Submitted:
20 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Collection and Quality Control
2.2. Genome Annotation and Architectural Characterization
2.3. Genomic Plasticity and Molecular Evolution Analysis
2.4. Niche-Specific Functional Divergence and Evolution
2.5. Statistical Analysis and Visualization
3. Results
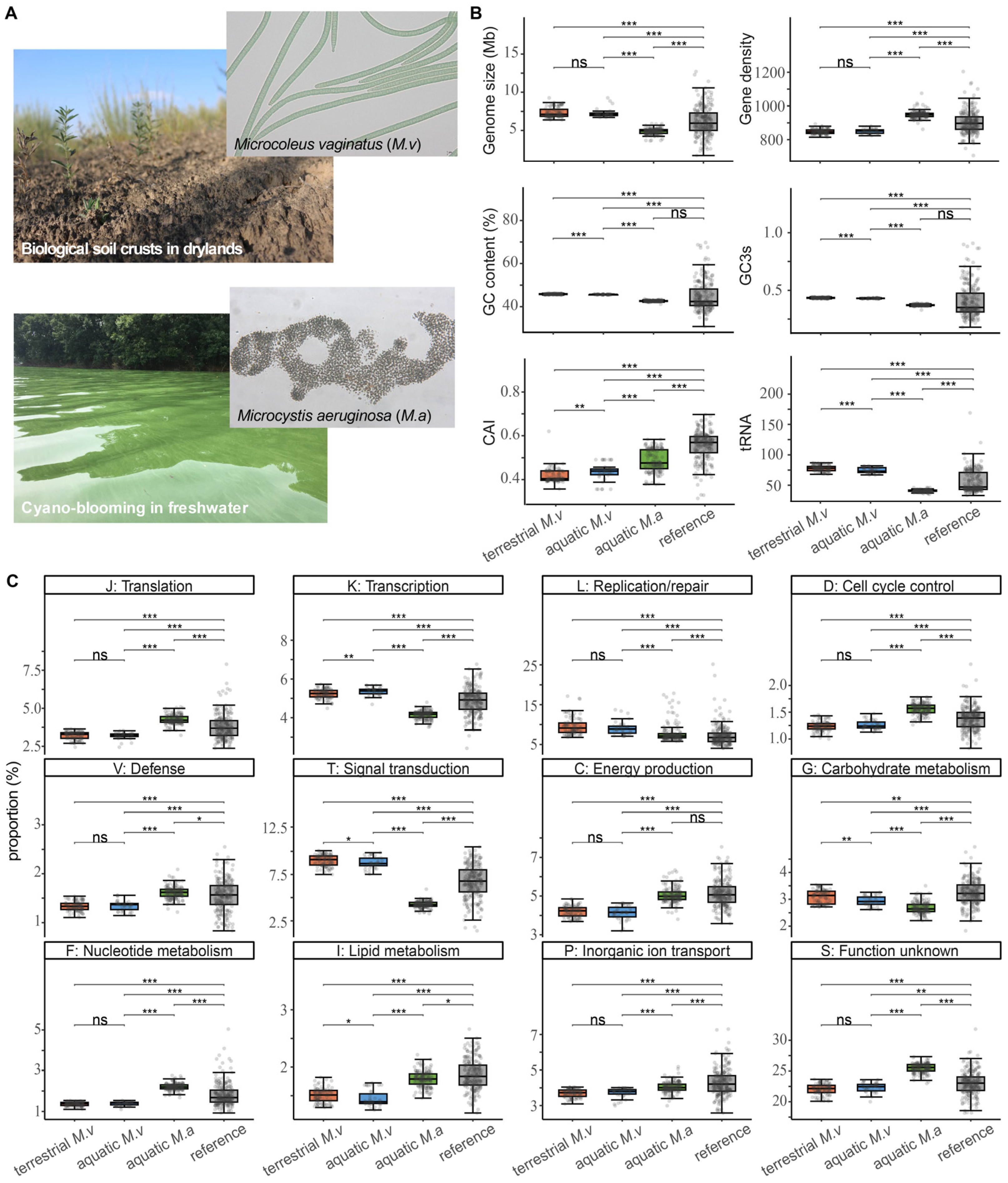
3.1. Divergence in Genomic Architecture and Functional Repertoires
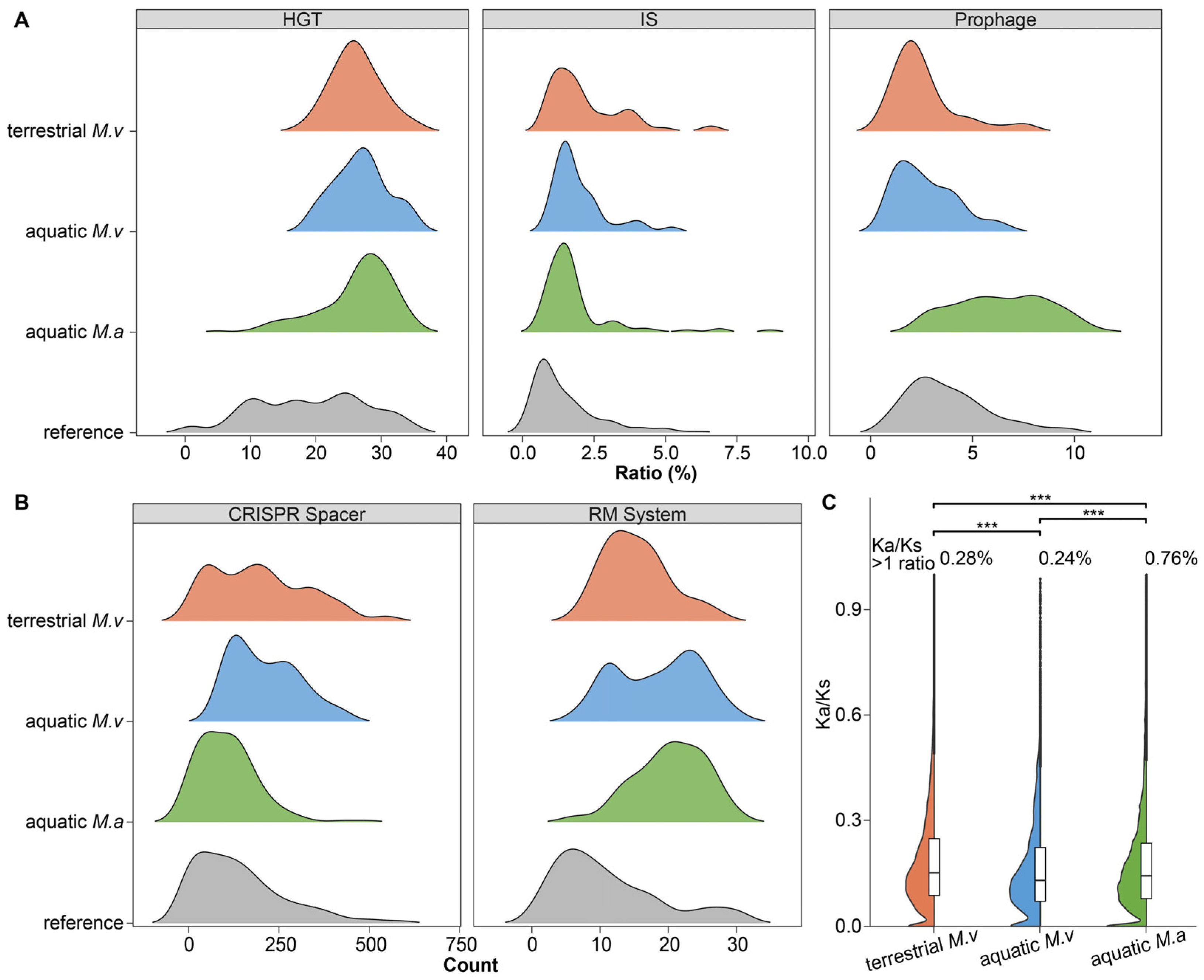
3.2. Genomic Plasticity and Defense System Trade-Offs
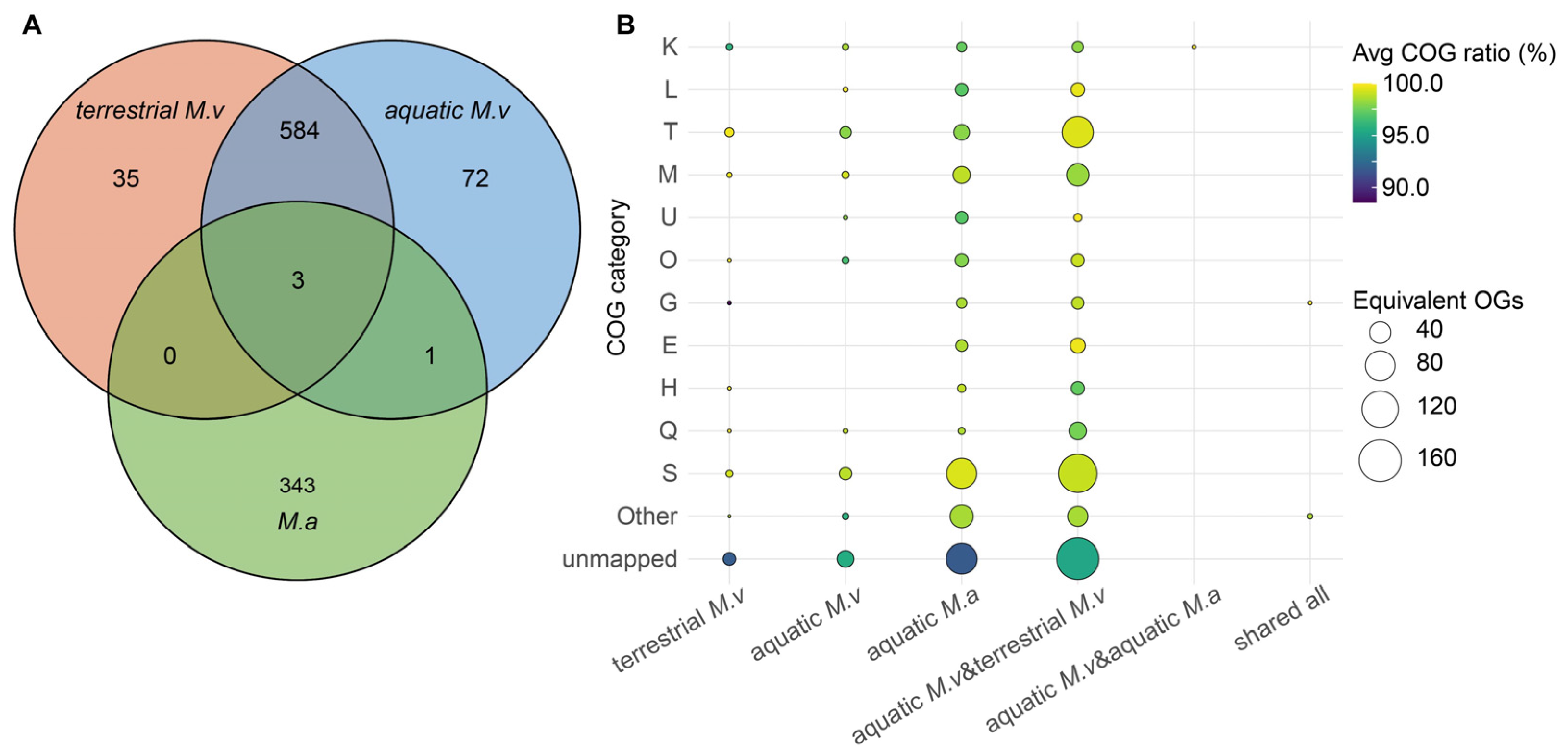
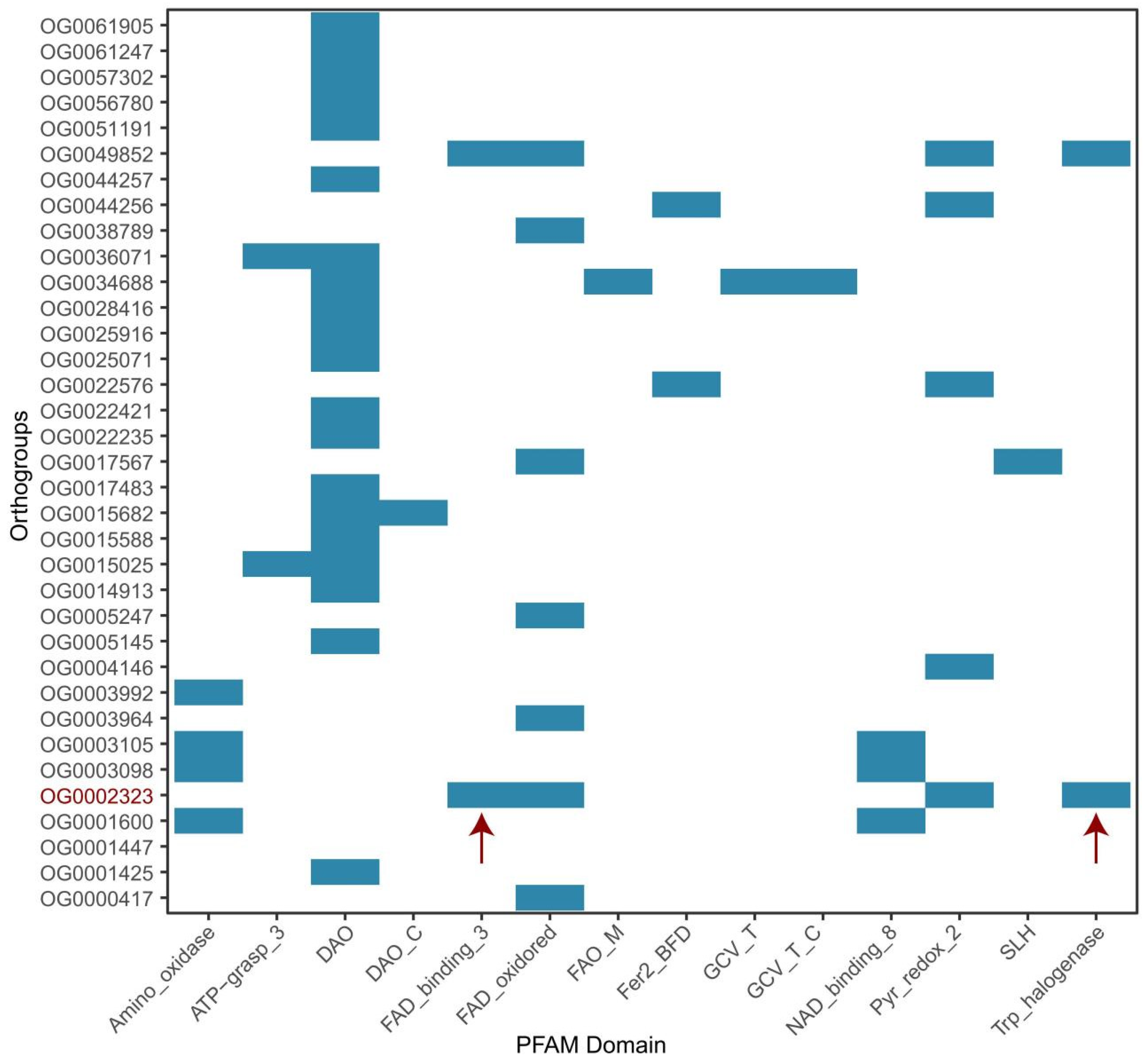
3.3. Niche-specific Functional Divergence and Adaptation
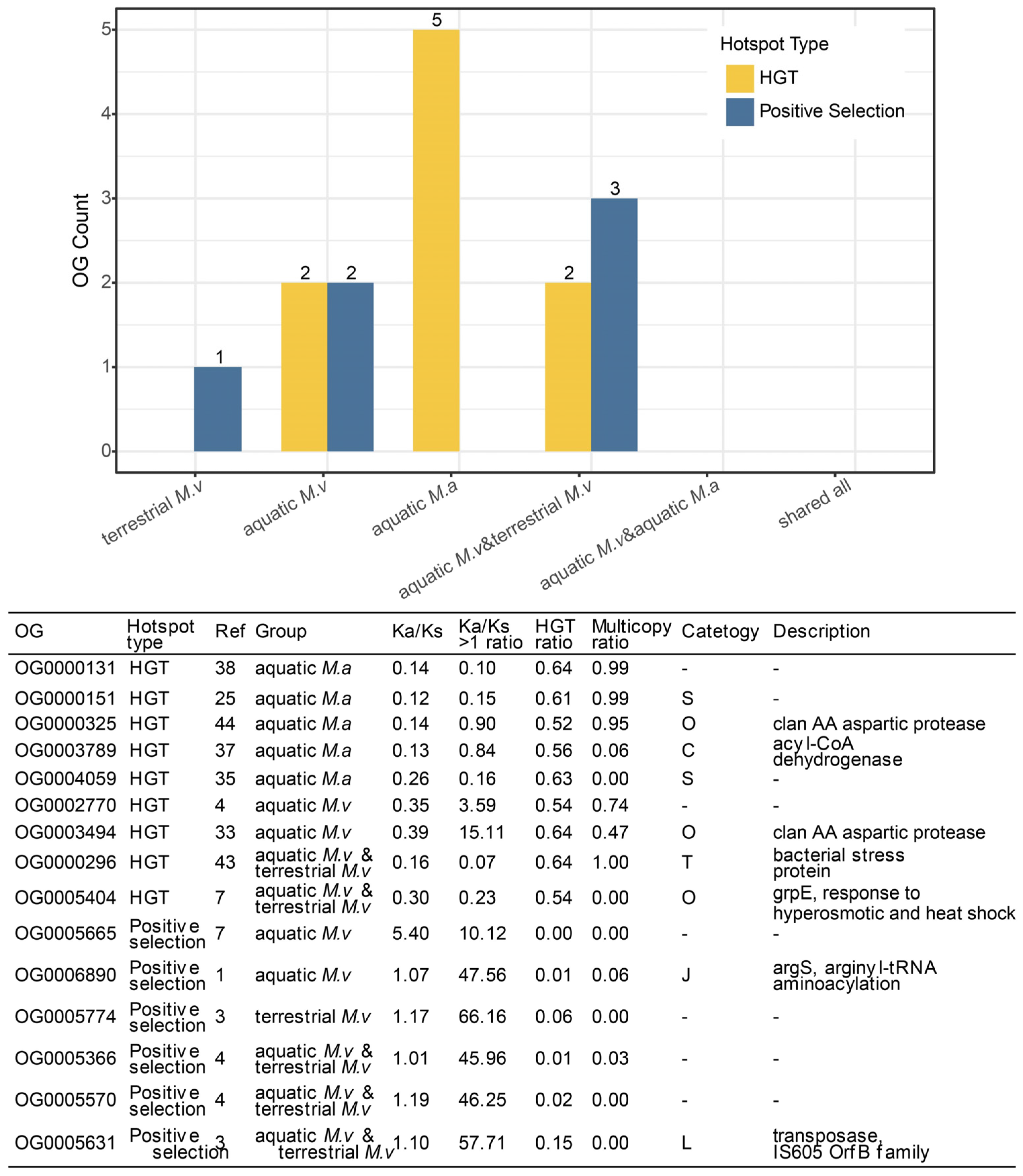
3.4. Evolutionary Drivers of Niche-Specific Genes
4. Discussion
4.1. Genomic Plasticity as a Prerequisite for Ecological Dominance
4.2. Universal retention Requirements Across Ecological Boundaries
4.3. Divergent Investment Upon Universal Chassis
4.4. Aquatic-Specific Adaptation and Gradual Ecological Transition
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sanchez-Baracaldo, P.; Hayes, P.K.; Blank, C.E. Morphological and habitat evolution in the Cyanobacteria using a compartmentalization approach. Review. Geobiology 2005, 3, 145–165. [Google Scholar] [CrossRef]
- Whitton, B.; Potts, M. The ecology of cyanobacteria. Their diversity in time and space.; Kluwer Academic Publishers: Dordrecht, 2000. [Google Scholar]
- Chen, M.Y.; Teng, W.K.; Zhao, L.; et al. Comparative genomics reveals insights into cyanobacterial evolution and habitat adaptation. ISME J. 2021, 15, 211–227. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Pichel, F.; Belnap, J.; Neuer, S.; Schanz, F.J.A.S. Estimates of global cyanobacterial biomass and its distribution. Algological Studies 2003, 109, 213–227. [Google Scholar] [CrossRef]
- Zehr, J.; Bench, S.; Carter, B.; et al. Globally Distributed Uncultivated Oceanic N2-Fixing Cyanobacteria Lack Oxygenic Photosystem II. Science 2008, 322, 1110–1112. [Google Scholar] [CrossRef] [PubMed]
- Bowker, M.A.; Maestre, F.T.; Eldridge, D.; et al. Biological soil crusts (biocrusts) as a model system in community, landscape and ecosystem ecology. Biodivers Conserv. 2014, 23, 1619–1637. [Google Scholar] [CrossRef]
- Stanojkovic, A.; Skoupy, S.; Johannesson, H.; Dvorak, P. The global speciation continuum of the cyanobacterium Microcoleus. Nat Commun. 2024, 15, 2122. [Google Scholar] [CrossRef]
- Li, H.; Huo, D.; Wang, W.; et al. Multifunctionality of biocrusts is positively predicted by network topologies consistent with interspecies facilitation. Mol Ecol. 2020, 29, 1560–1573. [Google Scholar] [CrossRef]
- Harke, M.J.; Steffen, M.M.; Gobler, C.J.; et al. A review of the global ecology, genomics, and biogeography of the toxic cyanobacterium, Microcystis spp. Harmful Algae 2016, 54, 4–20. [Google Scholar] [CrossRef]
- Huo, D.; Gan, N.; Geng, R.; et al. Cyanobacterial blooms in China: diversity, distribution, and cyanotoxins. Harmful Algae 2021, 109, 102106. [Google Scholar] [CrossRef]
- Lakshmikandan, M.; Li, M.; Pan, B. Cyanobacterial Blooms in Environmental Water: Causes and Solutions. Current Pollution Reports 2024, 10, 606–627. [Google Scholar] [CrossRef]
- Tatters, A.; Howard, M.; Nagoda, C.; Busse, L.; Gellene, A.; Caron, D. Multiple Stressors at the Land-Sea Interface: Cyanotoxins at the Land-Sea Interface in the Southern California Bight. Toxins 2017, 9, 95. [Google Scholar] [CrossRef]
- Shih, P.M.; Wu, D.; Latifi, A.; et al. Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. PNAS 2013, 110, 1053–1058. [Google Scholar] [CrossRef] [PubMed]
- Yamamichi, M. How does genetic architecture affect eco-evolutionary dynamics? A theoretical perspective. Phil Trans R Soc B 2022, 377, 20200504. [Google Scholar] [CrossRef]
- Murik, O.; Oren, N.; Shotland, Y.; et al. What distinguishes cyanobacteria able to revive after desiccation from those that cannot: the genome aspect. Environ Microbiol. 2017, 19, 535–550. [Google Scholar] [CrossRef]
- Chrismas, N.A.M.; Anesio, A.M.; Sanchez-Baracaldo, P. The future of genomics in polar and alpine cyanobacteria. FEMS Microbiol Ecol. 2018, 94. [Google Scholar] [CrossRef]
- Li, C.; Liao, H.; Xu, L.; et al. The adjustment of life history strategies drives the ecological adaptations of soil microbiota to aridity. Mol Ecol. 2022, 31, 2920–2934. [Google Scholar] [CrossRef] [PubMed]
- Muraille, E. Diversity Generator Mechanisms Are Essential Components of Biological Systems: The Two Queen Hypothesis. Front Microbiol. 2018, 9, 223. [Google Scholar] [CrossRef] [PubMed]
- Sriswasdi, S.; Yang, C.C.; Iwasaki, W. Generalist species drive microbial dispersion and evolution. Nat Commun. 2017, 8, 1162. [Google Scholar] [CrossRef]
- Ellegren, H.; Galtier, N. Determinants of genetic diversity. Nat Rev Genet. 2016, 17, 422–433. [Google Scholar] [CrossRef]
- Chu, X.; Li, S.; Wang, S.; Luo, D.; Luo, H. Gene loss through pseudogenization contributes to the ecological diversification of a generalist Roseobacter lineage. ISME J. 2021, 15, 489–502. [Google Scholar] [CrossRef]
- Wheatley, R.M.; MacLean, R.C. CRISPR-Cas systems restrict horizontal gene transfer in Pseudomonas aeruginosa. ISME J. 2021, 15, 1420–1433. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Pichel, F.; Wojciechowski, M. The Evolution of a Capacity to Build Supra-Cellular Ropes Enabled Filamentous Cyanobacteria to Colonize Highly Erodible Substrates. PLoS One 2009, 4, e7801. [Google Scholar] [CrossRef] [PubMed]
- Rajeev, L.; da Rocha, U.N.; Klitgord, N.; et al. Dynamic cyanobacterial response to hydration and dehydration in a desert biological soil crust. ISME J. 2013, 7, 2178–2191. [Google Scholar] [CrossRef]
- Couradeau, E.; Giraldo-Silva, A.; De Martini, F.; Garcia-Pichel, F. Spatial segregation of the biological soil crust microbiome around its foundational cyanobacterium, Microcoleus vaginatus, and the formation of a nitrogen-fixing cyanosphere. Microbiome 2019, 7, 55. [Google Scholar] [CrossRef]
- Xiao, M.; Li, M.; Reynolds, C. Colony formation in the cyanobacterium Microcystis. Biol Rev. 2018, 93, 1399–1420. [Google Scholar] [CrossRef]
- Yang, C.; Lin, F.; Li, Q.; Li, T.; Zhao, J. Comparative genomics reveals diversified CRISPR-Cas systems of globally distributed Microcystis aeruginosa, a freshwater bloom-forming cyanobacterium. Front Microbiol. 2015, 6, 394. [Google Scholar] [CrossRef]
- Jackrel, S.L.; White, J.D.; Evans, J.T.; et al. Genome evolution and host-microbiome shifts correspond with intraspecific niche divergence within harmful algal bloom-forming Microcystis aeruginosa. Mol Biol Evol. 2019, 28, 3994–4011. [Google Scholar] [CrossRef]
- Swan, B.K.; Tupper, B.; Sczyrba, A.; et al. Prevalent genome streamlining and latitudinal divergence of planktonic bacteria in the surface ocean. Proc Natl Acad Sci U S A 2013, 110, 11463–11468. [Google Scholar] [CrossRef]
- Parks, D.H.; Imelfort, M.; Skennerton, C.T.; Hugenholtz, P.; Tyson, G.W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015, 25, 1043–1055. [Google Scholar] [CrossRef]
- Bowers, R.; Kyrpides, N.; Stepanauskas, R.; et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat Biotechnol. 2017, 35, 725–731. [Google Scholar] [CrossRef] [PubMed]
- Bobay, L.; Ellis, B.; Ochman, H. ConSpeciFix: classifying prokaryotic species based on gene flow. Bioinformatics 2018, 34, 3738–3740. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Chan Patricia, P.; Lin Brian, Y.; Mak Allysia, J.; Lowe Todd, M. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 2021, 49, 9077–9096. [Google Scholar] [CrossRef]
- Ontiveros-Palacios, N.; Cooke, E.; Nawrocki Eric, P.; et al. Rfam 15: RNA families database in 2025. Nucleic Acids Res.;Nucleic Acids Research 2024, 53(D1), D258–D67. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional Annotation, Orthology Assignments, and Domain Prediction at the Metagenomic Scale. Mol Biol Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef] [PubMed]
- Machado, D.; Andrejev, S.; Tramontano, M.; Patil, K.R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Research 2018, 46, 7542–7553. [Google Scholar] [CrossRef] [PubMed]
- Ebrahim, A.; Lerman, J.A.; Palsson, B.O.; Hyduke, D.R. COBRApy: COnstraints-Based Reconstruction and Analysis for Python. BMC Syst Biol. 2013, 7, 74. [Google Scholar] [CrossRef]
- Emms, D.; Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef]
- Tettelin, H.; Riley, D.; Cattuto, C.; Medini, D. Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol. 2008, 11, 472–477. [Google Scholar] [CrossRef]
- Vernikos, G.S.; Parkhill, J. Interpolated variable order motifs for identification of horizontally acquired DNA: revisiting the Salmonella pathogenicity islands. Bioinformatics 2006, 22, 2196–2203. [Google Scholar] [CrossRef]
- Xie, Z.; Tang, H. ISEScan: automated identification of insertion sequence elements in prokaryotic genomes. Bioinformatics 2017, 33, 3340–3347. [Google Scholar] [CrossRef]
- Akhter, S.; Aziz, R.K.; Edwards, R.A. PhiSpy: a novel algorithm for finding prophages in bacterial genomes that combines similarity- and composition-based strategies. Nucleic Acids Res. 2012, 40, e126. e126. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Roberts, R.J.; Vincze, T.; Posfai, J.; Macelis, D. REBASE: a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Res. 2022, 51(D1), D629–D30. [Google Scholar] [CrossRef]
- Couvin, D.; Bernheim, A.; Toffano-Nioche, C.; et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 2018, 46(W1), W246–W51. [Google Scholar] [CrossRef] [PubMed]
- Nies, F.; Mielke, M.; Pochert, J.; Lamparter, T. Natural transformation of the filamentous cyanobacterium Phormidium lacuna. PLoS One 2020, 15, e0234440. [Google Scholar] [CrossRef]
- Averhoff, B.; Kirchner, L.; Pfefferle, K.; Yaman, D. Natural transformation in Gram-negative bacteria thriving in extreme environments: from genes and genomes to proteins, structures and regulation. Extremophiles 2021, 25(5-6), 425–36. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genomics Proteomics Bioinformatics 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Chen, H.; Zwaenepoel, A. Inference of Ancient Polyploidy from Genomic Data. In Polyploidy: Methods and Protocols; Van de Peer, Y., Ed.; Springer US, 2023; pp. 3–18. [Google Scholar]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res.;Nucleic Acids Research 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef]
- Perez-Carrascal, O.M.; Terrat, Y.; Giani, A.; et al. Coherence of Microcystis species revealed through population genomics. ISME J. 2019, 13, 2887–2900. [Google Scholar] [CrossRef]
- Kim, S.; Cho, C.-S.; Han, K.; Lee, J. Structural variation of AluElement and human disease. Genomics & informatics 2016, 14, 70–77. [Google Scholar]
- White, M.; Allers, T. DNA repair in the archaea-an emerging picture. FEMS Microbiol Rev. 2018, 42, 514–526. [Google Scholar] [CrossRef]
- Braus, S.A.G.; Short, F.L.; Holz, S.; Stedman, M.J.M.; Gossert, A.D.; Hospenthal, M.K. The molecular basis of FimT-mediated DNA uptake during bacterial natural transformation. Nat Commun. 2022, 13, 1065. [Google Scholar] [CrossRef] [PubMed]
- Hélaine, S.; Carbonnelle, E.; Prouvensier, L.; Beretti, J.; Nassif, X.; Pelicic, V. PilX, a pilus-associated protein essential for bacterial aggregation, is a key to pilus-facilitated attachment of Neisseria meningitidis to human cells. Mol Microbiol. 2005, 55, 65–77. [Google Scholar] [CrossRef] [PubMed]
- Sontheimer, E.; Davidson, A. Inhibition of CRISPR-Cas systems by mobile genetic elements. Curr Opin Microbiol. 2017, 37, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Middelboe, M.; Traving, S.; Castillo, D.; Kalatzis, P.; Glud, R. Prophage-encoded chitinase gene supports growth of its bacterial host isolated from deep-sea sediments. ISME J. 2025, 19, wraf004. [Google Scholar] [CrossRef]
- Koonin, E.; Makarova, K. Origins and evolution of CRISPR-Cas systems. Phil Trans R Soc B 2019, 374, 20180087. [Google Scholar] [CrossRef]
- Chen, T.; Xiong, Y.; Zhang, J.; et al. Temporal dynamics, microdiversity, and ecological functions of viral communities during cyanobacterial blooms in Lake Taihu. NPJ Biofilms Microbiomes 2025, 11, 178. [Google Scholar] [CrossRef]
- Trisolini, L.; Gambacorta, N.; Gorgoglione, R.; et al. FAD/NADH Dependent Oxidoreductases: From Different Amino Acid Sequences to Similar Protein Shapes for Playing an Ancient Function. Journal of Clinical Medicine 2019, 8, 2117. [Google Scholar] [CrossRef]
- Peng, W.; Xu, Y.; Yin, Y.; et al. Biological characteristics of manganese transporter MntP in Klebsiella pneumoniae. mSphere 2024, 9. [Google Scholar] [CrossRef]
- Eisenhut, M. Manganese Homeostasis in Cyanobacteria. Plants-BASEL 2020, 9, 18. [Google Scholar] [CrossRef]
- Bosma, E.; Rau, M.; van Gijtenbeek, L.; Siedler, S. Regulation and distinct physiological roles of manganese in bacteria. FEMS Microbiol Rev. 2021, 45, fuab028. [Google Scholar] [CrossRef]
- Matsuoka, Y.; Li, X.; Bennet, V. Adducin: structure, function and regulation. Cell Mol Life Sci. 2000, 57, 884–895. [Google Scholar] [CrossRef] [PubMed]
- Held, T.; Klemmer, D.; Lässig, M. Survival of the simplest in microbial evolution. Nat Commun. 2019, 10, 2472. [Google Scholar] [CrossRef] [PubMed]
- Hammer, P.E.; Hill, D.S.; Lam, S.T.; Van Pée, K.H.; Ligon, J.M. Four genes from Pseudomonas fluorescens that encode the biosynthesis of pyrrolnitrin. Appl Environ Microbiol. 1997, 63, 2147–2154. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Liu, H.C.; Zhou, Y.G.; Xin, Y.H. Microevolution and Adaptive Strategy of Psychrophilic Species Flavobacterium bomense sp. nov. Isolated From Glaciers. Front Microbiol. 2019, 10, 1069. [Google Scholar] [CrossRef]
- Dong, H.; Nilsson, L.; Kurland, C.G. Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J Mol Biol. 1996, 260, 649–663. [Google Scholar] [CrossRef]
- Weissman, J.L.; Hou, S.; Fuhrman, J.A. Estimating maximal microbial growth rates from cultures, metagenomes, and single cells via codon usage patterns. PNAS 2021, 118, e2016810118. [Google Scholar] [CrossRef]
- Chen, M.; Cui, R.; Hong, S.; et al. Broad-spectrum tolerance to disinfectant- mediated bacterial killing due to mutation of the PheS aminoacyl tRNA synthetase. PNAS 2025, 122, e2412871122. [Google Scholar] [CrossRef] [PubMed]
- Wagner, A. Adaptive evolvability through direct selection instead of indirect, second-order selection. Journal of Experimental Zoology Part B-Molecular and Developmental Evolution 2022, 338, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.J.; Lenski, R.E.; Zinser, E.R. The Black Queen Hypothesis: Evolution of Dependencies through Adaptive Gene Loss. Article. mBio 2012, 3, e00036-12. [Google Scholar] [CrossRef]
- Dick, G.J.; Duhaime, M.B.; Evans, J.T.; et al. The genetic and ecophysiological diversity of Microcystis. Environ Microbiol. 2021, 23, 7278–7313. [Google Scholar] [CrossRef]
- Chan, C.; Beiko, R.; Ragan, M. Lateral transfer of genes and gene fragments in Staphylococcus extends beyond mobile elements. J Bacteriol. 2011, 193, 3964–3977. [Google Scholar] [CrossRef]
- Koksharova, O.; Popova, A.; Plyuta, V.; Khmel, I. Four new genes of cyanobacterium Synechococcus elongatus PCC 7942 are responsible for sensitivity to 2-Nonanone. Microorganisms 2020, 8, 1234. [Google Scholar] [CrossRef]
- Monika, S.; Malgorzata, B.; Zbigniew, O. Contribution of Aspartic Proteases in Candida Virulence. Protease Inhibitors against Candida Infections. Review. Curr Protein Pept Sci. 2017, 18, 1050–1062. [Google Scholar] [CrossRef]
- Sharp, P.M.; Shields, D.C.; Wolfe, K.H.; Li, W.H. Chromosomal location and evolutionary rate variation in enterobacterial genes. Science 1989, 246, 808–810. [Google Scholar] [CrossRef]
- Rubin, I.N.; Ispolatov, Y.; Doebeli, M. Maximal ecological diversity exceeds evolutionary diversity in model ecosystems. Ecol Lett. 2023, 26, 384–397. [Google Scholar] [CrossRef]
- Fisher, K.A.; Yarwood, S.A.; James, B.R. Soil urease activity and bacterial ureC gene copy numbers: Effect of pH. Geoderma 2017, 285, 1–8. [Google Scholar] [CrossRef]
- Khandelwal, A.; Patel, A.; Tiwari, S.; Prasad, S.M. Tryptamine: a novel signaling molecule alleviating salt-induced toxicity by enhancing antioxidant defense and PSII photochemistry in Anabaena PCC7120. Arch Microbiol. 2025, 208, 64. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



