Submitted:
13 April 2026
Posted:
14 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
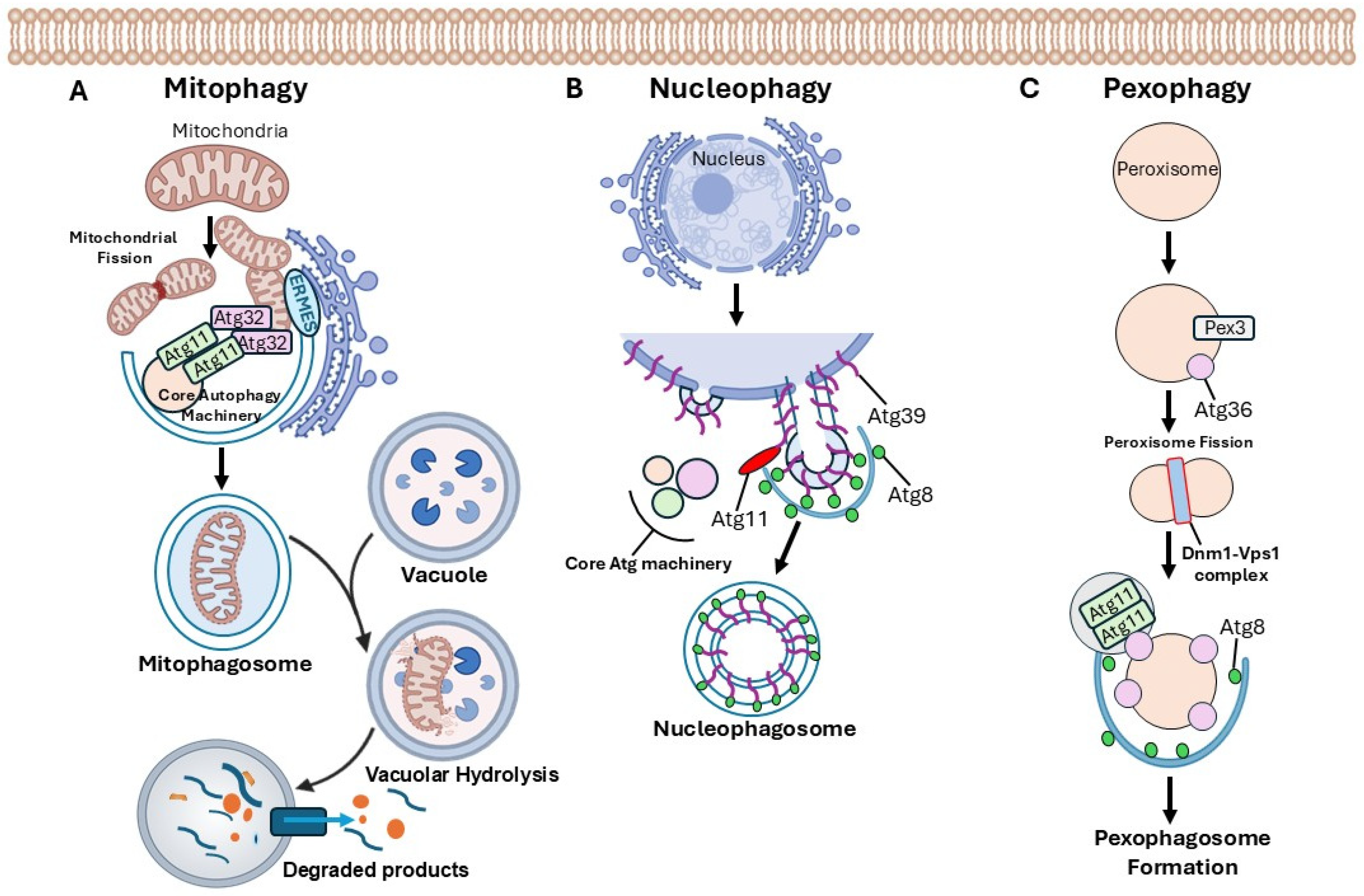
1.1. Selective Autophagy Mechanisms in Saccharomyces Cerevisiae
1.2. Autophagy Genes Regulate Multiple Processes in Plant Pathogenic Fungi
1.2.1. Magnaporthe Oryzae
1.2.2. Colletotrichum spp
1.2.3. Fusarium Graminearum
1.2.4. Ustilago Maydis
1.2.5. Botrytis Cinerea
1.2.6. Aspergillus Flavus
1.2.7. Sclerotinia Sclerotiorum
| Gene name | Organism | Growth/Conidiation | Pathogenesis | Reference |
| MoAtg1 | Magnaporthe oryzae | Reduced | Reduced | [23] |
| MoAtg2 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg3 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg4 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg5 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg5 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg7 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg8 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg9 | Magnaporthe oryzae | Reduced | Abolished | [27] |
| MoAtg10 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg13 | Magnaporthe oryzae | No effect | No effect | [27] |
| MoAtg13 | Magnaporthe oryzae | Reduced | Abolished | [22] |
| MoAtg14 | Magnaporthe oryzae | Reduced | Abolished | [60] |
| CoAtg8 | Colletotrichum orbiculare | Not explained | Abolished | [30] |
| CoAtg26 | Colletotrichum orbiculare | Not explained | Abolished | [30] |
| CfAtg4 | Colletotrichum fructicola | Reduced | Reduced | [32] |
| CfAtg8 | Colletotrichum fructicola | Unknown | Abolished | [31] |
| CfAtg9 | Colletotrichum fructicola | Unknown | Reduced | [31] |
| CsAtg8 | Colletotrichum scovillei | Reduced | Reduced | [33] |
| CgAtg16 | Colletotrichum gloeosporioides | Reduced | Reduced | [34] |
| ChAtg24 | Colletotrichum higginsianum | Reduced | Reduced | [35] |
| FgAtg1 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg2 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg3 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg4 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg5 | Fusarium graminearum | Reduced | No effect | [37] |
| FgAtg6 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg7 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg8 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg9 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg10 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg12 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg14 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg15 | Fusarium graminearum | Reduced | Reduced | [36] |
| FgAtg16 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg17 | Fusarium graminearum | No effect | Reduced | [37] |
| FgAtg18 | Fusarium graminearum | Reduced | Reduced | [37] |
| FgAtg27 | Fusarium graminearum | No effect | No effect | [39] |
| UmAtg1 | Ustilago maydis | No effect | Reduced | [42] |
| UmAtg8 | Ustilago maydis | No effect | Reduced | [42] |
| UmAtg11 | Ustilago maydis | Unknown | No effect | [44,45] |
| BcAtg1 | Botrytis cinerea | Reduced | Reduced | [47,61] |
| BcAtg2 | Botrytis cinerea | Reduced | Abolished | [48] |
| BcAtg3 | Botrytis cinerea | Reduced | Reduced | [49] |
| BcAtg6 | Botrytis cinerea | Reduced | Reduced | [62] |
| BcAtg7 | Botrytis cinerea | Reduced | Reduced | [49] |
| BcAtg8 | Botrytis cinerea | Reduced | Reduced | [49] |
| AfAtg8 | Aspergillus flavus | Reduced | Reduced mycotoxin | [63] |
| AoAtg4 | Aspergillus oryzae | Abolished | NA | [64] |
| AoAtg8 | Aspergillus oryzae | Abolished | NA | [65] |
| AoAtg11 | Aspergillus oryzae | Unknown | NA | [64] |
| AoAtg13 | Aspergillus oryzae | Reduced | NA | [64] |
| AoAtg15 | Aspergillus oryzae | No effect | NA | [66] |
| AoAtg26 | Aspergillus oryzae | Reduced | NA | [67] |
| AnAtg1 | Aspergillus niger | Reduced | NA | [68] |
| AnAtg8 | Aspergillus niger | Reduced | NA | [68] |
| AnAtg17 | Aspergillus niger | No effect | NA | [68] |
| BbAtg1 | Beauveria bassiana | Reduced | Reduced | [69] |
| BdAtg1 | Botryosphaeria dothidea | Reduced | Reduced | [70] |
| BdAtg3 | Botryosphaeria dothidea | Reduced | Reduced | [71] |
| CpAtg4 | Cryphonectria parasitica | Reduced | Reduced | [72] |
| FoAtg3 | Fusarium oxysporum | Reduced | Reduced | [73] |
| FoAtg8 | Fusarium oxysporum | Reduced | Reduced | [74] |
| PsAtg6a | Phytophthora sojae | Reduced | Reduced | [75] |
| PlAtg2 | Peronophythora litchii | Reduced | Reduced | |
| SmAtg12 | Sordaria macrospora | Reduced | Unknown | [76] |
| SsAtg1 | Sclerotinia sclerotiorum | Reduced | Reduced | [57] |
| SsAtg2 | Sclerotinia sclerotiorum | Unknown | Reduced | [57] |
| SsAtg4 | Sclerotinia sclerotiorum | Unknown | Reduced | [57] |
| SsAtg5 | Sclerotinia sclerotiorum | Unknown | Reduced | [57,58] |
| SsAtg9 | Sclerotinia sclerotiorum | Unknown | Reduced | [57] |
| SsAtg12 | Sclerotinia sclerotiorum | Unknown | Reduced | [58] |
| SsAtg17 | Sclerotinia sclerotiorum | Unknown | Reduced | [58] |
| UvAtg7 | Ustilaginoidea virens | Reduced | Reduced | [77,78] |
| UvAtg14 | Ustilaginoidea virens | Reduced | Reduced | [79] |
1.3. Industrially Useful Fungi
1.3.1. Aspergillus Oryzae
1.3.2. Aspergillus Niger
1.4. Autophagy in Human Fungal Pathogens
1.4.1. Cryptococcus Neoformans
1.4.2. Candida Albicans
1.4.3. Aspergillus Fumigatus
| Gene name | Organism | Conidiation | Pathogenesis | Reference |
| CnAtg8 | Cryptococcus neoformans | Unknown | Reduced | [89] |
| CnVps34 | Cryptococcus neoformans | Unknown | Reduced | [89] |
| CnAtg1 | Cryptococcus neoformans | Unknown | Reduced | [91] |
| CnAtg7 | Cryptococcus neoformans | Unknown | Reduced | [90,91] |
| CnAtg8 | Cryptococcus neoformans | Unknown | Reduced | [91] |
| CnAtg5 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg6 | Cryptococcus neoformans | Unknown | Abolished | [93] |
| CnAtg7 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg8 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg9 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg11 | Cryptococcus neoformans | Unknown | Abolished | [93] |
| CnAtg12 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg14-03 | Cryptococcus neoformans | Unknown | Abolished | [93] |
| CnAtg15 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg16 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg20 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg24 | Cryptococcus neoformans | Unknown | Reduced | [93] |
| CnAtg4 | Cryptococcus neoformans | Unknown | Unknown | [94] |
| CnApe4 | Cryptococcus neoformans | Unknown | Reduced | [92] |
| CaAtg1 | Candida albicans | Unknown | Unknown | [99] |
| CaAtg9 | Candida albicans | Unknown | Normal | [97] |
| CaAtg11 | Candida albicans | Reduced | Reduced | [98] |
| CaAtg13 | Candida albicans | Unknown | Unknown | [101] |
| CaAtg27 | Candida albicans | Unknown | Unknown | [101] |
| AfAtg1 | Aspergillus fumigatus | Reduced | Normal | [104] |
| AfAtg20 | Aspergillus fumigatus | Reduced | Reduced | [105] |
| AfAtg24 | Aspergillus fumigatus | Reduced | Reduced | [106] |
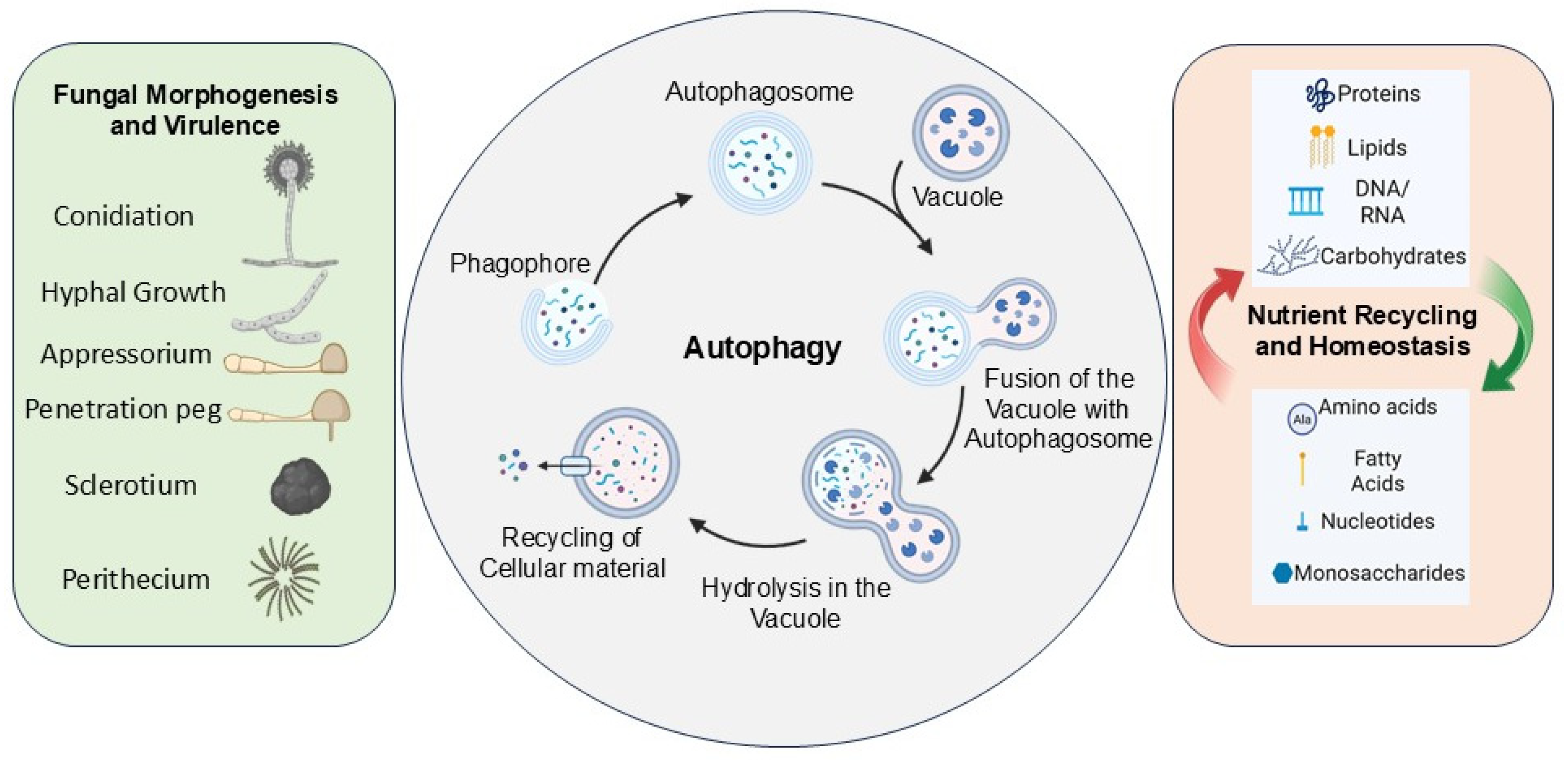
1.5. Multifunctional Roles for Autophagy in Fungi
1.5.1. Autophagy in Nutrient Recycling, Homeostasis, Cellular Differentiation, and Degradation
1.5.2. Autophagy in Antifungal Drug Resistance and Response Mechanisms
1.6. Regulation of Autophagy
1.6.1. Kinase-Phosphatase Modules Regulating Autophagy Machinery
1.6.2. Other Effectors Involved in the Regulation of Autophagy
2. Summary and Future Prospects
Acknowledgments
References
- Mizushima, N.; Klionsky, D. J. Protein turnover via autophagy: Implications for metabolism. Annu. Rev. Nutr. 2007, 27, 19–40. [Google Scholar] [CrossRef]
- Zaffagnini, G.; Martens, S. Mechanisms of Selective Autophagy. J. Mol. Biol. 2016, 428, 1714–1724. [Google Scholar] [CrossRef]
- Lamark, T.; Johansen, T. Mechanisms of Selective Autophagy. Annual Review of Cell and Developmental Biology Annu. Rev. Cell Dev. Biol. 2021 2026, 32, 33. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, H. , Z. S. and M. N. , Autophagy genes in biology and disease. Nat. Rev. Genet. 2023, 24, 382–400. [CrossRef]
- Zhu, X. M.; Li, L.; Wu, M.; Liang, S.; Shi, H. Bin; Liu, X. H.; Lin, F. C. Current opinions on autophagy in pathogenicity of fungi. Virulence 2019, 10, 481–489. [Google Scholar] [CrossRef]
- Parzych, K. R.; Klionsky, D. J. Vacuolar hydrolysis and efflux: current knowledge and unanswered questions. Autophagy 2019, 15, 212–227. [Google Scholar] [CrossRef]
- Kraft, C.; Martens, S. Mechanisms and regulation of autophagosome formation. Curr. Opin. Cell Biol. 2012, 24, 496–501. [Google Scholar] [CrossRef]
- Kanki, T.; Wang, K.; Cao, Y.; Baba, M.; Klionsky, D. J. Atg32 Is a Mitochondrial Protein that Confers Selectivity during Mitophagy. Dev. Cell 2009, 17, 98–109. [Google Scholar] [CrossRef]
- Kanki, T.; Klionsky, D. J. Mitophagy in yeast occurs through a selective mechanism. Journal of Biological Chemistry 2008, 283, 32386–32393. [Google Scholar] [CrossRef]
- Innokentev, A.; Kanki, T. Mitophagy in yeast: Molecular mechanism and regulation. Cells 2021, 10. [Google Scholar] [CrossRef]
- Mochida, K.; Oikawa, Y.; Kimura, Y.; Kirisako, H.; Hirano, H.; Ohsumi, Y.; Nakatogawa, H. Receptor-mediated selective autophagy degrades the endoplasmic reticulum and the nucleus. Nature 2015, 522, 359–362. [Google Scholar] [CrossRef]
- Mochida, K.; Otani, T.; Katsumata, Y.; Kirisako, H.; Kakuta, C.; Kotani, T.; Nakatogawa, H. Atg39 links and deforms the outer and inner nuclear membranes in selective autophagy of the nucleus. Journal of Cell Biology 2022, 221. [Google Scholar] [CrossRef]
- Lee, C. W.; Wilfling, F.; Ronchi, P.; Allegretti, M.; Mosalaganti, S.; Jentsch, S.; Beck, M.; Pfander, B. Selective autophagy degrades nuclear pore complexes. Nat. Cell Biol. 2020, 22, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Tomioka, Y.; Kotani, T.; Kirisako, H.; Oikawa, Y.; Kimura, Y.; Hirano, H.; Ohsumi, Y.; Nakatogawa, H. TORC1 inactivation stimulates autophagy of nucleoporin and nuclear pore complexes. Journal of Cell Biology 2020, 219. [Google Scholar] [CrossRef] [PubMed]
- Chandra, S.; Mannino, P. J.; Thaller, D. J.; Ader, N. R.; King, M. C.; Melia, T. J.; Lusk, C. P. Atg39 selectively captures inner nuclear membrane into lumenal vesicles for delivery to the autophagosome. Journal of Cell Biology 2021, 220. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Manandhar, L.; Kim, H.; Chhetri, A.; Hwang, J.; Jang, G.; Park, C.; Park, R. Pexophagy and Oxidative Stress: Focus on Peroxisomal Proteins and Reactive Oxygen Species (ROS) Signaling Pathways. Antioxidants 2025, 14. [Google Scholar] [CrossRef]
- Ding, L.; Sun, W.; Balaz, M.; He, A.; Klug, M.; Wieland, S.; Caiazzo, R.; Raverdy, V.; Pattou, F.; Lefebvre, P.; et al. Peroxisomal β-oxidation acts as a sensor for intracellular fatty acids and regulates lipolysis. Nat. Metab. 2021, 3, 1648–1661. [Google Scholar] [CrossRef]
- Liu, X.; Wen, X.; Klionsky, D. J. Endoplasmic Reticulum–Mitochondria Contacts Are Required for Pexophagy in Saccharomyces cerevisiae. Contact 2019, 2. [Google Scholar] [CrossRef]
- Yamaguchi, H.; Arakawa, S.; Kanaseki, T.; Miyatsuka, T.; Fujitani, Y.; Watada, H.; Tsujimoto, Y.; Shimizu, S. Golgi membrane-associated degradation pathway in yeast and mammals. EMBO J. 2016, 35, 1991–2007. [Google Scholar] [CrossRef]
- Noguchi, S.; Shimizu, S. Molecular mechanisms and biological roles of GOMED. FEBS Journal 2022, 289, 7213–7220. [Google Scholar] [CrossRef]
- Wilson, R. A.; Talbot, N. J. Under pressure: Investigating the biology of plant infection by Magnaporthe oryzae. Nat. Rev. Microbiol. 2009, 7, 185–195. [Google Scholar] [CrossRef]
- Kershaw, M. J.; Talbot, N. J.; Briggs, S. P. Genome-wide functional analysis reveals that infection-associated fungal autophagy is necessary for rice blast disease. Proceedings of the National Academy of Sciences 2009, 15, 15967–72. [Google Scholar] [CrossRef]
- Liu, X. H.; Lu, J. P.; Lin, F. C. Autophagy during conidiation, conidial germination and turgor generation in Magnaporthe grisea. Autophagy 2007, 3, 472–473. [Google Scholar] [CrossRef] [PubMed]
- Lu, J. P.; Liu, X. H.; Feng, X. X.; Min, H.; Lin, F. C. An autophagy gene, MgATG5, is required for cell differentiation and pathogenesis in Magnaporthe oryzae. Curr. Genet. 2009, 55, 461–473. [Google Scholar] [CrossRef] [PubMed]
- Liu, T. B.; Liu, X. H.; Lu, J. P.; Zhang, L.; Min, H.; Lin, F. C. The cysteine protease MoAtg4 interacts with MoAtg8 and is required for differentiation and pathogenesis in Magnaporthe oryzae. Autophagy 2010, 6, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Yi, Z. D.; Ramos-Pamplona, M.; Naqvi, N. I. Autophagy-assisted glycogen catabolism regulates asexual differentiation in Magnaporthe oryzae. Autophagy 2009, 5, 33–43. [Google Scholar] [CrossRef]
- Dong, B.; Liu, X. H.; Lu, J. P.; Zhang, F. S.; Gao, H. M.; Wang, H. K.; Lin, F. C. MgAtg9 trafficking in Magnaporthe oryzae. Autophagy 2009, 5, 946–953. [Google Scholar] [CrossRef]
- He, M.; Xu, Y.; Chen, J.; Luo, Y.; Lv, Y.; Su, J.; Kershaw, M. J.; Li, W.; Wang, J.; Yin, J.; et al. MoSnt2-dependent deacetylation of histone H3 mediates MoTor-dependent autophagy and plant infection by the rice blast fungus Magnaporthe oryzae. Autophagy 2018, 14, 1543–1561. [Google Scholar] [CrossRef]
- Kubo, Y.; Takano, Y. Dynamics of infection-related morphogenesis and pathogenesis in Colletotrichum orbiculare. Journal of General Plant Pathology 2013, 79, 233–242. [Google Scholar] [CrossRef]
- Takano, Y.; Asakura, M.; Sakai, Y. Atg26-mediated pexophagy and fungal phytopathogenicity. Autophagy 2009, 5, 1041–1042. [Google Scholar] [CrossRef]
- Zhang, S.; Guo, Y.; Li, S.; Li, H. Histone Acetyltransferase CfGcn5-Mediated Autophagy Governs the Pathogenicity of Colletotrichum fructicola. mBio 2022, 13. [Google Scholar]
- Guo, S.; Zhang, S. The Cysteine Protease CfAtg4 Interacts with CfAtg8 to Govern the Growth, Autophagy and Pathogenicity of Colletotrichum fructicola. Journal of Fungi 2024, 10. [Google Scholar]
- Lee, K. H.; Gumilang, A.; Fu, T.; Kang, S. W.; Kim, K. S. The Autophagy Protein CsATG8 is Involved in Asexual Development and Virulence in the Pepper Anthracnose Fungus Colletotrichum scovillei. Mycobiology 2022, 50, 467–474. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Huang, Z.; Xie, J.; He, E.; Wang, Q.; An, B.; He, C.; Luo, H. The CgATG16 Was Involved in Growth, Development and Virulence Through Autophagy Modulation in the Rubber Tree Anthracnose Fungus Colletotrichum gloeosporioides. Journal of Fungi 2025, 11. [Google Scholar] [CrossRef]
- Yan, Y.; Tang, J.; Yuan, Q.; Liu, C.; Chen, X.; Liu, H.; Huang, J.; Bao, C.; Hsiang, T.; Zheng, L. Mitochondrial prohibitin complex regulates fungal virulence via ATG24-assisted mitophagy. Commun. Biol. 2022, 5. [Google Scholar] [CrossRef]
- Nguyen, L. N.; Bormann, J.; Le, G. T. T.; Stärkel, C.; Olsson, S.; Nosanchuk, J. D.; Giese, H.; Schäfer, W. Autophagy-related lipase FgATG15 of Fusarium graminearum is important for lipid turnover and plant infection. Fungal Genetics and Biology 2011, 48, 217–224. [Google Scholar]
- Lv, W.; Wang, C.; Yang, N.; Que, Y.; Talbot, N. J.; Wang, Z. Genome-wide functional analysis reveals that autophagy is necessary for growth, sporulation, deoxynivalenol production and virulence in Fusarium graminearum. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Josefsen, L.; Droce, A.; Sondergaard, T. E.; Sørensen, J. L.; Bormann, J.; Schäfer, W.; Giese, H.; Olsson, S. Autophagy provides nutrients for nonassimilating fungal structures and is necessary for plant colonization but not for infection in the necrotrophic plant pathogen Fusarium graminearum. Autophagy 2012, 8, 326–337. [Google Scholar] [CrossRef]
- Wen, Y.; Wang, M.; Liu, X.; Yin, X.; Gong, S.; Yin, J. Deletion of FgAtg27 decreases the pathogenicity of Fusarium graminearum through influence autophagic process. Int. J. Biol. Macromol. 2025, 297, 139818. [Google Scholar] [CrossRef]
- Chai, X.; Liu, Y.; Ma, H.; Wang, S.; Niyitanga, E.; He, C. Effects of Macroautophagy and Mitophagy on the Pathogenicity of Fusarium graminearum. Phytopathology 2022, 112, 1928–1935. [Google Scholar]
- Nieto-Jacobo, F.; Pasch, D.; Basse, C. W. The mitochondrial Dnm1-like fission component is required for lga2-induced mitophagy but dispensable for starvation-induced mitophagy in Ustilago maydis. Eukaryot. Cell 2012, 11, 1154–1166. [Google Scholar] [CrossRef]
- Nadal, M.; Gold, S. E. The autophagy genes Atg8 and Atg1 affect morphogenesis and pathogenicity in Ustilago maydis. Mol. Plant Pathol. 2010, 11, 463–478. [Google Scholar] [CrossRef]
- Soberanes-Gutiérrez, C. V.; Vázquez-Carrada, M.; López-Villegas, E. O.; Vega-Arreguín, J. C.; Villa-Tanaca, L.; Ruiz-Herrera, J.; Valentina Soberanes-Gutiérrez, C. Autophagosomes accumulation in the vacuoles of the fungus Ustilago maydis and the role of proteases in their digestion. FEMS Microbiol. Lett. 2019. [Google Scholar] [CrossRef] [PubMed]
- Nieto-Jacobo, F.; Pasch, D.; Basse, C. W. The mitochondrial Dnm1-like fission component is required for lga2-induced mitophagy but dispensable for starvation-induced mitophagy in Ustilago maydis. Eukaryot. Cell 2012, 11, 1154–1166. [Google Scholar] [CrossRef] [PubMed]
- Wagner-Vogel, G.; Lämmer, F.; Kämper, J.; Basse, C. W. Uniparental mitochondrial DNA inheritance is not affected in Ustilago maydis Δatg11 mutants blocked in mitophagy. BMC Microbiol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Zhang, Z.; Chen, Y.; Li, B.; Tian, S. Botrytis cinerea. Current Biology 2023, 33, R460–R462. [Google Scholar] [CrossRef]
- Ren, W.; Zhang, Z.; Shao, W.; Yang, Y.; Zhou, M.; Chen, C. The autophagy-related gene BcATG1 is involved in fungal development and pathogenesis in Botrytis cinerea. Mol. Plant Pathol. 2017, 18, 238–248. [Google Scholar] [CrossRef]
- Liu, N.; Lian, S.; Li, B.; Ren, W. The autophagy protein BcAtg2 regulates growth, development and pathogenicity in the gray mold fungus Botrytis cinerea. Phytopathology Research 2022, 4. [Google Scholar] [CrossRef]
- Ren, W.; Sang, C.; Shi, D.; Song, X.; Zhou, M.; Chen, C. Ubiquitin-like activating enzymes BcAtg3 and BcAtg7 participate in development and pathogenesis of Botrytis cinerea. Curr. Genet. 2018, 64, 919–930. [Google Scholar] [CrossRef]
- Liu, N.; Zhou, S.; Li, B.; Ren, W. Involvement of the Autophagy Protein Atg6 in Development and Virulence in the Gray Mold Fungus Botrytis cinerea. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Liu, N.; Ren, W.; Li, F.; Chen, C.; Ma, Z. Involvement of the cysteine protease BcAtg4 in development and virulence of Botrytis cinerea. Curr. Genet. 2019, 65, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Ren, W.; Liu, N.; Sang, C.; Shi, D.; Zhou, M.; Chen, C.; Qin, Q.; Chen, W. The autophagy gene BcATG8 regulates the vegetative differentiation and pathogenicity of Botrytis cinerea. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- Klich, M. A. Aspergillus flavus: The major producer of aflatoxin. Mol. Plant Pathol. 2007, 8, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Geng, Q.; Hu, J.; Xu, P.; Sun, T.; Qiu, H.; Wang, S.; Song, F.; Shen, L.; Li, Y.; Liu, M.; et al. The Autophagy-Related Protein ATG8 Orchestrates Asexual Development and AFB1 Biosynthesis in Aspergillus flavus. Journal of Fungi 2024, 10. [Google Scholar] [CrossRef]
- Zhu, Z.; Yang, M.; Yang, G.; Zhang, B.; Cao, X.; Yuan, J.; Ge, F.; Wang, S. PP2C phosphatases Ptc1 and Ptc2 dephosphorylate PGK1 to regulate autophagy and aflatoxin synthesis in the pathogenic fungus Aspergillus flavus. mBio 2023, 14. [Google Scholar] [CrossRef]
- Jiao, W.; Yu, H.; Chen, X.; Xiao, K.; Jia, D.; Wang, F.; Zhang, Y.; Pan, H. The SsAtg1 Activating Autophagy Is Required for Sclerotia Formation and Pathogenicity in Sclerotinia sclerotiorum. Journal of Fungi 2022, 8, 1314. [Google Scholar] [CrossRef]
- Weerasinghe, T.; Li, J.; Chen, X.; Gao, J.; Tian, L.; Xu, Y.; Gong, Y.; Huang, W.; Zhang, Y.; Jiang, L.; et al. Autophagy-Related Proteins (ATGs) Are Differentially Required for Development and Virulence of Sclerotinia sclerotiorum. Journal of Fungi 2025, 11. [Google Scholar] [CrossRef]
- Zhu, G.; Zuo, Q.; Liu, S.; Zheng, P.; Zhang, Y.; Zhang, X.; Rollins, J. A.; Liu, J.; Pan, H. A FOX transcription factor phosphorylated for regulation of autophagy facilitates fruiting body development in Sclerotinia sclerotiorum. New Phytologist 2025, 2683–2701. [Google Scholar] [CrossRef]
- Zhang, H.; Li, Y.; Lai, W.; Huang, K.; Li, Y.; Wang, Z.; Chen, X.; Wang, A. SsATG8 and SsNBR1 mediated-autophagy is required for fungal development, proteasomal stress response and virulence in Sclerotinia sclerotiorum. Fungal Genetics and Biology 2021, 157, 103632. [Google Scholar] [CrossRef]
- Liu, X. H.; Zhao, Y. H.; Zhu, X. M.; Zeng, X. Q.; Huang, L. Y.; Dong, B.; Su, Z. Z.; Wang, Y.; Lu, J. P.; Lin, F. C. Autophagy-related protein MoAtg14 is involved in differentiation, development and pathogenicity in the rice blast fungus Magnaporthe oryzae. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Zhang, M.; Ma, L.; Lyu, Z.; Trushina, N. K.; Sharon, A. Autophagy and the Mitochondrial Lon1 Protease Are Necessary for Botrytis cinerea Heat Adaptation. Mol. Microbiol. 2025, 124, 358–369. [Google Scholar] [CrossRef]
- Liu, N.; Zhou, S.; Li, B.; Ren, W. Involvement of the Autophagy Protein Atg6 in Development and Virulence in the Gray Mold Fungus Botrytis cinerea. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Geng, Q.; Hu, J.; Xu, P.; Sun, T.; Qiu, H.; Wang, S.; Song, F.; Shen, L.; Li, Y.; Liu, M.; et al. The Autophagy-Related Protein ATG8 Orchestrates Asexual Development and AFB1 Biosynthesis in Aspergillus flavus. Journal of Fungi 2024, 10. [Google Scholar] [CrossRef]
- Kikuma, T.; Kitamoto, K. Analysis of autophagy in Aspergillus oryzae by disruption of Aoatg13, Aoatg4, and Aoatg15genes. FEMS Microbiol. Lett. 2011, 316, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Kikuma, T.; Ohneda, M.; Arioka, M.; Kitamoto, K. Functional analysis of the ATG8 homologue Aoatg8 and role of autophagy in differentiation and germination in Aspergillus oryzae. Eukaryot. Cell 2006, 5, 1328–1336. [Google Scholar] [CrossRef]
- Tadokoro, T.; Kikuma, T.; Kitamoto, K. Functional analysis of AoAtg11 in selective autophagy in the filamentous fungus Aspergillus oryzae. Fungal Biol. 2015, 119, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Kikuma, T.; Tadokoro, T.; Maruyama, J. I.; Kitamoto, K. AoAtg26, a putative sterol glucosyltransferase, is required for autophagic degradation of peroxisomes, mitochondria, and nuclei in the filamentous fungus Aspergillus oryzae. Biosci. Biotechnol. Biochem. 2017, 81, 384–395. [Google Scholar] [CrossRef]
- Nitsche, B. M.; Burggraaf-Van Welzen, A. M.; Lamers, G.; Meyer, V.; Ram, A. F. J. Autophagy promotes survival in aging submerged cultures of the filamentous fungus Aspergillus niger. Appl. Microbiol. Biotechnol. 2013, 97, 8205–8218. [Google Scholar] [CrossRef] [PubMed]
- Ying, S. H.; Liu, J.; Chu, X. L.; Xie, X. Q.; Feng, M. G. The autophagy-related genes BbATG1 and BbATG8 have different functions in differentiation, stress resistance and virulence of mycopathogen Beauveria bassiana. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Liu, N.; Zhu, M.; Zhang, Y.; Wang, Z.; Li, B.; Ren, W. Involvement of the Autophagy Protein Atg1 in Development and Virulence in Botryosphaeria dothidea. Journal of Fungi 2022, 8. [Google Scholar] [CrossRef]
- Han, W.; Han, Y.; Ma, Y.; Yu, G.; Jiang, X.; Yang, Q.; Liu, N.; Wang, C.; Li, B.; Lian, S.; et al. Involvement of the E2-like enzyme Atg3 in fungal development and virulence of Botryosphaeria dothidea. Front. Plant Sci. 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zhao, L.; Li, S.; Chen, F.; Qiu, J.; Bai, L.; Chen, B. The Autophagy-Related Gene CpAtg4 Is Required for Fungal Phenotypic Traits, Stress Tolerance, and Virulence in Cryphonectria parasitica. Phytopathology 2022, 112, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Rehman Khalid, A.; Lv, X.; Naeem, M.; Mehmood, K.; Shaheen, H.; Dong, P.; Qiu, D.; Ren, M. Autophagy related gene (ATG3) is a key regulator for cell growth, development, and virulence of Fusarium oxysporum. Genes (Basel). 2019, 10. [Google Scholar]
- Corral-Ramos, C.; Roca, M. G.; Di Pietro, A.; Roncero, M. I. G.; Ruiz-Roldán, C. Autophagy contributes to regulation of nuclear dynamics during vegetative growth and hyphal fusion in Fusarium oxysporum. Autophagy 2015, 11, 131–144. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, X.; Wang, W.; Geng, X.; Shi, Y.; Na, R.; Dou, D.; Li, H. Network and role analysis of autophagy in Phytophthora sojae. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Werner, A.; Herzog, B.; Frey, S.; Pöggeler, S. Autophagy-associated protein SmATG12 is required for fruiting-body formation in the filamentous ascomycete Sordaria macrospora. PLoS One 2016, 11. [Google Scholar] [CrossRef]
- Yu, J.; He, X.; Xu, C.; Yu, M.; Song, T.; Cao, H.; Pan, X.; Qi, Z.; Du, Y.; Zhang, R.; et al. Autophagy-related protein UvAtg7 contributes to mycelial growth, virulence, asexual reproduction and cell stress response in rice false smut fungus Ustilaginoidea virens. Fungal Genetics and Biology 2022, 159. [Google Scholar] [CrossRef]
- Yu, J.; He, X.; Xu, C.; Yu, M.; Song, T.; Cao, H.; Pan, X.; Qi, Z.; Du, Y.; Zhang, R.; Liang, D. Autophagy-related protein UvAtg7 contributes to mycelial growth, virulence, asexual reproduction and cell stress response in rice false smut fungus Ustilaginoidea virens. Fungal Genetics and Biology 2022, 159, 103668. [Google Scholar] [CrossRef]
- He, X.; Yu, J.; Pan, X.; Cao, H.; Yu, M.; Song, T.; Qi, Z.; Du, Y.; Zhang, R.; Liang, D.; et al. Autophagy-related protein UvAtg14 contributes to mycelial growth, asexual reproduction, virulence and cell stress response in rice false smut fungus Ustilaginoidea virens. Phytopathology Research 2022, 4. [Google Scholar] [CrossRef]
- Kobayashi, T.; Abe, K.; Asai, K.; Gomi, K.; Juvvadi, P. R.; Kato, M.; Kitamoto, K.; Takeuchi, M.; Machida, M. Genomics of Aspergillus oryzae. Biosci. Biotechnol. Biochem. 2007, 71, 646–670. [Google Scholar] [CrossRef]
- Kikuma, T.; Ohneda, M.; Arioka, M.; Kitamoto, K. Functional analysis of the ATG8 homologue Aoatg8 and role of autophagy in differentiation and germination in Aspergillus oryzae. Eukaryot. Cell 2006, 5, 1328–1336. [Google Scholar] [CrossRef]
- Nishio, J.; Takahashi, Y.; Kasahara, M.; Takeda, Y.; Kikuma, T. AeiA is a novel autophagy-related protein that promotes peroxisome degradation by pexophagy in Aspergillus oryzae. FEBS Lett. 2023, 597, 608–617. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, M.; Kimura, S.; Arioka, M. Nucleophagy in Aspergillus oryzae is Mediated by Autophagosome Formation and Vacuole-Mediated Degradation. Curr. Microbiol. 2024, 81. [Google Scholar] [CrossRef] [PubMed]
- Pel, H. J.; De Winde, J. H.; Archer, D. B.; Dyer, P. S.; Hofmann, G.; Schaap, P. J.; Turner, G.; De Vries, R. P.; Albang, R.; Albermann, K.; et al. Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513. Nat. Biotechnol. 2007, 25, 221–231. [Google Scholar] [CrossRef]
- Burggraaf, A. M.; Ram, A. F. J. Autophagy is dispensable to overcome ER stress in the filamentous fungus Aspergillus niger. Microbiologyopen 2016, 5, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Kaur, B.; Punekar, N. S. Autophagy is important to the acidogenic metabolism of Aspergillus niger. PLoS One 2019, 14. [Google Scholar] [CrossRef]
- Gibson, J. F.; Johnston, S. A. Immunity to Cryptococcus neoformans and C. gattii during cryptococcosis. Fungal Genetics and Biology 2015, 78, 76–86. [Google Scholar] [CrossRef]
- Hu, G.; Hacham, M.; Waterman, S. R.; Panepinto, J.; Shin, S.; Liu, X.; Gibbons, J.; Valyi-Nagy, T.; Obara, K.; Jaffe, H. A.; et al. PI3K signaling of autophagy is required for starvation tolerance and virulence of Cryptococcus neoformans. Journal of Clinical Investigation 2008, 118, 1186–1197. [Google Scholar] [CrossRef]
- Hu, G.; Hacham, M.; Waterman, S. R.; Panepinto, J.; Shin, S.; Liu, X.; Gibbons, J.; Valyi-Nagy, T.; Obara, K.; Jaffe, H. A.; et al. PI3K signaling of autophagy is required for starvation tolerance and virulence of Cryptococcus neoformans. Journal of Clinical Investigation 2008, 118, 1186–1197. [Google Scholar] [CrossRef]
- Oliveira, D. L.; Fonseca, F. L.; Zamith-Miranda, D.; Nimrichter, L.; Rodrigues, J.; Pereira, M. D.; Reuwsaat, J. C. V.; Schrank, A.; Staats, C.; Kmetzsch, L.; et al. The putative autophagy regulator Atg7 affects the physiology and pathogenic mechanisms of Cryptococcus neoformans. Future Microbiol. 2016, 11, 1405–1419. [Google Scholar] [CrossRef]
- Ding, H.; Caza, M.; Dong, Y.; Arif, A. A.; Horianopoulos, L. C.; Hu, G.; Johnson, P.; Kronstad, J. W. ATG genes influence the virulence of Cryptococcus neoformans through contributions beyond core autophagy functions. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef] [PubMed]
- De Assis Gontijo, F.; De Melo, A. T.; Pascon, R. C.; Fernandes, L.; Paes, H. C.; Alspaugh, J. A.; Vallim, M. A. The role of Aspartyl aminopeptidase (Ape4) in Cryptococcus neoformans virulence and authophagy. PLoS One 2017, 12. [Google Scholar]
- Zhao, X.; Feng, W.; Zhu, X.; Li, C.; Ma, X.; Li, X.; Zhu, X.; Wei, D. Conserved Autophagy Pathway Contributes to Stress Tolerance and Virulence and Differentially Controls Autophagic Flux Upon Nutrient Starvation in Cryptococcus neoformans. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Roberto, T. N.; Lima, R. F.; Pascon, R. C.; Idnurm, A.; Vallim, M. A. Biological functions of the autophagy-related proteins Atg4 and Atg8 in Cryptococcus neoformans. PLoS One 2020, 15. [Google Scholar] [CrossRef]
- Sobel, J. D. Recurrent vulvovaginal candidiasis. Am. J. Obstet. Gynecol. 2016, 214, 15–21. [Google Scholar] [CrossRef]
- De Repentigny, L.; Lewandowski, D.; Jolicoeur, P. Immunopathogenesis of oropharyngeal candidiasis in human immunodeficiency virus infection. Clin. Microbiol. Rev. 2004, 17, 729–759. [Google Scholar] [CrossRef]
- Palmer, G. E.; Kelly, M. N.; Sturtevant, J. E. Autophagy in the pathogen Candida albicans. Microbiology (N. Y). 2007, 153, 51–58. [Google Scholar] [CrossRef]
- Cui, L.; Zhao, H.; Yin, Y.; Liang, C.; Mao, X.; Liu, Y.; Yu, Q.; Li, M. Function of Atg11 in non-selective autophagy and selective autophagy of Candida albicans. Biochem. Biophys. Res. Commun. 2019, 516, 1152–1158. [Google Scholar] [CrossRef]
- Yu, Q.; Jia, C.; Dong, Y.; Zhang, B.; Xiao, C.; Chen, Y.; Wang, Y.; Li, X.; Wang, L.; Zhang, B.; et al. Candida albicans autophagy, no longer a bystander: Its role in tolerance to ER stress-related antifungal drugs. Fungal Genetics and Biology 2015, 81, 238–249. [Google Scholar] [CrossRef]
- Huang, Y.; Yu, S.; Liu, S.; Zhao, X.; Chen, X.; Wei, X. Autophagy Activated by Atg1 Interacts With Atg9 Promotes Biofilm Formation and Resistance of Candida albicans. J. Basic Microbiol.; 65 2025, e2400603. [Google Scholar] [CrossRef]
- Liu, S.; Jiang, L.; Miao, H.; Lv, Y.; Zhang, Q.; Ma, M.; Duan, W.; Huang, Y.; Wei, X. Autophagy regulation of ATG13 and ATG27 on biofilm formation and antifungal resistance in Candida albicans. Biofouling 38AD, 926–939. [CrossRef] [PubMed]
- Mao, X.; Yang, L.; Yu, D.; Ma, T.; Ma, C.; Wang, J.; Yu, Q.; Li, M. The Vacuole and Mitochondria Patch (vCLAMP) Protein Vam6 is Crucial for Autophagy in Candida albicans. Mycopathologia 2021, 186, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Sugui, J. A.; Kwon-Chung, K. J.; Juvvadi, P. R.; Latgé, J. P.; Steinbach, W. J. Aspergillus fumigatus and related species. Cold Spring Harb. Perspect. Med. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Richie, D. L.; Fuller, K. K.; Fortwendel, J.; Miley, M. D.; McCarthy, J. W.; Feldmesser, M.; Rhodes, J. C.; Askew, D. S. Unexpected link between metal ion deficiency and autophagy in Aspergillus fumigatus. Eukaryot. Cell 2007, 6, 2437–2447. [Google Scholar] [CrossRef]
- Shwab, E. K.; Juvvadi, P. R.; Shaheen, S. K.; Allen, J.; Waitt, G.; Soderblom, E. J.; Asfaw, Y. G.; Moseley, M. A.; Steinbach, W. J. Protein Kinase A Regulates Autophagy-Associated Proteins Impacting Growth and Virulence of Aspergillus fumigatus. Journal of Fungi 2022, 8. [Google Scholar] [CrossRef]
- Keats Shwab, E.; Juvvadi, P. R.; Waitt, G.; Shaheen, S.; Allen, J.; Soderblom, E. J.; Bobay, B. G.; Asfaw, Y. G.; Moseley, M. A.; Steinbach, W. J. The protein kinase a-dependent phosphoproteome of the human pathogen Aspergillus fumigatus reveals diverse virulence-associated kinase targets. mBio 2020, 11, 1–22. [Google Scholar]
- Vaneault-Fourrey, C.; Barooah, M.; Egan, M.; Wakley, G.; Talbot, N. J. Autophagic fungal cell death is necessary for infection by the rice blast fungus. Science (1979). 2006, 312, 580–583. [Google Scholar] [CrossRef]
- Klionsky, D. J.; Emr, S. D. Autophagy as a Regulated Pathway of Cellular Degradation; 2000; Vol. 290. [Google Scholar]
- Pollack, J.K.; Li, Z.J.; Marten, M.R. Fungal mycelia show lag time before re-growth on endogenous carbon. Biotechnol. Bioeng. 2008, 458–465. [Google Scholar] [CrossRef]
- Liu, X. H.; Chen, S. M.; Gao, H. M.; Ning, G. A.; Shi, H. Bin; Wang, Y.; Dong, B.; Qi, Y. Y.; Zhang, D. M.; Lu, G. D.; et al. The small GTPase MoYpt7 is required for membrane fusion in autophagy and pathogenicity of Magnaporthe oryzae. Environ. Microbiol. 2015, 17, 4495–4510. [Google Scholar] [CrossRef]
- Pinan-Lucarré, B.; Paoletti, M.; Dementhon, K.; Coulary-Salin, B.; Clavé, C. Autophagy is induced during cell death by incompatibility and is essential for differentiation in the filamentous fungus Podospora anserina. Mol. Microbiol. 2003, 47, 321–333. [Google Scholar] [CrossRef]
- Pinan-Lucarré, B.; Balguerie, A.; Clavé, C. Accelerated cell death in Podospora autophagy mutants. Eukaryot. Cell 2005, 4, 1765–1774. [Google Scholar] [CrossRef] [PubMed]
- White, S.; McIntyre, M.; Berry, D. R.; McNeil, B. The Autolysis of Industrial Filamentous Fungi; 2002 2002, 22(1), 1–4.
- Puumala, E.; Fallah, S.; Robbins, N.; Cowen, L. E. Advancements and challenges in antifungal therapeutic development. Clin. Microbiol. Rev. 2024, 37. [Google Scholar] [CrossRef] [PubMed]
- Bashiri, H.; Tabatabaeian, H. Autophagy: A Potential Therapeutic Target to Tackle Drug Resistance in Multiple Myeloma. Int. J. Mol. Sci. 2023, 24. [Google Scholar] [CrossRef]
- Kroemer, G.; Mariño, G.; Levine, B. Autophagy and the Integrated Stress Response. Mol. Cell 2010, 40, 280–293. [Google Scholar] [CrossRef]
- Perlin, D. S.; Shor, E.; Zhao, Y. Update on Antifungal Drug Resistance. Curr. Clin. Microbiol. Rep. 2015, 2, 84–95. [Google Scholar] [CrossRef]
- Schubert, S.; Barker, K. S.; Znaidi, S.; Schneider, S.; Dierolf, F.; Dunkel, N.; Aïd, M.; Boucher, G.; Rogers, P. D.; Raymond, M.; et al. Regulation of efflux pump expression and drug resistance by the transcription factors Mrr1, Upc2, and Cap1 in Candida albicans. Antimicrob. Agents Chemother. 2011, 55, 2212–2223. [Google Scholar] [CrossRef]
- Shen, J.; Ma, M.; Duan, W.; Huang, Y.; Shi, B.; Wu, Q.; Wei, X. Autophagy Alters the Susceptibility of Candida albicans Biofilms to Antifungal Agents. Microorganisms 2023, 11. [Google Scholar] [CrossRef]
- Huang, Y.; Miao, H.; Lv, Y.; Wang, Y.; Yu, S.; Wei, X. Aspirin Combined with Antifungal Drugs Suppresses Candida albicans Biofilm by Activating Autophagy. J. Microbiol. Biotechnol. 2025, 35. [Google Scholar] [CrossRef]
- Robke, L.; Laraia, L.; Carnero Corrales, M.A.; Konstantinidis, G.; Muroi, M.; Richters, A.; Winzker, M.; Engbring, T.; Tomassi, S.; Watanabe, N.; Osada, H. Phenotypic identification of a novel autophagy inhibitor chemotype targeting lipid kinase VPSAngew. Chem. Int. Ed. 2017, 56, 8153–8157. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, Q.; Ma, K.; Shi, P.; Liu, W.; Huang, Z. Fluconazole inhibits cellular ergosterol synthesis to confer synergism with berberine against yeast cells. J. Glob. Antimicrob. Resist. 2018, 13, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Kamada, Y.; Funakoshi, T.; Shintani, T.; Nagano, K.; Ohsumi, M.; Ohsumi. Tor-mediated induction of autophagy via an Apg1 protein kinase complex. The Journal of Cell. Biol 2000, 150, 1507–13. [Google Scholar] [CrossRef] [PubMed]
- Kamada, Y.; Yoshino, K.; Kondo, C.; Kawamata, T.; Oshiro, N.; Yonezawa, K.; Ohsumi, Y. Tor Directly Controls the Atg1 Kinase Complex To Regulate Autophagy. Mol. Cell. Biol. 2010, 30, 1049–1058. [Google Scholar] [CrossRef] [PubMed]
- Yeh, Y. Y.; Wrasman, K.; Herman, P. K. Autophosphorylation within the Atg1 activation loop is required for both kinase activity and the induction of autophagy in Saccharomyces cerevisiae. Genetics 2010, 185, 871–882. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, B.; Jang, H. S.; Huh, W. K. Atg1-dependent phosphorylation of Vps34 is required for dynamic regulation of the phagophore assembly site and autophagy in Saccharomyces cerevisiae. Autophagy 2023, 19, 2428–2442. [Google Scholar] [CrossRef]
- Papinski, D.; Schuschnig, M.; Reiter, W.; Wilhelm, L.; Barnes, C. A.; Maiolica, A.; Hansmann, I.; Pfaffenwimmer, T.; Kijanska, M.; Stoffel, I.; et al. Early Steps in Autophagy Depend on Direct Phosphorylation of Atg9 by the Atg1 Kinase. Mol. Cell 2014, 53, 471–483. [Google Scholar] [CrossRef]
- Sánchez-Wandelmer, J.; Kriegenburg, F.; Rohringer, S.; Schuschnig, M.; Gómez-Sánchez, R.; Zens, B.; Abreu, S.; Hardenberg, R.; Hollenstein, D.; Gao, J.; et al. Atg4 proteolytic activity can be inhibited by Atg1 phosphorylation. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef]
- Hu, Y.; Reggiori, F. The yeast dynamin-like GTPase Vps1 mediates Atg9 transport to the phagophore assembly site in Saccharomyces cerevisiae. Autophagy Reports 2023, 2. [Google Scholar] [CrossRef]
- Gatica, D.; Wen, X.; Cheong, H.; Klionsky, D. J. Vac8 determines phagophore assembly site vacuolar localization during nitrogen starvation-induced autophagy. Autophagy 2021, 17, 1636–1648. [Google Scholar] [CrossRef]
- Chen, X.; Liang, Y. Vtc4 Promotes the Entry of Phagophores into Vacuoles in the Saccharomyces cerevisiae Snf7 Mutant Cell. Journal of Fungi 2023, 9. [Google Scholar] [CrossRef]
- Shi, H. Bin; Chen, N.; Zhu, X. M.; Su, Z. Z.; Wang, J. Y.; Lu, J. P.; Liu, X. H.; Lin, F. C. The casein kinase MoYck1 regulates development, autophagy, and virulence in the rice blast fungus. Virulence 2019, 10, 719–733. [Google Scholar] [CrossRef]
- Memisoglu, G.; Eapen, V. V.; Yang, Y.; Klionsky, D. J.; Haber, J. E. PP2C phosphatases promote autophagy by dephosphorylation of the Atg1 complex. Proc. Natl. Acad. Sci. U. S. A. 2019, 116, 1613–1620. [Google Scholar] [CrossRef]
- Kondo, A.; Mostofa, M. G.; Miyake, K.; Terasawa, M.; Nafisa, I.; Yeasmin, A. M. S. T.; Waliullah, T. M.; Kanki, T.; Ushimaru, T. Cdc14 Phosphatase Promotes TORC1-Regulated Autophagy in Yeast. J. Mol. Biol. 2018, 430, 1671–1684. [Google Scholar] [CrossRef]
- Wu, Z.; Shi, H.; Li, Y.; Yan, F.; Sun, Z.; Lin, C.; Xu, M.; Lin, F.; Kou, Y.; Tao, Z. Transcriptional Regulation of Autophagy-Related Genes by Sin3 Negatively Modulates Autophagy in Magnaporthe oryzae. Microbiol. Spectr. 2023, 11. [Google Scholar] [CrossRef]
- Zhong, K.; Li, X.; Le, X.; Kong, X.; Zhang, H.; Zheng, X.; Wang, P.; Zhang, Z. MoDnm1 Dynamin Mediating Peroxisomal and Mitochondrial Fission in Complex with MoFis1 and MoMdv1 Is Important for Development of Functional Appressorium in Magnaporthe oryzae. PLoS Pathog. 2016, 12. [Google Scholar] [CrossRef]
- Ohsumi, Y. Historical landmarks of autophagy research. Cell Res. 2014, 24, 9–23. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




