Submitted:
20 April 2026
Posted:
22 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
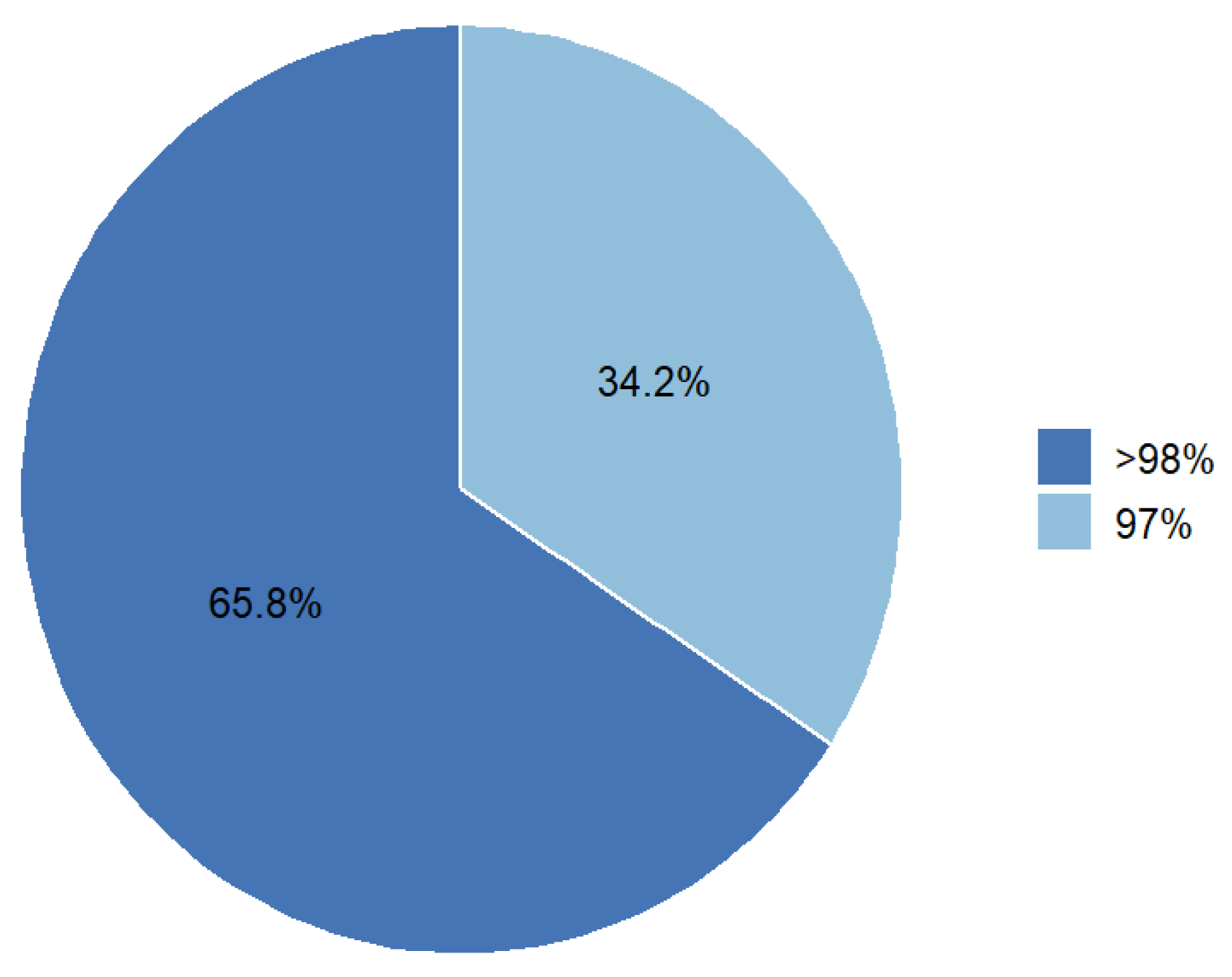
2.1. SNP Markers Distribution and Quality
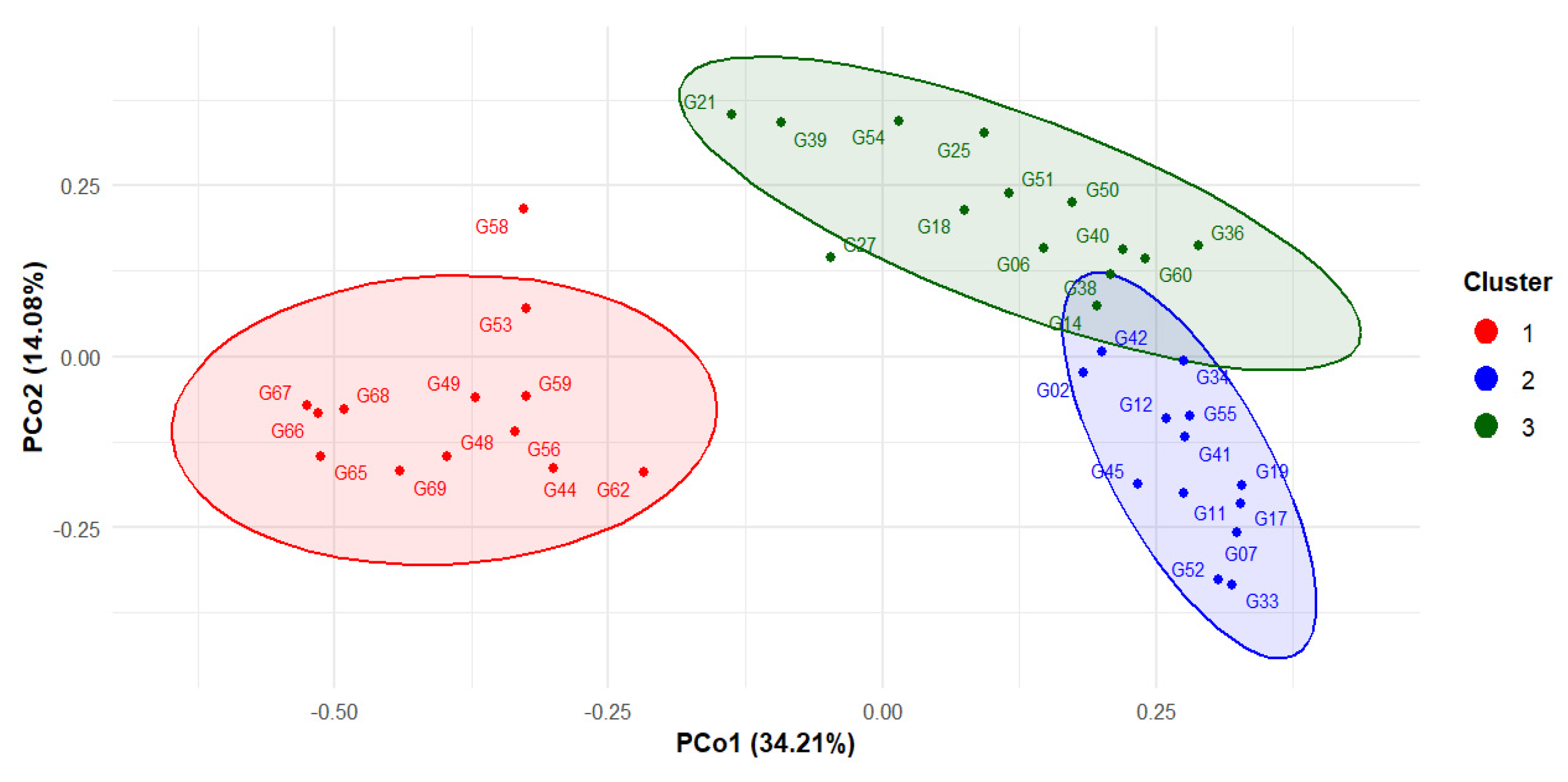
2.2. Genetic Diversity Parameters and Visualization
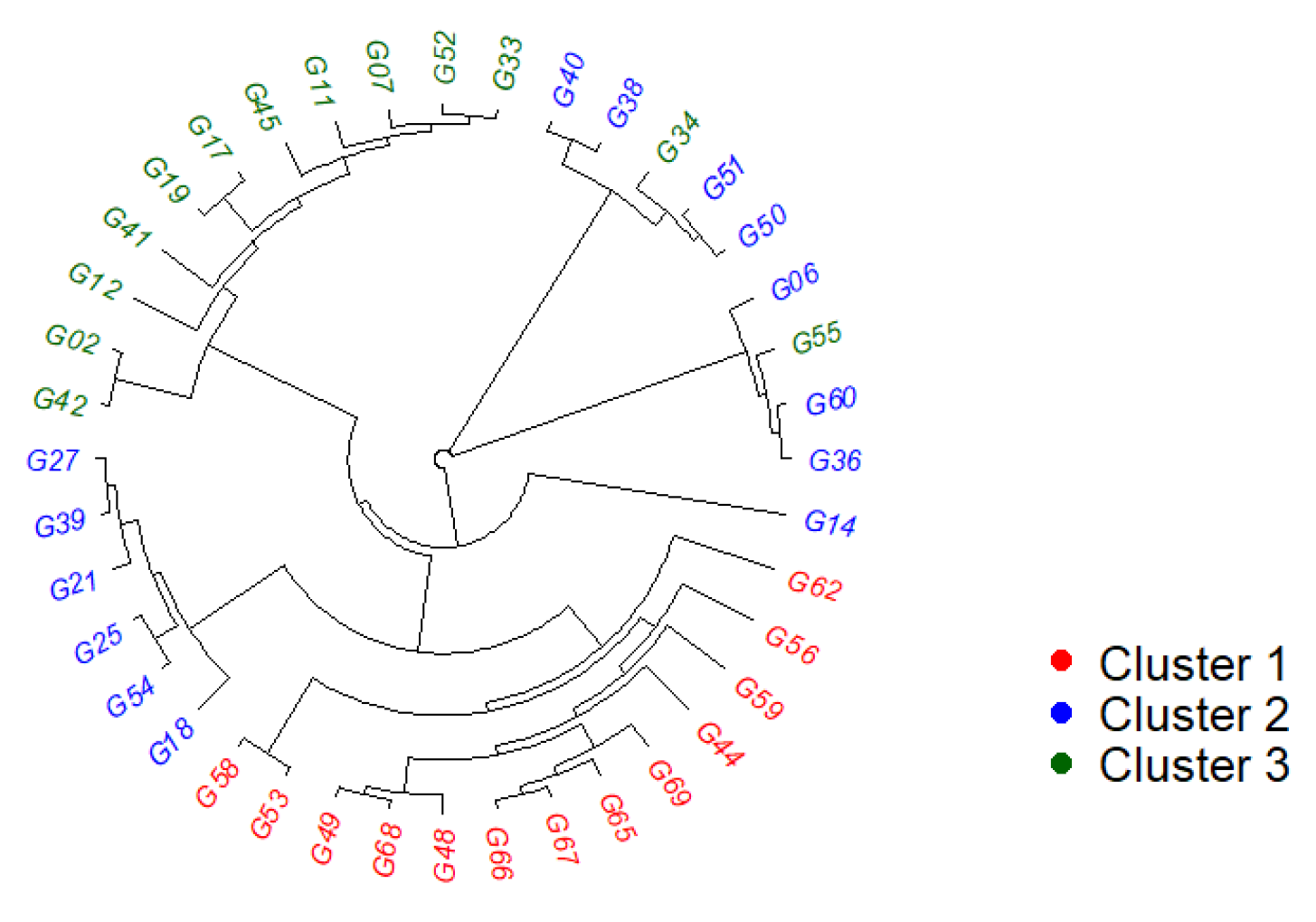
- Principal coordinate analysis (PCoA) and Neighbor-joining (NJ) tree
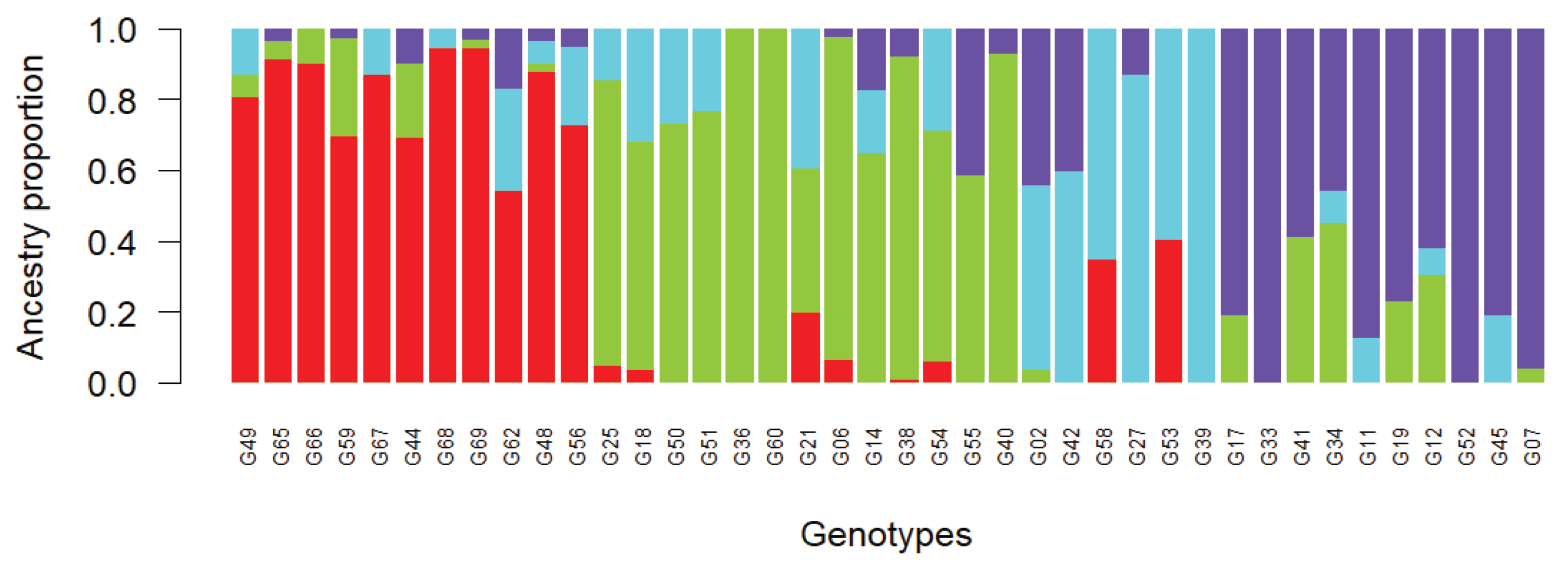
2.3. Population Structure Analysis
- Admixture
- Population genetic diversity parameters
- Pairwise fixation index (FST) and Nei’s genetic distance
- Analysis of molecular variance
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Genotyping
- Sample Preparation and DNA Extraction
- Library preparation and sequencing
- SNP calling and alignment to rice reference genome
- SNP filtering and quality control
4.3. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| SNP | Single Nucleotide Polymorphism |
| AMOVA | Analysis of Molecular Variance |
| Mb | Megabase |
References
- Zilberman, D.; Goetz, R.; Garrido, A.; Otsuka, K.; Mano, Y.; Takahashi, K. Natural Resource Management and Policy Series Editors: Rice Green Revolution in Sub-Saharan Africa.; Otsuka, K., Mano, Y., Takahashi, K., Eds.; Springer Nature: Singapore, 2023; Vol. 56, ISBN 978-981-19-8045-9. [Google Scholar] [CrossRef]
- Traoré, F.; Debucquet, D.L.; Del Prete, D.; Sánchez, M.; Diop, I. The Rice Value Chain in Africa . In Africa Agriculture Trade Monitor 2025; Chapter 3; Odjo, Sunday, Traore, Fousseini, Zaki, Chahir, Eds.; Kigali, 2025; pp. Pp. 85–118. [Google Scholar] [CrossRef]
- Arouna, A.; Fatognon, I.A.; Saito, K.; Futakuchi, K. Moving toward Rice Self-Sufficiency in Sub-Saharan Africa by 2030: Lessons Learned from 10 Years of the Coalition for African Rice Development. World Dev. Perspect. 2021, 21. [Google Scholar] [CrossRef]
- FAOSTAT Statistical Database. Food and Agriculture Organization of the United Nations, Rome. Available online: https://www.fao.org/faostat/en/#data/QCL (accessed on 10 January 2026).
- Adjah, K.L.; Asante, M.D.; Toure, A.; Aziadekey, M.; Amoako-Andoh, F.O.; Frei, M.; Diallo, Y.; Agboka, K. Improvement of Rice Production under Drought Conditions in West Africa: Application of QTLs in Breeding for Drought Resistance. Rice Sci. 2022, 29, 512–521. [Google Scholar] [CrossRef]
- Ismael, F.; Ndayiragije, A.; Fangueiro, D. New Fertilizer Strategies Combining Manure and Urea for Improved Rice Growth in Mozambique. 2021. [Google Scholar] [CrossRef]
- Kajisa, K.; Vu, T.T. The Importance of Farm Management Training for the African Rice Green Revolution: Experimental Evidence from Rainfed Lowland Areas in Mozambique. Food Policy 2023, 114. [Google Scholar] [CrossRef]
- Saito, K.; Senthilkumar, K.; Dossou-Yovo, E.R.; Ali, I.; Johnson, J.M.; Mujawamariya, G.; Rodenburg, J. Status Quo and Challenges of Rice Production in Sub-Saharan Africa. Plant Prod. Sci. 2023, 26, 320–333. [Google Scholar] [CrossRef]
- Kodama, W.; Pede, V.O.; Mishra, A.K.; Cuevas, R.P.O.; Ndayiragije, A.; Cabrera, E.R.; Langa, M.; Ali, J. Assessing the Benefits of Green Super Rice in Sub-Saharan Africa: Evidence from Mozambique. Q Open 2022, 2. [Google Scholar] [CrossRef]
- Kehinde, B.O.; Xie, L.; Song, B.-K.; Zheng, X.; Fan, L. African Cultivated, Wild and Weedy Rice (Oryza Spp.): Anticipating Further Genomic Studies. Biology (Basel). 2024, 13, 697. [Google Scholar] [CrossRef]
- Wambugu, P.W.; Furtado, A.; Le Waters, D.; Nyamongo, D.O.; Henry, R.J. Conservation and Utilization of African Oryza Genetic Resources. Rice 2013, 6, 1–13. [Google Scholar] [CrossRef]
- Fan, J.; Venuprasad, R.; Xia, S.; Yang, Z.; Zheng, X.; Chen, F. Strengthening Global Rice Germplasm Sharing: Insights from the INGER Platform . 2025. [Google Scholar] [CrossRef] [PubMed]
- IRRI International Rice Research Institute (IRRI). AfricaRice Collaborate on Developing and Delivering Improved and Localized Rice Varieties to Smallholder Farmers in Africa. Available online: https://www.irri.org/news-and-events/news/irri-and-africarice-collaborate-developing-and-delivering-improved-and? (accessed on 18 March 2026).
- Nangombe, S.; Macharia, M.; Solemanegy, M.; Caproni, L.; Takele, R.; Munisse, P.; Amane, M.; Buizza, R. Report on the Characterization of Climate-Ready Rice and Cowpea Varieties . In Research and Innovation Action; CSIR, 2023; p. 66 pages. [Google Scholar]
- APPSA Facilitates Collection and Characterization of Rice Landraces as Genetic Resources for Enhancing Resilience to Climate Change and Biotic Stresses in Mozambique. Available online: https://www.ccardesa.org/appsa-facilitates-collection-and-characterization-rice-landraces-genetic-resources-enhancing? (accessed on 18 March 2026).
- Mropes, S.K.M.N.; Moiana, L.D.; Alberto, L.A.; Andreque, J.M.; Abade, H. Morphological Characterization of 18 Varieties of Local Rice in Mozambique. Aust. J. Crop Sci. 2026, 20, 188–197. [Google Scholar] [CrossRef]
- Warioba, K.G.; Macandza, C.M.; Moiana, L.D. Screening Rice (Oryza Sativa L.) Genotypes for Seedling-Stage Drought Tolerance. Stresses 2026, 6, 13. [Google Scholar] [CrossRef]
- Bidyananda, N.; Jamir, I.; Nowakowska, K.; Varte, V.; Vendrame, W.A.; Devi, R.S.; Nongdam, P. Plant Genetic Diversity Studies: Insights from DNA Marker Analyses. International Journal of Plant Biology 2024, 15, 607–640. [Google Scholar] [CrossRef]
- Sarif, H.M.; Rafii, M.Y.; Ramli, A.; Oladosu, Y.; Musa, H.M.; Rahim, H.A.; Zuki, Z.M.; Chukwu, S.C. Genetic Diversity and Variability among Pigmented Rice Germplasm Using Molecular Marker and Morphological Traits. Biotechnology and Biotechnological Equipment 2020, 34, 747–762. [Google Scholar] [CrossRef]
- Yu, R.; Liu, J.; Niu, Y.; Han, X.; Wang, X.; Yang, Y. A Simple and Flexible Approach for Detecting Small Numbers of SNPs. Front. Plant Sci. 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Berdugo-Cely, J.A.; Pérez-Pazos, J. V.; Perez-Cantero, S.P.; Morales-Angulo, J.G.; Romero-Ferrer, J.L. Genetic Diversity and Population Structure of Regional Rice Genotypes from Colombia’s Caribbean and Pacific Regions: Differentiation and Ancestry in Relation to the 3000 Rice Genomes Project. Genet. Resour. Crop Evol. 2025. [Google Scholar] [CrossRef]
- Raza, Q.; Riaz, A.; Saher, H.; Bibi, A.; Raza, M.A.; Ali, S.S.; Sabar, M. Grain Fe and Zn Contents Linked SSR Markers Based Genetic Diversity in Rice. PLoS One 2020, 15. [Google Scholar] [CrossRef] [PubMed]
- Gouda, A.C.; Sangare, J.R.; Gnikoua, K.; Wambugu, P.; Huggins, T.D.; Ndjiondjop, M.N. Genetic Variation and Population Structure of the Rice Accessions Maintained in the AfricaRice Genebank Using DArTseq. Crop Sci. 2024. [Google Scholar] [CrossRef]
- Ndjiondjop, M.N.; Gouda, A.C.; Eizenga, G.C.; Warburton, M.L.; Kpeki, S.B.; Wambugu, P.W.; Gnikoua, K.; Tia, D.D.; Bachabi, F. Genetic Variation and Population Structure of Oryza Sativa Accessions in the AfricaRice Collection and Development of the AfricaRice O. Sativa Core Collection. Crop Sci. 2023, 63, 724–739. [Google Scholar] [CrossRef]
- Ndjiondjop, M.N.; Semagn, K.; Sow, M.; Manneh, B.; Gouda, A.C.; Kpeki, S.B.; Pegalepo, E.; Wambugu, P.; Sié, M.; Warburton, M.L. Assessment of Genetic Variation and Population Structure of Diverse Rice Genotypes Adapted to Lowland and Upland Ecologies in Africa Using SNPs. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Kimwemwe, P.K.; Bukomarhe, C.B.; Mamati, E.G.; Githiri, S.M.; Civava, R.M.; Mignouna, J.; Kimani, W.; Fofana, M. Population Structure and Genetic Diversity of Rice (Oryza Sativa L.) Germplasm from the Democratic Republic of Congo (DRC) Using DArTseq-Derived Single Nucleotide Polymorphism (SNP). Agronomy 2023, 13. [Google Scholar] [CrossRef]
- Serrote, C.M.L.; Reiniger, L.R.S.; Silva, K.B.; Rabaiolli, S.M. dos S.; Stefanel, C.M. Determining the Polymorphism Information Content of a Molecular Marker. Gene 2020, 726. [Google Scholar] [CrossRef]
- Razak, S.A.; Azman, N.H.E.N.; Kamaruzaman, R.; Saidon, S.A.; Yusof, M.F.M.; Ismail, S.N.; Jaafar, M.A.; Abdullah, N. Genetic Diversity of Released Malaysian Rice Varieties Based on Single Nucleotide Polymorphism Markers. Czech Journal of Genetics and Plant Breeding 2020, 56, 62–70. [Google Scholar] [CrossRef]
- Hodgins, K.A.; Yeaman, S. Mating System Impacts the Genetic Architecture of Adaptation to Heterogeneous Environments. New Phytologist 2019, 224, 1201–1214. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.Y.; Purugganan, M.D. Multiple Origin but Single Domestication Led to Oryza Sativa. G3: Genes, Genomes, Genetics 2018, 8, 797–803. [Google Scholar] [CrossRef]
- Choudhury, D.R.; Kumar, R.; Vimala Devi, S.; Singh, K.; Singh, N.K.; Singh, R. Identification of a Diverse Core Set Panel of Rice From the East Coast Region of India Using SNP Markers. Front. Genet. 2021, 12. [Google Scholar] [CrossRef]
- Mvuyekure, S.M.; Sibiya, J.; Derera, J.; Nzungize, J.; Nkima, G. Assessment of Genetic Diversity of Rice Based on SNP Markers for Selection of Parents for Sheath Rot (Sarocladium Oryzae) Resistance Breeding. South African Journal of Plant and Soil 2018, 35, 51–59. [Google Scholar] [CrossRef]
- Baksh, S.K.Y.; Donde, R.; Kumar, J.; Mukherjee, M.; Meher, J.; Behera, • Lambodar; Sushanta, •; Dash, K. Genetic Relationship, Population Structure Analysis and Pheno-Molecular Characterization of Rice (Oryza Sativa L.) Cultivars for Bacterial Leaf Blight Resistance and Submergence Tolerance Using Trait Specific STS Markers. Physiology and Molecular Biology of Plants 2021, 27, 543–562. [Google Scholar] [CrossRef] [PubMed]
- Aesomnuk, W.; Ruengphayak, S.; Ruanjaichon, V.; Sreewongchai, T.; Malumpong, C.; Vanavichit, A.; Toojinda, T.; Wanchana, S.; Arikit, S. Estimation of the Genetic Diversity and Population Structure of Thailand’s Rice Landraces Using Snp Markers. Agronomy 2021, 11. [Google Scholar] [CrossRef]
- Skotte, L.; Korneliussen, T.S.; Albrechtsen, A. Estimating Individual Admixture Proportions from next Generation Sequencing Data. Genetics 2013, 195, 693–702. [Google Scholar] [CrossRef]
- Ghazy, M.I.; EL-Naem, S.A.; Hefeina, A.G.; Sallam, A.; Eltaher, S. Genome-Wide Association Study of Rice Diversity Panel Reveals New QTLs for Tolerance to Water Deficit Under the Egyptian Conditions. Rice 2024, 17. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, D.R.; Kumar, R.; Maurya, A.; Semwal, D.P.; Rathi, R.S.; Gautam, R.K.; Trivedi, A.K.; Bishnoi, S.K.; Ahlawat, S.P.; Singh, K.; et al. SSR and SNP Marker-Based Investigation of Indian Rice Landraces in Relation to Their Genetic Diversity, Population Structure, and Geographical Isolation. Agriculture (Switzerland) 2023, 13. [Google Scholar] [CrossRef]
- Kumar, K.P.; Pushpam, R.; Manonmani, S.; Raveendran, M.; Santhiya, S.; Senthil, A. Enhancing Stress Resilience in Rice (Oryza Sativa L.) through Profiling Early-Stage Morpho-Physiological and Molecular Responses to Multiple Abiotic Stress Tolerance. Front. Plant Sci. 2024, 15. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Peng, J.; Lv, Q.; Ding, J.; Chen, S.; Duan, M.; He, Q.; Wu, J.; Tian, Y.; Yu, D.; et al. Dissecting the Genetic Basis of Heterosis in Elite Super-Hybrid Rice. Plant Physiol. 2023, 192, 307–325. [Google Scholar] [CrossRef] [PubMed]
- McCouch, S.R.; Wright, M.H.; Tung, C.W.; Maron, L.G.; McNally, K.L.; Fitzgerald, M.; Singh, N.; DeClerck, G.; Agosto-Perez, F.; Korniliev, P.; et al. Open Access Resources for Genome-Wide Association Mapping in Rice. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef]
- Gouda, A.C.; Ndjiondjop, M.N.; Djedatin, G.L.; Warburton, M.L.; Goungoulou, A.; Kpeki, S.B.; N’Diaye, A.; Semagn, K. Comparisons of Sampling Methods for Assessing Intra- and Inter-Accession Genetic Diversity in Three Rice Species Using Genotyping by Sequencing. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef]
- Fumagalli, M. Assessing the Effect of Sequencing Depth and Sample Size in Population Genetics Inferences. PLoS One 2013, 8. [Google Scholar] [CrossRef]
- Aguirre-Liguori, J.A.; Luna-Sánchez, J.A.; Gasca-Pineda, J.; Eguiarte, L.E. Evaluation of the Minimum Sampling Design for Population Genomic and Microsatellite Studies: An Analysis Based on Wild Maize. Front. Genet. 2020, 11. [Google Scholar] [CrossRef]
- Wright, S. Evolution and the Genetics of Populations: Variability within and among Natural Populations; University of Chicago Press, 1978; Vol. 4. [Google Scholar]
- Thant, A.A.; Zaw, H.; Kalousova, M.; Singh, R.K.; Lojka, B. Genetic Diversity and Population Structure of Myanmar Rice (Oryza Sativa L.) Varieties Using DArTseq-Based SNP and SilicoDArT Markers. Plants 2021, 10. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.K.; Kumar, D.; Gemmati, D.; Ellur, R.K.; Singh, A.; Tisato, V.; Dwivedi, D.K.; Singh, S.K.; Kumar, K.; Khan, N.A.; et al. Investigating Genetic Diversity and Population Structure in Rice Breeding from Association Mapping of 116 Accessions Using 64 Polymorphic SSR Markers. Crops 2024, 4, 180–194. [Google Scholar] [CrossRef]
- Suvi, W.T.; Shimelis, H.; Laing, M.; Mathew, I.; Shayanowako, A.I.T. Assessment of the Genetic Diversity and Population Structure of Rice Genotypes Using SSR Markers. Acta Agric. Scand. B Soil Plant Sci. 2020, 70, 76–86. [Google Scholar] [CrossRef]
- Hartfield, M.; Bataillon, T.; Glémin, S. The Evolutionary Interplay between Adaptation and Self-Fertilization. Trends in Genetics 2017, 33, 420–431. [Google Scholar] [CrossRef] [PubMed]
- Anwar, A.; Tabassum, J.; Ahmad, S.; Ashfaq, M.; Hussain, A.; Ullah, M.A.; Saad, N.S.B.M.; Ghazy, A.I.; Javed, M.A. Screening and Assessment of Genetic Diversity of Rice (Oryza Sativa L.) Germplasm in Response to Soil Salinity Stress at Germination Stage. Agronomy 2025, 15. [Google Scholar] [CrossRef]
- Kilian, A.; Wenzl, P.; Huttner, E.; Carling, J.; Xia, L.; Blois, H.; Caig; Hok, P.; Uszynski, G. Diversity Arrays Technology: A Generic Genome Profiling Technology on Open Platforms. In Data Production and Analysis in Population Genomics; Pompanon, F., Bonin, A., Eds.; 2012; Vol. 888, pp. 67–89. [Google Scholar] [CrossRef]
- Knaus, B.J.; Grünwald, N.J. VCFR: A Package to Manipulate and Visualize Variant Call Format Data in R. Mol. Ecol. Resour. 2017, 17, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Nei, M. Estimation of Average Heterozygosity and Genetic Distance from a Small Number of Individuals. Genetics 1978, 583–590. [Google Scholar] [CrossRef]
- Jombart, T.; Ahmed, I. Adegenet 1.3-1: New Tools for the Analysis of Genome-Wide SNP Data. Bioinformatics 2011. [Google Scholar] [CrossRef]
- Torgerson, W.S. Multidimensional Scaling: I. Theory and Method. Psychometrika 1952, 17, 401–419. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol. Biol. Evol. 1987. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Frichot, E.; Francois, O. LEA: Package for Landscape and Ecological Association Studies. Methods in Ecology and Evolution. 2015. Available online: https://membres-timc.imag.fr/Olivier.Francois/lea.html.
- Weir, B.S.; Cockerham, C.C. Estimating F -Statistics for the Analysis of Population Structure. Evolution (N. Y). 1984, 38, 1358–1370. [Google Scholar] [CrossRef]
- Nei, M. Molecular Evolutionary Genetics; Columbia University Press: New York; ISBN 0231063544, 1987. [Google Scholar]
- Goudet, J.; Jombart, T. Hierfstat: Estimation and Tests of Hierarchical F-Statistics . 2022. Available online: https://CRAN.R-project.org/package=hierfsta.
- Nei, M.; Li, W.H. Mathematical Model for Studying Genetic Variation in Terms of Restriction Endonucleases. Proceedings of the National Academy of Sciences 1979, 76, 5269–5273. [Google Scholar] [CrossRef] [PubMed]
- Paradis, E. Pegas: An R Package for Population Genetics with an Integrated–Modular Approach. Bioinformatics 2010, 26, 419–420. [Google Scholar] [CrossRef] [PubMed]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of Molecular Variance Inferred from Metric Distances among DNA Haplotypes: Application to Human Mitochondrial DNA Restriction Data. Genetics 1992, 131, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Kamvar, Z.N.; Brooks, J.C.; Grünwald, N.J. Novel R Tools for Analysis of Genome-Wide Population Genetic Data with Emphasis on Clonality. Front. Genet. 2015, 6. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing (Version 4.5.1). R Foundation for Statistical Computing, Vienna, Austria. Available online: https://www.R-project.org/ (accessed on 19 October 2025).
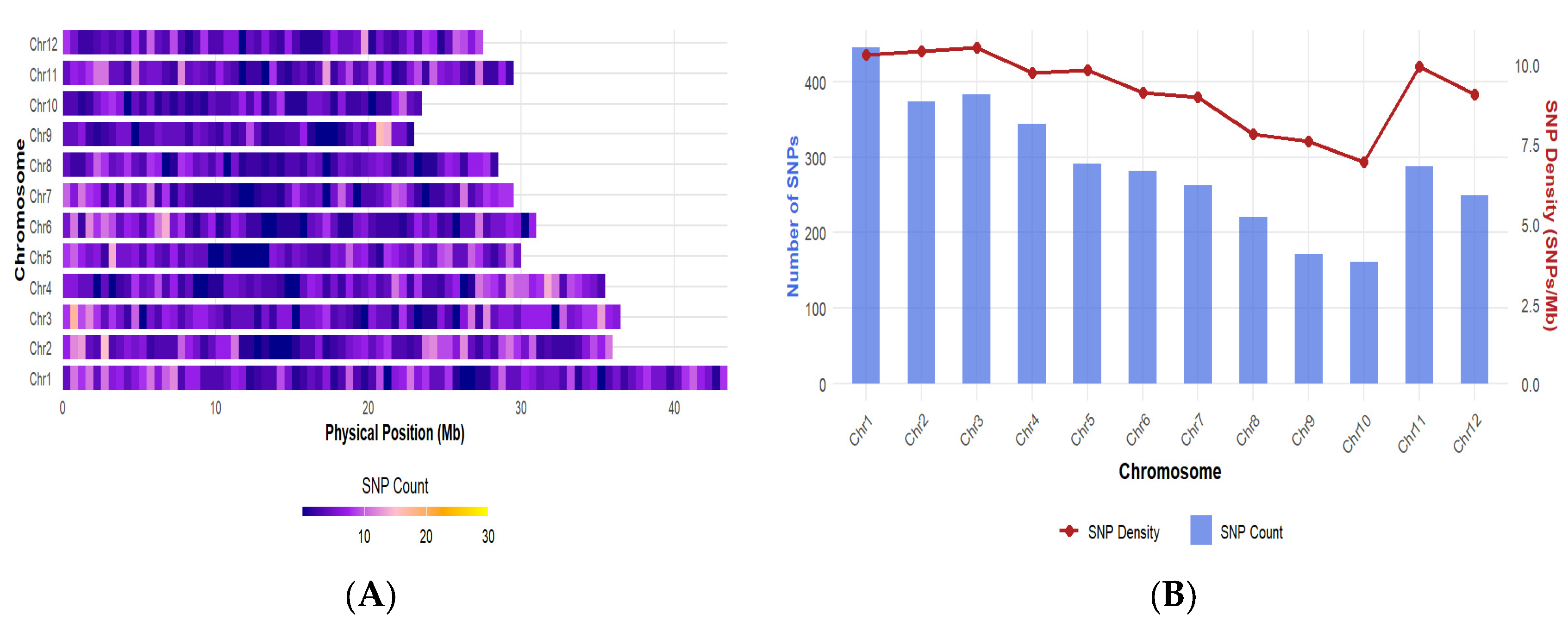
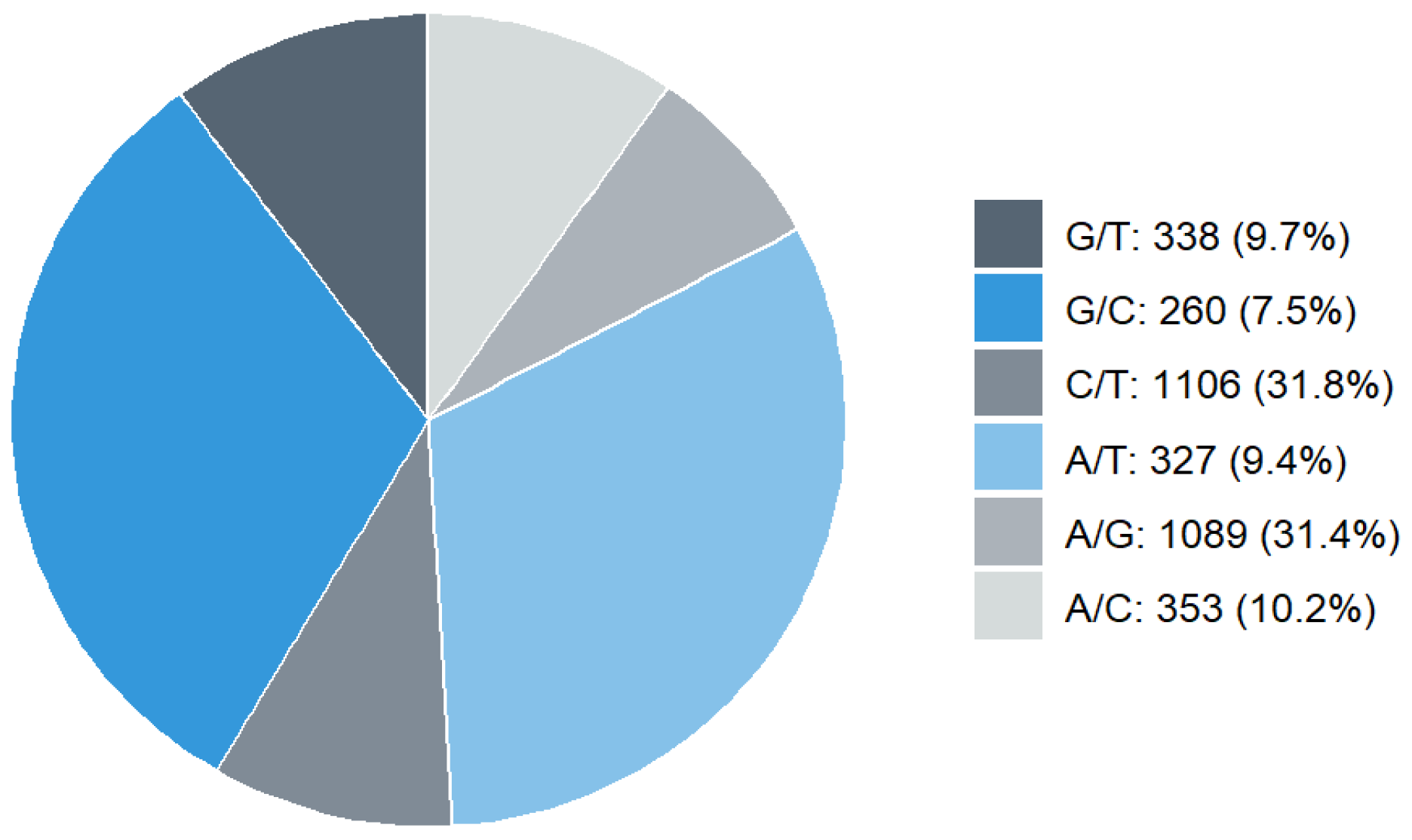
| Parameter | Minimum | Maximum | Mean ± SE |
| Observed heterozygosity (Ho) | 0.000 | 0.750 | 0.125 ± 0.001 |
| Unbiased expected heterozygosity (uHe) | 0.000 | 0.506 | 0.314 ± 0.003 |
| Polymorphic information content (PIC) | 0.000 | 0.375 | 0.250 ± 0.002 |
| Total number of SNP | 3473 | ||
| % of polymorphic loci | 98.04 | ||
| Parameter | n | Ho | uHe | PIC | %P | FIS | π |
| Subpop I | 10 | 0.120 ± 0.002 | 0.283 ± 0.003 | 0.161 ± 0.002 | 78.52 ± 0.70 | 0.528 ± 0.006 | 0.283 ± 0.003 |
| Subpop II | 11 | 0.081 ± 0.002 | 0.226 ± 0.004 | 0.130 ± 0.002 | 65.65 ± 0.81 | 0.551 ± 0.006 | 0.226 ± 0.004 |
| Subpop III | 3 | 0.320 ± 0.005 | 0.313 ± 0.004 | 0.156 ± 0.002 | 67.81 ± 0.79 | -0.064 ± 0.008 | 0.313 ± 0.004 |
| Subpop IV | 8 | 0.061 ± 0.002 | 0.132 ± 0.003 | 0.082 ± 0.002 | 45. 93 ± 0.85 | 0.414 ± 0.008 | 0.132 ± 0.003 |
| Admixed | 8 | 0.183 ± 0.003 | 0.309 ± 0.003 | 0.172 ± 0.002 | 81.57 ± 0.66 | 0.355 ± 0.006 | 0.309 ± 0.003 |
| Overall | 40 | 0.153 ± 0.003 | 0.253 ± 0.004 | 0.140 ± 0.002 | 67.90 ± 0.76 | 0.357 ± 0.007 | 0.253 ± 0.003 |
| Subpopulation | Nei’s genetic distance | |||||
| I | II | III | IV | Admixed | ||
| FST | I | 0 | 0.215 | 0.155 | 0.284 | 0.146 |
| II | 0.336 | 0 | 0.131 | 0.148 | 0.058 | |
| III | 0.170 | 0.184 | 0 | 0.229 | 0.094 | |
| IV | 0.458 | 0.347 | 0.443 | 0 | 0.077 | |
| Admixed | 0.200 | 0.077 | 0.056 | 0.163 | 0 | |
| Source | df | SS | MS | Est. var | % variation | Φ-statistics | P-value |
| Among subpopulations | 4 | 12208.30 | 3052.07 | 311.36 | 32.90 | 0.329 | 0.001 |
| Within subpopulations | 35 | 22228.81 | 635.11 | 635.11 | 67.10 | ||
| Total | 39 | 34437.10 | 883.00 | 946.47 | 100 |
| Entry No. | Genotype Name | Origin | Type | Notes |
| G02 | Fardamento | IIAM | Landrace | Rainfed lowland |
| G06 | Mpulo | IIAM | Landrace | Rainfed lowland |
| G07 | Mamima | IIAM | Landrace | Rainfed lowland |
| G11 | Mucabo | IIAM | Landrace | Rainfed lowland |
| G12 | Nhacungo | IIAM | Landrace | Rainfed lowland |
| G14 | Muindeia | IIAM | Landrace | Rainfed lowland |
| G17 | Paulo | IIAM | Landrace | Rainfed lowland |
| G18 | Chinchurica | IIAM | Landrace | Rainfed lowland |
| G19 | Ercidji | IIAM | Landrace | Rainfed lowland |
| G21 | Muluabo | IIAM | Landrace | Rainfed lowland |
| G25 | Sabuadigae | IIAM | Landrace | Rainfed lowland |
| G27 | Mucamba | IIAM | Landrace | Rainfed lowland |
| G33 | Nene | IIAM | Landrace | Rainfed lowland |
| G34 | Canduacafri | IIAM | Landrace | Rainfed lowland |
| G36 | Angelo | IIAM | Landrace | Rainfed lowland |
| G38 | Mucandra | IIAM | Landrace | Rainfed lowland |
| G39 | Nasoco | IIAM | Landrace | Rainfed lowland |
| G40 | Nasaia | IIAM | Landrace | Rainfed lowland |
| G41 | Mwenhe | IIAM | Landrace | Rainfed lowland |
| G42 | Mutanzania | IIAM | Landrace | Rainfed lowland |
| G44 | Mexoeira | IIAM | Landrace | Rainfed lowland |
| G45 | Bridhan P-14 | IIAM | Landrace | Rainfed lowland |
| G48 | Sinabibi | IIAM | Landrace | Rainfed lowland |
| G49 | Simao | IIAM | Landrace | Rainfed lowland |
| G50 | Namapupa | IIAM | Landrace | Rainfed lowland |
| G51 | Tacabina | IIAM | Landrace | Rainfed lowland |
| G52 | Chupa | IIAM | Landrace | Rainfed lowland |
| G53 | Agulha | IIAM | Landrace | Rainfed lowland |
| G54 | Carrungo | IIAM | Landrace | Rainfed lowland |
| G55 | Indamula | IIAM | Landrace | Rainfed lowland |
| G56 | Balachao | IIAM | Landrace | Rainfed lowland |
| G58 | Vitinho | IIAM | Landrace | Rainfed lowland |
| G59 | Aviao Branco | IIAM | Landrace | Rainfed lowland |
| G60 | Namurawani | IIAM | Landrace | Rainfed lowland |
| G62 | B1P15 | Africa rice | Line | Rainfed lowland |
| G65 | B1P02 | Africa rice | Line | Rainfed lowland |
| G66 | B1P11 | Africa rice | Line | Rainfed lowland |
| G67 | B1P01 | Africa rice | Line | Rainfed lowland |
| G68 | IRB1P21 | IRRI | Line | Rainfed lowland |
| G69 | IRB1P26 | IRRI | Line | Rainfed lowland |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




