Background: Lung cancer causes more deaths than any other malignancy worldwide, accounting for 2.2 million new cases and 1.8 million deaths in 2020. Extracting structured clinical knowledge from unstructured French-language oncology records remains methodologically unresolved in Tunisian and Francophone healthcare systems, where validated natural language processing tools do not yet exist. This study examined the effectiveness of transformer-based named entity recognition for automated clinical annotation of Tunisian lung cancer reports.
Aim: The study aimed to (i) benchmark four transformer-based models on a publicly available thoracic radiology dataset, (ii) evaluate five models, including a French biomedical specialist, on a newly constructed Tunisian clinical corpus, and (iii) demonstrate prototype deployment feasibility for structured clinical decision support.
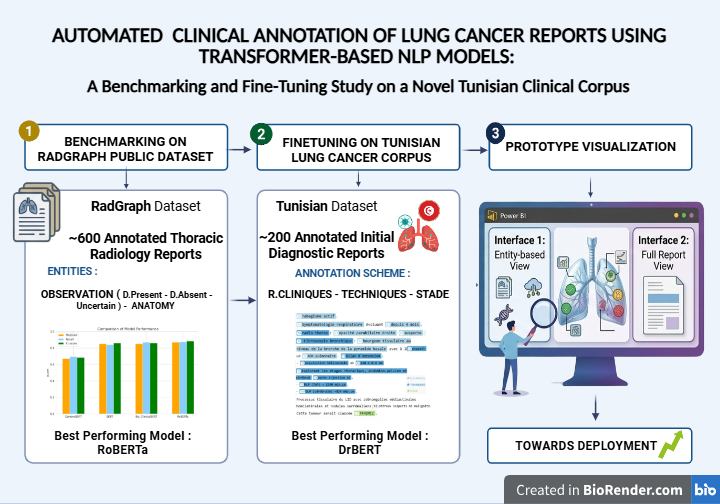
Methods: A benchmarking study evaluated BERT, RoBERTa, BioClinicalBERT, and CamemBERT on the RadGraph dataset (600 annotated thoracic radiology reports). Five models were subsequently fine-tuned on 200 manually annotated initial diagnostic reports from Mami Pneumo-Phthisiology Hospital, Tunis. All models were trained for a maximum of 10 epochs, with a learning rate of 5x10-5, a batch size of 16, and an 80/10/10 train-validation-test split, and evaluated using precision, recall, and F1-score.
Results: On RadGraph, RoBERTa achieved the highest F1-score of 0.873 (precision: 0.869, recall: 0.877), followed by BioClinicalBERT (F1: 0.868) and BERT (F1: 0.857). CamemBERT achieved an F1 score of 0.682 on this English dataset. On the Tunisian corpus, DrBERT outperformed all models with an F1-score of 0.811, compared to RoBERTa at 0.79. A prototype interface generated structured clinical summaries encompassing prior conditions, imaging modalities, and TNM staging.
Conclusion: Language- and domain-adapted transformer models effectively extract structured clinical entities from French-language Tunisian lung cancer reports. DrBERT's precision advantage confirms that biomedical pretraining in the target language is the primary driver of performance in specialized French oncology text. This work establishes foundational infrastructure for NLP-driven oncology data management in Tunisia and comparable Francophone settings.