Submitted:
20 April 2026
Posted:
21 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Population and Sample Collection
2.2. Clinical Data Collection and Symptom Questionnaires
2.3. Microbiome Analysis
2.3.1. DNA Extraction and 16S rRNA Sequencing
2.3.2. Bioinformatics
2.4. Statistical Analysis
3. Results
3.1. Participants Overview
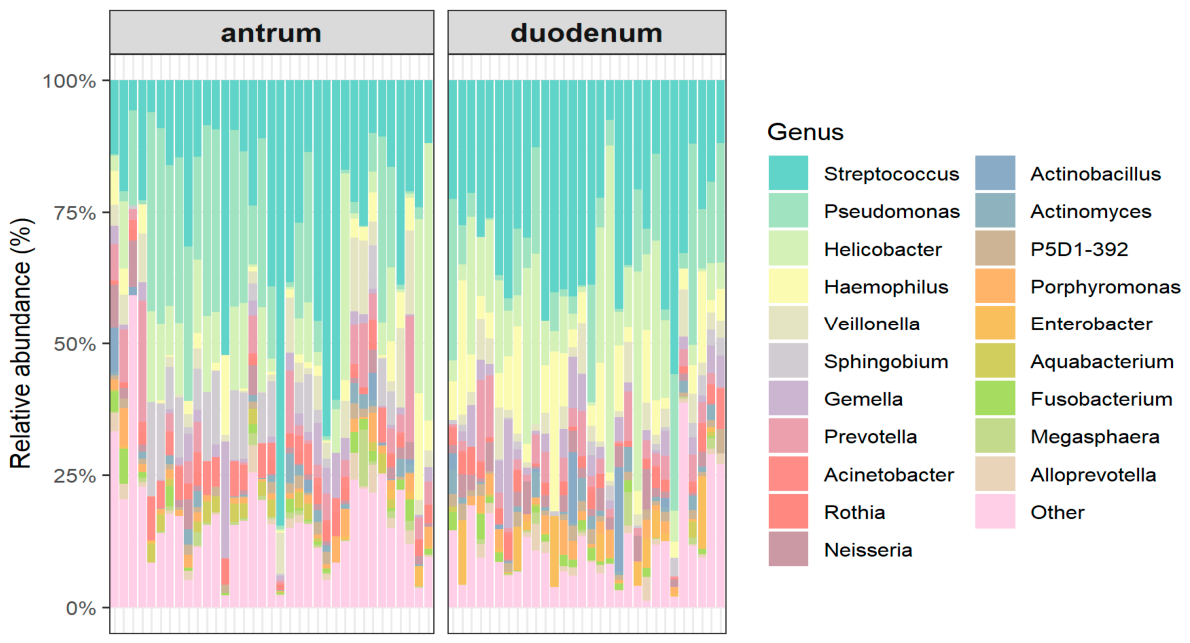
3.2. Site-Specific Variation in Gastric and Duodenal Microbial Communities
3.3. Microbial Differences across GERD Phenotypes
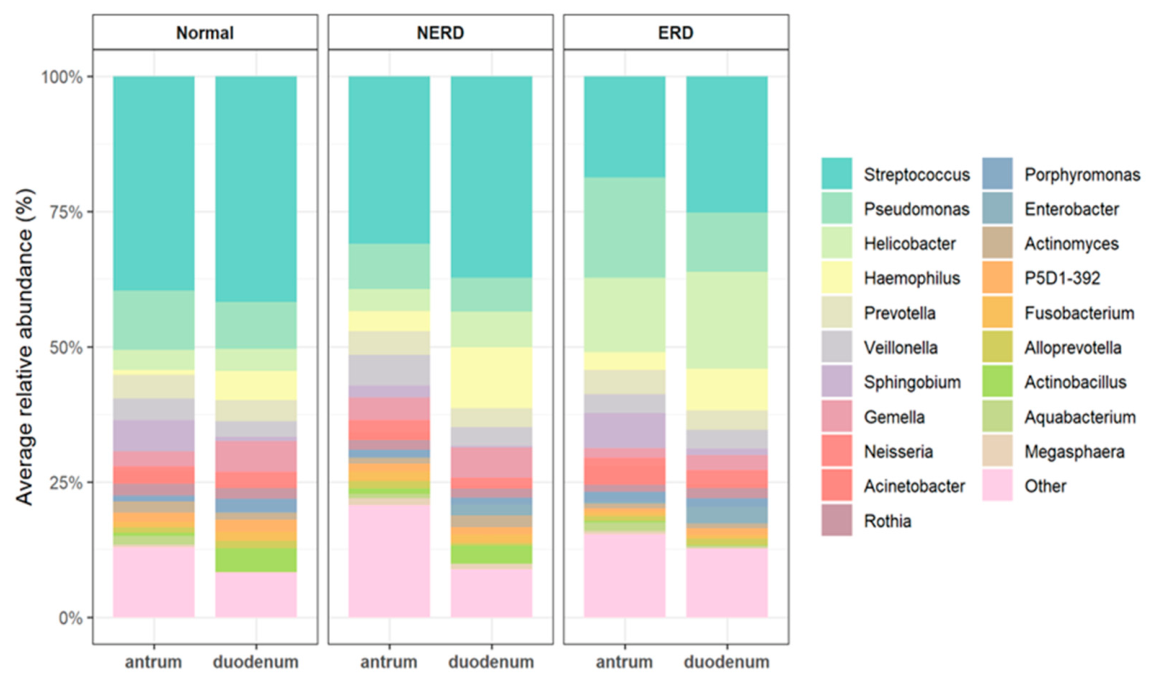
3.3.1. Microbial Composition across GERD Phenotypes in the Antrum and Duodenum
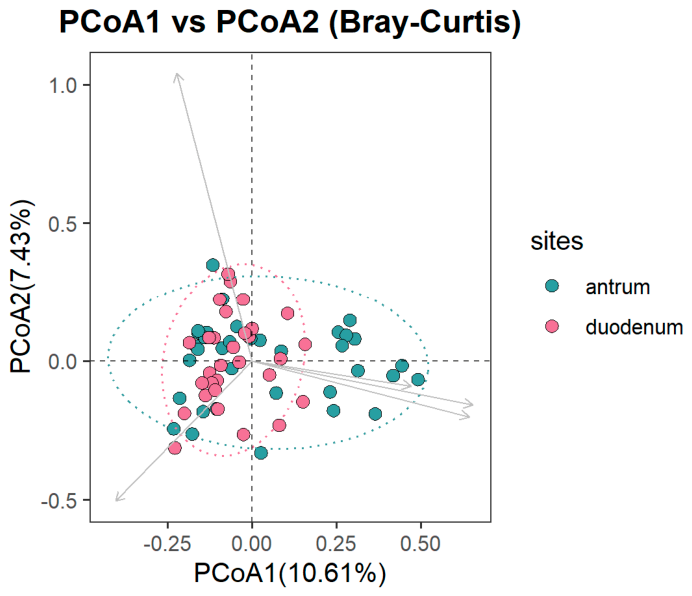
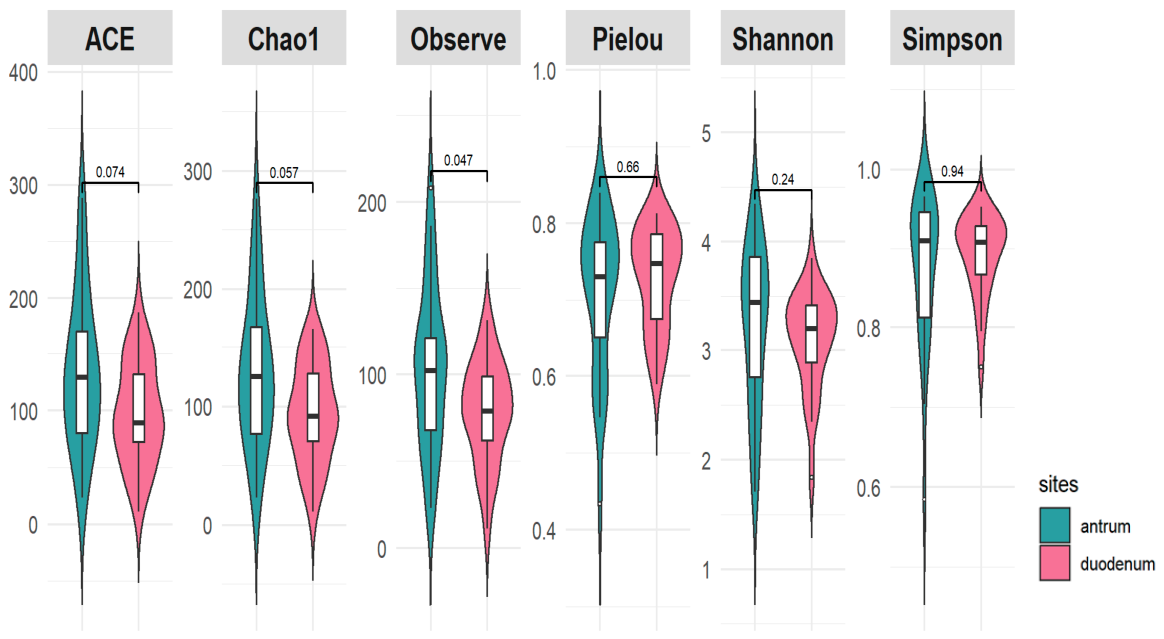
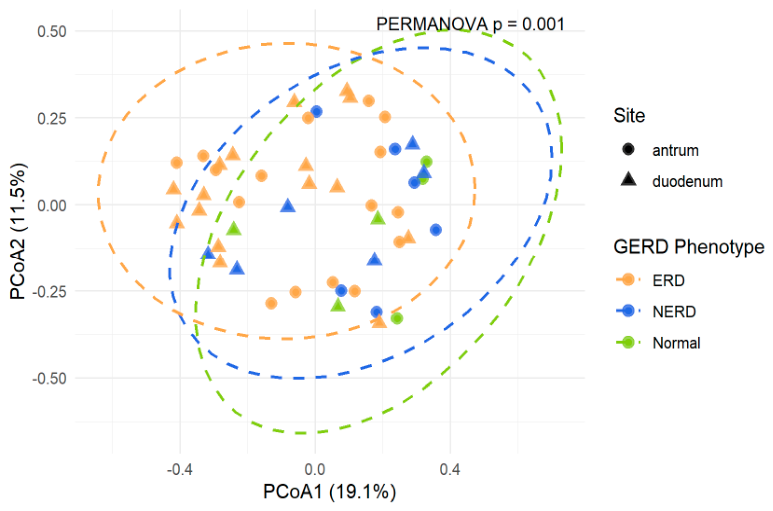
3.3.2. Alpha and Beta Diversity Differences across GERD Phenotypes by Anatomical Site
3.3.3. Associations between Microbial Profiles and Symptom Severity Scores
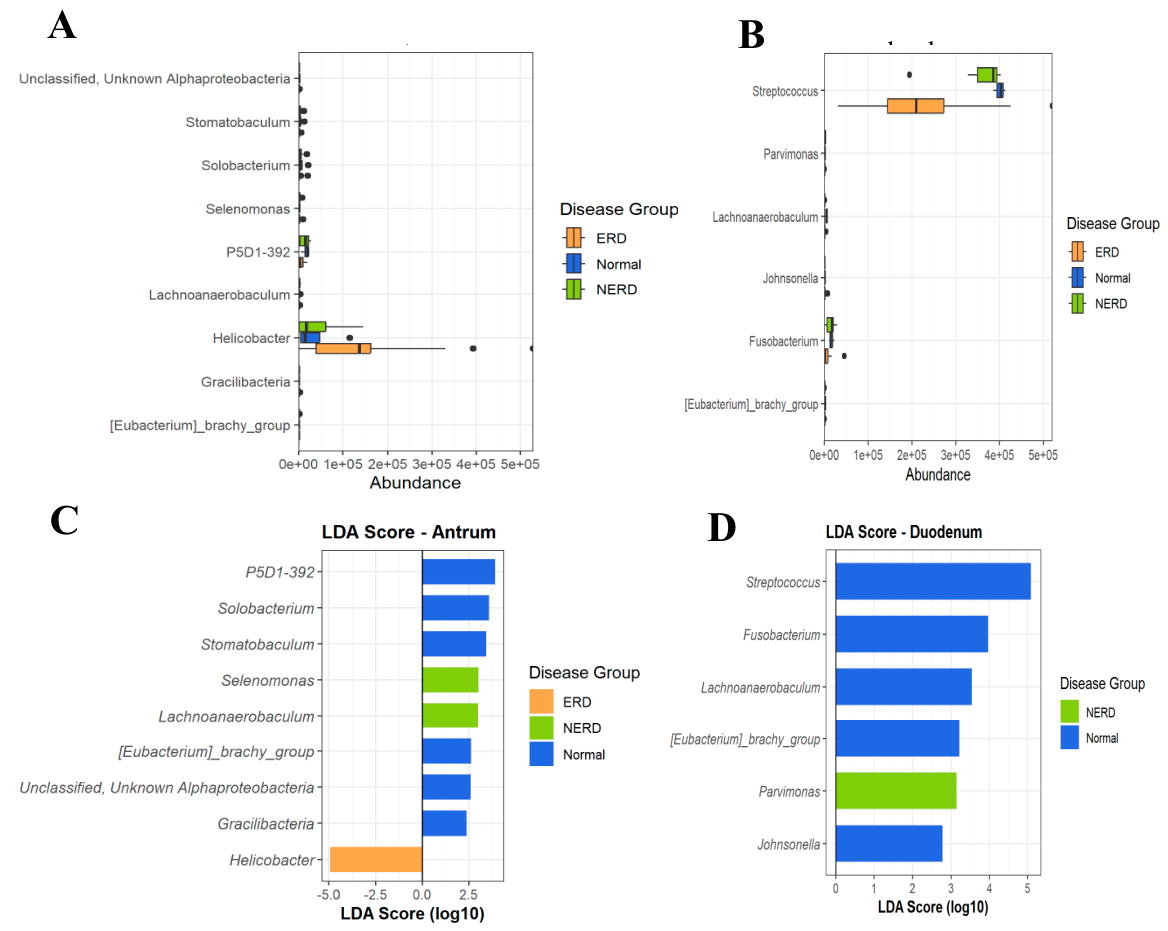
3.4. Differential Taxa and Biomarker Identification
3.5. Paired Analysis of Gastric and Duodenal Microbiota
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Deshpande, N.P.; Riordan, S.M.; Castaño-Rodríguez, N.; Wilkins, M.R.; Kaakoush, N.O. Signatures within the Esophageal Microbiome Are Associated with Host Genetics, Age, and Disease. Microbiome 2018, 6. [Google Scholar] [CrossRef]
- Eusebi, L.H.; Ratnakumaran, R.; Yuan, Y.; Solaymani-Dodaran, M.; Bazzoli, F.; Ford, A.C. Global Prevalence of, and Risk Factors for, Gastro-Oesophageal Reflux Symptoms: A Meta-Analysis. Gut 2018, 67, 430–440. [Google Scholar] [CrossRef]
- Jung, H.-K. Epidemiology of Gastroesophageal Reflux Disease in Asia: A Systematic Review. J. Neurogastroenterol. Motil. 2011, 17, 14–27. [Google Scholar] [CrossRef]
- Vakil, N.; Van Zanten, S.V.; Kahrilas, P.; Dent, J.; Jones, R. the Global Consensus Group The Montreal Definition and Classification of Gastroesophageal Reflux Disease: A Global Evidence-Based Consensus. Am. J. Gastroenterol. 2006, 101, 1900–1920. [Google Scholar] [CrossRef]
- Syam, A.F.; Sobur, C.S.; Hapsari, F.C.P.; Abdullah, M.; Makmun, D. Prevalence and Risk Factors of GERD in Indonesian Population—An Internet-Based Study. Adv. Sci. Lett. 2017, 23, 6734–6738. [Google Scholar] [CrossRef]
- Argüero, J.; Sifrim, D. Pathophysiology of Gastro-Oesophageal Reflux Disease: Implications for Diagnosis and Management. Nat. Rev. Gastroenterol. Hepatol. 2024, 21, 282–293. [Google Scholar] [CrossRef]
- Gyawali, C.P.; Kahrilas, P.J.; Savarino, E.; Zerbib, F.; Mion, F.; Smout, A.J.P.M.; Vaezi, M.; Sifrim, D.; Fox, M.R.; Vela, M.F.; et al. Modern Diagnosis of GERD: The Lyon Consensus. Gut 2018, 67, 1351–1362. [Google Scholar] [CrossRef] [PubMed]
- Davis, T.A.; Gyawali, C.P. Refractory Gastroesophageal Reflux Disease: Diagnosis and Management. J. Neurogastroenterol. Motil. 2024, 30, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Labenz, J.; Labenz, G.; Stephan, D.; Willeke, F.; 9. LOPA-Studiengruppe. Unzureichende Symptomkontrolle unter Langzeittherapie mit PPI bei GERD — Fakt oder Fiktion? MMW—Fortschritte Med. 2016, 158, 7–11. [Google Scholar] [CrossRef]
- Bonazzi, E.; Lorenzon, G.; Maniero, D.; De Barba, C.; Bertin, L.; Barberio, B.; Salvador, R.; Valmasoni, M.; Zingone, F.; Ghisa, M.; et al. The Esophageal Microbiota in Esophageal Health and Disease. Gastroenterol. Insights 2024, 15, 998–1013. [Google Scholar] [CrossRef]
- Guan, Y.; Cheng, H.; Zhang, N.; Cai, Y.; Zhang, Q.; Jiang, X.; Wang, A.; Zeng, H.; Jia, B. The Role of the Esophageal and Intestinal Microbiome in Gastroesophageal Reflux Disease: Past, Present, and Future. Front. Immunol. 2025, 16. [Google Scholar] [CrossRef]
- Nakae, H.; Tsuda, A.; Matsuoka, T.; Mine, T.; Koga, Y. Gastric Microbiota in the Functional Dyspepsia Patients Treated with Probiotic Yogurt. BMJ Open Gastroenterol. 2016, 3, e000109. [Google Scholar] [CrossRef]
- Wang, Z.-T.; Tan, W.-T.; Huang, J.-L.; Zhang, P.-F.; Li, Q.; Wang, M.-M.; Meng, M.-M.; Su, H.; Guo, C.-M.; Liu, H. Correlation Between Gastroesophageal Reflux Disease and Small Intestinal Bacterial Overgrowth: Analysis of Intestinal Microbiome and Metabolic Characteristics. J. Inflamm. Res. 2025, Volume 18, 33–51. [Google Scholar] [CrossRef]
- Bik, E.M.; Eckburg, P.B.; Gill, S.R.; Nelson, K.E.; Purdom, E.A.; Francois, F.; Perez-Perez, G.; Blaser, M.J.; Relman, D.A. Molecular Analysis of the Bacterial Microbiota in the Human Stomach. Proc. Natl. Acad. Sci. 2006, 103, 732–737. [Google Scholar] [CrossRef]
- Igarashi, M.; Nakae, H.; Matsuoka, T.; Takahashi, S.; Hisada, T.; Tomita, J.; Koga, Y. Alteration in the Gastric Microbiota and Its Restoration by Probiotics in Patients with Functional Dyspepsia. BMJ Open Gastroenterol. 2017, 4, e000144. [Google Scholar] [CrossRef]
- Zhang, X.; Li, Q.; Xia, S.; He, Y.; Liu, Y.; Yang, J.; Xiao, X. Proton Pump Inhibitors and Oral–Gut Microbiota: From Mechanism to Clinical Significance. Biomedicines 2024, 16 12, 2271. [Google Scholar] [CrossRef]
- Revaiah, P.C.; Kochhar, R.; Rana, S.V.; Berry, N.; Ashat, M.; Dhaka, N.; Rami Reddy, Y.; Sinha, S.K. Risk of Small Intestinal Bacterial Overgrowth in Patients Receiving Proton Pump Inhibitors versus Proton Pump Inhibitors plus Prokinetics. JGH Open 2018, 2, 47–53. [Google Scholar] [CrossRef]
- Wei, L.; Singh, R.; Ro, S.; Ghoshal, U.C. Gut Microbiota Dysbiosis in Functional Gastrointestinal Disorders: Underpinning the Symptoms and Pathophysiology. JGH Open 2021, 5, 976–987. [Google Scholar] [CrossRef] [PubMed]
- Lundell, L.R.; Dent, J.; Bennett, J.R.; Blum, A.L.; Armstrong, D.; Galmiche, J.P.; Johnson, F.; Hongo, M.; Richter, J.E.; Spechler, S.J.; et al. Endoscopic Assessment of Oesophagitis: Clinical and Functional Correlates and Further Validation of the Los Angeles Classification. Gut 1999, 45, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Saragih, R.H.; Rey, I. FSSG Scale System in Comparison with GERD Questionnaires in Predicting Endoscopic. Indones. J. Gastroenterol. Hepatol. Dig. Endosc. 2012, 14, 136. [Google Scholar]
- Simadibrata, M.; Rani, A.; Adi, P.; Djumhana, A.; Abdullah, M. The Gastro-Esophageal Reflux Disease Questionnaire Using Indonesian Language: A Language Validation Survey. Med. J. Indones. 2011, 125. [Google Scholar] [CrossRef]
- Sugihartono, T.; Fauzia, K.A.; Miftahussurur, M.; Waskito, L.A.; Rejeki, P.S.; I’tishom, R.; Alfaray, R.I.; Doohan, D.; Amalia, R.; Savitri, C.M.A.; et al. Analysis of Gastric Microbiota and Helicobacter Pylori Infection in Gastroesophageal Reflux Disease. Gut Pathog. 2022, 22 14. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Robeson, M.S.; O’Rourke, D.R.; Kaehler, B.D.; Ziemski, M.; Dillon, M.R.; Foster, J.T.; Bokulich, N.A. RESCRIPt: Reproducible Sequence Taxonomy Reference Database Management. PLOS Comput. Biol. 2021, 24 17, e1009581. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2—Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K. MAFFT Version 5: Improvement in Accuracy of Multiple Sequence Alignment. Nucleic Acids Res. 2005, 33, 511–518. [Google Scholar] [CrossRef]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Pavoine, S.; Dufour, A.-B.; Chessel, D. From Dissimilarities among Species to Dissimilarities among Communities: A Double Principal Coordinate Analysis. J. Theor. Biol. 2004, 228, 523–537. [Google Scholar] [CrossRef] [PubMed]
- Ramette, A. Multivariate Analyses in Microbial Ecology. FEMS Microbiol. Ecol. 2007, 62, 142–160. [Google Scholar] [CrossRef]
- Kelly, B.J.; Gross, R.; Bittinger, K.; Sherrill-Mix, S.; Lewis, J.D.; Collman, R.G.; Bushman, F.D.; Li, H. Power and Sample-Size Estimation for Microbiome Studies Using Pairwise Distances and PERMANOVA. Bioinformatics 2015, 31, 2461–2468. [Google Scholar] [CrossRef]
- Yuan, L.-Z.; Yi, P.; Wang, G.-S.; Tan, S.-Y.; Huang, G.-M.; Qi, L.-Z.; Jia, Y.; Wang, F. Lifestyle Intervention for Gastroesophageal Reflux Disease: A National Multicenter Survey of Lifestyle Factor Effects on Gastroesophageal Reflux Disease in China. Ther. Adv. Gastroenterol. 2019, 12. [Google Scholar] [CrossRef] [PubMed]
- Katzka, D.A.; Kahrilas, P.J. Advances in the Diagnosis and Management of Gastroesophageal Reflux Disease. BMJ 2020, m3786. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Yu, F.; Zhou, J.; Zhao, C.; Wu, J.; Ni, X. Analysis of the Gut Microbiota in Children with Gastroesophageal Reflux Disease Using Metagenomics and Metabolomics. Front. Cell. Infect. Microbiol. 2023, 13. [Google Scholar] [CrossRef]
- Vikram Singh, A.; Sharma, P.; Das, K.; Bhaskar, Y.; Goel, I.; Singh, H.; Das, R. Alteration in the Gastric Microbiota of Erosive Reflux Disease and Non-Erosive Reflux Disease Patients: An Indicator for Disease Progression. Biomedicine 2024, 34 44, 79–88. [Google Scholar] [CrossRef]
- Martinez-Guryn, K.; Leone, V.; Chang, E.B. Regional Diversity of the Gastrointestinal Microbiome. Cell Host Microbe 2019, 26, 314–324. [Google Scholar] [CrossRef]
- Han, H.S.; Lee, S.-Y.; Oh, S.Y.; Moon, H.W.; Cho, H.; Kim, J.-H. Correlations of the Gastric and Duodenal Microbiota with Histological, Endoscopic, and Symptomatic Gastritis. J. Clin. Med. 2019, 8, 312. [Google Scholar] [CrossRef]
- Zhou, J.; Shrestha, P.; Qiu, Z.; Harman, D.G.; Teoh, W.-C.; Al-Sohaily, S.; Liem, H.; Turner, I.; Ho, V. Distinct Microbiota Dysbiosis in Patients with Non-Erosive Reflux Disease and Esophageal Adenocarcinoma. J. Clin. Med. 2020, 9, 2162. [Google Scholar] [CrossRef]
- Wang, K.; Wang, S.; Chen, Y.; Lu, X.; Wang, D.; Zhang, Y.; Pan, W.; Zhou, C.; Zou, D. Causal Relationship between Gut Microbiota and Risk of Gastroesophageal Reflux Disease: A Genetic Correlation and Bidirectional Mendelian Randomization Study. Front. Immunol. 2024, 15. [Google Scholar] [CrossRef]
- Park, C.H.; Seo, S.I.; Kim, J.S.; Kang, S.H.; Kim, B.J.; Choi, Y.J.; Byun, H.J.; Yoon, J.-H.; Lee, S.K. Treatment of Non-Erosive Reflux Disease and Dynamics of the Esophageal Microbiome: A Prospective Multicenter Study. Sci. Rep. 2020, 10, 15154. [Google Scholar] [CrossRef] [PubMed]
- Park, C.H.; Lee, S.K. Exploring Esophageal Microbiomes in Esophageal Diseases: A Systematic Review. J. Neurogastroenterol. Motil. 2020, 26, 171–179. [Google Scholar] [CrossRef]
- Chen, S.; Jiang, D.; Zhuang, Q.; Hou, X.; Jia, X.; Chen, J.; Lin, H.; Zhang, M.; Tan, N.; Xiao, Y. Esophageal Microbial Dysbiosis Impairs Mucosal Barrier Integrity via Toll-like Receptor 2 Pathway in Patients with Gastroesophageal Reflux Symptoms. J. Transl. Med. 2024, 22, 1145. [Google Scholar] [CrossRef] [PubMed]
- Corning, B.; Copland, A.P.; Frye, J.W. The Esophageal Microbiome in Health and Disease. Curr. Gastroenterol. Rep. 2018, 20, 39. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Chen, L.; Zhang, T.; Gabo, R.; Wang, Q.; Zhong, Z.; Yao, M.; Wei, W.; Su, X. Duodenal Microbiota Dysbiosis in Functional Dyspepsia and Its Potential Role of the Duodenal Microbiota in Gut–Brain Axis Interaction: A Systematic Review. Front. Microbiol. 2024, 15, 1409280. [Google Scholar] [CrossRef]
- Oshima, T. Functional Dyspepsia: Current Understanding and Future Perspective. Digestion 2024, 105, 26–33. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



