Submitted:
15 April 2026
Posted:
16 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. DNA Methylation and Epigenetic Inheritance
1.2. Evolutionary Implications of 5mC
1.3. The Kingdom of Fungi as a Natural Experiment
1.4. Recent Advances and Scope
2. Cytosine Methylation: Mechanisms and Functions
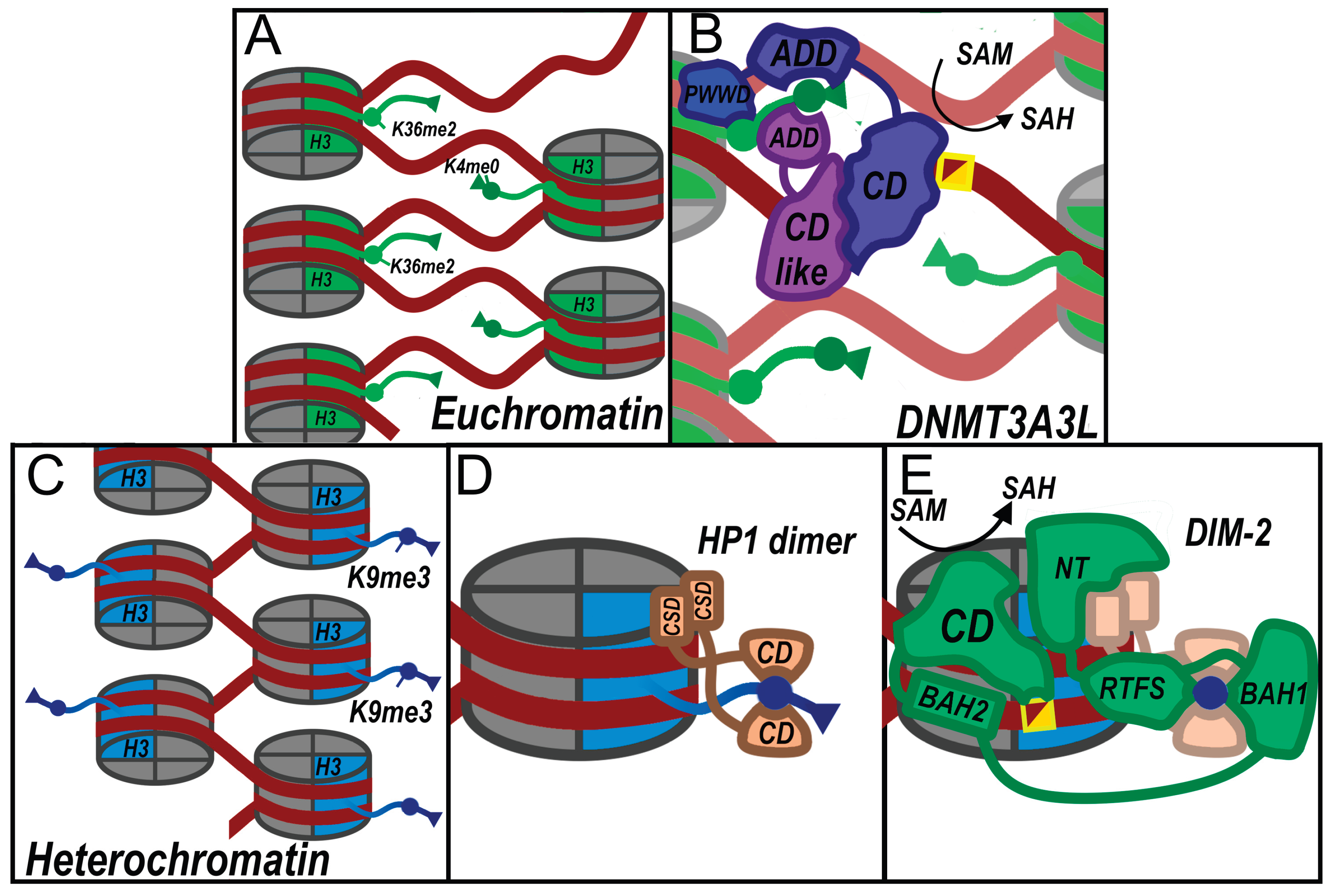
2.1. Transposon Silencing, RIP, and the De Novo DIM-2 Methylation Cascade
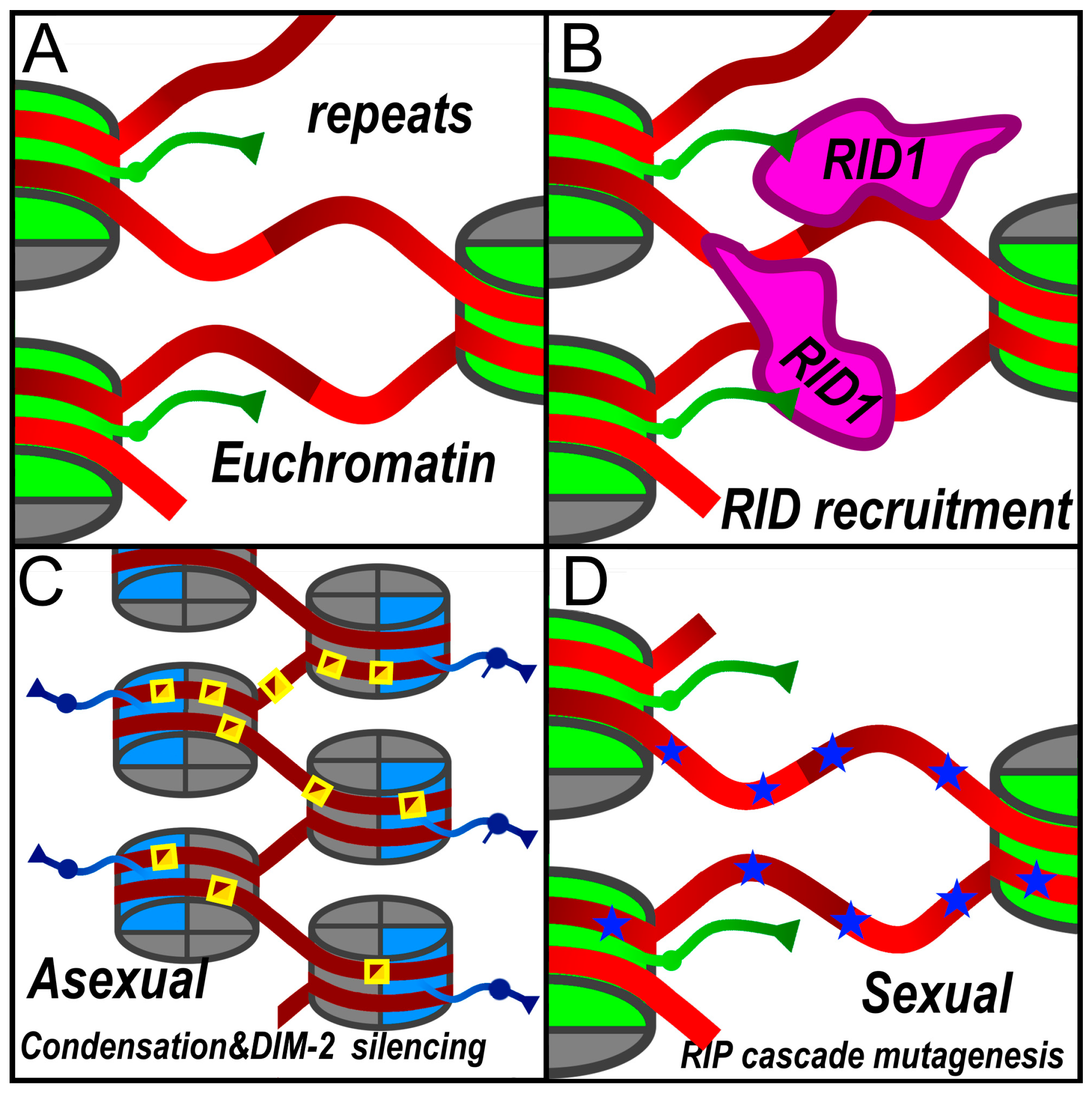
2.2. RID: Bifunctional Initiator of RIP and Heterochromatin Formation
2.3. Antiviral Protection: Methylation in Vegetative Hyphae Directed by RNA Interference
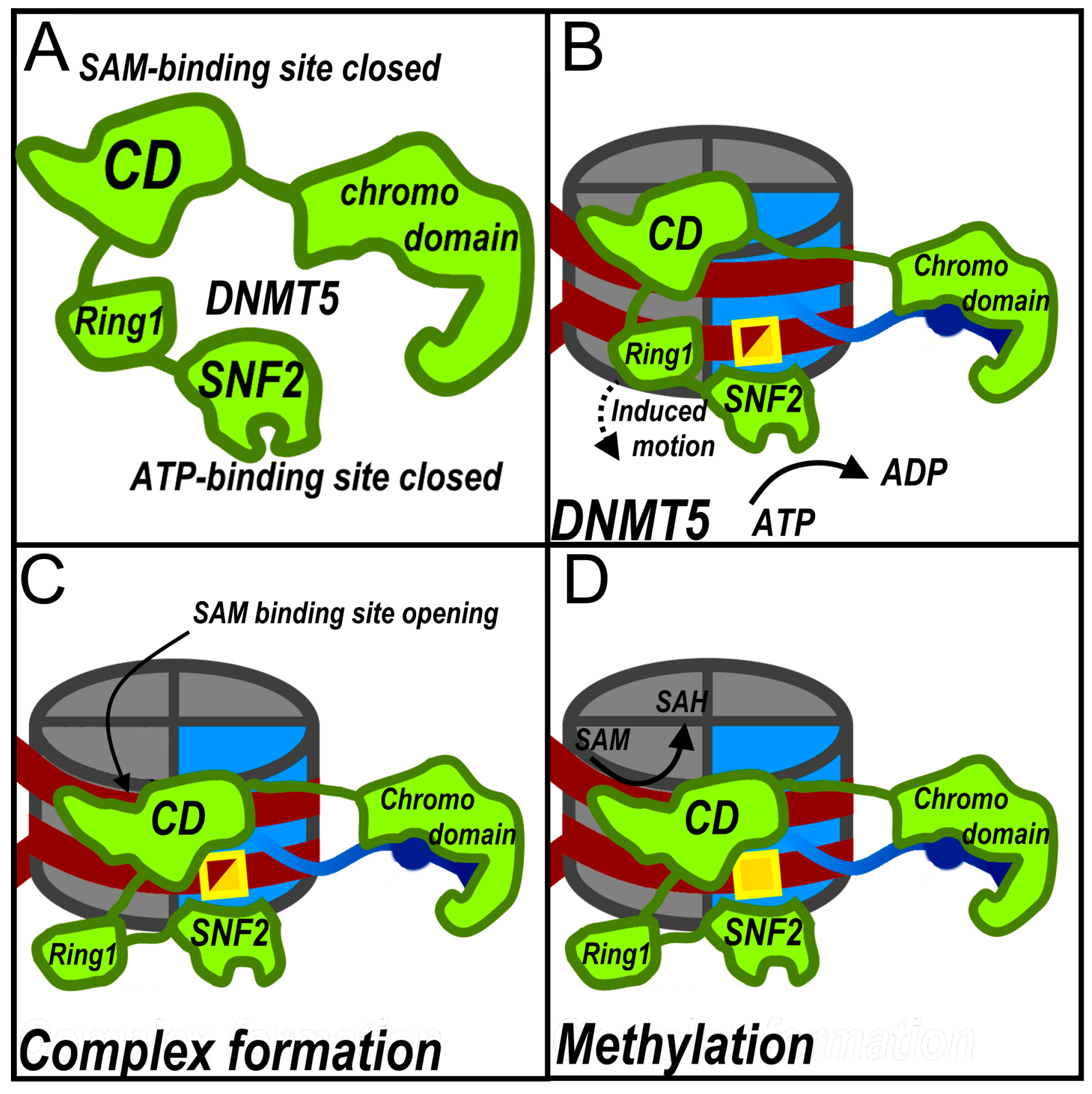
2.4. DNMT5: A High-Fidelity ATP-Dependent Methyltransferase
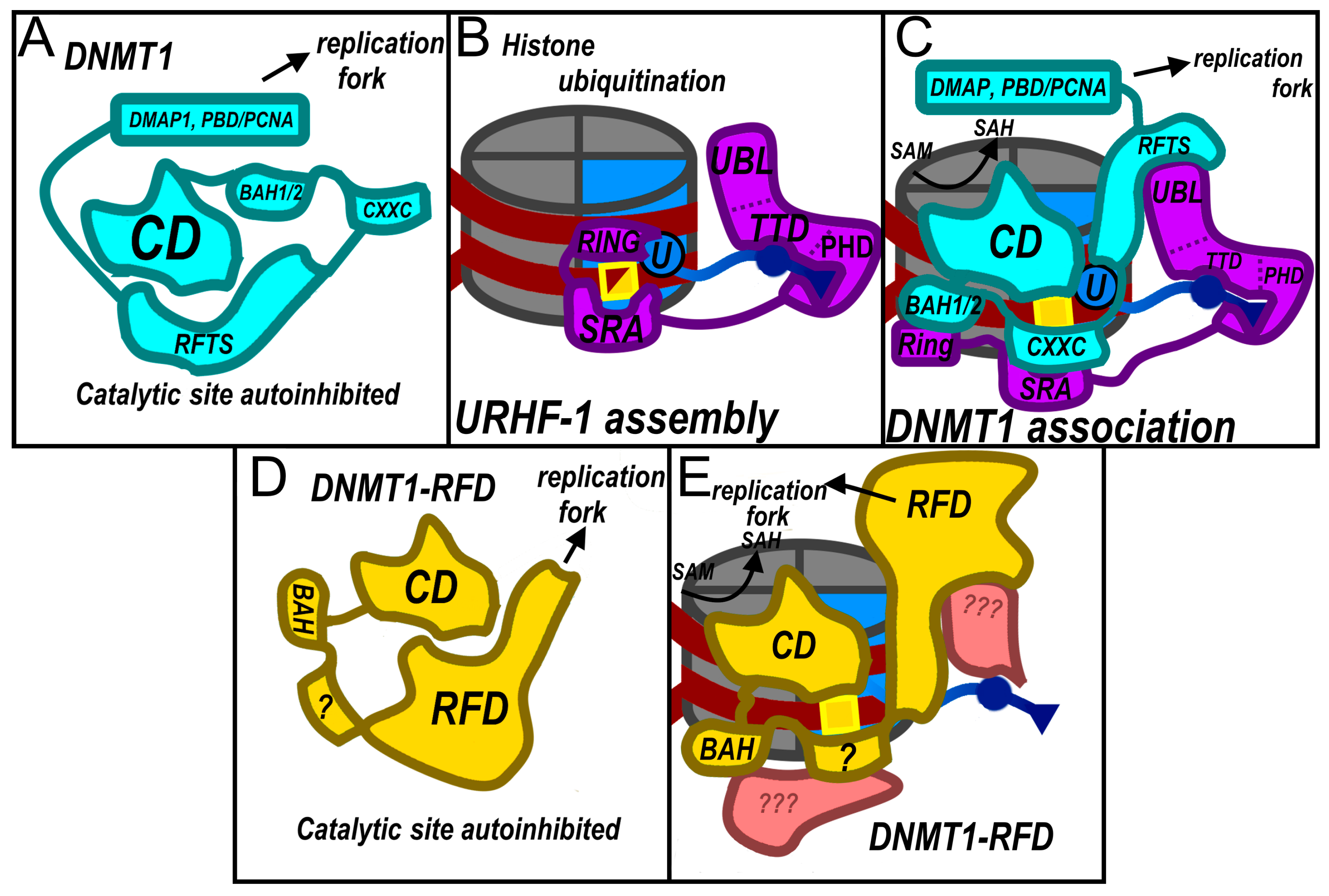
2.5. DNMT1-RFD: A CpG Maintenance Methyltransferase in Basidiomycota
2.6. Methylation in Development, Pathogenesis, and Secondary Metabolism
2.7. Methylation Plasticity and Reversible Phenotypic Adaptation
3. Diversity and Evolutionary Dynamics
3.1. Methyltransferase Repertoires Across Fungal Phyla
3.2. Methylation Levels, TE Content, and Developmental Variation
3.3. Intraspecific DNMT Polymorphism: Gene Loss as an Ongoing Process
4. 6-Methyladenine: Writers, Erasers, and Functions
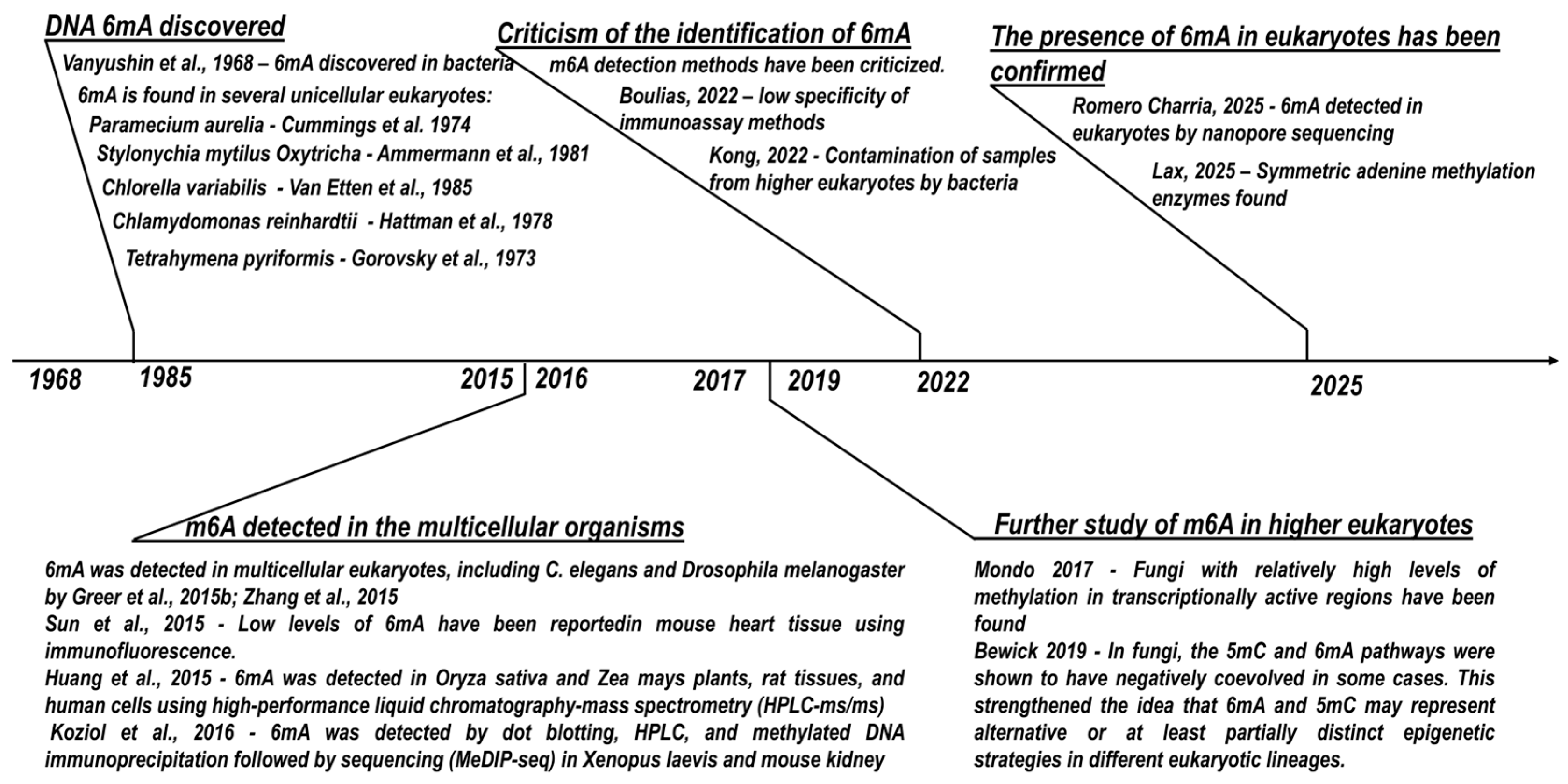
4.1. Discovery, Prevalence, and the Contamination Debate
4.2. Symmetric and Asymmetric 6mA
4.3. Biological Functions of 6mA in Fungi
4.3.1. 6mA in Transcriptional Regulation
4.3.2. 6mA in Life Cycle Regulation
5. Methylation Detection and Functional Analysis
5.1. Quantification and Mapping of 5mC
5.2. Detection and Mapping of 6mA
5.3. Genetic, Enzymatic, and Structural Approaches to Studying DNA MTase Function
6. Conclusions and Open Questions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 5mC | 5-methylcytosine |
| 6mA | N6-methyladenine |
| BAH1 | Bromo-Adjacent Homology 1 domain |
| BAH2 | Bromo-Adjacent Homology 2 domain |
| BGC | biosynthetic gene cluster |
| ChIP-seq | chromatin immunoprecipitation sequencing |
| DAP-seq | DNA affinity purification sequencing |
| DNMT | DNA methyltransferase |
| EDF | early-diverging fungi |
| H3K36me2 | histone H3 lysine 36 dimethylation |
| H3K9me3 | histone H3 lysine 9 trimethylation |
| HDA-1 | histone deacetylase 1 |
| HP1 | heterochromatin protein 1 |
| IPD | inter-pulse duration |
| LC-MS/MS | liquid chromatography–tandem mass spectrometry |
| LINE | long interspersed nuclear element |
| LT50 | median lethal time |
| MAC | methylated adenine cluster |
| MeDIP-seq | methylated DNA immunoprecipitation sequencing |
| MIP | methylation induced premeiotically |
| MSAP | methylation-sensitive amplification polymorphism |
| MTase | methyltransferase |
| ONT | Oxford Nanopore Technologies |
| RdDM | RNA-directed DNA methylation |
| RFD | replication focus domain |
| RFTS | replication foci targeting sequence |
| RIP | repeat-induced point mutation |
| SAM | S-adenosyl-L-methionine |
| SMRT | single-molecule real-time (sequencing) |
| SNF2 | sucrose non-fermenting 2 ATPase domain |
| ssDNA | single-stranded DNA |
| TE | transposable element |
| TET | Ten-Eleven Translocation dioxygenase |
| TSS | transcription start site |
| UHRF1 | ubiquitin-like with PHD and RING finger domains 1 |
| WGBS | whole-genome bisulfite sequencing |
References
- Chen, T.; Li, E. Structure and Function of Eukaryotic DNA Methyltransferases. Curr. Top. Dev. Biol. 2004, 60, 55–89. [CrossRef]
- Del Castillo Falconi, V.M.; Torres-Arciga, K.; Matus-Ortega, G.; Díaz-Chávez, J.; Herrera, L.A. DNA Methyltransferases: From Evolution to Clinical Applications. Int. J. Mol. Sci. 2022, 23. [CrossRef]
- O’Brown, Z.K.; Greer, E.L. N6-Methyladenine: A Conserved and Dynamic DNA Mark. Adv. Exp. Med. Biol. 2016, 945, 213–246. [CrossRef]
- Boulias, K.; Greer, E.L. Means, Mechanisms and Consequences of Adenine Methylation in DNA. Nat. Rev. Genet. 2022, 23, 411–428. [CrossRef]
- Nan, B.; Gao, S. 5mC and 6mA DNA Methylation in Eukaryotic Genomes. Trends Biochem. Sci. 2026, 51, 261–274. [CrossRef]
- Riggs, A.D. X Inactivation, Differentiation, and DNA Methylation. Cytogenet. Cell Genet. 1975, 14, 9–25. [CrossRef]
- Holliday, R.; Pugh, J.E. DNA Modification Mechanisms and Gene Activity during Development. Science 1975, 187, 226–232.
- Jurkowska, R.Z.; Jurkowski, T.P.; Jeltsch, A. Structure and Function of Mammalian DNA Methyltransferases. Chembiochem 2011, 12, 206–222. [CrossRef]
- Cheng, T.; Zhang, J.; Li, H.; Diao, J.; Zhang, W.; Niu, J.; Kawaguchi, T.; Nakayama, J.I.; Kataoka, K.; Gao, S. Identification and Characterization of the de Novo Methyltransferases for Eukaryotic N6-Methyladenine (6mA). Sci. Adv. 2025, 11. [CrossRef]
- Wang, Y.; Nan, B.; Ye, F.; Zhang, Z.; Yang, W.; Pan, B.; Wei, F.; Duan, L.; Li, H.; Niu, J.; et al. Dual Modes of DNA N6-Methyladenine Maintenance by Distinct Methyltransferase Complexes. Proc. Natl. Acad. Sci. U. S. A. 2025, 122. [CrossRef]
- Cooper, D.N.; Krawczak, M. Cytosine Methylation and the Fate of CpG Dinucleotides in Vertebrate Genomes. Hum. Genet. 1989, 83, 181–188. [CrossRef]
- Martienssen, R.A.; Colot, V. DNA Methylation and Epigenetic Inheritance in Plants and Filamentous Fungi. Science 2001, 293, 1070–1074. [CrossRef]
- Aramayo, R.; Selker, E.U. Neurospora Crassa, a Model System for Epigenetics Research. Cold Spring Harb. Perspect. Biol. 2013, 5. [CrossRef]
- Gladyshev, E. Repeat-Induced Point Mutation and Other Genome Defense Mechanisms in Fungi. Microbiol. Spectr. 2017, 5. [CrossRef]
- Zemach, A.; McDaniel, I.E.; Silva, P.; Zilberman, D. Genome-Wide Evolutionary Analysis of Eukaryotic DNA Methylation. Science 2010, 328, 916–919. [CrossRef]
- Bewick, A.J.; Schmitz, R.J. Gene Body DNA Methylation in Plants. Curr. Opin. Plant Biol. 2017, 36, 103–110. [CrossRef]
- Bewick, A.J.; Hofmeister, B.T.; Powers, R.A.; Mondo, S.J.; Grigoriev, I. V; James, T.Y.; Stajich, J.E.; Schmitz, R.J. Diversity of Cytosine Methylation across the Fungal Tree of Life. Nat. Ecol. Evol. 2019, 3, 479–490. [CrossRef]
- de Mendoza, A.; Lister, R.; Bogdanovic, O. Evolution of DNA Methylome Diversity in Eukaryotes. J. Mol. Biol. 2020, 432, 1687–1705. [CrossRef]
- Nai, Y.-S.; Huang, Y.-C.; Yen, M.-R.; Chen, P.-Y. Diversity of Fungal DNA Methyltransferases and Their Association With DNA Methylation Patterns. Front. Microbiol. 2020, 11, 616922. [CrossRef]
- Capuano, F.; Mülleder, M.; Kok, R.; Blom, H.J.; Ralser, M. Cytosine DNA Methylation Is Found in Drosophila Melanogaster but Absent in Saccharomyces Cerevisiae, Schizosaccharomyces Pombe, and Other Yeast Species. Anal. Chem. 2014, 86, 3697–3702. [CrossRef]
- Lax, C.; Mondo, S.J.; Osorio-Concepción, M.; Muszewska, A.; Corrochano-Luque, M.; Gutiérrez, G.; Riley, R.; Lipzen, A.; Guo, J.; Hundley, H.; et al. Symmetric and Asymmetric DNA N6-Adenine Methylation Regulates Different Biological Responses in Mucorales. Nat. Commun. 2024, 15, 6066. [CrossRef]
- He, Z.; Wu, N.; Yao, R.; Tan, H.; Sun, Y.; Chen, J.; Xue, L.; Chen, X.; Yang, S.; Hurst, L.D.; et al. RID Is Required for Both Repeat-Induced Point Mutation and Nucleation of a Novel Transitional Heterochromatic State for Euchromatic Repeats. Nucleic Acids Res. 2025, 53. [CrossRef]
- Wang, J.; Catania, S.; Wang, C.; de la Cruz, M.J.; Rao, B.; Madhani, H.D.; Patel, D.J. Structural Insights into DNMT5-Mediated ATP-Dependent High-Fidelity Epigenome Maintenance. Mol. Cell 2022, 82, 1186-1198.e6. [CrossRef]
- Freitag, M. Histone Methylation by SET Domain Proteins in Fungi. Annu. Rev. Microbiol. 2017, 71, 413–439. [CrossRef]
- Mondo, S.J.; Dannebaum, R.O.; Kuo, R.C.; Louie, K.B.; Bewick, A.J.; LaButti, K.; Haridas, S.; Kuo, A.; Salamov, A.; Ahrendt, S.R.; et al. Widespread Adenine N6-Methylation of Active Genes in Fungi. Nat. Genet. 2017, 49, 964–968. [CrossRef]
- Romero Charria, P.; Navarrete, C.; Ovchinnikov, V.; Xu, L.; Sarre, L.A.; Shabardina, V.; Ksiezopolska, E.; Casacuberta, E.; Lara-Astiaso, D.; Sebé-Pedrós, A.; et al. Adenine DNA Methylation Associated with Transcriptionally Permissive Chromatin Is Widespread across Eukaryotes. Nat. Genet. 2025, 57, 3126–3136. [CrossRef]
- He, C.; Zhang, Z.; Li, B.; Tian, S. The Pattern and Function of DNA Methylation in Fungal Plant Pathogens. Microorganisms 2020, 8. [CrossRef]
- Shao, Z.; Lu, J.; Khudaverdyan, N.; Song, J. Multi-Layered Heterochromatin Interaction as a Switch for DIM2-Mediated DNA Methylation. Nat. Commun. 2024, 15, 6815. [CrossRef]
- Möller, M.; Habig, M.; Lorrain, C.; Feurtey, A.; Haueisen, J.; Fagundes, W.C.; Alizadeh, A.; Freitag, M.; Stukenbrock, E.H. Recent Loss of the Dim2 DNA Methyltransferase Decreases Mutation Rate in Repeats and Changes Evolutionary Trajectory in a Fungal Pathogen. PLoS Genet. 2021, 17, e1009448. [CrossRef]
- Wang, Y.; Chen, W.; Zhang, L.; Wang, S.; Xu, J.-R.; Guo, L. Methylation of Mycovirus DNA Is Mediated by the RNAi Machinery in Vegetative Hyphae of Fusarium Graminearum. Nucleic Acids Res. 2025, 53, gkaf478. [CrossRef]
- Lax, C.; Mondo, S.J.; Martínez, J.F.; Muszewska, A.; Baumgart, L.A.; Pérez-Ruiz, J.A.; Carrillo-Marín, P.; LaButti, K.; Lipzen, A.; Zhang, Y.; et al. Symmetric Adenine Methylation Is an Essential DNA Modification in the Early-Diverging Fungus Rhizopus Microsporus. Nat. Commun. 2025, 16, 3843. [CrossRef]
- Osorio-Concepción, M.; Lax, C.; Navarro, E.; Nicolás, F.E.; Garre, V. DNA Methylation on N6-Adenine Regulates the Hyphal Development during Dimorphism in the Early-Diverging Fungus Mucor Lusitanicus. J. Fungi (Basel) 2021, 7, 738. [CrossRef]
- Mu, Y.; Zhang, L.; Hu, J.; Zhou, J.; Lin, H.W.; He, C.; Chen, H.Z.; Zhang, L. A Fungal Dioxygenase CcTet Serves as a Eukaryotic 6mA Demethylase on Duplex DNA. Nat. Chem. Biol. 2022, 18, 733–741. [CrossRef]
- Castanera, R.; López-Varas, L.; Borgognone, A.; LaButti, K.; Lapidus, A.; Schmutz, J.; Grimwood, J.; Pérez, G.; Pisabarro, A.G.; Grigoriev, I. V.; et al. Transposable Elements versus the Fungal Genome: Impact on Whole-Genome Architecture and Transcriptional Profiles. PLoS Genet. 2016, 12. [CrossRef]
- Lewis, Z.A.; Adhvaryu, K.K.; Honda, S.; Shiver, A.L.; Selker, E.U. Identification of DIM-7, a Protein Required to Target the DIM-5 H3 Methyltransferase to Chromatin. Proc. Natl. Acad. Sci. U. S. A. 2010, 107, 8310–8315. [CrossRef]
- Klocko, A.D.; Summers, C.A.; Glover, M.L.; Parrish, R.; Storck, W.K.; McNaught, K.J.; Moss, N.D.; Gotting, K.; Stewart, A.; Morrison, A.M.; et al. Selection and Characterization of Mutants Defective in DNA Methylation in Neurospora Crassa. Genetics 2020, 216, 671–688. [CrossRef]
- Honda, S.; Bicocca, V.T.; Gessaman, J.D.; Rountree, M.R.; Yokoyama, A.; Yu, E.Y.; Selker, J.M.L.; Selker, E.U. Dual Chromatin Recognition by the Histone Deacetylase Complex HCHC Is Required for Proper DNA Methylation in Neurospora Crassa. Proc. Natl. Acad. Sci. U. S. A. 2016, 113, E6135–E6144. [CrossRef]
- Jeltsch, A.; Jurkowska, R.Z. Allosteric Control of Mammalian DNA Methyltransferases - a New Regulatory Paradigm. Nucleic Acids Res. 2016, 44, 8556–8575. [CrossRef]
- Yan, Y.; Zhou, X.E.; Thomas, S.L.; Liu, M.; Lai, G.-Q.; Worden, E.J.; Jones, P.A.; Xu, T.-H. Mechanisms of DNMT3A–3L-Mediated de Novo DNA Methylation on Chromatin. Nat. Struct. Mol. Biol. 2026, 33, 171–183. [CrossRef]
- Kouzminova, E.; Selker, E.U. Dim-2 Encodes a DNA Methyltransferase Responsible for All Known Cytosine Methylation in Neurospora. EMBO J. 2001, 20, 4309–4323. [CrossRef]
- Grayburn, W.S.; Selker, E.U. A Natural Case of RIP: Degeneration of the DNA Sequence in an Ancestral Tandem Duplication. Mol. Cell. Biol. 1989, 9, 4416–4421. [CrossRef]
- Galagan, J.E.; Selker, E.U. RIP: The Evolutionary Cost of Genome Defense. Trends in Genetics 2004, 20, 417–423. [CrossRef]
- Goyon, C.; Rossignol, J.L.; Faugeron, G. Native DNA Repeats and Methylation in Ascobolus. Nucleic Acids Res. 1996, 24, 3348–3356. [CrossRef]
- Amselem, J.; Lebrun, M.-H.; Quesneville, H. Whole Genome Comparative Analysis of Transposable Elements Provides New Insight into Mechanisms of Their Inactivation in Fungal Genomes. BMC Genomics 2015, 16, 141. [CrossRef]
- Dang, Y.; Li, L.; Guo, W.; Xue, Z.; Liu, Y. Convergent Transcription Induces Dynamic DNA Methylation at DisiRNA Loci. PLoS Genet. 2013, 9, e1003761. [CrossRef]
- Huang, R.; Ding, Q.; Xiang, Y.; Gu, T.; Li, Y. Comparative Analysis of DNA Methyltransferase Gene Family in Fungi: A Focus on Basidiomycota. Front. Plant Sci. 2016, 7, 1556. [CrossRef]
- Grognet, P.; Timpano, H.; Carlier, F.; Aït-Benkhali, J.; Berteaux-Lecellier, V.; Debuchy, R.; Bidard, F.; Malagnac, F. A RID-like Putative Cytosine Methyltransferase Homologue Controls Sexual Development in the Fungus Podospora Anserina. PLoS Genet. 2019, 15, e1008086. [CrossRef]
- Yang, K.; Liang, L.; Ran, F.; Liu, Y.; Li, Z.; Lan, H.; Gao, P.; Zhuang, Z.; Zhang, F.; Nie, X.; et al. The DmtA Methyltransferase Contributes to Aspergillus Flavus Conidiation, Sclerotial Production, Aflatoxin Biosynthesis and Virulence. Sci. Rep. 2016, 6, 23259. [CrossRef]
- Liu, S.-Y.; Lin, J.-Q.; Wu, H.-L.; Wang, C.-C.; Huang, S.-J.; Luo, Y.-F.; Sun, J.-H.; Zhou, J.-X.; Yan, S.-J.; He, J.-G.; et al. Bisulfite Sequencing Reveals That Aspergillus Flavus Holds a Hollow in DNA Methylation. PLoS One 2012, 7, e30349. [CrossRef]
- Jovanska, L.; Lin, I.C.; Yao, J.S.; Chen, C.L.; Liu, H.C.; Li, W.C.; Chuang, Y.C.; Chuang, C.N.; Yu, A.C.H.; Lin, H.N.; et al. DNA Cytosine Methyltransferases Differentially Regulate Genome-Wide Hypermutation and Interhomolog Recombination in Trichoderma Reesei Meiosis. Nucleic Acids Res. 2024, 52, 9551. [CrossRef]
- Matzke, M.A.; Mosher, R.A. RNA-Directed DNA Methylation: An Epigenetic Pathway of Increasing Complexity. Nat. Rev. Genet. 2014, 15, 394–408. [CrossRef]
- Ko, Y.-H.; So, K.-K.; Chun, J.; Kim, D.-H. Distinct Roles of Two DNA Methyltransferases from Cryphonectria Parasitica in Fungal Virulence, Responses to Hypovirus Infection, and Viral Clearance. mBio 2021, 12, 1–16. [CrossRef]
- Li, R.; Zhou, S.; Li, Y.; Shen, X.; Wang, Z.; Chen, B. Comparative Methylome Analysis Reveals Perturbation of Host Epigenome in Chestnut Blight Fungus by a Hypovirus. Front. Microbiol. 2018, 9, 1026. [CrossRef]
- Catania, S.; Dumesic, P.A.; Pimentel, H.; Nasif, A.; Stoddard, C.I.; Burke, J.E.; Diedrich, J.K.; Cook, S.; Shea, T.; Geinger, E.; et al. Evolutionary Persistence of DNA Methylation for Millions of Years after Ancient Loss of a De Novo Methyltransferase. Cell 2020, 180, 263-277.e20. [CrossRef]
- Coelho, M.A.; David-Palma, M.; Shea, T.; Bowers, K.; McGinley-Smith, S.; Mohammad, A.W.; Gnirke, A.; Yurkov, A.M.; Nowrousian, M.; Sun, S.; et al. Comparative Genomics of the Closely Related Fungal Genera Cryptococcus and Kwoniella Reveals Karyotype Dynamics and Suggests Evolutionary Mechanisms of Pathogenesis. PLoS Biol. 2024, 22. [CrossRef]
- Schotanus, K.; Yadav, V.; Heitman, J. Epigenetic Dynamics of Centromeres and Neocentromeres in Cryptococcus Deuterogattii. PLoS Genet. 2021, 17, e1009743. [CrossRef]
- Tam, R.; Möller, M.; Luo, R.; Luo, Z.; Jones, A.; Periyannan, S.; Rathjen, J.P.; Schwessinger, B. Long-Read Genomics Reveal Extensive Nuclear-Specific Evolution and Allele-Specific Expression in a Dikaryotic Fungus. Genome Res. 2025, 35, 1364. [CrossRef]
- Kramer, H.M.; Cook, D.E.; van den Berg, G.C.M.; Seidl, M.F.; Thomma, B.P.H.J. Three Putative DNA Methyltransferases of Verticillium Dahliae Differentially Contribute to DNA Methylation That Is Dispensable for Growth, Development and Virulence. Epigenetics Chromatin 2021, 14, 21. [CrossRef]
- Jeon, J.; Choi, J.; Lee, G.-W.; Park, S.-Y.; Huh, A.; Dean, R.A.; Lee, Y.-H. Genome-Wide Profiling of DNA Methylation Provides Insights into Epigenetic Regulation of Fungal Development in a Plant Pathogenic Fungus, Magnaporthe Oryzae. Sci. Rep. 2015, 5, 8567. [CrossRef]
- Shi, J.; Liu, J.; Li, H.; Tang, Y.; Liu, S.; Sun, Z.; Yu, Z.; Ji, X. DNA Methylation Plays Important Roles in Lifestyle Transition of Arthrobotrys Oligospora. IET Syst. Biol. 2024, 18, 92–102. [CrossRef]
- Bondzie-Quaye, P.; Sossah, F.L.; Swallah, M.S.; Fetisoa, M.R.; Bashir, M.A.; Huang, Q. Genome-Wide Identification and Expression Analysis of Epigenetic Regulator Gene Families in the Medicinal Mushroom Ganoderma Lucidum. Arch. Microbiol. 2025, 207, 121. [CrossRef]
- Li, G.; Feng, D.; Li, K.; Han, S.; Lv, Y.; Deng, Z.; Zeng, G.; Qin, X.; Shen, X.; Liu, S. Integrated Transcriptome and DNA Methylome Analysis Reveal the Browning Mechanism in Agaricus Bisporus. Gene 2025, 955, 149437. [CrossRef]
- Mishra, P.K.; Baum, M.; Carbon, J. DNA Methylation Regulates Phenotype-Dependent Transcriptional Activity in Candida Albicans. Proc. Natl. Acad. Sci. U. S. A. 2011, 108, 11965–11970. [CrossRef]
- Dubey, A.; Jeon, J. Epigenetic Regulation of Development and Pathogenesis in Fungal Plant Pathogens. Mol. Plant Pathol. 2017, 18, 887–898. [CrossRef]
- Chen, Y.-Y.; Zhu, C.; Zhao, J.-H.; Liu, T.; Gao, F.; Zhang, Y.-C.; Duan, C.-G. DNA Methylation-Dependent Epigenetic Regulation of Verticillium Dahliae Virulence in Plants. aBIOTECH 2023, 4, 185–201. [CrossRef]
- Zhang, Z.; He, C.; Chen, Y.; Li, B.; Tian, S. DNA Methyltransferases Regulate Pathogenicity of Botrytis Cinerea to Horticultural Crops. J. Fungi (Basel) 2021, 7, 659. [CrossRef]
- Miao, Z.; Wang, G.; Shen, H.; Wang, X.; Gabriel, D.W.; Liang, W. BcMettl4-Mediated DNA Adenine N6-Methylation Is Critical for Virulence of Botrytis Cinerea. Front. Microbiol. 2022, 13, 925868. [CrossRef]
- Meng, L.; Wang, J.; Li, S.; Xu, X.; Zhang, Z.; Shi, X.; Song, H. DNA Methylation Is Involved in the Regulation of the Pectin Depolymerase Gene of Colletotrichum Gloeosporioides and Accelerates the Infection of Mango Fruit. J. Plant Biochem. Biotechnol. 2023, 32, 253–264. [CrossRef]
- Bonner, C.; Sproule, A.; Rowland, O.; Overy, D.; Subramaniam, R. DNA Methylation Is Responsive to the Environment and Regulates the Expression of Biosynthetic Gene Clusters, Metabolite Production, and Virulence in Fusarium Graminearum. Frontiers in fungal biology 2020, 1, 614633. [CrossRef]
- Zhang, Y.; Yu, W.; Lu, Y.; Wu, Y.; Ouyang, Z.; Tu, Y.; He, B. Epigenetic Regulation of Fungal Secondary Metabolism. J. Fungi (Basel) 2024, 10. [CrossRef]
- Zutz, C.; Gacek, A.; Sulyok, M.; Wagner, M.; Strauss, J.; Rychli, K. Small Chemical Chromatin Effectors Alter Secondary Metabolite Production in Aspergillus Clavatus. Toxins (Basel). 2013, 5, 1723–1741. [CrossRef]
- Kumar, R. Epigenetic Regulation of Fungicide Resistance in Plant-Pathogenic Fungi. Plant Pathol. 2025, 74, 1495–1503. [CrossRef]
- Vogt, G. Environmental Adaptation of Genetically Uniform Organisms with the Help of Epigenetic Mechanisms-An Insightful Perspective on Ecoepigenetics. Epigenomes 2022, 7. [CrossRef]
- So, K.-K.; Ko, Y.-H.; Chun, J.; Bal, J.; Jeon, J.; Kim, J.-M.; Choi, J.; Lee, Y.-H.; Huh, J.H.; Kim, D.-H. Global DNA Methylation in the Chestnut Blight Fungus Cryphonectria Parasitica and Genome-Wide Changes in DNA Methylation Accompanied with Sectorization. Front. Plant Sci. 2018, 9, 103. [CrossRef]
- Breen, J.; Mur, L.A.J.; Sivakumaran, A.; Akinyemi, A.; Wilkinson, M.J.; Rodriguez Lopez, C.M. Botrytis Cinerea Loss and Restoration of Virulence during In Vitro Culture Follows Flux in Global DNA Methylation. Int. J. Mol. Sci. 2022, 23, 3034. [CrossRef]
- Tini, F.; Beccari, G.; Marconi, G.; Porceddu, A.; Sulyok, M.; Gardiner, D.M.; Albertini, E.; Covarelli, L. Identification of Putative Virulence Genes by DNA Methylation Studies in the Cereal Pathogen Fusarium Graminearum. Cells 2021, 10. [CrossRef]
- Wang, Y.; Wang, T.; Qiao, L.; Zhu, J.; Fan, J.; Zhang, T.; Wang, Z.-X.; Li, W.; Chen, A.; Huang, B. DNA Methyltransferases Contribute to the Fungal Development, Stress Tolerance and Virulence of the Entomopathogenic Fungus Metarhizium Robertsii. Appl. Microbiol. Biotechnol. 2017, 101, 4215–4226. [CrossRef]
- Li, W.; Wang, Y.; Zhu, J.; Wang, Z.; Tang, G.; Huang, B. Differential DNA Methylation May Contribute to Temporal and Spatial Regulation of Gene Expression and the Development of Mycelia and Conidia in Entomopathogenic Fungus Metarhizium Robertsii. Fungal Biol. 2017, 121, 293–303. [CrossRef]
- Hu, S.; Bidochka, M.J. DNA Methyltransferase Implicated in the Recovery of Conidiation, through Successive Plant Passages, in Phenotypically Degenerated Metarhizium. Appl. Microbiol. Biotechnol. 2020, 104, 5371–5383. [CrossRef]
- Xin, X.; Yin, J.; Zhang, B.; Li, Z.; Zhao, S.; Gui, Z. Genome-Wide Analysis of DNA Methylation in Subcultured Cordyceps Militaris. Arch. Microbiol. 2019, 201, 369–375. [CrossRef]
- Danner, C.; Mello de Sousa, T.M.; Mach, R.L.; Mach-Aigner, A.R. The Impact of DNA Methylation in Trichoderma Reesei on Cellulase Production and Strain Degeneration. Microorganisms 2025, 13. [CrossRef]
- Borgognone, A.; Castanera, R.; Morselli, M.; López-Varas, L.; Rubbi, L.; Pisabarro, A.G.; Pellegrini, M.; Ramírez, L. Transposon-Associated Epigenetic Silencing during Pleurotus Ostreatus Life Cycle. DNA Research 2018, 25, 451–464. [CrossRef]
- Hua, S.; Qi, B.; Fu, Y.P.; Li, Y. DNA Methylation Changes in Pleurotus Eryngii Subsp. Tuoliensis (Bailinggu) in Response to Low Temperature Stress. Int. J. Agric. Biol. 2017, 19, 328–334. [CrossRef]
- Qiu, J.; Zang, H.; Zhang, K.; Fan, N.; Yang, Y.; Yue, H.; Chen, D.; Guo, R. Whole-Genome Identification and Investigation of DNA Methylation Sites in Nosema Ceranae. Biology (Basel). 2026, 15, 299. [CrossRef]
- Zeng, Z.; Wu, J.; Kovalchuk, A.; Raffaello, T.; Wen, Z.; Liu, M.; Asiegbu, F.O. Genome-Wide DNA Methylation and Transcriptomic Profiles in the Lifestyle Strategies and Asexual Development of the Forest Fungal Pathogen Heterobasidion Parviporum. Epigenetics 2019, 14, 16–40. [CrossRef]
- Russell, P.J.; Rodland, K.D.; Rachlin, E.M.; McCloskey, J.A. Diferential DNA Methylation during the Vegetative Life Cycle of Neurospora Crassa. J. Bacteriol. 1987, 169, 2902–2905. [CrossRef]
- Hosseini, S.; Meunier, C.; Nguyen, D.; Reimegård, J.; Johannesson, H. Comparative Analysis of Genome-Wide DNA Methylation in Neurospora. Epigenetics 2020, 15, 972–987. [CrossRef]
- Sbaraini, N.; Bellini, R.; Penteriche, A.B.; Guedes, R.L.M.; Garcia, A.W.A.; Gerber, A.L.; Vainstein, M.H.; de Vasconcelos, A.T.R.; Schrank, A.; Staats, C.C. Genome-Wide DNA Methylation Analysis of Metarhizium Anisopliae during Tick Mimicked Infection Condition. BMC Genomics 2019, 20, 836. [CrossRef]
- Li, Y.-H.; Chang, J.-C.; Yen, M.-R.; Huang, Y.-F.; Chen, T.-H.; Chen, L.-H.; Nai, Y.-S. Whole-Genome DNA Methylome Analysis of Different Developmental Stages of the Entomopathogenic Fungus Beauveria Bassiana NCHU-157 by Nanopore Sequencing. Front. Genet. 2023, 14, 1085631. [CrossRef]
- Montanini, B.; Chen, P.-Y.; Morselli, M.; Jaroszewicz, A.; Lopez, D.; Martin, F.; Ottonello, S.; Pellegrini, M. Non-Exhaustive DNA Methylation-Mediated Transposon Silencing in the Black Truffle Genome, a Complex Fungal Genome with Massive Repeat Element Content. Genome Biol. 2014, 15, 411. [CrossRef]
- Liu, Y.; Wang, Y.; Bao, D.; Chen, H.; Gong, M.; Sun, S.; Zou, G. Cross-Kingdom DNA Methylation Dynamics: Comparative Mechanisms of 5mC/6mA Regulation and Their Implications in Epigenetic Disorders. Biology (Basel). 2025, 14, 461. [CrossRef]
- Dhillon, B.; Cavaletto, J.R.; Wood, K. V; Goodwin, S.B. Accidental Amplification and Inactivation of a Methyltransferase Gene Eliminates Cytosine Methylation in Mycosphaerella Graminicola. Genetics 2010, 186, 67–77. [CrossRef]
- Ikeda, K.; Van Vu, B.; Kadotani, N.; Tanaka, M.; Murata, T.; Shiina, K.; Chuma, I.; Tosa, Y.; Nakayashiki, H. Is the Fungus Magnaporthe Losing DNA Methylation? Genetics 2013, 195, 845–855. [CrossRef]
- Kong, Y.; Cao, L.; Deikus, G.; Fan, Y.; Mead, E.A.; Lai, W.; Zhang, Y.; Yong, R.; Sebra, R.; Wang, H.; et al. Critical Assessment of DNA Adenine Methylation in Eukaryotes Using Quantitative Deconvolution. Science 2022, 375, 515–522. [CrossRef]
- Vanyushin, B.F.; Belozersky, A.N.; Kokurina, N.A.; Kadirova, D.X. 5-Methylcytosine and 6-Methylamino-Purine in Bacterial DNA. Nature 1968, 218, 1066–1067. [CrossRef]
- Gorovsky, M.A.; Hattman, S.; Pleger, G.L. ( 6 N)Methyl Adenine in the Nuclear DNA of a Eucaryote, Tetrahymena Pyriformis. J. Cell Biol. 1973, 56, 697–701. [CrossRef]
- Cummings, D.J.; Tait, A.; Goddard, J.M. Methylated Bases in DNA from Paramecium Aurelia. Biochim. Biophys. Acta 1974, 374, 1–11. [CrossRef]
- Hattman, S.; Kenny, C.; Berger, L.; Pratt, K. Comparative Study of DNA Methylation in Three Unicellular Eucaryotes. J. Bacteriol. 1978, 135, 1156–1157. [CrossRef]
- Ammermann, D.; Steinbrück, G.; Baur, R.; Wohlert, H. Methylated Bases in the DNA of the Ciliate Stylonychia Mytilus. Eur. J. Cell Biol. 1981, 24, 154–156.
- Van Etten, J.L.; Schuster, A.M.; Girton, L.; Burbank, D.E.; Swinton, D.; Hattman, S. DNA Methylation of Viruses Infecting a Eukaryotic Chlorella-like Green Alga. Nucleic Acids Res. 1985, 13, 3471–3478. [CrossRef]
- Greer, E.L.; Blanco, M.A.; Gu, L.; Sendinc, E.; Liu, J.; Aristizábal-Corrales, D.; Hsu, C.H.; Aravind, L.; He, C.; Shi, Y. DNA Methylation on N6-Adenine in C. Elegans. Cell 2015, 161, 868–878. [CrossRef]
- Zhang, G.; Huang, H.; Liu, D.; Cheng, Y.; Liu, X.; Zhang, W.; Yin, R.; Zhang, D.; Zhang, P.; Liu, J.; et al. N6-Methyladenine DNA Modification in Drosophila. Cell 2015, 161, 893–906. [CrossRef]
- Sun, Q.; Huang, S.; Wang, X.; Zhu, Y.; Chen, Z.; Chen, D. N6-Methyladenine Functions as a Potential Epigenetic Mark in Eukaryotes. Bioessays 2015, 37, 1155–1162. [CrossRef]
- Huang, W.; Xiong, J.; Yang, Y.; Liu, S.-M.; Yuan, B.-F.; Feng, Y.-Q. Determination of DNA Adenine Methylation in Genomes of Mammals and Plants by Liquid Chromatography/Mass Spectrometry. RSC Adv. 2015, 5, 64046–64054. [CrossRef]
- Koziol, M.J.; Bradshaw, C.R.; Allen, G.E.; Costa, A.S.H.; Frezza, C.; Gurdon, J.B. Identification of Methylated Deoxyadenosines in Vertebrates Reveals Diversity in DNA Modifications. Nat. Struct. Mol. Biol. 2016, 23, 24–30. [CrossRef]
- Wang, Y.; Sheng, Y.; Liu, Y.; Zhang, W.; Cheng, T.; Duan, L.; Pan, B.; Qiao, Y.; Liu, Y.; Gao, S. A Distinct Class of Eukaryotic MT-A70 Methyltransferases Maintain Symmetric DNA N6-Adenine Methylation at the ApT Dinucleotides as an Epigenetic Mark Associated with Transcription. Nucleic Acids Res. 2019, 47, 11771–11789. [CrossRef]
- Sheng, Y.; Wang, Y.; Yang, W.; Wang, X.Q.; Lu, J.; Pan, B.; Nan, B.; Liu, Y.; Ye, F.; Li, C.; et al. Semiconservative Transmission of DNA N 6-Adenine Methylation in a Unicellular Eukaryote. Genome Res. 2024, 34, 740–756. [CrossRef]
- Zhang, L.; Mu, Y.; Li, T.; Hu, J.; Lin, H.; Zhang, L. Molecular Basis of an Atypical DsDNA 5mC/6mA Bifunctional Dioxygenase CcTet from Coprinopsis Cinerea in Catalyzing DsDNA 5mC Demethylation. Nucleic Acids Res. 2024, 52, 3886–3895. [CrossRef]
- Lax, C.; Baumgart, L.A.; Tahiri, G.; Nicolás-Muñoz, N.; Zhang, Y.; Blaby, I.K.; Mondo, S.J.; Kamel, B.; O’Malley, R.C.; Ng, V.; et al. Unveiling a Pervasive DNA Adenine Methylation Regulatory Network in the Early-Diverging Fungus Rhizopus Microsporus. Nat. Commun. 2025, 16, 10277. [CrossRef]
- Kisil, O.; Sergeev, A.; Bacheva, A.; Zvereva, M. Methods for Detection and Mapping of Methylated and Hydroxymethylated Cytosine in DNA. Biomolecules 2024, 14, 1346. [CrossRef]
- Adamczyk, S.; Chojak-Koźniewska, J.; Poimala, A.; Velmala, S.; Adamczyk, B. Fast and Reliable Method to Estimate Global DNA Methylation in Plants and Fungi with High-Pressure Liquid Chromatography (HPLC)-Ultraviolet Detection and Even More Sensitive One with HPLC-Mass Spectrometry. J. Biotechnol. 2023, 374, 1–4. [CrossRef]
- Yang, K.; Zhuang, Z.; Zhang, F.; Song, F.; Zhong, H.; Ran, F.; Yu, S.; Xu, G.; Lan, F.; Wang, S. Inhibition of Aflatoxin Metabolism and Growth of Aspergillus Flavus in Liquid Culture by a DNA Methylation Inhibitor. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2015, 32, 554–563. [CrossRef]
- Poças-Fonseca, M.J.; Cabral, C.G.; Manfrão-Netto, J.H.C. Epigenetic Manipulation of Filamentous Fungi for Biotechnological Applications: A Systematic Review. Biotechnol. Lett. 2020, 42, 885–904. [CrossRef]
- Kojima, K.K. Heterogeneous Lineages of DNA Transposons Encode a TET/JBP Dioxygenase in Fungi. Biology (Basel). 2025, 14. [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



