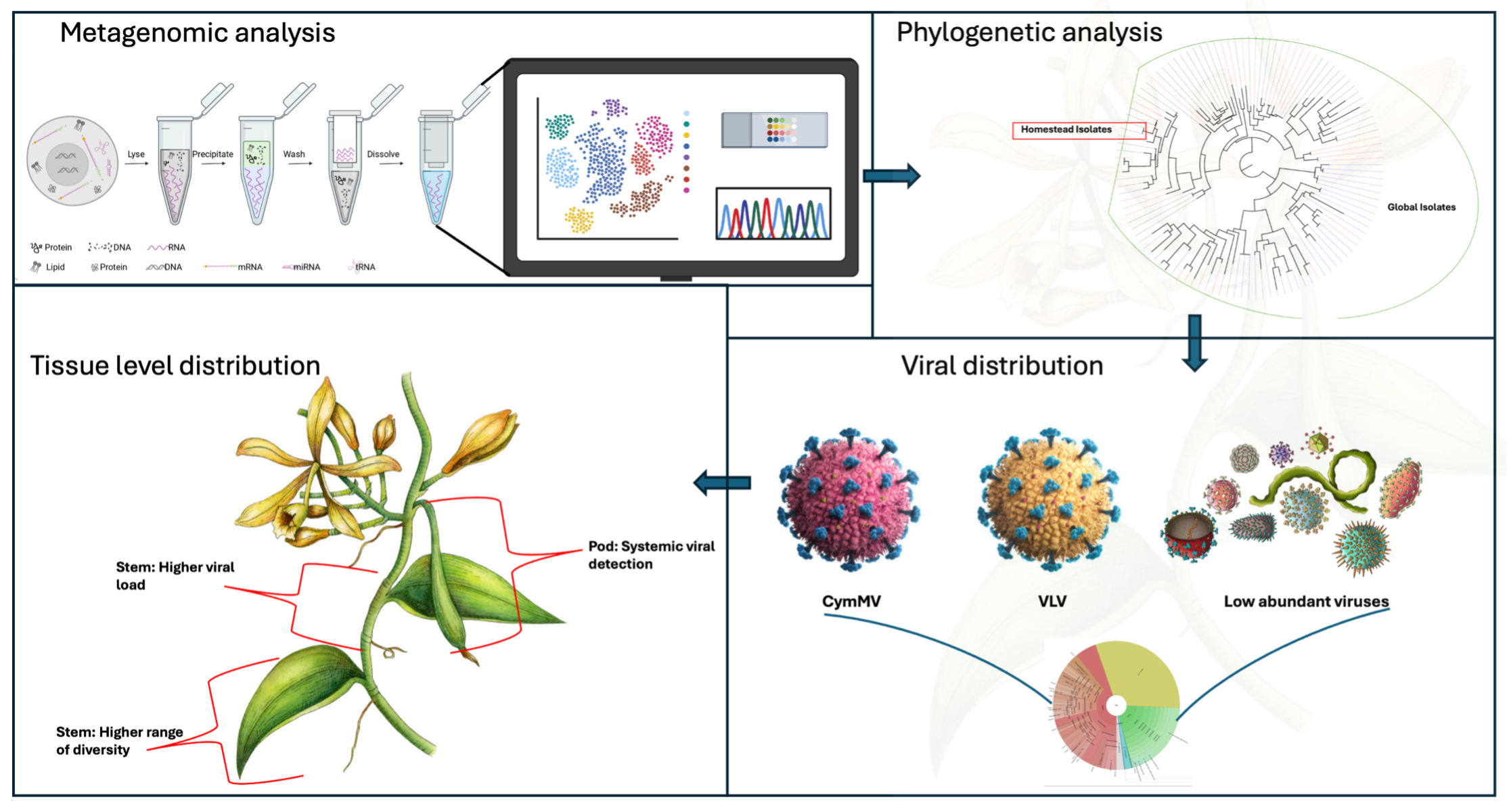
Vanilla planifolia, a high-value tropical orchid, is significantly impacted by viral pathogens that threaten its cultivation and productivity. This study employs metagenomic technique to detect and characterize the viral communities associated with V. planifolia in south Florida. Using high-throughput RNA sequencing, the Cymbidium mosaic virus (CymMV) and Vanilla latent virus (VLV) were prevalent in the plant system, with CymMV being the dominant viral species. Phylogenetic analysis of the CymMV coat protein gene revealed notable genetic divergence in the Homestead isolate, forming a distinct clade from global reference strains, suggesting local adaptation or host-specific evolution. Viral distribution across plant system revealed higher viral loads in stem tissue, consistent with their role in systemic transport, whereas leaves exhibited greater viral diversity, likely due to in-creased environmental exposure. The low abundance of other viral species, including Garlic viruses and Senna severe yellow mosaic virus, highlights the complex viral ecology associated with V. planifolia. This study underscores the value of metagenomic approaches for uncovering both well-characterized and novel viruses in plant systems and highlight the need for continuous viral surveillance to guide disease management strategies in economically important crops such as vanilla.