Submitted:
16 April 2026
Posted:
18 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sequence Retrieval of Protein
2.2. Epitope Mapping
2.3. Bovine Homology
2.4. Epitope Conservancy
2.5. Assessment of B Cell and T Cell Epitopes
2.6. Assembly of the Final Multi-Epitope mRNA Vaccine
2.7. Profile of Physicochemical and Immunological Properties of the Construct
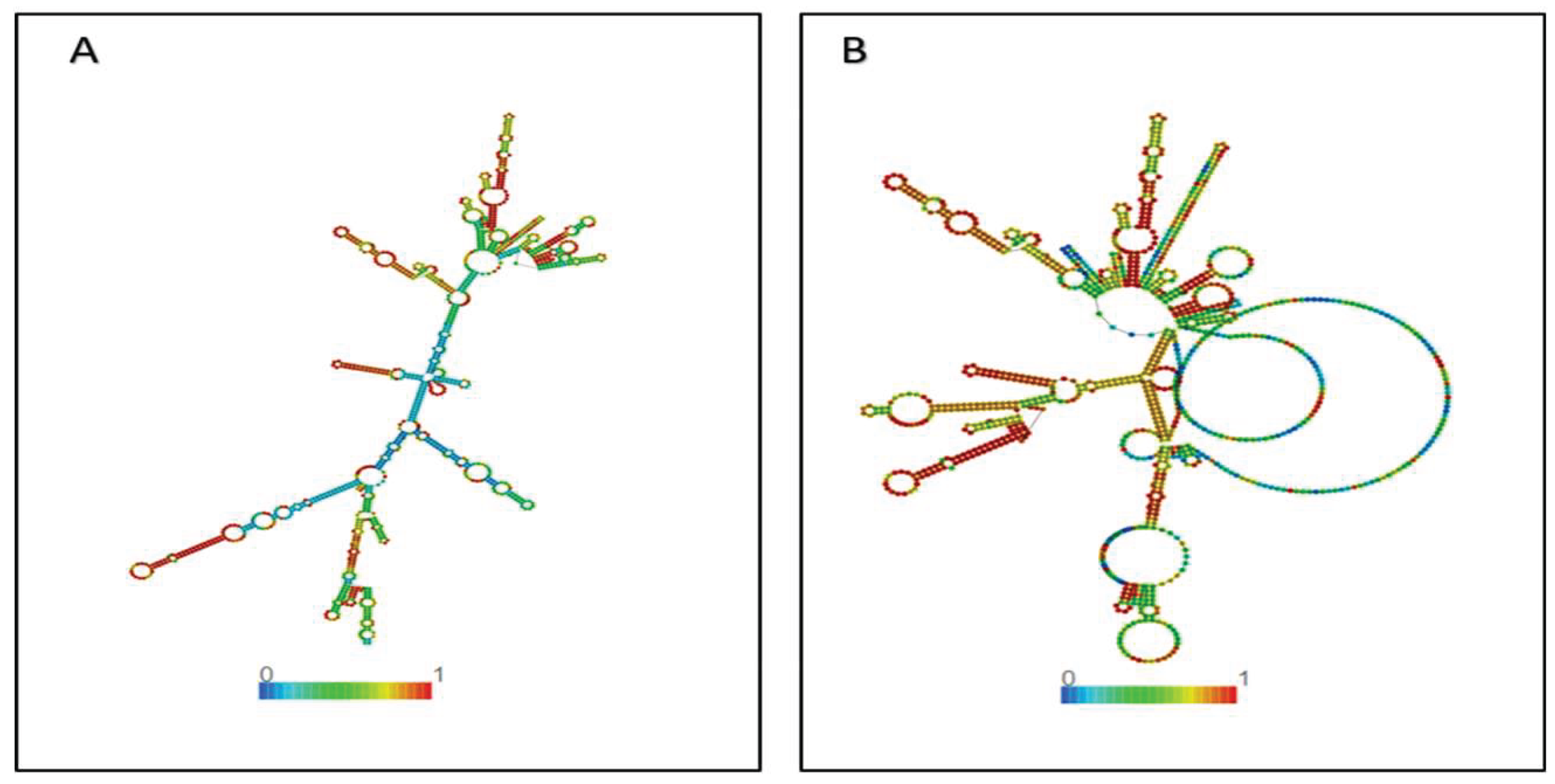
2.8. Prediction of Secondary Structure (2D)
2.9. Prediction of Tertiary Structure (3D)
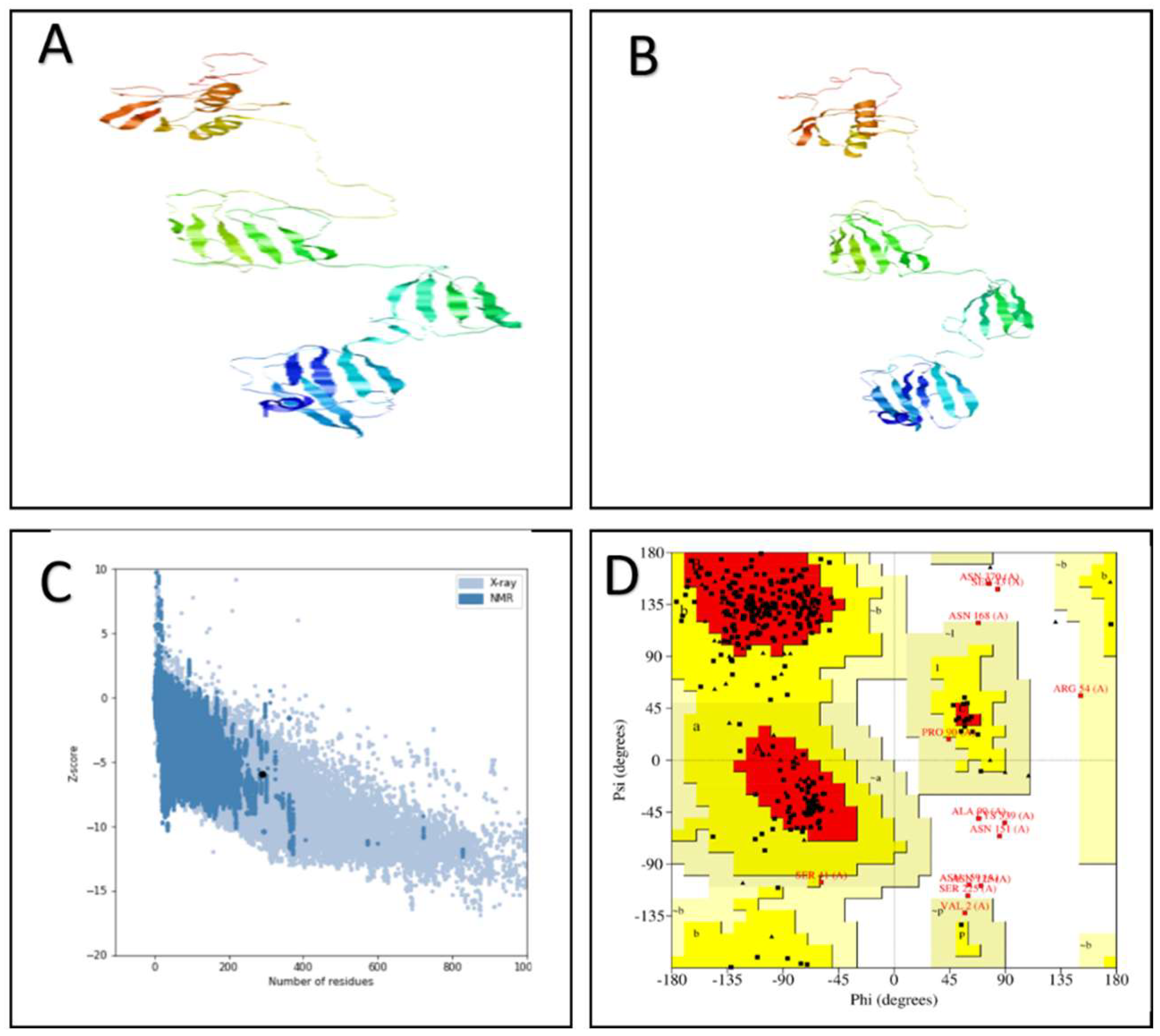
2.10. Validation and Refinement of the Vaccine Model
2.11. Molecular Docking of the Vaccine Candidate with Immune Receptors
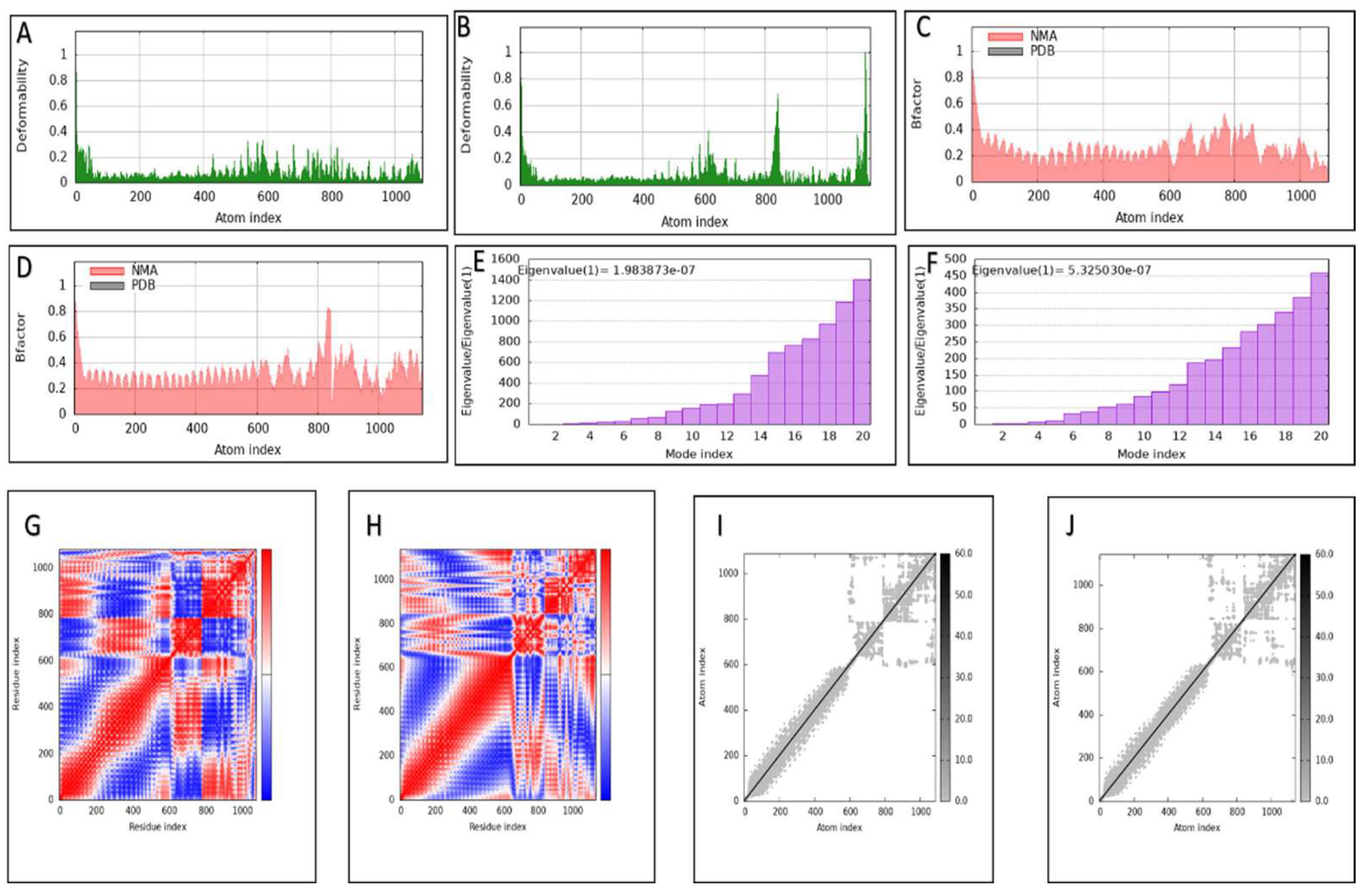
2.12. Molecular Dynamic Simulations (MDs)
2.13. Immune Simulation of the Vaccine Candidate
3. Results
3.1. Retrieval and Analysis of Selected Proteins Sequences
3.2. Perdition and Evaluation of T and B Cell Epitopes
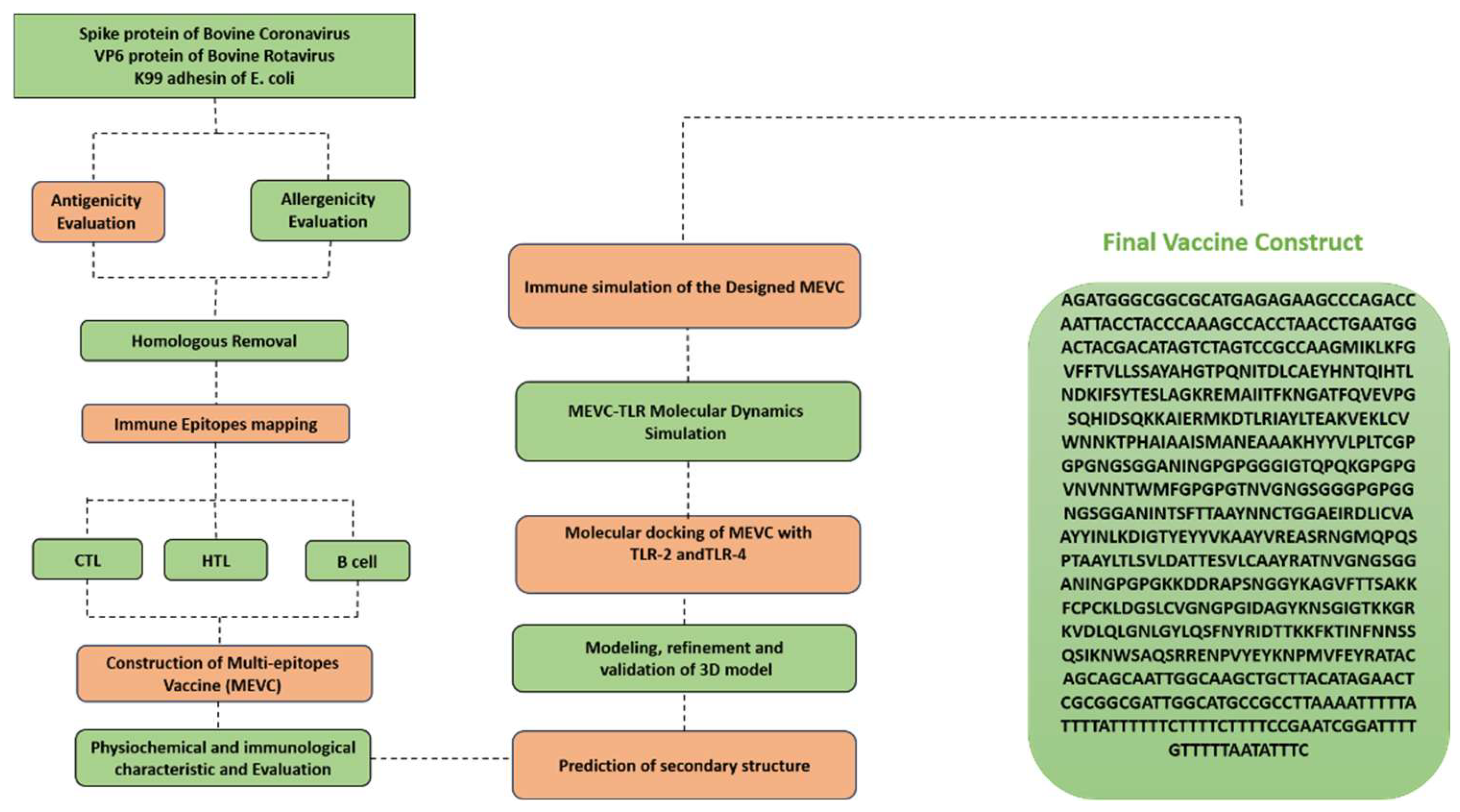
3.3. Construction of the Multi-Epitope mRNA Vaccine
3.4. Physicochemical and Immunological Analysis of the Proposed Vaccine
| Characteristic | Measurement | Remark |
|---|---|---|
| Total number of amino acids | 377 | Suitable |
| Molecular weight | 39070.24 | Appropriate |
| Theoretical pI | 9.26 | Basic |
| Formula | C1702H2625N499O540S11 | ------- |
| Predicted half-life (Escherichia coli, in vivo) | 10h | ----- |
| Predicted half-life (mammalian reticulocytes, in vitro) | 1.9h | ------ |
| Predicted half-life (yeast-cells, in vivo) | 20h | ----- |
| Grand average of hydropathicity (GRAVY) | -0.479 | Hydrophilic |
| Instability index of vaccine | 25.07 | stable |
| Antigenicity | 1.1056 | antigenic |
| Allergenicity | Non-allergic | Non-allergic |
| Toxicity | Nontoxic | Nontoxic |
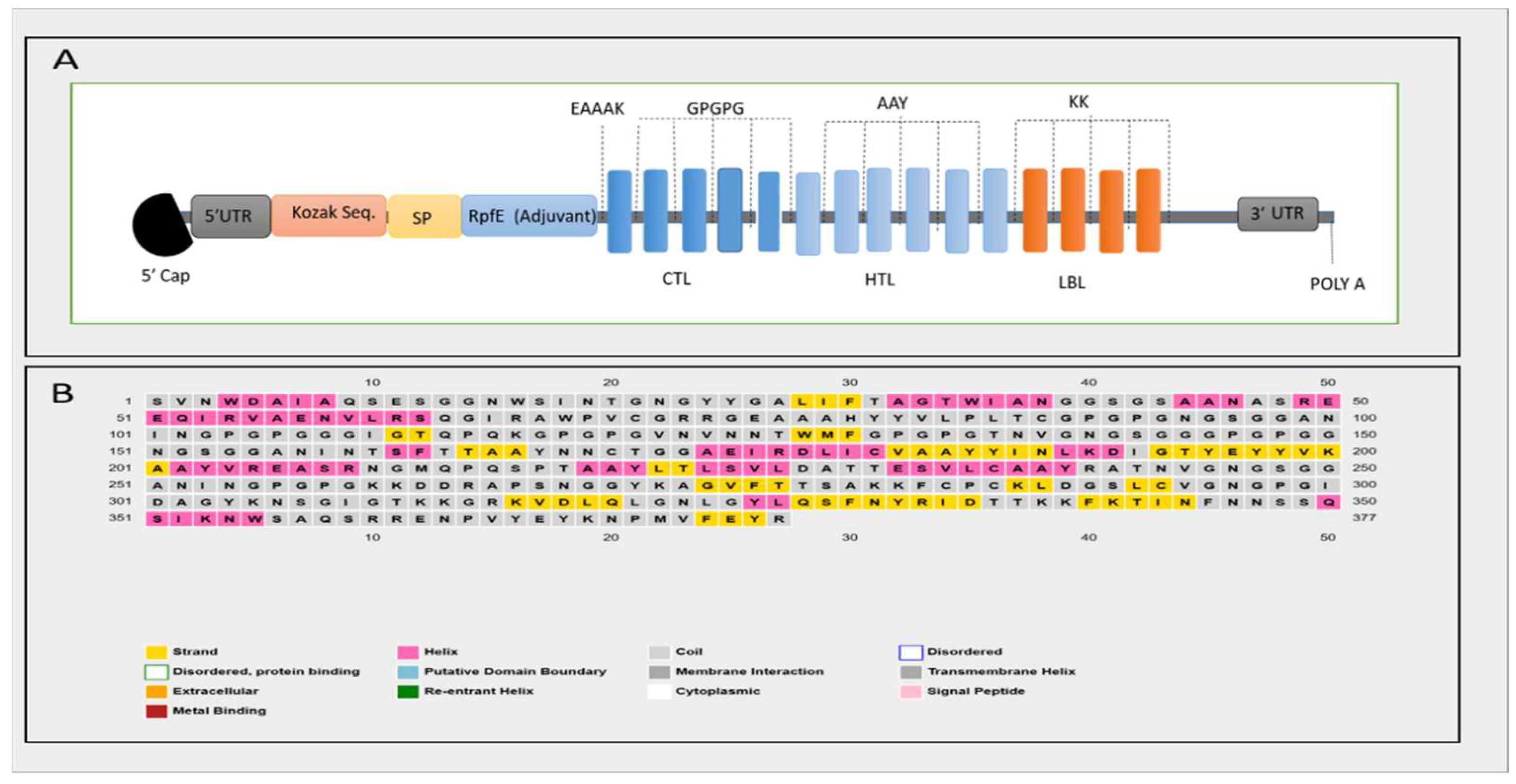
3.5. Secondary and 3D Structure of the Proposed Vaccine Construct
3.6. Prediction of Discontinuous B-Cell Epitopes
| No | Residues | Number of residues | score |
|---|---|---|---|
| 1 | A:N354, A:W355, A:S356, A:A357, A:Q358, A:S359, A:R360, A:R361, A:E362, A:N363, A:P364, A:V365 | 12 | 0.875 |
| 2 | A:G111, A:T112, A:Q113, A:P114, A:Q115, A:K116, A:V124, A:N125, A:N126, A:T127, A:W128, A:P134, A:G135, A:T136, A:N137, A:V138, A:G139, A:N140, A:G141, A:S142, A:G143, A:G144, A:G145, A:P146, A:G147, A:P148, A:G149, A:G150, A:N151, A:G152, A:S153, A:G154, A:G155, A:A156, A:N157, A:I158, A:N159, A:T160, A:S161, A:F162, A:T163, A:T164, A:A165, A:A166, A:Y167, A:N168, A:N169, A:C170 | 48 | 0.729 |
| 3 | A:S1, A:V2, A:N3, A:W4, A:D5, A:A6, A:I7, A:A8, A:Q9, A:S10, A:E11, A:S12, A:G13, A:G14, A:N15, A:W16, A:S17, A:I18, A:N19, A:T20, A:A27, A:I29, A:F30, A:T31, A:A32, A:G33, A:T34, A:G39, A:G40, A:S41, A:G42, A:S43, A:A44, A:A45, A:N46, A:A47, A:S48, A:R49, A:Q52, A:V55, A:A56, A:E57, A:N58, A:V59, A:L60, A:R61, A:S62, A:Q63, A:G64, A:I65, A:A67, A:W68, A:P69, A:V70, A:C71, A:G72, A:G75, A:E76, A:T87, A:C88, A:G89, A:P90, A:G91 | 63 | 0.691 |
| 4 | A:K282, A:F283, A:C284, A:G295, A:N296, A:G297, A:P298, A:D301, A:A302, A:G303, A:Y304, A:N306, A:S307, A:G308, A:I309, A:G310, A:T311, A:K312, A:K313, A:G314, A:R315, A:K316, A:V317, A:D318, A:L319, A:Q320, A:L321, A:G322, A:L324, A:G325, A:L327, A:Q328, A:S329, A:F330, A:N331, A:Y332, A:R333, A:I334, A:D335, A:T336, A:T337, A:K338, A:K341, A:T342, A:I343, A:N344, A:F345, A:N346, A:N347, A:S348, A:S349, A:Q350, A:S351, A:I352, A:Y366, A:E367, A:Y368, A:K369, A:N370, A:P371, A:M372, A:V373 | 62 | 0.675 |
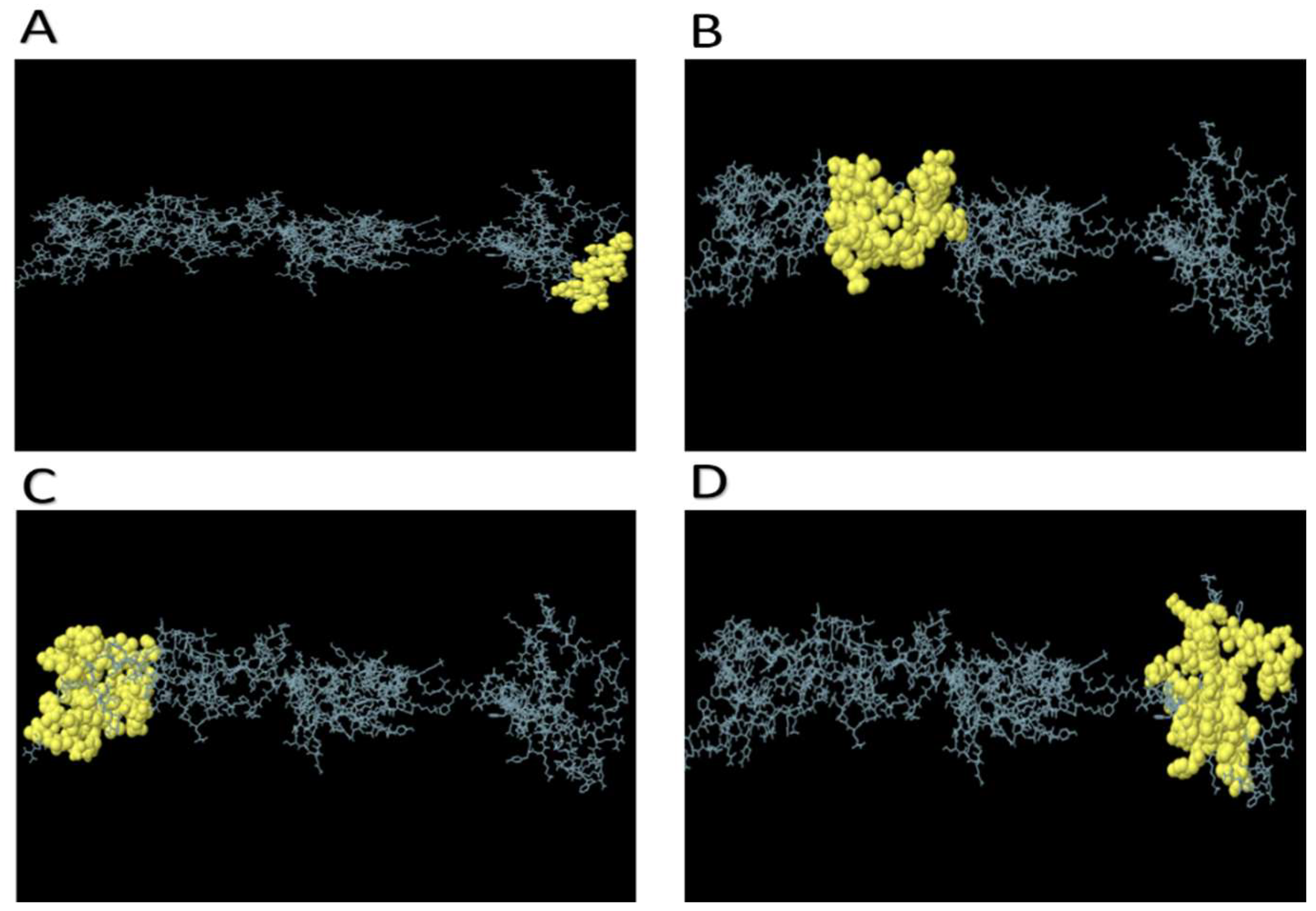
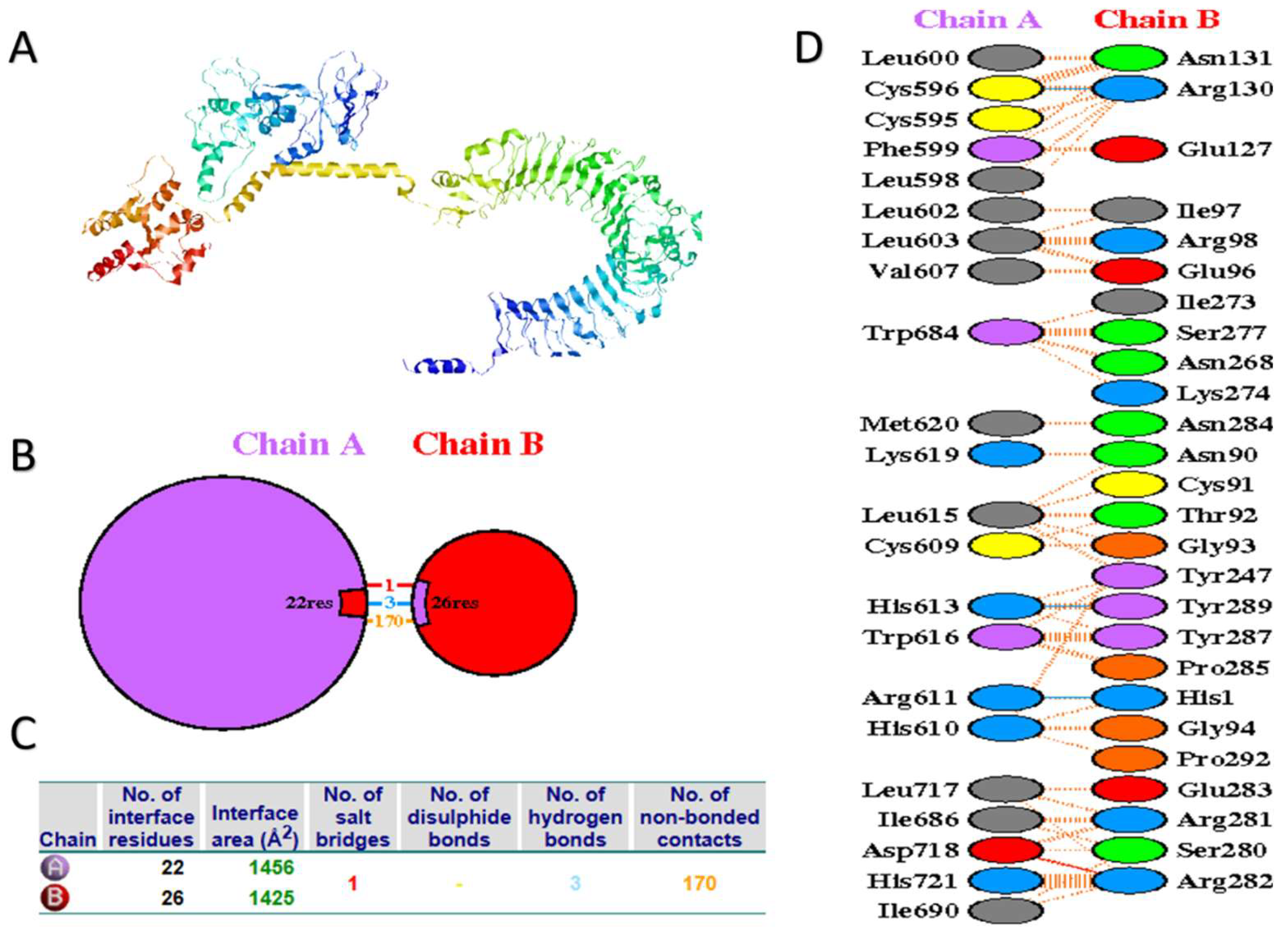
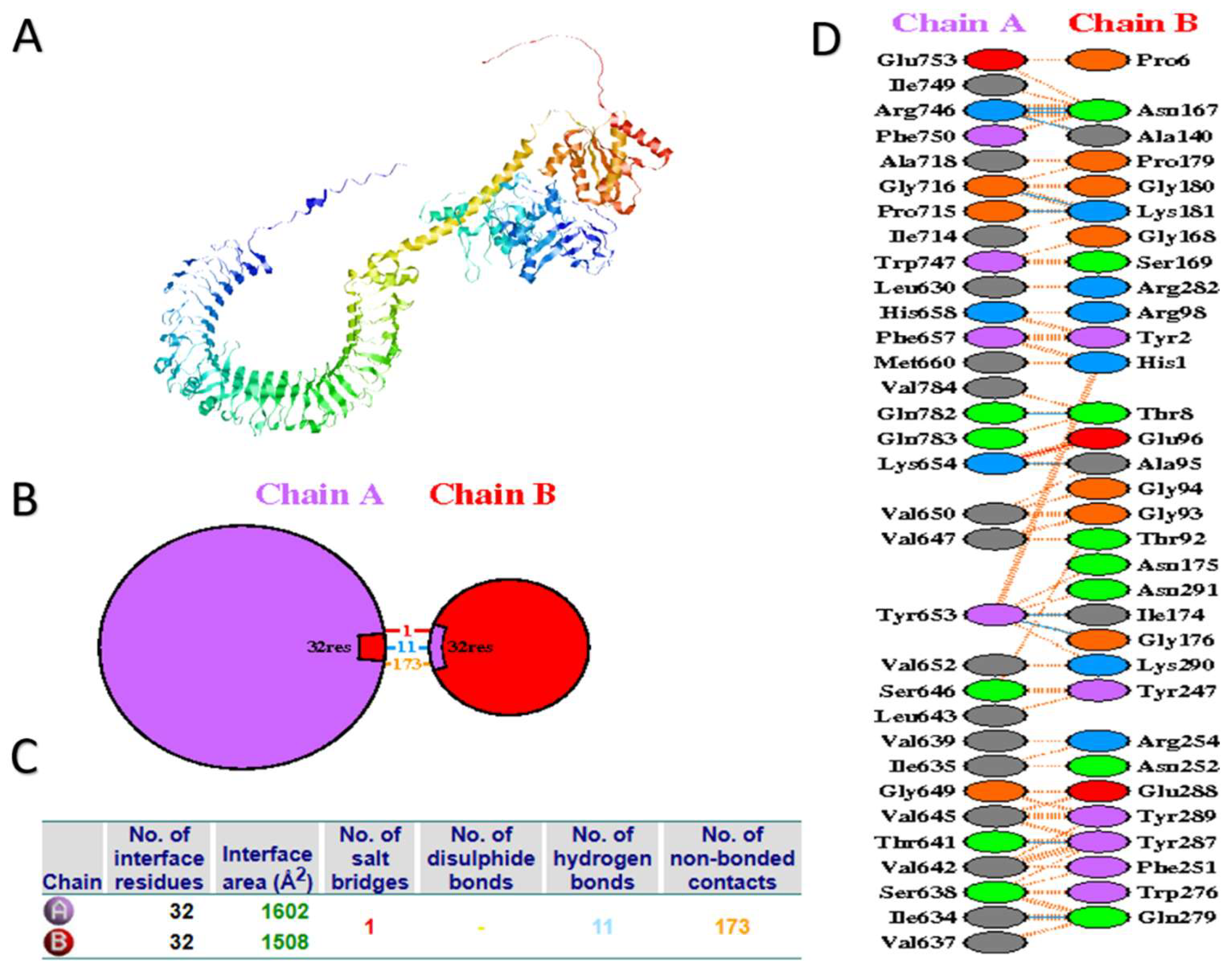
3.7. Molecular Docking of the Vaccine Candidate with Bovine TLR2 and TRL4
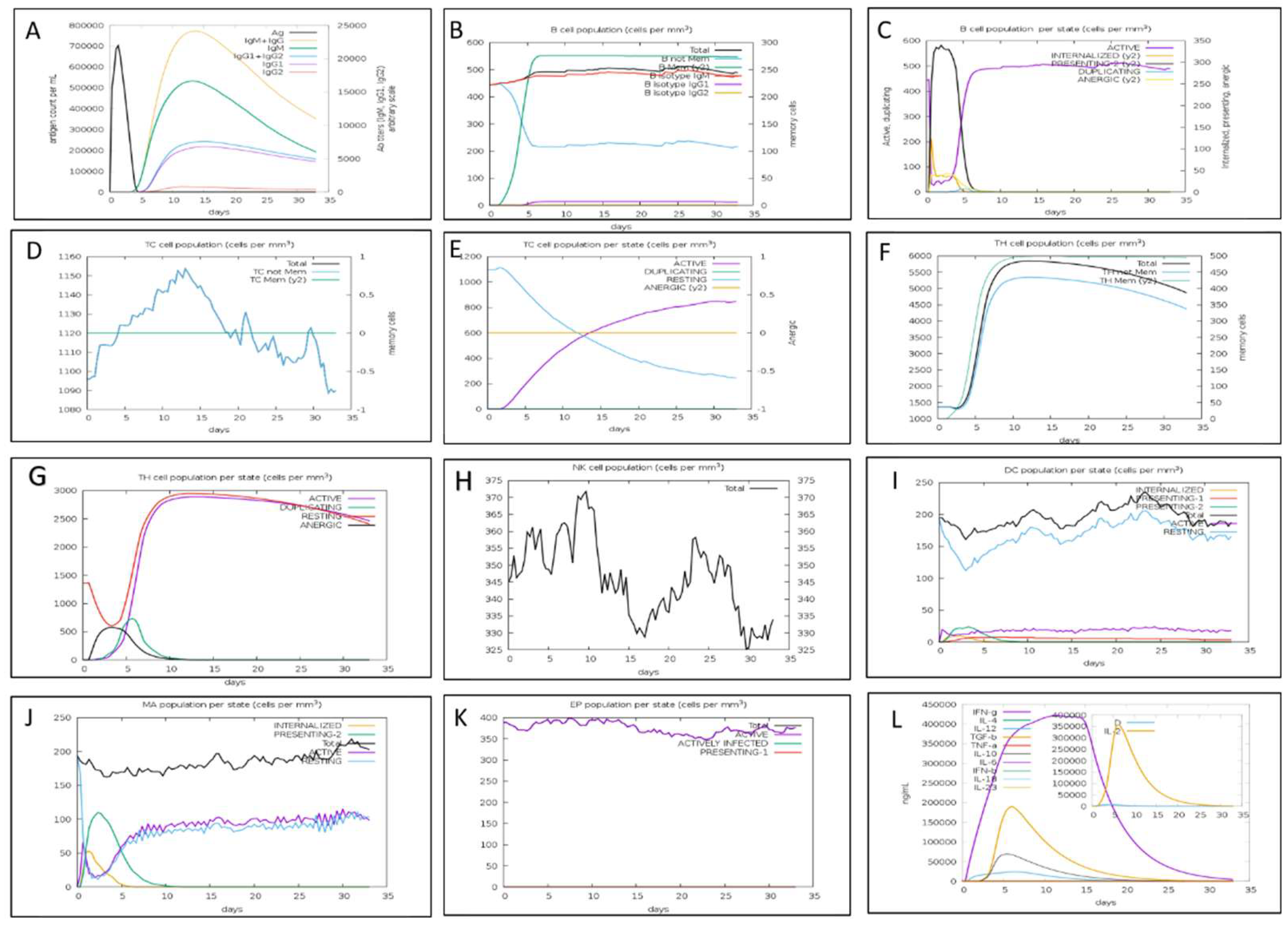
3.8. Immune Simulation of the Proposed Vaccine
3.9. mRNA Vaccine Structure and In Silico Codon Optimization
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Foster, D.M.; Smith, G.W. Pathophysiology of diarrhea in calves. Vet Clin North Am Food Anim Pract 2009, 25(1), 13–36. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.I.; Yoon, K.J. An overview of calf diarrhea – infectious etiology, diagnosis, and intervention. J Vet Sci. 2014, 15(1), 1–17. [Google Scholar] [CrossRef]
- Kostanić, V.; Kunić, V.; Prišlin Šimac, M.; Lolić, M.; Sukalić, T.; Brnić, D. Comparative insights into acute gastroenteritis in cattle caused by bovine rotavirus A and bovine coronavirus. Vet Sci. 2024, 11(12), 671. [Google Scholar] [CrossRef]
- Maier, G.U.; Breitenbuecher, J.; Gomez, J.P.; Samah, F.; Fausak, E.; Van Noord, M. Vaccination for the prevention of neonatal calf diarrhea in cow-calf operations: a scoping review. Vet Anim Sci. 2022, 15, 100238. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines — a new era in vaccinology. Nat Rev Drug Discov. 2018, 17(4), 261–79. [Google Scholar] [CrossRef]
- Maier, G.U.; Breitenbuecher, J.; Gomez, J.P.; Samah, F.; Fausak, E.; Van Noord, M. Vaccination for the prevention of neonatal calf diarrhea in cow-calf operations: A scoping review. Vet Anim Sci. 2022, 15, 100238. [Google Scholar] [CrossRef]
- Pumchan, A.; Proespraiwong, P.; Sawatdichaikul, O.; Phurahong, T.; Hirono, I.; Unajak, S. Computational design of novel chimeric multiepitope vaccine against bacterial and viral disease in tilapia (Oreochromis sp.). Scientific Reports 2024, 14(1), 14048. [Google Scholar] [CrossRef]
- Leong, K.Y.; Tham, S.K.; Poh, C.L. Revolutionizing immunization: a comprehensive review of mRNA vaccine technology and applications. Virol J. 2025, 22(1), 71. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yuan, X.; He, Y.; Chen, Y.; Hu, C.; Chen, J.; Zhang, L.; Chen, X.; Guo, A. A novel multiepitope fusion antigen as a vaccine candidate for the prevention of enterotoxigenic E. coli-induced calf diarrhea. Vaccines 2024, 12(5), 457. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, P.; Kim, Y.; Haste-Andersen, P.; Beaver, J.; Bourne, P.E.; Bui, H.H.; Buus, S.; Frankild, S.; Greenbaum, J.; Lund, O. Immune epitope database analysis resource (IEDB-AR). Nucleic acids research 2008, 36 (suppl_2), W513–8. [Google Scholar] [CrossRef]
- Vita, R.; Mahajan, S.; Overton, J.A.; Dhanda, S.K.; Martini, S.; Cantrell, J.R.; Wheeler, D.K.; Sette, A.; Peters, B. The immune epitope database (IEDB): 2018 update. Nucleic acids research 2019, 47(D1), D339–43. [Google Scholar] [CrossRef] [PubMed]
- Bui, H.H.; Sidney, J.; Dinh, K.; Southwood, S.; Newman, M.J.; Sette, A. Predicting population coverage of T-cell epitope-based diagnostics and vaccines. BMC bioinformatics 2006, 7(1), 153. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC bioinformatics 2007, 8(1), 4. [Google Scholar] [CrossRef]
- Dimitrov, I.; Bangov, I.; Flower, D.R.; Doytchinova, I. AllerTOP v. 2—a server for in silico prediction of allergens. Journal of molecular modeling 2014, 20(6), 2278. [Google Scholar] [CrossRef]
- Chen, X.; Zaro, J.L.; Shen, W.C. Fusion protein linkers: property, design and functionality. Advanced drug delivery reviews 2013, 65(10), 1357–69. [Google Scholar] [CrossRef]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.E.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein identification and analysis tools on the ExPASy server. In InThe proteomics protocols handbook; Humana press: Totowa, NJ, 22 Mar 2005; pp. 571–607. [Google Scholar]
- McGuffin, L.J.; Bryson, K.; Jones, D.T. The PSIPRED protein structure prediction server. Bioinformatics 2000, 16(4), 404–5. [Google Scholar] [CrossRef]
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nature protocols 2010, 5(4), 725–38. [Google Scholar] [CrossRef]
- Heo, L.; Park, H.; Seok, C. GalaxyRefine: Protein structure refinement driven by side-chain repacking. Nucleic acids research 2013, 41(W1), W384–8. [Google Scholar] [CrossRef]
- Lovell, S.C.; Davis, I.W.; Arendall, W.B., III; De Bakker, P.I.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins: Structure, Function, and Bioinformatics 2003, 50(3), 437–50. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Wang, X.; Hu, J.; Zhang, B.; Zhao, L.; Zhang, G.; Qi, J.; Wei, Z.; Bao, Y.; Tian, M.; Wang, S. In silico design of a multi-epitope vaccine against Mycobacterium avium subspecies paratuberculosis. Frontiers in Immunology 2025, 16, 1505313. [Google Scholar] [CrossRef] [PubMed]
- Rapin, N.; Lund, O.; Bernaschi, M.; Castiglione, F. Computational immunology meets bioinformatics: the use of prediction tools for molecular binding in the simulation of the immune system. PloS one 2010, 5(4), e9862. [Google Scholar] [CrossRef]
- Brunauer, Michael; Roch, Franz-Ferdinand; Conrady, Beate. Prevalence of worldwide neonatal calf diarrhoea caused by bovine rotavirus in combination with bovine coronavirus, Escherichia coli K99 and Cryptosporidium spp.: a meta-analysis. Animals 2021, 11(no. 4), 1014. [Google Scholar] [CrossRef] [PubMed]
- Alawadi, Z.I.; Alsultan, A.; Alsallami, D.; Alnomasy, S.F.; Alqasmi, M.; Almufarriji, F.M.; Alotaibi, B.S.; Mazhari, B.B.; Alenazy, R. Designing a multiepitope mRNA-based vaccine against enterotoxigenic Escherichia coli infection in calves: immunoinformatics and molecular modeling approach. Open Veterinary Journal 2024, 14(6), 1417. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Li, Y.; Chen, L.; Jia, S. Designing a broad-spectrum multi-epitope vaccine against influenza A and Mycoplasma pneumoniae: an immunoinformatics approach. Frontiers in Public Health 2026, 14, 1671035. [Google Scholar] [CrossRef] [PubMed]
- Mohammadipour, S.; Tavakkoli, H.; Fatemi, S.N.; Sharifi, A.; Mahmoudi, P. Designing a multi-epitope universal vaccine for concurrent infections of SARS-CoV-2 and influenza viruses using an immunoinformatics approach. BMC Infectious Diseases 2025, 25(1), 688. [Google Scholar] [CrossRef]
- Chaudhary, N.; Weissman, D.; Whitehead, K.A. mRNA vaccines for infectious diseases: principles, delivery and clinical translation. Nature reviews Drug discovery 2021, 20(11), 817–38. [Google Scholar] [CrossRef]
| Epitopes | Antigenicity score | Toxicity | Allergenicity | conservancy | Protein |
|---|---|---|---|---|---|
| NGSGGANIN |
2.5205 |
No | None | 100% | K99 |
| TNVGNGSGG | 2.8300 |
No | None | 100% | K99 |
| VNVNNTWMF |
1.0835 | No | None | 100% | spike |
| HYYVLPLTC |
0.9452 |
No | None | 100% | spike |
| GGIGTQPQK | 1.8178 | No | None | 100% | VP6 |
| Epitopes | Antigenicity score | Toxicity | Allergenicity | conservancy | pathogen |
|---|---|---|---|---|---|
| NNCTGGAEIRDLICV |
1.0165 |
No | None | 100% | Spike |
| YINLKDIGTYEYYVK |
1.0400 |
No | None | 100% | Spike |
| LTLSVLDATTESVLC |
0.9570 |
No | None | 100% | Vp6 |
| VREASRNGMQPQSPT | 0.4824 |
No | None | 100% | VP6 |
| GNGSGGANINTSFTT |
1.9647 |
No | None | 100% | K99 |
| RATNVGNGSGGANIN |
2.0509 |
No | None | 100% | K99 |
| Epitopes | Antigenicity score | Toxicity | Allergenicity | conservancy | Protein |
|---|---|---|---|---|---|
| KKDDRAPSNGGYKAGVFTTSA | 0.8975 | No | None | 100% | K99 |
| FCPCKLDGSLCVGNGPGIDAGYKNSGIGT |
0.5114 |
No | None | 100% | Spike |
| GRKVDLQLGNLGYLQSFNYRIDTT |
1.2781 |
No | None | 100% | Spike |
| FKTINFNNSSQSIKNWSAQSRRENPVYEYKNPMVFEYR | 0.9673 |
No | None | 100% | Vp6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




