Submitted:
15 April 2026
Posted:
17 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
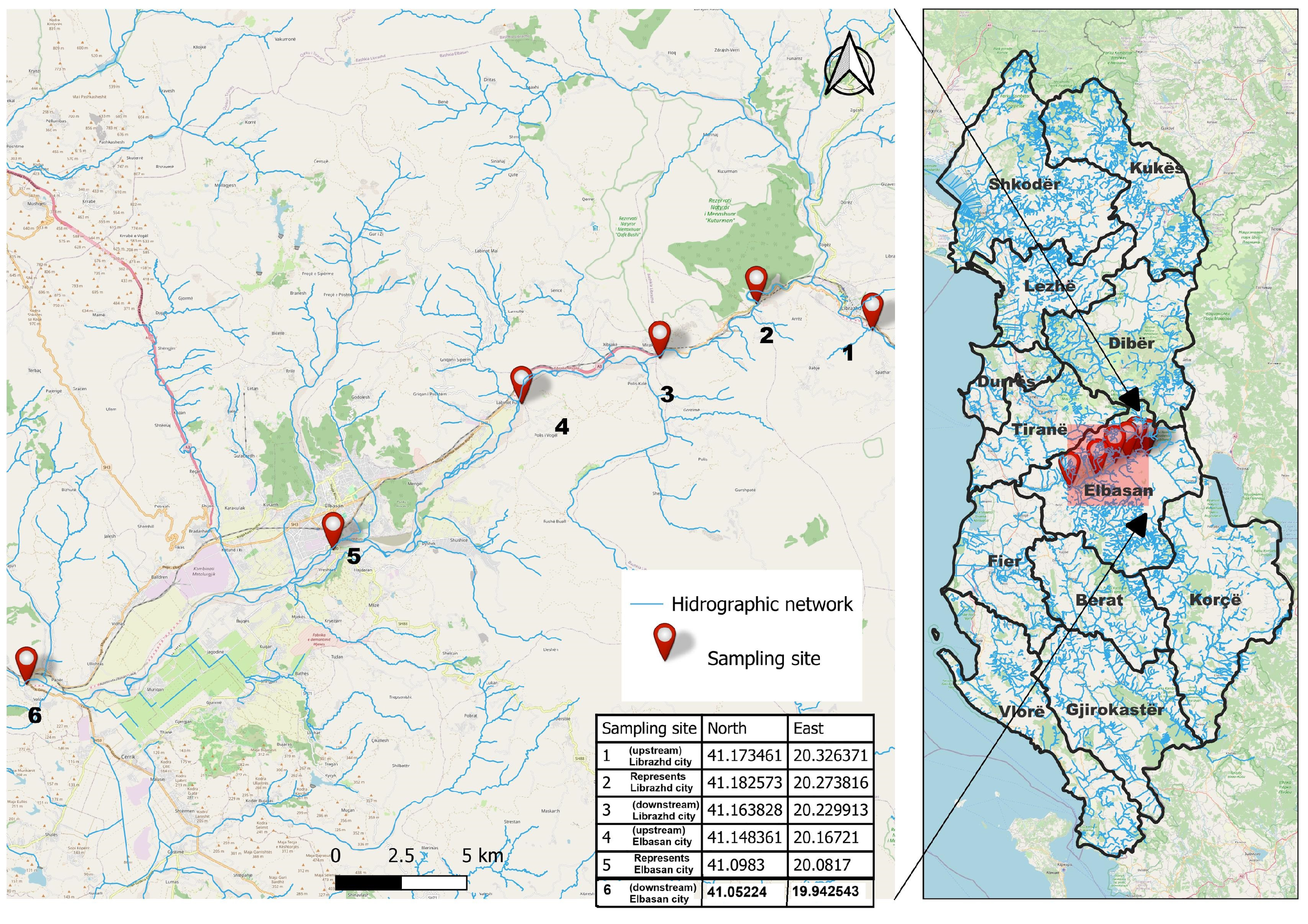
2.1. Description of the Study Location and Sample Collection
2.2. Collection of Isolates
2.3. Antimicrobial Susceptibility Profiles of ESBL-Ec
2.4. Assessment of Biofilm Formation Ability of ESBL-Ec
2.5. Pulsed-Field Gel Electrophoresis Genotyping
2.6. Detection of Resistance Genes and Phylogenetic Typing
3. Results
3.1. Prevalence of ESBL-Ec
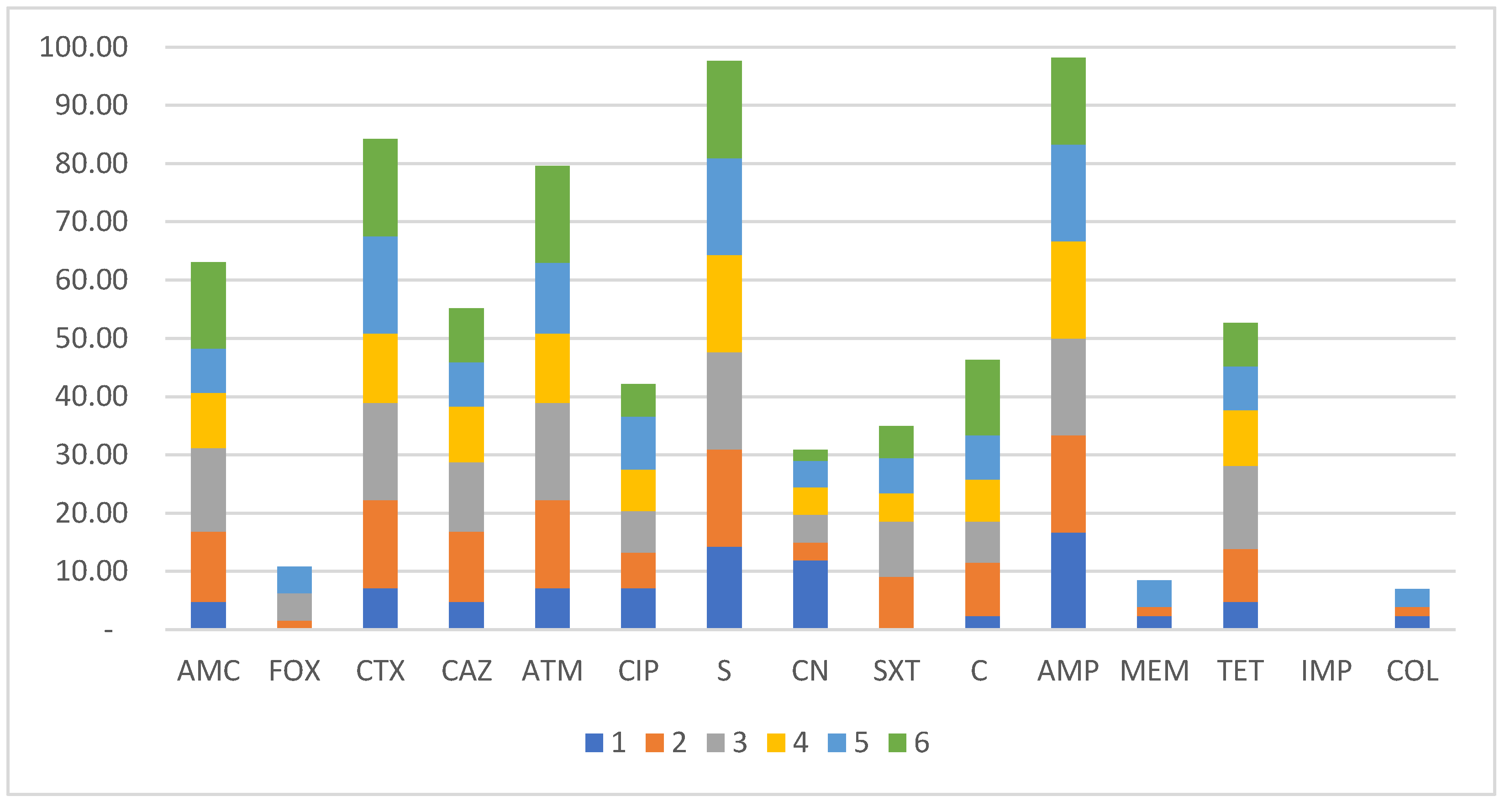
3.2. Phenotypic Characterization of ESBL-Ec-Antibiotic Resistance Profiling and Biofilm Formation Ability
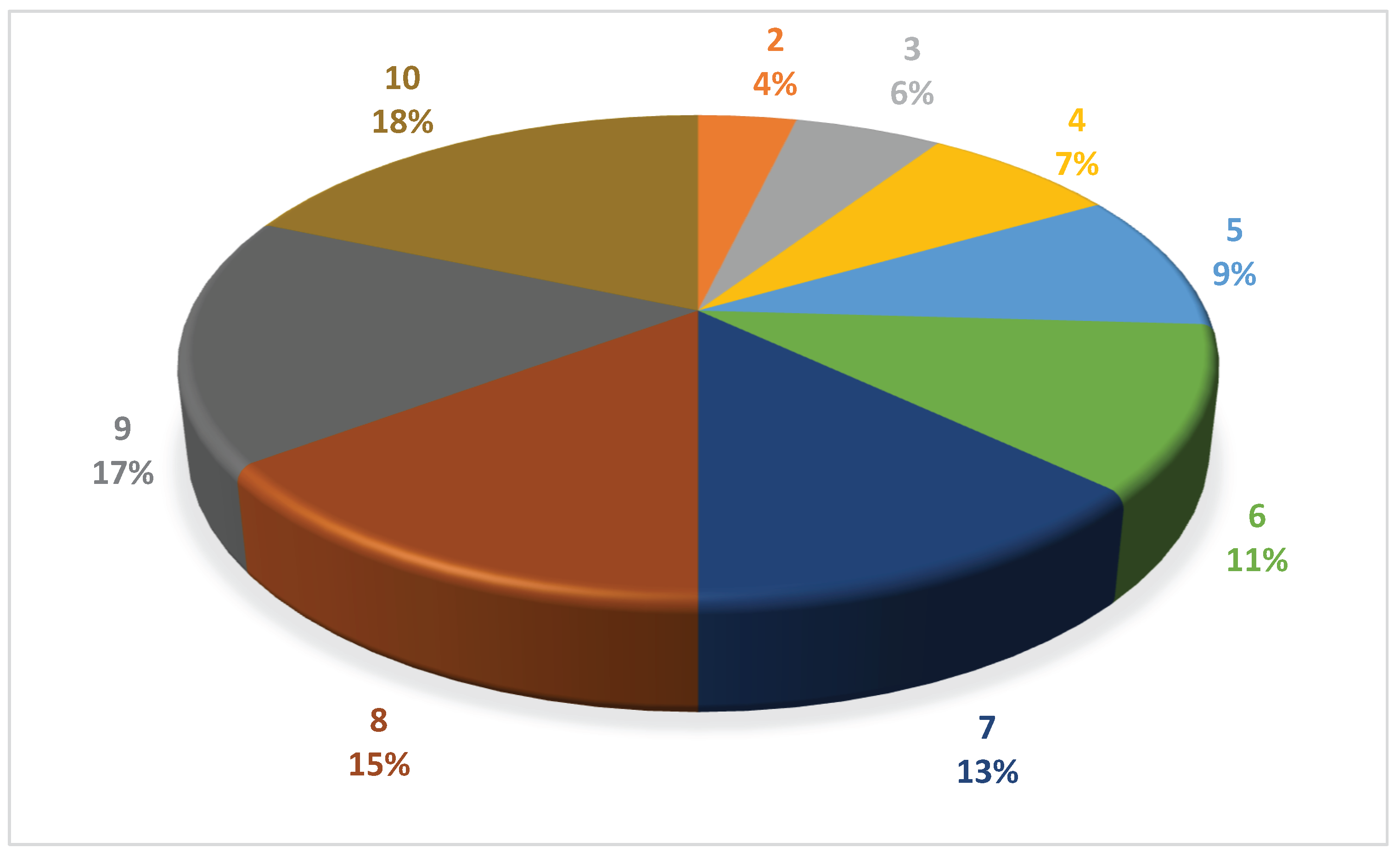
3.3. Distribution of ESBL Genotypes and Phylogenetic Groups
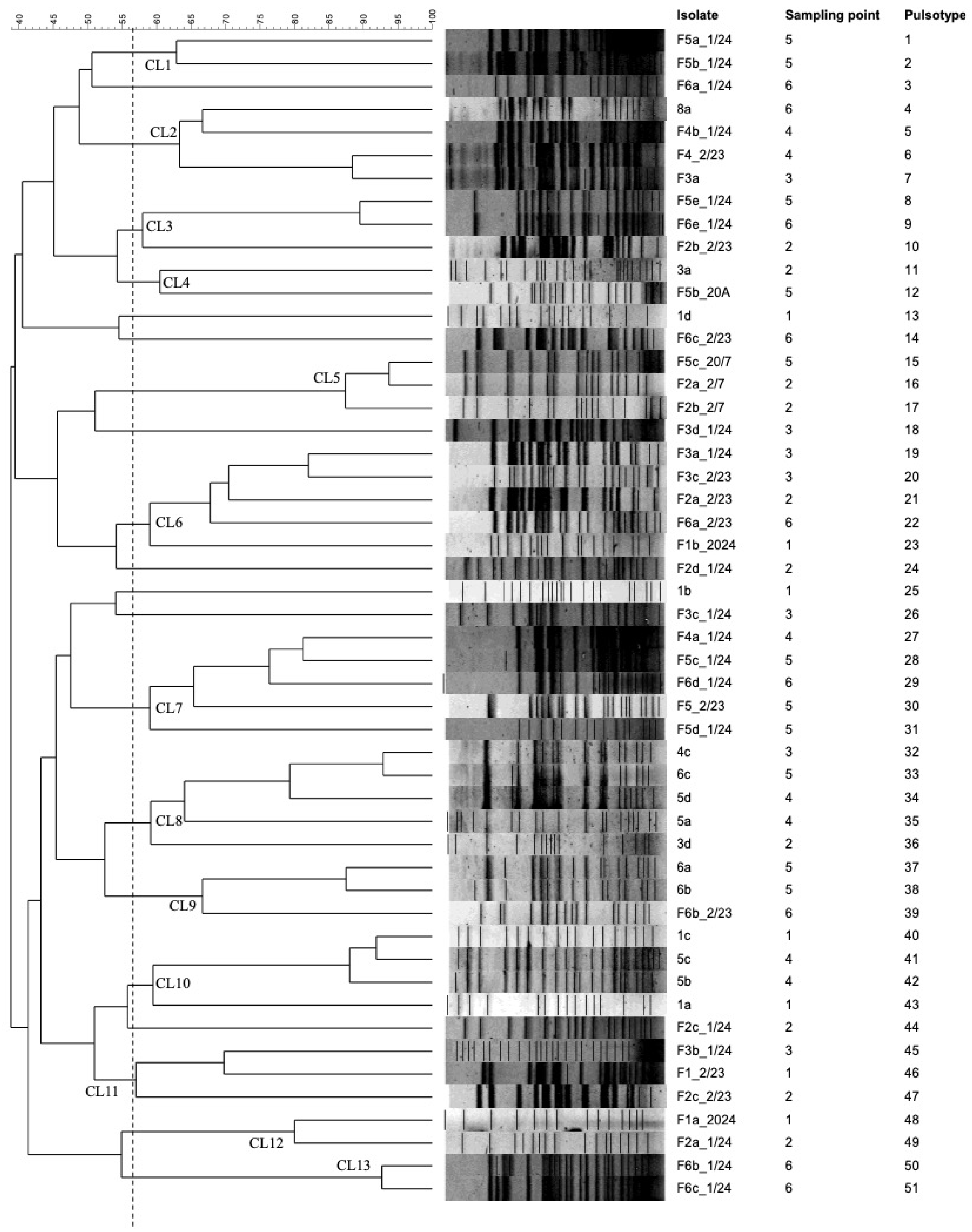
3.4. ESBL-Ec Strains Genetic Relatedness
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Naghavi, M.; Vollset, S.E.; Ikuta, K.S.; Swetschinski, L.R.; Gray, A.P.; Wool, E.E.; Robles Aguilar, G.; Mestrovic, T.; Smith, G.; Han, C.; et al. Global Burden of Bacterial Antimicrobial Resistance 1990–2021: A Systematic Analysis with Forecasts to 2050. The Lancet 2024, 404, 1199–1226. [Google Scholar] [CrossRef]
- Hooban, B.; Joyce, A.; Fitzhenry, K.; Chique, C.; Morris, D. The Role of the Natural Aquatic Environment in the Dissemination of Extended Spectrum Beta-Lactamase and Carbapenemase Encoding Genes: A Scoping Review. Water Res. 2020, 180. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31, 1–61. [Google Scholar] [CrossRef]
- Husna, A.; Rahman, M.M.; Badruzzaman, A.T.M.; Sikder, M.H.; Islam, M.R.; Rahman, M.T.; Alam, J.; Ashour, H.M. Extended-Spectrum β-Lactamases (ESBL): Challenges and Opportunities. Biomedicines 2023, 11. [Google Scholar] [CrossRef] [PubMed]
- Lindblom, A.; Kiszakiewicz, C.; Kristiansson, E.; Yazdanshenas, S.; Kamenska, N.; Karami, N.; Åhrén, C. The Impact of the ST131 Clone on Recurrent ESBL-Producing E. Coli Urinary Tract Infection: A Prospective Comparative Study. Sci. Rep. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Peirano, G.; Endimiani, A.; Pitout, J.D.D. CTX-M-Producing Escherichia Coli: History, Molecular Epidemiology and Laboratory Detection. Infect. Drug Resist. 2025, 18, 6549–6560. [Google Scholar] [CrossRef]
- MacNair, C.R.; Stokes, J.M.; Carfrae, L.A.; Fiebig-Comyn, A.A.; Coombes, B.K.; Mulvey, M.R.; Brown, E.D. Overcoming Mcr-1 Mediated Colistin Resistance with Colistin in Combination with Other Antibiotics. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef]
- Dadashi, M.; Sameni, F.; Bostanshirin, N.; Yaslianifard, S.; Khosravi-Dehaghi, N.; Nasiri, M.J.; Goudarzi, M.; Hashemi, A.; Hajikhani, B. Global Prevalence and Molecular Epidemiology of Mcr-Mediated Colistin Resistance in Escherichia Coli Clinical Isolates: A Systematic Review. J. Glob. Antimicrob. Resist. 2022, 29, 444–461. [Google Scholar] [CrossRef]
- Gharaibeh, M.H.; Shatnawi, S.Q. An Overview of Colistin Resistance, Mobilized Colistin Resistance Genes Dissemination, Global Responses, and the Alternatives to Colistin: A Review. Vet. World 2019, 12, 1735–1746. [Google Scholar] [CrossRef]
- Touati, A.; Ibrahim, N.A.; Mairi, A.; Kirat, H.; Basher, N.S.; Idres, T. One Health at Risk: Plasmid-Mediated Spread of Mcr-1 Across Clinical, Agricultural, and Environmental Ecosystems. Antibiotics 2025, 14. [Google Scholar] [CrossRef]
- Mmatli, M.; Mbelle, N.M.; Osei Sekyere, J. Global Epidemiology, Genetic Environment, Risk Factors and Therapeutic Prospects of Mcr Genes: A Current and Emerging Update. Front. Cell. Infect. Microbiol. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Maric, L.; Rupnik, M.; Janezic, S. Diversity of ESBL-Producing E. Coli in Various Water and Sediment Types. PLoS One 2025, 20. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, L.F.; Nespolo, N.M.; Rossi, G.A.M.; Fairbrother, J.M. Exploring Extended-Spectrum Beta-Lactamase (ESBL)-Producing Escherichia Coli in Food-Producing Animals and Animal-Derived Foods. Pathogens 2024, 13. [Google Scholar] [CrossRef] [PubMed]
- Zurfluh, K.; Hächler, H.; Nüesch-Inderbinen, M.; Stephan, R. Characteristics of Extended-Spectrum β-Lactamase- and Carbapenemase-Producing Enterobacteriaceae Isolates from Rivers and Lakes in Switzerland. Appl. Environ. Microbiol. 2013, 79, 3021–3026. [Google Scholar] [CrossRef]
- Kutilova, I.; Medvecky, M.; Leekitcharoenphon, P.; Munk, P.; Masarikova, M.; Davidova-Gerzova, L.; Jamborova, I.; Bortolaia, V.; Pamp, S.J.; Dolejska, M. Extended-Spectrum Beta-Lactamase-Producing Escherichia Coli and Antimicrobial Resistance in Municipal and Hospital Wastewaters in Czech Republic: Culture-Based and Metagenomic Approaches. Environ. Res. 2021, 193. [Google Scholar] [CrossRef]
- Cho, S.; Jackson, C.R.; Frye, J.G. Freshwater Environment as a Reservoir of Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae. J. Appl. Microbiol. 2023, 134. [Google Scholar] [CrossRef]
- Meena, P.R.; Priyanka, P.; Singh, A.P. Extraintestinal Pathogenic Escherichia Coli (ExPEC) Reservoirs, and Antibiotics Resistance Trends: A One-Health Surveillance for Risk Analysis from “Farm-to-Fork.”. Lett. Appl. Microbiol. 2023, 76. [Google Scholar] [CrossRef]
- Chen, P.A.; Hung, C.H.; Huang, P.C.; Chen, J.R.; Huang, I.F.; Chen, W.L.; Chiou, Y.H.; Hung, W.Y.; Wang, J.L.; Cheng, M.F. Characteristics of CTX-M Extendedspectrum β-Lactamase-Producing Escherichia Coli Strains Isolated from Multiple Rivers in Southern Taiwan. Appl. Environ. Microbiol. 2016, 82, 1889–1897. [Google Scholar] [CrossRef]
- Rozwandowicz, M.; Brouwer, M.S.M.; Fischer, J.; Wagenaar, J.A.; Gonzalez-Zorn, B.; Guerra, B.; Mevius, D.J.; Hordijk, J. Plasmids Carrying Antimicrobial Resistance Genes in Enterobacteriaceae. Journal of Antimicrobial Chemotherapy 2018, 73, 1121–1137. [Google Scholar] [CrossRef]
- WHO Integrated Global Surveillance on ESBL-Producing E. coli Using a “One Health” Approach. Available online: https://www.who.int/publications/i/item/who-integrated-global-surveillance-on-esbl-producing-e.-coli-using-a-one-health-approach (accessed on 20 February 2026).
- The European Committee on Antimicrobial Susceptibility Testing. EUCAST guidelines for detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance, Version 2.0. Available online: https://www.eucast.org/bacteria/important-additional-information/resistance-detection/ (accessed on 20 February 2026).
- The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters, Version 16.0. Available online: http://www.eucast.org/clinical_breakpoints<b>/</b> (accessed on 23 February 2026).
- Clinical and Laboratory Standards Institute. M07-A9: Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard, 9th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2018; ISBN 1-56238-783-9. [Google Scholar]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-Resistant, Extensively Drug-Resistant and Pandrug-Resistant Bacteria: An International Expert Proposal for Interim Standard Definitions for Acquired Resistance. Clinical Microbiology and Infection 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Stepanovic, S.; Vukovic, D.; Hola, V.; DI Bonaventura, G.; Djukic, S.; Ruzicka, F.; Bonaventura, D.G. The Authors Printed in Denmark. All rights reserved Journal Compilation C 2007, 115, 891–900. [Google Scholar]
- Borges, S.; Silva, J.; Teixeira, P. Survival and Biofilm Formation by Group B Streptococci in Simulated Vaginal Fluid at Different PHs. Antonie van Leeuwenhoek, International Journal of General and Molecular Microbiology 2012, 101, 677–682. [Google Scholar] [CrossRef] [PubMed]
- Sonnei, Shigella; Flexneri, Shigella; CDC PulseNet. Standard Operating Procedure for PulseNet PFGE of Escherichia coli O157: H7, Escherichia coli non-O157 (STEC), Salmonella Serotypes; PNL05 Centers for Disease Control and Prevention: Atlanta, GA, USA, 2017; Volume 157, pp. 1–16. [Google Scholar]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia Coli Phylo-Typing Method Revisited: Improvement of Specificity and Detection of New Phylo-Groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Dallenne, C.; da Costa, A.; Decré, D.; Favier, C.; Arlet, G. Development of a Set of Multiplex PCR Assays for the Detection of Genes Encoding Important β-Lactamases in Enterobacteriaceae. Journal of Antimicrobial Chemotherapy 2010, 65, 490–495. [Google Scholar] [CrossRef]
- Rebelo, A.R.; Bortolaia, V.; Kjeldgaard, J.S.; Pedersen, S.K.; Leekitcharoenphon, P.; Hansen, I.M.; Guerra, B.; Malorny, B.; Borowiak, M.; Hammerl, J.A.; et al. Multiplex PCR for Detection of Plasmid-Mediated Colistin Resistance Determinants, Mcr-1, Mcr-2, Mcr-3, Mcr-4 and Mcr-5 for Surveillance Purposes. Eurosurveillance 2018, 23. [Google Scholar] [CrossRef]
- Bekteshi, L.; Hoxha, B.; Gega, N.; Karamelo, P. THE IMPACT OF URBAN AND INDUSTRIAL ACTIVITY IN THE POLLUTION OF SHKUMBINI RIVER. International Conference on Applied Engineering and Natural Sciences 2023, 1, 1066–1070. [Google Scholar] [CrossRef]
- Sulce, S.; Rroco, E.; Malltezi, J.; Shallari, S.; Libohova, Z.; Sinaj, S.; Qafoku, N. Water Quality in Albania: An Overview of Sources of Contamination and Controlling Factors. Albanian j. agric. sci. Special edition – Proceedings of ICOALS 2018, 2, 279–297. [Google Scholar]
- Hoxha, I.; Malaj, A.; Kraja, B.; Bino, S.; Oluka, M.; Marković-Peković, V.; Godman, B. Are Pharmacists’ Good Knowledge and Awareness on Antibiotics Taken for Granted? The Situation in Albania and Future Implications across Countries. J. Glob. Antimicrob. Resist. 2018, 13, 240–245. [Google Scholar] [CrossRef]
- Fagerström, A.; Mölling, P.; Khan, F.A.; Sundqvist, M.; Jass, J.; Söderquist, B. Comparative Distribution of Extended-Spectrum Beta-Lactamase–Producing Escherichia Coli from Urine Infections and Environmental Waters. PLoS One 2019, 14. [Google Scholar] [CrossRef]
- Robertson, J.; Iwamoto, K.; Hoxha, I.; Ghazaryan, L.; Abilova, V.; Cvijanovic, A.; Pyshnik, H.; Darakhvelidze, M.; Makalkina, L.; Jakupi, A.; et al. Antimicrobial Medicines Consumption in Eastern Europeand Central Asia – An Updated Cross-National Study and Assessment of QuantitativeMetrics for Policy Action. Front. Pharmacol. 2019, 9. [Google Scholar] [CrossRef]
- Aguilar-Salazar, A.; Martínez-Vázquez, A.V.; Aguilera-Arreola, G.; de Jesus de Luna-Santillana, E.; Cruz-Hernández, M.A.; Escobedo-Bonilla, C.M.; Lara-Ramírez, E.; Sánchez-Sánchez, M.; Guerrero, A.; Rivera, G.; et al. Prevalence of ESKAPE Bacteria in Surface Water and Wastewater Sources: Multidrug Resistance and Molecular Characterization, an Updated Review. Water (Switzerland) 2023, 15. [Google Scholar] [CrossRef]
- Van Duijkeren, E.; Schwarz, C.; Bouchard, D.; Catry, B.; Pomba, C.; Baptiste, K.E.; Moreno, M.A.; Rantala, M.; Ružauskas, M.; Sanders, P.; et al. The Use of Aminoglycosides in Animals within the EU: Development of Resistance in Animals and Possible Impact on Human and Animal Health: A Review. Journal of Antimicrobial Chemotherapy 2019, 74, 2480–2496. [Google Scholar] [CrossRef] [PubMed]
- Sati, H.; Carrara, E.; Savoldi, A.; Hansen, P.; Garlasco, J.; Campagnaro, E.; Boccia, S.; Castillo-Polo, J.A.; Magrini, E.; Garcia-Vello, P.; et al. The WHO Bacterial Priority Pathogens List 2024: A Prioritisation Study to Guide Research, Development, and Public Health Strategies against Antimicrobial Resistance. Lancet Infect. Dis. 2025, 25, 1033–1043. [Google Scholar] [CrossRef]
- van Duin, D.; Doi, Y. The Global Epidemiology of Carbapenemase-Producing Enterobacteriaceae. Virulence 2017, 8, 460–469. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile Genetic Elements Associated with Antimicrobial Resistance. Clin. Microbiol. Rev. 2018, 31. [Google Scholar] [CrossRef]
- Dantas Palmeira, J.; do Arte, I.; Ragab Mersal, M.M.; Carneiro da Mota, C.; Ferreira, H.M.N. KPC-Producing Enterobacterales from Douro River, Portugal—Persistent Environmental Contamination by Putative Healthcare Settings. Antibiotics 2023, 12. [Google Scholar] [CrossRef]
- Nobili, G.; Cocomazzi, A.; Basanisi, M.G.; Damato, A.M.; Coppola, R.; Cariglia, M.G.; Franconieri, I.; Stallone, A.; Notarangelo, M.; Scirocco, T.; et al. Occurrence and Genomic Characterization of ESBL-, AmpC-, and Carbapenemase-Producing Escherichia Coli and Klebsiella Pneumoniae Isolated from Surface Water in Southern Italy, 2023–2024. Microorganisms 2026, 14. [Google Scholar] [CrossRef]
- Falgenhauer, L.; zur Nieden, A.; Harpel, S.; Falgenhauer, J.; Domann, E. Clonal CTX-M-15-Producing Escherichia Coli ST-949 Are Present in German Surface Water. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Çomo, E.; Hasimi, A.; Tako, E.; Ormeni, R. Evaluation of Seasonal Distribution of Heavy Metals in the Water of Shkumbini River, Albania. International Journal of Environmental Science and Development 2024, 15, 140–145. [Google Scholar] [CrossRef]
- Shyti, B.; Bekteshi, L.; Paralloj, S.; Hila, E. Remodeling of the WQI Index for the Evaluation of the Shkumbini River’s Water Quality in Albania Using the Statistical Method. Ecologia Balkanica 2024, 16, 58–67. [Google Scholar]
- Alcaine, S.D.; Molla, L.; Nugen, S.R.; Kruse, H. Results of a Pilot Antibiotic Resistance Survey of Albanian Poultry Farms. J. Glob. Antimicrob. Resist. 2016, 4, 60–64. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.J.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. The Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef] [PubMed]
- Parascandalo, F.A.; Zarb, P.; Tartari, E.; Lacej, D.; Bitincka, S.; Manastirliu, O.; Nika, D.; Borg, M.A. Carriage of Multidrug-Resistant Organisms in a Tertiary University Hospital in Albania-a Point Prevalence Survey. Antimicrob. Resist. Infect. Control 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Girlich, D.; Bonnin, R.A.; Naas, T. Occurrence and Diversity of CTX-M-Producing Escherichia Coli From the Seine River. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef]
- Zarfel, G.; Lipp, M.; Gürtl, E.; Folli, B.; Baumert, R.; Kittinger, C. Troubled Water under the Bridge: Screening of River Mur Water Reveals Dominance of CTX-M Harboring Escherichia Coli and for the First Time an Environmental VIM-1 Producer in Austria. Science of the Total Environment 2017, 593–594, 399–405. [Google Scholar] [CrossRef]
- Kotzamanidis, C.; Malousi, A.; Paraskeva, A.; Vafeas, G.; Giantzi, V.; Hatzigiannakis, E.; Dalampakis, P.; Kinigopoulou, V.; Vrouhakis, I.; Zouboulis, A.; et al. River Waters in Greece: A Reservoir for Clinically Relevant Extended-Spectrum-β-Lactamases-Producing Escherichia Coli. Science of the Total Environment 2024, 941. [Google Scholar] [CrossRef]
- Berendonk, T.U.; Manaia, C.M.; Merlin, C.; Fatta-Kassinos, D.; Cytryn, E.; Walsh, F.; Bürgmann, H.; Sørum, H.; Norström, M.; Pons, M.N.; et al. Tackling Antibiotic Resistance: The Environmental Framework. Nat. Rev. Microbiol. 2015, 13, 310–317. [Google Scholar] [CrossRef]
- Gambero, M.L.; Blarasin, M.; Bettera, S.; Albo, J.G. Genetic Diversity of Escherichia Coli Isolates from Surface Water and Groundwater in a Rural Environment. J. Water Health 2017, 15, 757–765. [Google Scholar] [CrossRef]
- Franz, E.; Veenman, C.; Van Hoek, A.H.A.M.; Husman, A.D.R.; Blaak, H. Pathogenic Escherichia Coli Producing Extended-Spectrum β-Lactamases Isolated from Surface Water and Wastewater. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef]
- Clermont, O.; Gordon, D.; Denamur, E. Guide to the Various Phylogenetic Classification Schemes for Escherichia Coli and the Correspondence among Schemes. Microbiology (United Kingdom) 2015, 161, 980–988. [Google Scholar] [CrossRef]
- Abe, K.; Nomura, N.; Suzuki, S. Biofilms: Hot Spots of Horizontal Gene Transfer (HGT) in Aquatic Environments, with a Focus on a New HGT Mechanism. FEMS Microbiol. Ecol. 2021, 96. [Google Scholar] [CrossRef]
- Soto, S.M. Importance of Biofilms in Urinary Tract Infections: New Therapeutic Approaches. Adv. Biol. 2014, 2014, 1–13. [Google Scholar] [CrossRef]
- Chandran, A.; Mazumder, A. Pathogenic Potential, Genetic Diversity, and Population Structure of Escherichia Coli Strains Isolated from a Forest-Dominated Watershed (Comox Lake) in British Columbia, Canada. Appl. Environ. Microbiol. 2015, 81, 1788–1798. [Google Scholar] [CrossRef]
| Strain | ARP typea | Sampling point 1 (n=7) |
Sampling point 2 (n=11) |
Sampling point 3 (n=7) |
Sampling point 4 (n=7) |
Sampling point 5 (n=11) |
Sampling point 6 (n=9) |
| 5c | S AMP | 1 (14.3%) | |||||
| 1b | CIP CN AMP | 1 (14.3%) | |||||
| 1c, 1d | S CN AMP | 2 (28.6%) | |||||
| 1a | S CN AMP MEM IMP | 1 (14.3%) | |||||
| 3a | ATM S CN AMP | 1 (9.1%) | |||||
| 5b | CAZ S AMP TET | 1 (14.3%) | |||||
| 6a | CTX S CN AMP | 1 (9.1%) | |||||
| F5b_1/24 | CTX CIP S AMP | 1 (9.1%) | |||||
| F5c_1/24 | CTX ATM S AMP | 1 (9.1%) | |||||
| 3d | AMC CTX CAZ S AMP | 1 (9.1%) | |||||
| 8a | CTX ATM S CN TET | 1 (11,1%) | |||||
| F2b_1/24, F3a_1/24, F6c_1/24 | AMC CTX ATM S AMP | 1 (9.1%) | 1 (14.3%) | 1 (11,1%) | |||
| F2c_1/24 | CTX ATM S SXT AMP | 1 (9.1%) | |||||
| F5d_1/24, F6a_1/24, F6e_1/24 | AMC CTX ATM S C AMP | 1 (9.1%) | 2 (22.2%) | ||||
| F5e_1/24 | CTX CAZ ATM S AMP TET | 1 (9.1%) | |||||
| F4a_1/24 | AMC CTX CAZ ATM S AMP | 1 (14.3%) | |||||
| 6b | CTX CAZ ATM S AMP MEM | 1 (9.1%) | |||||
| 6c | CTX CIP S C AMP TET | 1 (9.1%) | |||||
| F1b_2024 | AMC CTX CAZ ATM CIP S AMP | 1 (14.3%) | |||||
| F3b_1/24 | CTX CAZ ATM S SXT AMP TET | 1 (14.3%) | |||||
| F4b_1/24 | AMC CTX CAZ ATM S CN AMP | 1 (14.3%) | |||||
| F6d_1/24 | AMC CTX CAZ ATM S C AMP TET | 1 (11,1%) | |||||
| 4c | AMC CTX ATM CIP S C AMP TET | 1 (14.3%) | |||||
| 5d | CTX ATM CIP S CN C AMP TET | 1 (14.3%) | |||||
| F1_2/23 | CTX CAZ ATM S C AMP TET COL | 1 (14.3%) | |||||
| F2a_2/23 | AMC CTX CAZ ATM CIP S C AMP | 1 (9.1%) | |||||
| F2c_2/23 | CTX CAZ ATM S SXT C AMP TET | 1 (9.1%) | |||||
| F1a_2024 | AMC CTX ATM CIP S CN AMP TET | 1 (14.3%) | |||||
| F6b_1/24 | AMC CTX CAZ ATM S C AMP TET | 1 (11,1%) | |||||
| 5a | AMC CTX ATM CIP S SXT C AMP TET | 1 (14.3%) | |||||
| F6a_2/23 | AMC CTX CAZ ATM CIP S SXT C AMP | 1 (11,1%) | |||||
| F3a | AMC CTX CAZ ATM CIP S SXT AMP TET | 1 (14.3%) | |||||
| F2a_1/24 | AMC CTX CAZ ATM S SXT C AMP TET | 1 (9.1%) | |||||
| F2d_1/24 | AMC CTX CAZ ATM CIP S SXT AMP TET | 1 (9.1%) | |||||
| F2b_2/23 | AMC CTX CAZ ATM S CN SXT C AMP TET | 1 (9.1%) | |||||
| F3c_2/23, F5_2/23, F6b_2/23, F6c_2/23, F4_2/23, F2a_2/7 |
AMC CTX CAZ ATM CIP S SXT C AMP TET | 1 (9.1%) | 1 (14.3%) | 1 (14.3%) | 1 (9.1%) | 2 (22.2%) | |
| F3c_1/24 | AMC FOX CTX CAZ ATM S CN SXT AMP TET | 1 (14.3%) | |||||
| F3d_1/24 | AMC FOX CTX CAZ ATM S CN C AMP TET | 1 (14.3%) | |||||
| F5a_1/24 | AMC FOX CTX CAZ ATM CIP S CN SXT AMP | 1 (9.1%) | |||||
| F2b_2/7 | AMC FOX CTX CAZ ATM CIP S C AMP MEM TET COL | 1 (9.1%) | |||||
| F5c_20/7 | AMC FOX CTX ATM CIP S SXT C AMP MEM TET COL | 1 (9.1%) | |||||
| F5b_20A | AMC FOX CTX CAZ ATM CIP S CN SXT C AMP MEM TET COL | 1 (9.1%) |
| Sampling point | Period of sampling | Isolates | Antimicrobial resistance profilea | Biofilmb | Phylogenetic groupc | Antimicrobial resistance genes (PCR)d | |||||||||||||||||||||
| AMC | FOX | CTX | CAZ | ATM | CIP | S | CN | SXT | C | AMP | MEM | TET | IMP | COL | OXA family | SHV family | TEM family | CTX-M group 1 | CTX-M group 2 | CTX-M group 9 | CTX-M group 8/25 | mcr-1,-2,-3,-4,-5 | |||||
| 1 | September2022 | 1a | S | S | S | S | S | S | R | R | S | S | R | R | S | R | S | M | A | ||||||||
| 1b | S | S | S | S | S | R | S | R | S | S | R | S | S | S | S | W | A | ||||||||||
| 1c | S | S | S | S | S | S | R | R | S | S | R | S | S | S | S | W | C | ||||||||||
| 1d | S | S | S | S | S | S | R | R | S | S | R | S | S | S | S | W | A | ||||||||||
| 2 | 3a | S | S | S | S | R | S | R | R | S | S | R | S | S | S | S | W | A | |||||||||
| 3d | R | S | R | R | S | S | R | S | S | S | R | S | S | S | S | NB | A | ||||||||||
| 3 | 4c | R | S | R | S | R | R | R | S | S | R | R | S | R | S | S | W | C | |||||||||
| 4 | 5a | R | S | R | S | R | R | R | S | R | R | R | S | R | S | S | W | C | |||||||||
| 5b | S | S | S | R | S | S | R | S | S | S | R | S | R | S | S | NB | A | ||||||||||
| 5c | S | S | S | S | S | S | R | S | S | S | R | S | S | S | S | NB | A | ||||||||||
| 5d | S | S | R | S | R | R | R | R | S | R | R | S | R | S | S | NB | A | ||||||||||
| 5 | 6a | S | S | R | S | S | S | R | R | S | s | R | S | S | S | S | NB | A | |||||||||
| 6b | S | S | R | R | R | S | R | S | S | S | R | R | S | S | S | W | A | ||||||||||
| 6c | S | S | R | S | S | R | R | S | S | R | R | S | R | S | S | W | A | ||||||||||
| 6 | 8a | S | S | R | S | R | S | R | R | S | S | S | S | R | S | S | NB | B2 | |||||||||
| 1 | February 2023 | F1_2/23 | S | S | R | R | R | S | R | S | S | R | R | S | R | S | R | W | D | ||||||||
| 2 | F2a_2/23 | R | S | R | R | R | R | R | S | S | R | R | S | S | S | S | NB | B2 | |||||||||
| F2b_2/23 | R | S | R | R | R | S | R | R | R | R | R | S | R | S | S | NB | B2 | ||||||||||
| F2c_2/23 | S | S | R | R | R | S | R | S | R | R | R | S | R | S | S | NB | A | ||||||||||
| 3 | F3c_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | W | B2 | |||||||||
| 5 | F5_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | NB | A | |||||||||
| 6 | F6a_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | S | S | S | W | B2 | |||||||||
| F6b_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | W | A | ||||||||||
| F6c_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | NB | A | ||||||||||
| 4 | F4_2/23 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | W | B2 | |||||||||
| 5 | F5b_20A | R | R | R | R | R | R | R | R | R | R | R | R | R | S | R | W | A | |||||||||
| 2 | June 2023 | F2a_2/7 | R | S | R | R | R | R | R | S | R | R | R | S | R | S | S | W | D | ||||||||
| F2b_2/7 | R | R | R | R | R | R | R | S | S | R | R | R | R | S | R | NB | D | ||||||||||
| 3 | F3a | R | S | R | R | R | R | R | S | R | S | R | S | R | S | S | W | B2 | |||||||||
| 5 | F5c_20/7 | R | R | R | S | R | R | R | S | R | R | R | R | R | S | R | M | D | |||||||||
| 1 | February 2024 | F1a_2024 | R | S | R | S | R | R | R | R | S | S | R | S | R | S | S | W | A | ||||||||
| F1b_2024 | R | S | R | R | R | R | R | S | S | S | R | S | S | S | S | NB | B2 | ||||||||||
| 2 | F2a_1/24 | R | S | R | R | R | S | R | S | R | R | R | S | R | S | S | NB | A | |||||||||
| F2b_1/24 | R | S | R | S | R | S | R | S | S | S | R | S | S | S | S | NB | B1 | ||||||||||
| F2c_1/24 | S | S | R | S | R | S | R | S | R | S | R | S | S | S | S | W | D | ||||||||||
| F2d_1/24 | R | S | R | R | R | R | R | S | R | S | R | S | R | S | S | W | E | ||||||||||
| 3 | F3a_1/24 | R | S | R | S | R | S | R | S | S | S | R | S | S | S | S | W | B1 | |||||||||
| F3b_1/24 | S | S | R | R | R | S | R | S | R | S | R | S | R | S | S | W | A | ||||||||||
| F3c_1/24 | R | R | R | R | R | S | R | R | R | S | R | S | R | S | S | W | A | ||||||||||
| F3d_1/24 | R | R | R | R | R | S | R | R | S | R | R | S | R | S | S | NB | E | ||||||||||
| 4 | F4a_1/24 | R | S | R | R | R | S | R | S | S | S | R | S | S | S | S | M | A | |||||||||
| F4b_1/24 | R | S | R | R | R | S | R | R | S | S | R | S | S | S | S | W | B2 | ||||||||||
| 5 | F5a_1/24 | R | R | R | R | R | R | R | R | R | S | R | S | S | S | S | NB | B2 | |||||||||
| F5b_1/24 | S | S | R | S | S | R | R | S | S | S | R | S | S | S | S | W | B2 | ||||||||||
| F5c_1/24 | S | S | R | S | R | S | R | S | S | S | R | S | S | S | S | NB | A | ||||||||||
| F5d_1/24 | R | S | R | S | R | S | R | S | S | R | R | S | S | S | S | W | B2 | ||||||||||
| F5e_1/24 | S | S | R | R | R | S | R | S | S | S | R | S | R | S | S | NB | A | ||||||||||
| 6 | F6a_1/24 | R | S | R | S | R | S | R | S | S | R | R | S | S | S | S | NB | B1 | |||||||||
| F6b_1/24 | R | S | R | R | R | S | R | S | S | R | R | S | R | S | S | NB | B1 | ||||||||||
| F6c_1/24 | R | S | R | S | R | S | R | S | S | S | R | S | S | S | S | M | B1 | ||||||||||
| F6d_1/24 | R | S | R | R | R | S | R | S | S | R | R | S | S | S | S | NB | A | ||||||||||
| F6e_1/24 | R | S | R | S | R | S | R | S | S | R | R | S | S | S | S | NB | D | ||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




