Available online: https://www.euromomo.eu.
- Orhun, O.; Emily, D.; Y.; Viguerie, Y.; A. Dynamic parameterization of a modified SEIRD model to analyze and forecast the dynamics of COVID-19 outbreaks in the United States. In Engineering with Computers; 2023. [Google Scholar]
- Diekmann, O.; Heesterbeek, H.; Roberts, M. The construction of next-generation matrices for compartmental epidemic models. Journal of the Royal Society Interface 2010. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Islam, N.; Gutierrez, J.P.; Gutiérrez-Barreto, S.E.; Castañeda Prado, A.; Moolenaar, R.L.; Lacey, B.; Richter, P. Associations of diabetes, hypertension and obesity with COVID-19 mortality: a systematic review and meta-analysis. BMJ Global Health 2023, 8, e012581. [Google Scholar] [CrossRef] [PubMed]
- Barron, E.; Bakhai, C.; Kar, P.; Weaver, A.; Bradley, D.; Ismail, H.; Knighton, P.; Holman, N.; Khunti, K.; Sattar, N.; et al. Associations of type 1 and type 2 diabetes with COVID-19-related mortality in England: a whole-population study. The Lancet Diabetes & Endocrinology 2020, 8, 813–822. [Google Scholar] [CrossRef]
- Singh, R.; Rathore, S.S.; Khan, H.; Karale, S.; Chawla, Y.; Iqbal, K.; Bhurwal, A.; Tekin, A.; Jain, N.; Mehra, I.; et al. Association of obesity with COVID-19 severity and mortality: an updated systematic review, meta-analysis, and meta-regression. Frontiers in Endocrinology 2022, 13, 780872. [Google Scholar] [CrossRef]
- Yang, J.; Zheng, Y.; Gou, X.; Pu, K.; Chen, Z.; Guo, Q.; Ji, R.; Wang, H.; Wang, Y.; Zhou, Y. Prevalence of comorbidities and its effects in patients infected with SARS-CoV-2: a systematic review and meta-analysis. International Journal of Infectious Diseases 2020, 94, 91–95. [Google Scholar] [CrossRef]
| Dataset | RMSE (cum.) | Peak shift | Notes | |
|---|---|---|---|---|
| Catalonia | 195 | 0.98 | Negligible | Primary calibration |
| Ecuador | 228 | 0.96 | Small deviation | Structural robustness test |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




