Submitted:
13 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Design and Study Protocol
2.2. Search Strategy
2.3. Eligibility Criteria
2.4. Study Selection Process
2.5. Data Extraction
2.6. Methodological Quality Assessment
2.7. Evidence Synthesis
3. Results
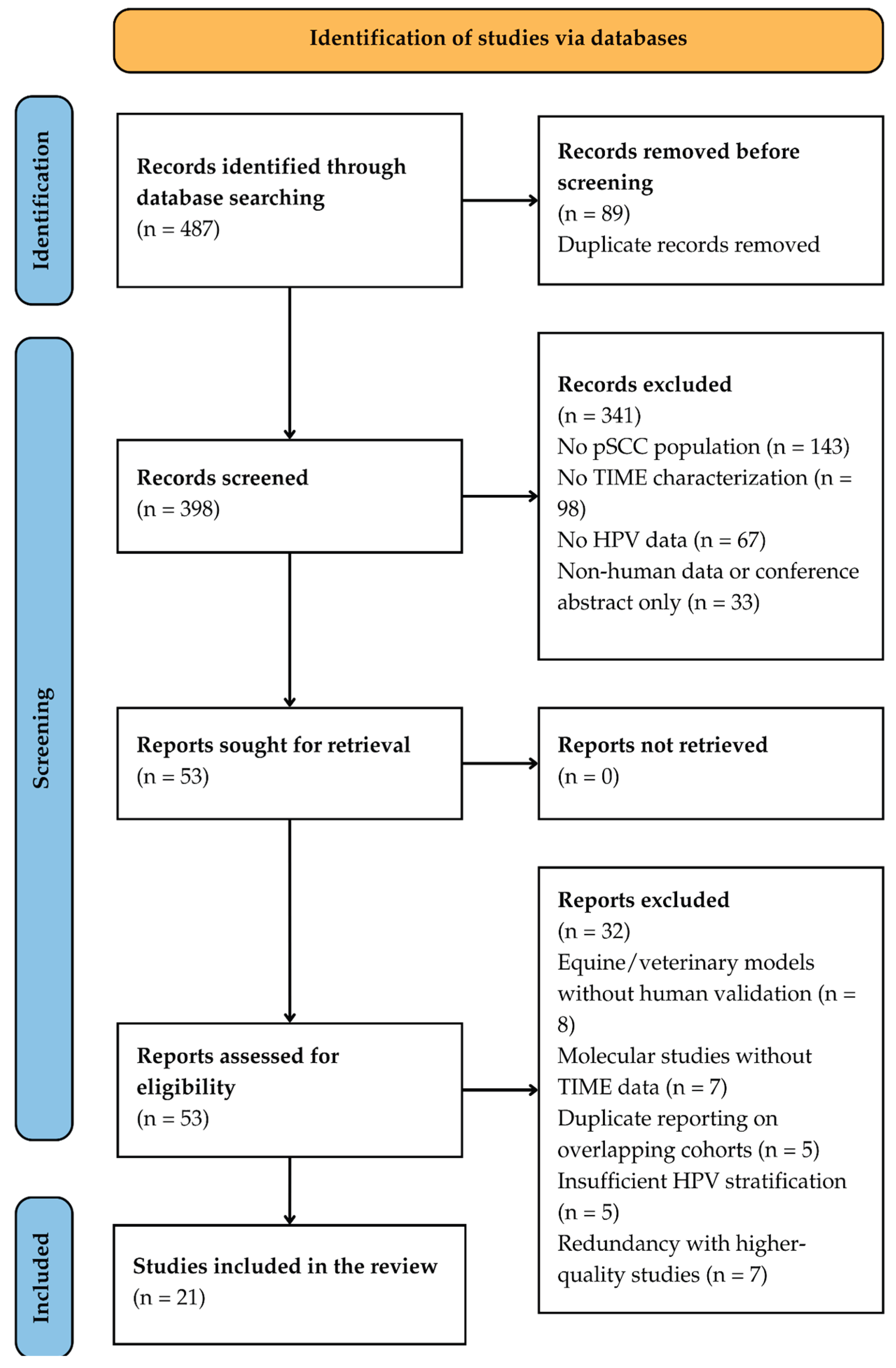
3.1. Study Selection
3.2. Characteristics of the Included Studies
3.3. Molecular Foundations of Time Divergence: The Two Carcinogenic Pathways
3.4. Tumor-Infiltrating Lymphocytes: Differential Density and Phenotype by HPV Status
3.5. The PD-1/PD-L1 Axis: Differential Expression and Prognostic Significance
3.6. Regulatory T Cells and Immunosuppressive Macrophages
3.7. Alternative Immune Checkpoints: TIM-3, LAG-3, and TIGIT
3.8. Spatial and Single-Cell Resolution of TIME Architecture
3.9. Clinical Immunotherapy Evidence and Predictive Biomarkers
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Study (first author, year) | D1 | D2 | D3 | D4 | D5 | Global judgment | Tool |
|---|---|---|---|---|---|---|---|
| Ahmed et al., 2020 [3] | NI | NI | M | B | M | Moderate | ROBINS-I |
| Baweja & Mar, 2020 [24] | M | NI | NI | B | NI | High | ROBINS-I |
| Cañete-Portillo et al., 2024 [21] | B | B | M | B | M | Moderate | ROBINS-I |
| Cao et al., 2021 [16] | M | M | M | B | M | Moderate | ROBINS-I |
| Chahoud et al., 2021 [6] | B | B | M | B | M | Moderate | ROBINS-I |
| Chu et al., 2020 [20] | B | M | B | B | M | Moderate | ROBINS-I |
| de Vries et al., 2023 [11] | B | B | B | B | B | Low | RoB 2 |
| Hladek et al., 2022 [8] | B | B | B | B | M | Low | ROBINS-I |
| Lü et al., 2022 [7] | B | B | B | B | B | Low | AMSTAR-2 |
| Canto et al., 2022 [18] | B | M | M | B | M | Moderate | ROBINS-I |
| Müller et al., 2021 [9] | B | M | M | B | M | Moderate | ROBINS-I |
| Mumba et al., 2024 [1] | B | M | M | B | M | Moderate | ROBINS-I |
| Ribera-Cortada et al., 2022 [5] | B | B | B | B | B | Low | AMSTAR-2 |
| Sangkhamanon et al., 2023 [22] | M | M | M | B | M | Moderate | ROBINS-I |
| Song et al., 2025 [10] | B | B | M | B | B | Low | ROBINS-I |
| Starita et al., 2022 [17] | B | M | M | B | M | Moderate | ROBINS-I |
| Taghizadeh & Fajković, 2025 [23] | B | B | B | B | B | Low | AMSTAR-2 |
| Tang et al., 2022 [19] | NI | B | M | B | M | Moderate | ROBINS-I |
| Vanthoor et al., 2020 [4] | NI | NI | B | B | B | Moderate | ROBINS-I |
| Winkelmann et al., 2024 [25] | B | M | M | B | M | Moderate | ROBINS-I |
| Zhu et al., 2025 [2] | B | B | B | B | B | Low | ROBINS-I |
References
- Mumba, C; Muhimbe, Z; Mapulanga, V; Kawimbe, M; Mutale, K; Hamasuku, A; et al. The effects of HIV and oncogenic human papillomavirus on the tumor immune microenvironment of penile squamous cell carcinoma. PLoS One [Internet] Available from. 2024, 19, e0300729. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L; Shen, D; Zhou, J; Cheng, C; Xu, Z; Liang, Y; et al. Single-cell transcriptomic profiling reveals distinct tumor microenvironments in HPV-associated penile squamous cell carcinoma. Precis Clin Med [Internet] Available from. 2025, 8, pbaf013. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, ME; Falasiri, S; Hajiran, A; Chahoud, J; Spiess, PE. The immune microenvironment in penile cancer and rationale for immunotherapy. J Clin Med [Internet] 2020, 9, 3334. Available online: https://www.mdpi.com/2077-0383/9/10/3334. [CrossRef] [PubMed]
- Vanthoor, J; Vos, G; Albersen, M. Penile cancer: potential target for immunotherapy? World J Urol [Internet] Available from. 2021, 4 39, 1405–11. [Google Scholar] [CrossRef]
- Ribera-Cortada, I; Guerrero-Pineda, J; Trias, I; Veloza, L; Garcia, A; Marimon, L; et al. Pathogenesis of penile squamous cell carcinoma: Molecular update and systematic review. Int J Mol Sci [Internet] Available from. 2021, 5 23, 251. [Google Scholar] [CrossRef]
- Chahoud, J; Gleber-Netto, FO; McCormick, BZ; Rao, P; Lu, X; Guo, M; et al. Whole-exome sequencing in penile squamous cell carcinoma uncovers novel prognostic categorization and drug targets similar to head and neck squamous cell carcinoma. Clin Cancer Res [Internet] Available from. 2021, 27, 2560–70. [Google Scholar] [CrossRef]
- Lu, Y; Wang, Y; Su, H; Li, H. PD-L1 is associated with the prognosis of penile cancer: A systematic review and meta-analysis. Front Oncol [Internet] Available from. 2022, 7 12, 1013806. [Google Scholar] [CrossRef]
- Hladek, L; Bankov, K; von der Grün, J; Filmann, N; Demes, M; Vallo, S; et al. Tumor-associated immune cell infiltrate density in penile squamous cell carcinomas. Virchows Arch [Internet] Available from. 2022, 8 480, 1159–69. [Google Scholar] [CrossRef]
- Müller, T; Demes, M; Lehn, A; Köllermann, J; Vallo, S; Wild, PJ; et al. The peri- and intratumoral immune cell infiltrate and PD-L1 status in invasive squamous cell carcinomas of the penis. Clin Transl Oncol [Internet] Available from. 2022, 24, 331–41. [Google Scholar] [CrossRef]
- Song, H; Tong, Z; Xie, G; Li, Y; Zhao, Y; Fan, F; et al. Single-cell and spatial transcriptomic profiling of penile squamous cell carcinoma reveals dynamics of tumor differentiation and immune microenvironment. Adv Sci (Weinh) [Internet] Available from. 2025, 10 12, e00216. [Google Scholar] [CrossRef]
- de Vries, HM; Rafael, TS; Gil-Jimenez, A; de Feijter, JM; Bekers, E; van der Laan, E; et al. Atezolizumab with or without radiotherapy for advanced squamous cell carcinoma of the penis (the PERICLES study): A phase II trial. J Clin Oncol [Internet] Available from. 2023, 11 41, 4872–80. [Google Scholar] [CrossRef] [PubMed]
- Whittemore, R; Knafl, K. The integrative review: updated methodology. J Adv Nurs [Internet] Available from. 2005 [cited 2025 Nov 27], 12 52, 546–53. [Google Scholar] [CrossRef] [PubMed]
- Page, MJ; McKenzie, JE; Bossuyt, PM; Boutron, I; Hoffmann, TC; Mulrow, CD; et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ [Internet] Available from. 2021, 13 372, n71. [Google Scholar] [CrossRef] [PubMed]
- Sterne, JA; Hernán, MA; Reeves, BC; Savović, J; Berkman, ND; Viswanathan, M; et al. ROBINS-I: a tool for assessing risk of bias in non-randomised studies of interventions. BMJ [Internet] Available from. 2016, 14 355, i4919. [Google Scholar] [CrossRef]
- Sterne, JAC; Savović, J; Page, MJ; Elbers, RG; Blencowe, NS; Boutron, I; et al.; 15 RoB 2: a revised tool for assessing risk of bias in randomised trials. BMJ [Internet] Available from. 2019, 366, l4898. [Google Scholar] [CrossRef]
- Cao, J; Yang, C-H; Han, W-Q; Xie, Y; Liu, Z-Z; Jiang, S-S. Correlation between the evolution of somatic alterations during lymphatic metastasis and clinical outcome in penile squamous cell carcinoma. Front Oncol [Internet] Available from. 2021, 16 11, 641869. [Google Scholar] [CrossRef]
- Starita, N; Pezzuto, F; Sarno, S; Losito, NS; Perdonà, S; Buonaguro, L; et al. Mutations in the telomerase reverse transcriptase promoter and PIK3CA gene are common events in penile squamous cell carcinoma of Italian and Ugandan patients. Int J Cancer [Internet] Available from. 2022, 17 150, 1879–88. [Google Scholar] [CrossRef]
- Canto, LM; da Silva, JM; Castelo-Branco, PV; da Silva, IM; Nogueira, L; Fonseca-Alves, CE; et al. Mutational signature and integrative genomic analysis of human Papillomavirus-associated penile squamous cell carcinomas from Latin American patients. Cancers (Basel) [Internet] Available from. 2022, 18 14, 3514. [Google Scholar] [CrossRef]
- Tang, Y; Hu, X; Wu, K; Li, X. Immune landscape and immunotherapy for penile cancer. Front Immunol [Internet] Available from. 2022, 19 13, 1055235. [Google Scholar] [CrossRef]
- Chu, C; Yao, K; Lu, J; Zhang, Y; Chen, K; Lu, J; et al. Immunophenotypes based on the tumor immune microenvironment allow for unsupervised penile cancer patient stratification. Cancers (Basel) [Internet] 2020, 20 12, 1796. Available online: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC7407624”. [CrossRef]
- Canete-Portillo, S; Cubilla, AL; Netto, GJ; Chaux, A. Interplay of PD-L1, FOXP3, and CD8 in the immune microenvironment of penile squamous cell carcinoma: Expression profiles and correlations [Internet]. medRxiv Available from. 2024, 21. [Google Scholar] [CrossRef]
- Sangkhamanon, S; Kotano, N; Sirithanaphol, W; Rompsaithong, U; Kiatsopit, P; Sookprasert, A; et al. Programmed death-ligand 1 expression in tumor cells and tumor-infiltrating lymphocytes are associated with depth of tumor invasion in penile cancer. Biomed Rep [Internet] Available from. 2023, 22 19, 44. [Google Scholar] [CrossRef]
- Taghizadeh, H; Fajkovic, H. Immunotherapy in the management of penile cancer-A systematic review. Cancers (Basel) [Internet] Available from. 2025, 23 17, 883. [Google Scholar] [CrossRef]
- Baweja, A; Mar, N. Metastatic penile squamous cell carcinoma with dramatic response to combined checkpoint blockade with ipilimumab and nivolumab. J Oncol Pharm Pract [Internet] Available from. 2021, 24. 27, 212–5. [Google Scholar] [CrossRef]
- Winkelmann, R; Becker, N; Leichner, R; Wild, PJ; Demes, M; Banek, S; et al. Gene expression profiling of the peritumoral immune cell infiltrate of penile squamous cell carcinomas. Int J Mol Sci [Internet] Available from. 2024, 25 25, 12142. [Google Scholar] [CrossRef]
| Characteristic | n (%) or Value | Range/Detail |
|---|---|---|
| Number of included studies | 21 | — |
| Total participants | > 4,500 | — |
| Geographical Distribution | ||
| Asian | 7 (33.3%) | — |
| European | 5 (23.8%) | — |
| Multinational | 4 (19.0%) | — |
| North American | 4 (19.0%) | — |
| African | 1 (4.8%) | — |
| Study Design | ||
| Immunohistochemical cohort study | 7 (33.3%) | Retrospective |
| Systematic review/meta-analysis | 4 (19.0%) | — |
| Whole-exome/genomic sequencing | 4 (19.0%) | — |
| Single-cell/spatial transcriptomics | 2 (9.5%) | — |
| Narrative review | 2 (9.5%) | — |
| Phase II clinical trial | 1 (4.8%) | — |
| Case report | 1 (4.8%) | — |
| TIME Components Evaluated | ||
| TIL subsets (CD3, CD8, CD20) | 14 (66.7%) | Immunohistochemistry |
| PD-L1/PD-1 | 13 (61.9%) | IHC or transcriptomics |
| FOXP3+ Tregs | 5 (23.8%) | IHC or scRNA-seq |
| TAM markers (CD68, CD163) | 4 (19.1%) | IHC or scRNA-seq |
| TIM-3/LAG-3/TIGIT | 3 (14.3%) | IHC or scRNA-seq |
| HPV Detection Method | ||
| PCR (genotyping) | 18 (85.7%) | — |
| p16 immunohistochemistry | 8 (38.1%) | Surrogate marker |
| In situ hybridization | 1 (4.8%) | — |
| Methodological Quality | ||
| Low risk of bias | 7 (33.3%) | — |
| Moderate risk of bias | 13 (61.9%) | — |
| High risk of bias | 1 (4.8%) | — |
| Publication Period | ||
| 2020–2021 | 7 (33.3%) | — |
| 2022–2023 | 8 (38.1%) | — |
| 2024–2026 | 6 (28.6%) | — |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




