Submitted:
13 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Methods: Oligomer vs. Fibril Formation as Function of Aβ42 Concentration
2.1.1. Aβ42 Solution Preparation
2.1.2. Aβ42 Aggregation Kinetics
2.1.3. Data Analysis
2.2. Computational Methods
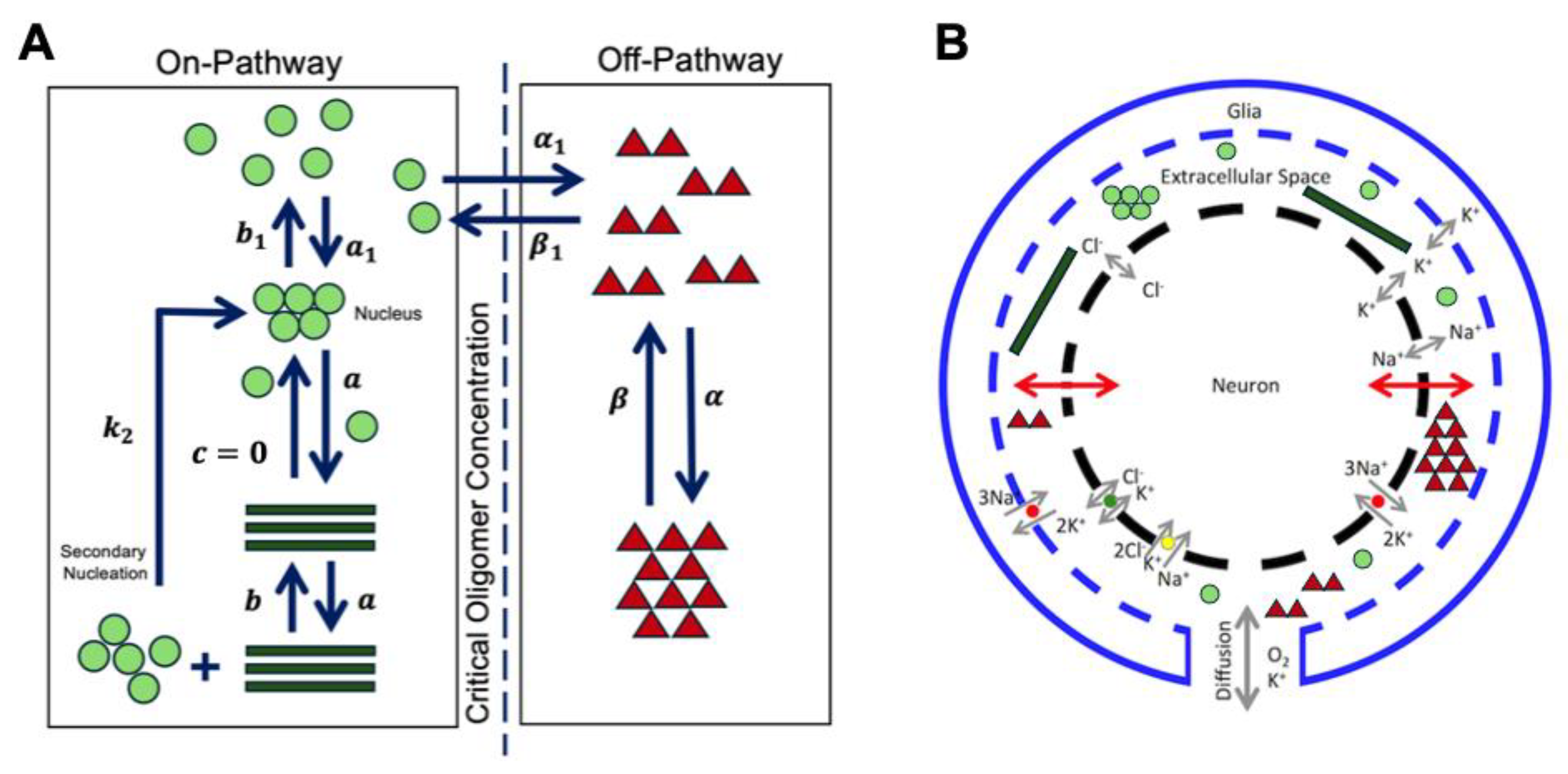
2.2.1. Aβ42 Aggregation Kinetics
2.2.2. Neuronal Model
2.2.3. Modeling Neuronal Membrane Potential
2.2.4. Dynamics of Extra- and Intracellular Ion Concentrations
2.2.5. Volume Regulation
3. Results
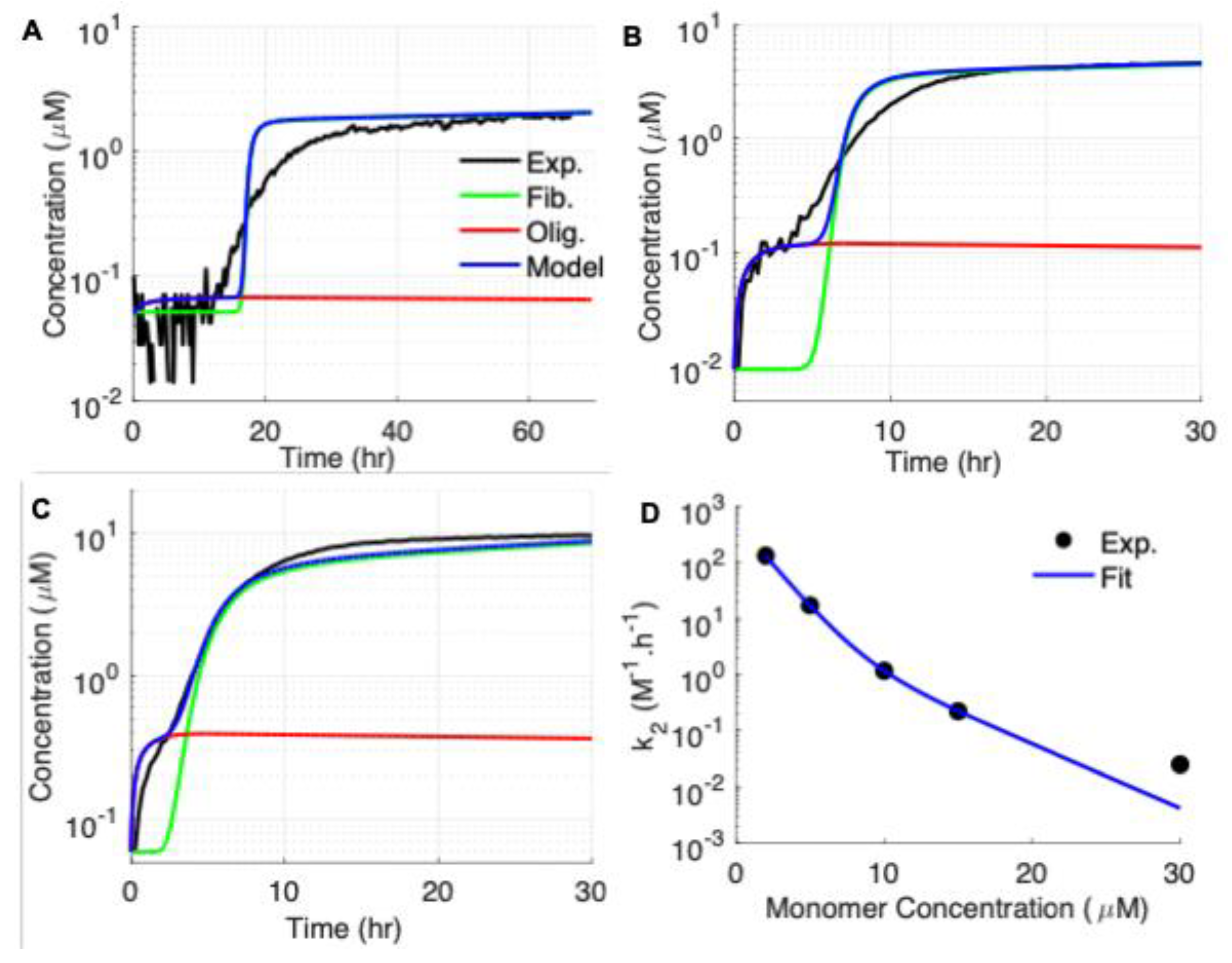
3.1. Experimental Results: ThT Kinetics of Fibril vs Oligomer Formation by Aβ42
3.2. Modeling the Aggregation Kinetics of Aβ42
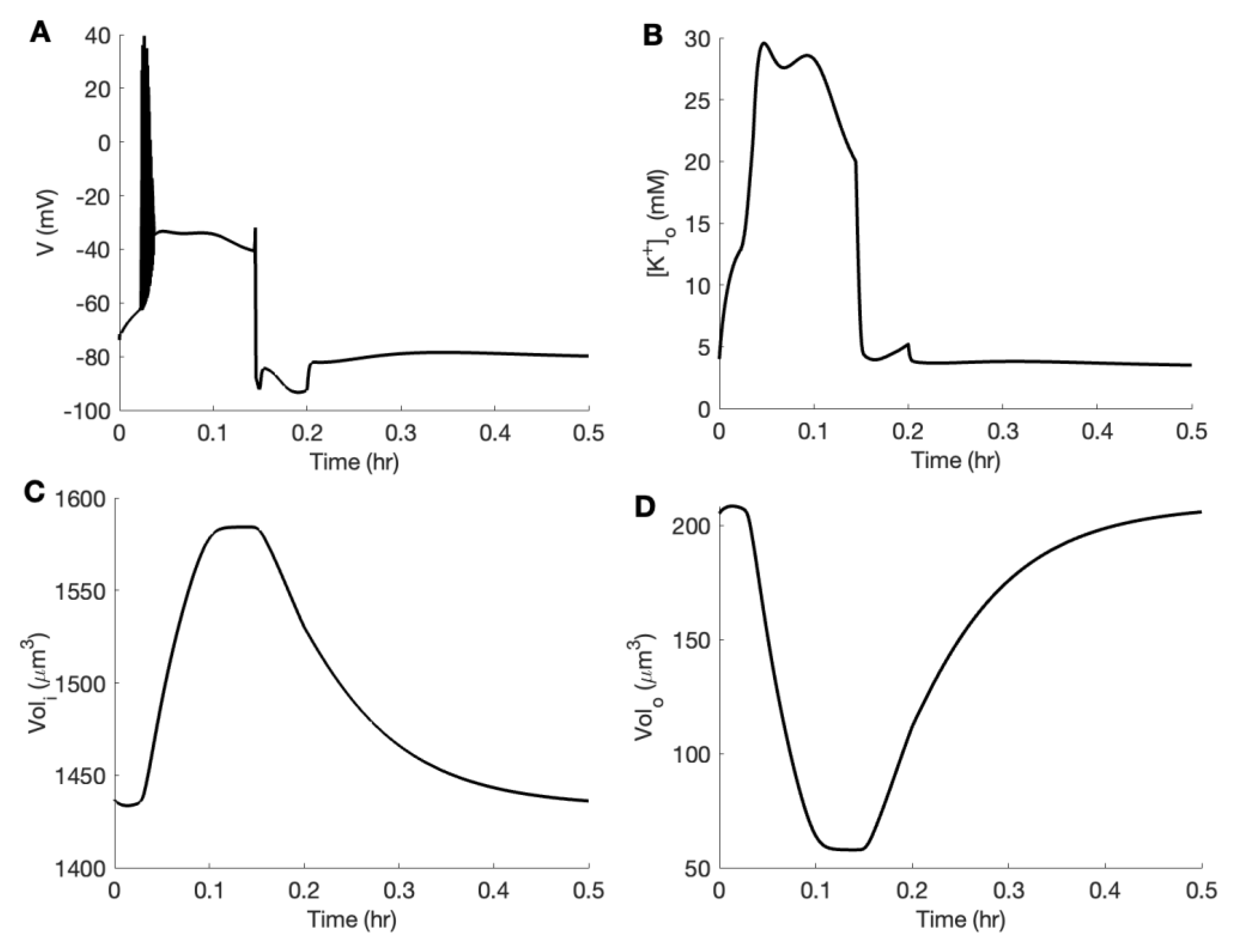
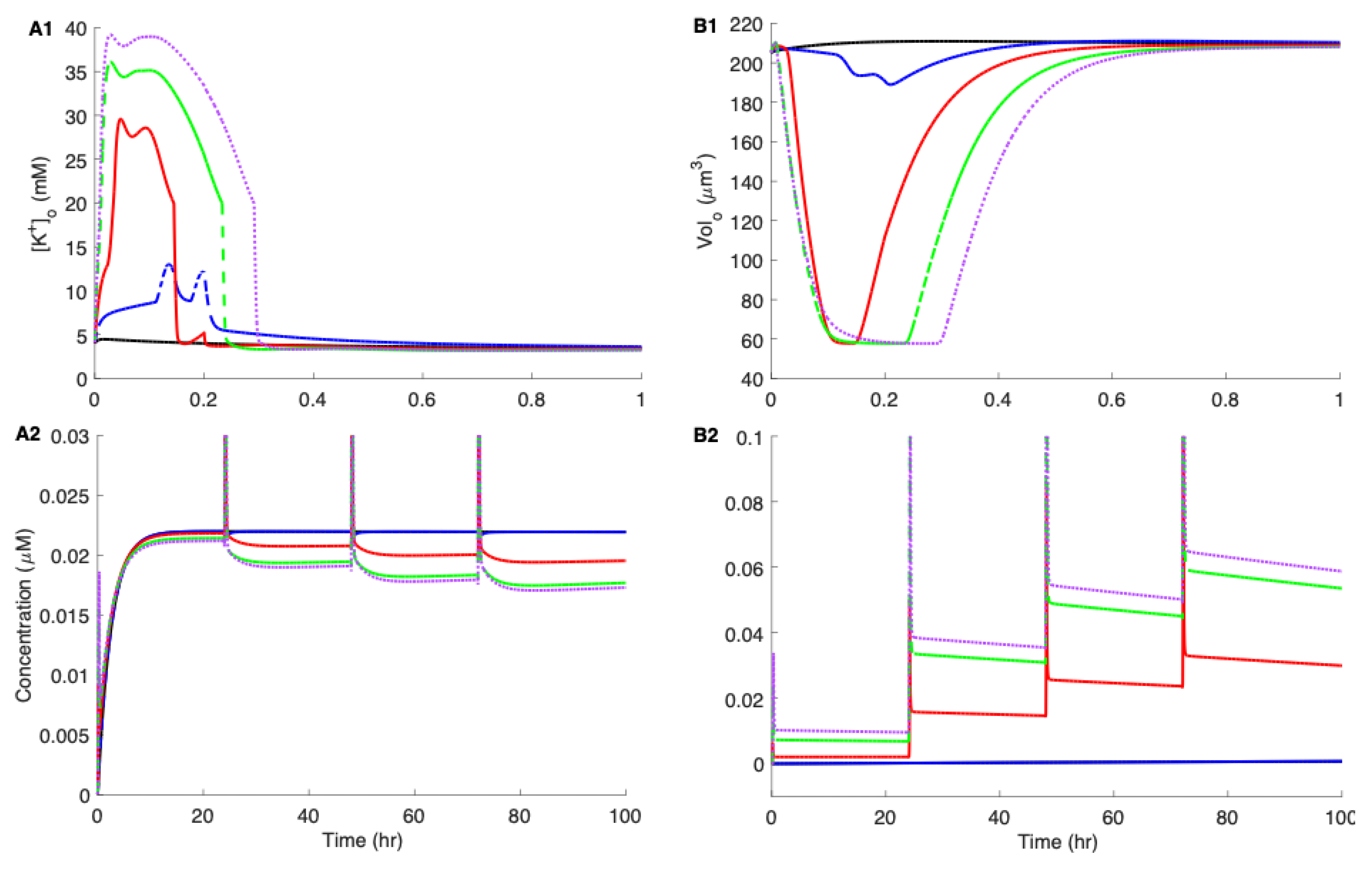
3.3. Linking Neuronal Ion Homeostasis and Swelling to the Aggregation Kinetics of Aβ42
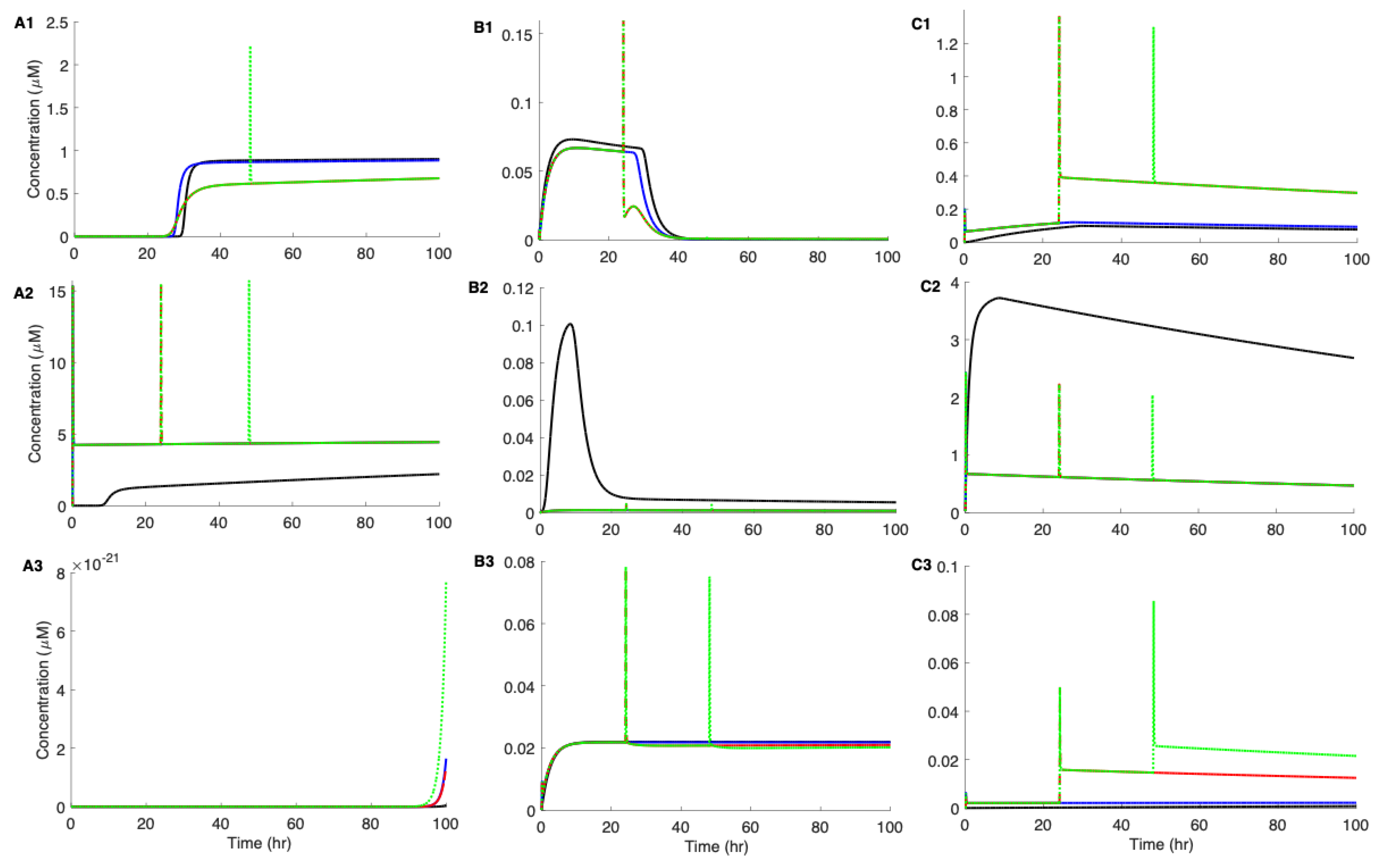
3.4. The Effect of Intensity and Duration of Metabolic Stress on the Aggregation Kinetics of Aβ42
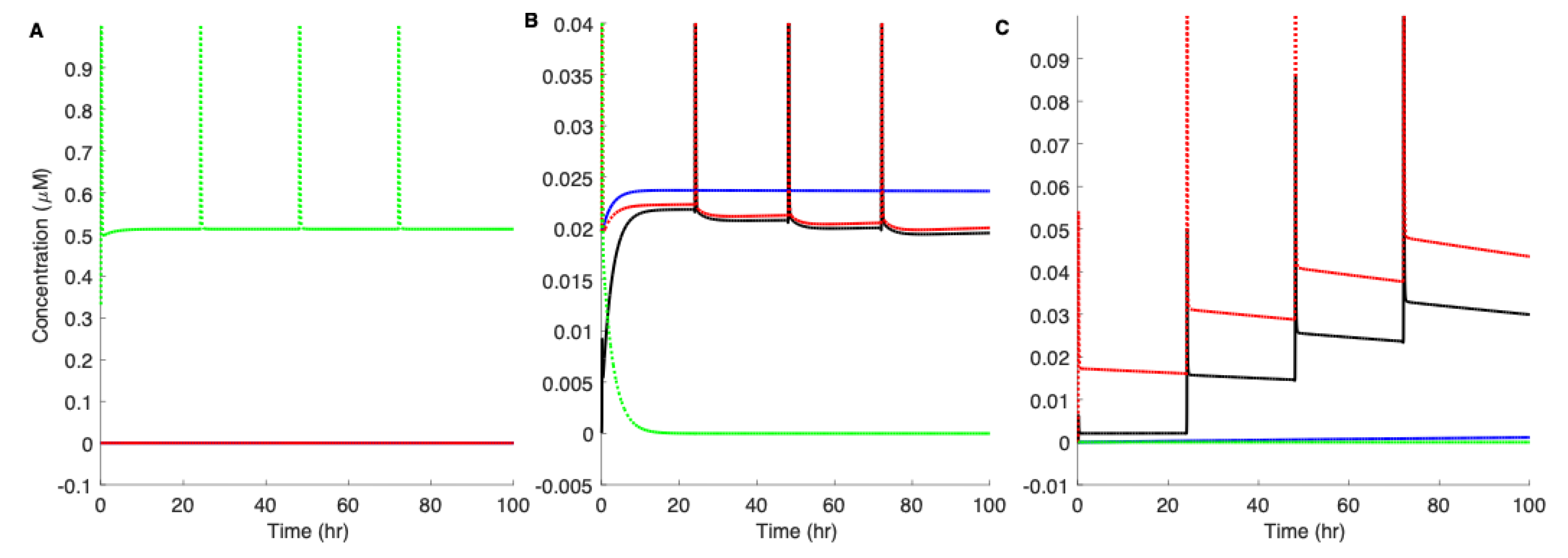
3.5. Pre-Existing Nuclei or Off-Pathway Dimers Significantly Accelerate the Aggregation Kinetics of Aβ42
4. Discussion
References
- Hardy, J.; Selkoe, D.J. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. science 2002, 297(5580), 353–356. [Google Scholar] [CrossRef]
- Hardy, J.A.; Higgins, G.A. Alzheimer’s disease: the amyloid cascade hypothesis . Science 1992, 256(5054), 184–185. [Google Scholar] [CrossRef]
- Brisson, C.D.; Andrew, R.D. A neuronal population in hypothalamus that dramatically resists acute ischemic injury compared to neocortex. Journal of Neurophysiology 2012, 108(2), 419–430. [Google Scholar] [CrossRef]
- Glabe, C. Intracellular mechanisms of amyloid accumulation and pathogenesis in Alzheimer’s disease. Journal of Molecular Neuroscience 2001, 17(2), 137–145. [Google Scholar] [CrossRef] [PubMed]
- Cole, S.L.; Vassar, R. The Alzheimer’s disease β-secretase enzyme, BACE1 . Molecular neurodegeneration 2007, 2(1), 1–25. [Google Scholar] [CrossRef]
- Schützmann, M.P. Endo-lysosomal Aβ concentration and pH trigger formation of Aβ oligomers that potently induce Tau missorting. Nature Communications 2021, 12(1), 1–14. [Google Scholar] [CrossRef] [PubMed]
- Stine, W.B. In vitro characterization of conditions for amyloid-β peptide oligomerization and fibrillogenesis. Journal of Biological Chemistry 2003, 278(13), 11612–11622. [Google Scholar] [CrossRef]
- Atwood, C.S. Dramatic aggregation of Alzheimer Aβ by Cu (II) is induced by conditions representing physiological acidosis. Journal of Biological Chemistry 1998, 273(21), 12817–12826. [Google Scholar] [CrossRef] [PubMed]
- Campos-Ramírez, A. Effect of ionic strength on the aggregation kinetics of the amidated amyloid beta peptide Aβ (1-40) in aqueous solutions. Biophysical Chemistry 2017, 228, 98–107. [Google Scholar] [CrossRef]
- Gorman, P.M. Alternate aggregation pathways of the Alzheimer β-amyloid peptide: Aβ association kinetics at endosomal pH. Journal of molecular biology 2003, 325(4), 743–757. [Google Scholar] [CrossRef]
- Silvers, R. Aggregation and fibril structure of AβM01–42 and Aβ1–42 . Biochemistry 2017, 56(36), 4850–4859. [Google Scholar] [CrossRef] [PubMed]
- Perez, C. Mechanism of fibril and soluble oligomer formation in amyloid beta and hen egg white lysozyme proteins. The Journal of Physical Chemistry B 2019, 123(27), 5678–5689. [Google Scholar] [CrossRef] [PubMed]
- Hasecke, F. Origin of metastable oligomers and their effects on amyloid fibril self-assembly. Chemical science 2018, 9(27), 5937–5948. [Google Scholar] [CrossRef]
- Dreier, J.P.; Reiffurth, C. The stroke-migraine depolarization continuum . Neuron 2015, 86(4), 902–922. [Google Scholar] [CrossRef]
- Thapaliya, P. Modeling the heterogeneity of sodium and calcium homeostasis between cortical and hippocampal astrocytes and its impact on bioenergetics. Frontiers in cellular neuroscience 2023, 17, 1035553. [Google Scholar] [CrossRef]
- Meisl, G. Molecular mechanisms of protein aggregation from global fitting of kinetic models. Nature protocols 2016, 11(2), 252–272. [Google Scholar] [CrossRef]
- Cohen, S.I. Proliferation of amyloid-β42 aggregates occurs through a secondary nucleation mechanism . Proceedings of the National Academy of Sciences 2013, 110(24), 9758–9763. [Google Scholar] [CrossRef]
- Ayata, C.; Lauritzen, M. Spreading depression, spreading depolarizations, and the cerebral vasculature. Physiological reviews 2015, 95(3), 953–993. [Google Scholar] [CrossRef] [PubMed]
- Somjen, G.G. Mechanisms of spreading depression and hypoxic spreading depression-like depolarization. Physiological reviews 2001, 81(3), 1065–1096. [Google Scholar] [CrossRef]
- Guglielmotto, M. The up-regulation of BACE1 mediated by hypoxia and ischemic injury: role of oxidative stress and HIF1α. Journal of neurochemistry 2009, 108(4), 1045–1056. [Google Scholar] [CrossRef]
- Gren, M. Blood biomarkers indicate mild neuroaxonal injury and increased amyloid β production after transient hypoxia during breath-hold diving. Brain injury 2016, 30(10), 1226–1230. [Google Scholar] [CrossRef]
- Zhang, X. Hypoxia-inducible factor 1α (HIF-1α)-mediated hypoxia increases BACE1 expression and β-amyloid generation. Journal of Biological Chemistry 2007, 282(15), 10873–10880. [Google Scholar] [CrossRef]
- Kramer, D.R. Cortical spreading depolarization: pathophysiology, implications, and future directions. Journal of Clinical Neuroscience 2016, 24, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Lauritzen, M. Clinical relevance of cortical spreading depression in neurological disorders: migraine, malignant stroke, subarachnoid and intracranial hemorrhage, and traumatic brain injury. Journal of Cerebral Blood Flow & Metabolism 2011, 31(1), 17–35. [Google Scholar]
- Hartings, J.A. The continuum of spreading depolarizations in acute cortical lesion development: examining Leao’s legacy. Journal of Cerebral Blood Flow & Metabolism 2017, 37(5), 1571–1594. [Google Scholar]
- Dreier, J.P. The role of spreading depression, spreading depolarization and spreading ischemia in neurological disease. Nature medicine 2011, 17(4), 439–447. [Google Scholar] [CrossRef] [PubMed]
- Blasko, I. Experimental traumatic brain injury in rats stimulates the expression, production and activity of Alzheimer’s disease β-secretase (BACE-1). Journal of neural transmission 2004, 111(4), 523–536. [Google Scholar] [CrossRef]
- Pluta, R. Sporadic Alzheimer’s disease begins as episodes of brain ischemia and ischemically dysregulated Alzheimer’s disease genes. Molecular neurobiology 2013, 48(3), 500–515. [Google Scholar] [CrossRef]
- Morton, R.E.; John, P.D. St.; Tyas, S.L. Migraine and the risk of all-cause dementia, Alzheimer’s disease, and vascular dementia: A prospective cohort study in community-dwelling older adults. International journal of geriatric psychiatry 2019, 34(11), 1667–1676. [Google Scholar] [CrossRef]
- Roe, C.M. Amyloid imaging and CSF biomarkers in predicting cognitive impairment up to 7.5 years later. Neurology 2013, 80(19), 1784–1791. [Google Scholar] [CrossRef]
- Vos, S.J. Preclinical Alzheimer’s disease and its outcome: a longitudinal cohort study. The Lancet Neurology 2013, 12(10), 957–965. [Google Scholar] [CrossRef]
- Knopman, D. Short-term clinical outcomes for stages of NIA-AA preclinical Alzheimer disease. Neurology 2012, 78(20), 1576–1582. [Google Scholar] [CrossRef] [PubMed]
- Tarawneh, R. Visinin-like protein-1: diagnostic and prognostic biomarker in Alzheimer disease. Annals of neurology 2011, 70(2), 274–285. [Google Scholar] [CrossRef] [PubMed]
- Nordström, A.; Nordström, P. Traumatic brain injury and the risk of dementia diagnosis: A nationwide cohort study. PLoS medicine 2018, 15(1), e1002496. [Google Scholar] [CrossRef]
- Association, A.s. 2022. Available online: https://www.alz.org/alzheimers-dementia/what-is-dementia/related_conditions/traumatic-brain-injury.
- Li, W. Traumatic brain injury and age at onset of cognitive impairment in older adults. Journal of neurology 2016, 263(7), 1280–1285. [Google Scholar] [CrossRef]
- Rostowsky, K.A.; Irimia, A. Acute cognitive impairment after traumatic brain injury predicts the occurrence of brain atrophy patterns similar to those observed in Alzheimer’s disease. GeroScience 2021, 43(4), 2015–2039. [Google Scholar] [CrossRef]
- Association, A.s. Alzheimer’s disease facts and figures. Alzheimer’s & dementia 2019, 15(3), 321–387. [Google Scholar]
- Mies, G.; Paschen, W. Regional changes of blood flow, glucose, and ATP content determined on brain sections during a single passage of spreading depression in rat brain cortex. Experimental neurology 1984, 84(2), 249–258. [Google Scholar] [CrossRef] [PubMed]
- Hansen, A.J.; Zeuthen, T. Extracellular ion concentrations during spreading depression and ischemia in the rat brain cortex. Acta Physiologica Scandinavica 1981, 113(4), 437–445. [Google Scholar] [CrossRef]
- Andrew, R.D. Questioning glutamate excitotoxicity in acute brain damage: the importance of spreading depolarization. Neurocritical care 2022, 37 (Suppl 1), 11–30. [Google Scholar] [CrossRef]
- Andrew, R.D. The critical role of spreading depolarizations in early brain injury: consensus and contention. In Neurocritical Care; 2022; pp. 1–19. [Google Scholar]
- Rungta, R.L. The cellular mechanisms of neuronal swelling underlying cytotoxic edema. Cell 2015, 161(3), 610–621. [Google Scholar] [CrossRef]
- Sonn, J.; Mayevsky, A. Effects of brain oxygenation on metabolic, hemodynamic, ionic and electrical responses to spreading depression in the rat. Brain research 2000, 882(1-2), 212–216. [Google Scholar] [CrossRef]
- Lauritzen, M. Regional cerebral blood flow during cortical spreading depression in rat brain: increased reactive hyperperfusion in low-flow states. Acta Neurologica Scandinavica 1987, 75(1), 1–8. [Google Scholar] [CrossRef] [PubMed]
- Piper, R.D.; Lambert, G.A.; Duckworth, J.W. Cortical blood flow changes during spreading depression in cats . American Journal of Physiology-Heart and Circulatory Physiology 1991, 261(1), H96–H102. [Google Scholar] [CrossRef] [PubMed]
- Jing, J.; Aitken, P.G.; Somjen, G.G. Interstitial volume changes during spreading depression (SD) and SD-like hypoxic depolarization in hippocampal tissue slices. Journal of Neurophysiology 1994, 71(6), 2548–2551. [Google Scholar] [CrossRef]
- Chesler, M. Regulation and modulation of pH in the brain . Physiological reviews 2003, 83(4), 1183–1221. [Google Scholar] [CrossRef]
- Chesler, M. Failure and function of intracellular pH regulation in acute hypoxic-ischemic injury of astrocytes. Glia 2005, 50(4), 398–406. [Google Scholar] [CrossRef]
- Menyhárt, Á. Spreading depolarization remarkably exacerbates ischemia-induced tissue acidosis in the young and aged rat brain. Scientific reports 2017, 7(1), 1–13. [Google Scholar] [CrossRef]
- Rogers, M.L. Simultaneous monitoring of potassium, glucose and lactate during spreading depolarization in the injured human brain–Proof of principle of a novel real-time neurochemical analysis system, continuous online microdialysis. Journal of Cerebral Blood Flow & Metabolism 2017, 37(5), 1883–1895. [Google Scholar]
- Taylor, D.L.; Obrenovitch, T.P.; Symon, L. Changes in extracellular acid-base homeostasis in cerebral ischemia . Neurochemical research 1996, 21(9), 1013–1021. [Google Scholar] [CrossRef]
- Pérez-Pinzón, M.; Tao, L.; Nicholson, C. Extracellular potassium, volume fraction, and tortuosity in rat hippocampal CA1, CA3, and cortical slices during ischemia. Journal of neurophysiology 1995, 74(2), 565–573. [Google Scholar] [CrossRef]
- Song, M.; Yu, S.P. Ionic regulation of cell volume changes and cell death after ischemic stroke. Translational stroke research 2014, 5(1), 17–27. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Somjen, G.G. Na+ and K+ concentrations, extra-and intracellular voltages, and the effect of TTX in hypoxic rat hippocampal slices. Journal of neurophysiology 2000, 83(2), 735–745. [Google Scholar] [CrossRef]
- Mutch, W.; Hansen, A. Extracellular pH changes during spreading depression and cerebral ischemia: mechanisms of brain pH regulation. Journal of Cerebral Blood Flow & Metabolism 1984, 4(1), 17–27. [Google Scholar]
- Powers, E.T.; Powers, D.L. Mechanisms of protein fibril formation: nucleated polymerization with competing off-pathway aggregation. Biophysical journal 2008, 94(2), 379–391. [Google Scholar] [CrossRef]
- Hodgkin, A.L.; Huxley, A.F. A quantitative description of membrane current and its application to conduction and excitation in nerve. The Journal of physiology 1952, 117(4), 500. [Google Scholar] [CrossRef]
- Hasecke, F. Protofibril–Fibril Interactions Inhibit Amyloid Fibril Assembly by Obstructing Secondary Nucleation. Angewandte Chemie 2021, 133(6), 3053–3058. [Google Scholar] [CrossRef]
- Miti, T. Stable, metastable, and kinetically trapped amyloid aggregate phases. Biomacromolecules 2015, 16(1), 326–335. [Google Scholar] [CrossRef]
- Larsen, B.R. Glutamate transporter activity promotes enhanced Na+/K+-ATPase-mediated extracellular K+ management during neuronal activity. The Journal of physiology 2016, 594(22), 6627–6641. [Google Scholar] [CrossRef] [PubMed]
- Larsen, B.R. Contributions of the Na+/K+-ATPase, NKCC1, and Kir4. 1 to hippocampal K+ clearance and volume responses. Glia 2014, 62(4), 608–622. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Ullah, G.; Schiff, S.J. Unification of neuronal spikes, seizures, and spreading depression . Journal of Neuroscience 2014, 34(35), 11733–11743. [Google Scholar] [CrossRef]
- Syková, E.; Nicholson, C. Diffusion in brain extracellular space . Physiological reviews 2008, 88(4), 1277–1340. [Google Scholar] [CrossRef]
- Yang, L.-B. Elevated β-secretase expression and enzymatic activity detected in sporadic Alzheimer disease. Nature medicine 2003, 9(1), 3–4. [Google Scholar] [CrossRef]
- Sun, X. Hypoxia facilitates Alzheimer’s disease pathogenesis by up-regulating BACE1 gene expression . Proceedings of the National Academy of Sciences 2006, 103(49), 18727–18732. [Google Scholar] [CrossRef]
- Holsinger, R.D. Increased expression of the amyloid precursor β-secretase in Alzheimer’s disease . Annals of neurology: Official journal of the American neurological association and the child neurology society 2002, 51(6), 783–786. [Google Scholar] [CrossRef] [PubMed]
- Risher, W.C.; Croom, D.; Kirov, S.A. Persistent astroglial swelling accompanies rapid reversible dendritic injury during stroke-induced spreading depolarizations. Glia 2012, 60(11), 1709–1720. [Google Scholar] [CrossRef] [PubMed]
- Ries, M.; Sastre, M. Mechanisms of Aβ clearance and degradation by glial cells . Frontiers in aging neuroscience 2016, 8, 160. [Google Scholar] [CrossRef]
- Liu, C.-C. Astrocytic LRP1 mediates brain Aβ clearance and impacts amyloid deposition. Journal of Neuroscience 2017, 37(15), 4023–4031. [Google Scholar] [CrossRef]
- Durand, D. Unraveling the β-amyloid clearance by astrocytes: Involvement of metabotropic glutamate receptor 3, sAPPα, and class-A scavenger receptor. Neurochemistry International 2019, 131, 104547. [Google Scholar] [CrossRef]
- Sword, J. Evolution of neuronal and astroglial disruption in the peri-contusional cortex of mice revealed by in vivo two-photon imaging. Brain 2013, 136(5), 1446–1461. [Google Scholar] [CrossRef] [PubMed]
| Primary nucleation rate | ||
| Dissociation rate of primary nuclei | ||
| RFs formation rate | ||
| RFs dissociation rate | ||
| Dimer formation rate | ||
| Dimer dissociation rate | ||
| gOs formation rate | ||
| gOs dissociation rate |
| Membrane capacitance | 1 | |
| Maximal conductance of sodium current | 30 | |
| Maximal conductance of potassium current | 25 | |
| Conductance of leak sodium current | 0.0247 | |
| Conductance of leak potassium current | 0.05 | |
| Conductance of leak chloride current | 0.1 | |
| Ratio of the initial intra-/extracellular volume | 7 | |
| Maximal glial uptake strength of potassium | 5 | |
| Maximal potassium diffusion rate | 0.25 | |
| Normal bath potassium concentration | 3.5 | |
| Oxygen diffusion rate | 0.17 | |
| Conversion factor | 5.3 | |
| Maximal KCC2 cotransporter strength | 0.3 | |
| Maximal NKCC1 cotransporter strength | 0.1 | |
| Normal bath oxygen concentration | 32 | |
| Maximal Na/K pump rate | 0.8 | |
| Sodium concentration in glia | 18 | |
| Intracellular impermeable anions | 132 | |
| Extracellular impermeable anions | 18 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




