Submitted:
10 April 2026
Posted:
14 April 2026
You are already at the latest version
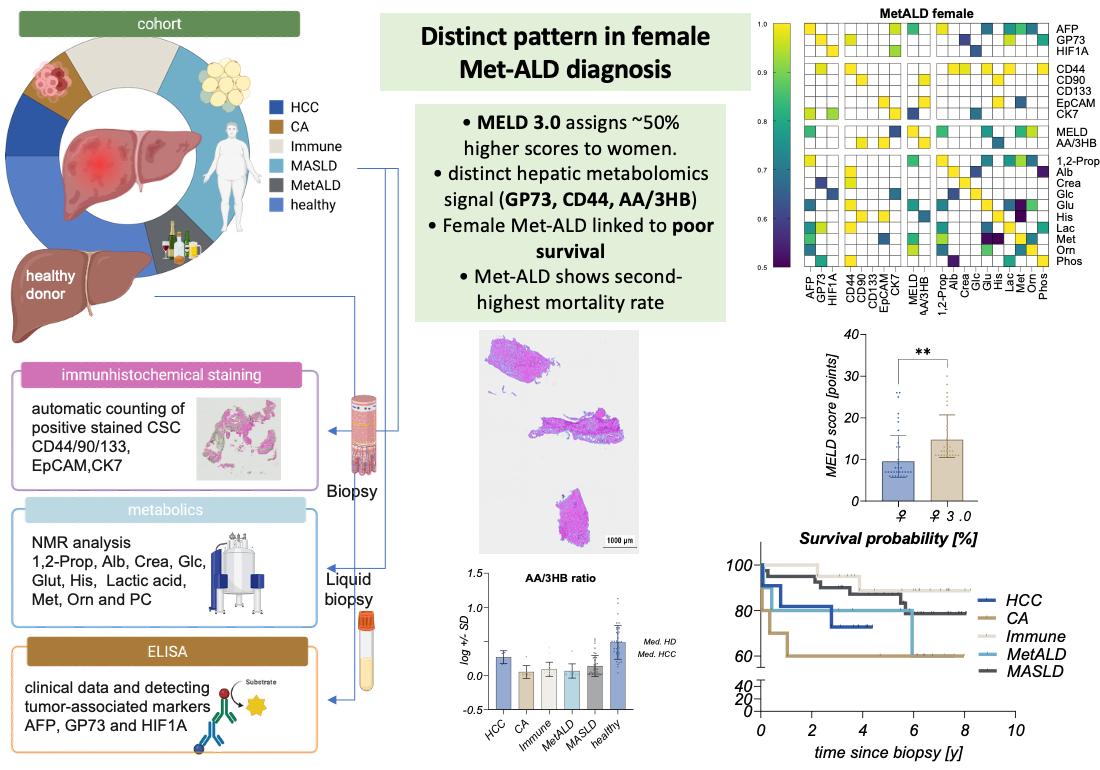
Abstract
Keywords:
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 1,2-Propanediol | 1,2-Propanediol |
| AA/3HB | Acetoacetate/3-Hydroxybutyrate |
| AFP | α-Fetoprotein |
| AKBR | Arterial ketone body ratio |
| ALB | Albumin |
| ALT | Alanin-Aminotransferase |
| ANOVA | analysis of variants |
| CD | Cluster of differentiation |
| CK | Cytokeratin |
| CPT | Child-Torcotte-Pugh |
| Crea | Creatine |
| CSC | Cancer stem cell |
| ELISA | enzyme-linked immunosorbent assay |
| EPCAM | Epithelial Cell Adhesion Molecule |
| Glc | Glucose |
| Glu | Glutamine |
| GP73 | Golgi protein 73 |
| HBV, HCV | hepatitis B, C virus |
| HCC | hepatocellular carcinoma |
| Hif1α | Hypoxinduced factor 1 alpha |
| His | Histidine |
| Lac | Lactic acid |
| MASLD | metabolic dysfunction-associated steatotic liver disease |
| Met | Methionine |
| MetALD | metabolic dysfunction-associated alcohol-related liver disease |
| MELD | Model of endstage liver disease |
| NMR | Nuclear magnetic resonance (spectroscopy) |
| Orn | Ornithine |
| Phos | Phosphatidylcholine |
Appendix A
| Category in manuscript | Clinical term | description |
|---|---|---|
| HCC | Hepatocellular carcinoma | Primary malignant liver tumour arising from hepatocytes |
| CA (Cancerogenic liver disease) |
Lesion suspicious for hepatocellular carcinoma | Cancerogenic hepatic lesion of extrahepatic origin (e.g., metastasis) arising from cholangiocarcinoma or from primary tumours of the breast, colon, or lung |
| Immune | Immune-mediated liver disease | liver injury primarily caused by an aberrant immune response against hepatic or biliary mainly autoimmune origin |
| MetALD | metabolic dysfunction-associated alcohol-related liver disease | Chronic liver disease characterized by hepatic steatosis in the presence of metabolic dysfunction and significant alcohol consumption. |
| MASLD | metabolic dysfunction-associated steatotic liver disease |
Chronic liver disease associated with metabolic dysfunction and hepatic steatosis in the absence of significant alcohol intake |
References
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J Clinicians. 2021 May;71(3):209–49. [CrossRef]
- Rumgay H, Arnold M, Ferlay J, Lesi O, Cabasag CJ, Vignat J, et al. Global burden of primary liver cancer in 2020 and predictions to 2040. Journal of Hepatology. 2022 Dec;77(6):1598–606. [CrossRef]
- Chan SL, Sun HC, Xu Y, Zeng H, El-Serag HB, Lee JM, et al. The Lancet Commission on addressing the global hepatocellular carcinoma burden: comprehensive strategies from prevention to treatment. The Lancet. 2025 Aug;406(10504):731–78. [CrossRef]
- Bowie K. Majority of global liver cancer cases “preventable,” Lancet commission finds. BMJ. 2025 Jul 29;390:r1587. [CrossRef]
- Henry-Blake C, Balachandrakumar V, Kassab M, Devonport J, Matthews C, Fox J, et al. Lower hepatocellular carcinoma surveillance in metabolic dysfunction-associated steatotic liver disease: Impact on treatment eligibility. J of Gastro and Hepatol. 2024 Dec;39(12):2817–25. [CrossRef]
- Qiu S, Cai J, Yang Z, He X, Xing Z, Zu J, et al. Trends in Hepatocellular Carcinoma Mortality Rates in the US and Projections Through 2040. JAMA Netw Open. 2024 Nov 18;7(11):e2445525. [CrossRef]
- Berzigotti A, Tsochatzis E, Boursier J, Castera L, Cazzagon N, Friedrich-Rust M, et al. EASL Clinical Practice Guidelines on non-invasive tests for evaluation of liver disease severity and prognosis – 2021 update. Journal of Hepatology. 2021 Sep;75(3):659–89. [CrossRef]
- Galle PR, Forner A, Llovet JM, Mazzaferro V, Piscaglia F, Raoul JL, et al. EASL Clinical Practice Guidelines: Management of hepatocellular carcinoma. Journal of Hepatology. 2018 Jul;69(1):182–236. [CrossRef]
- Elderkin J, Al Hallak N, Azmi AS, Aoun H, Critchfield J, Tobon M, et al. Hepatocellular Carcinoma: Surveillance, Diagnosis, Evaluation and Management. Cancers. 2023 Oct 24;15(21):5118. [CrossRef]
- Strandberg R, Åberg F, Asteljoki JV, Luukkonen PK, Salomaa V, Jula A, et al. Use of new CORE risk score to predict 10 year risk of liver cirrhosis in general population: population based cohort study. BMJ. 2025 Sep 29;390:e083182. [CrossRef]
- Graupera I, Thiele M, Serra-Burriel M, Caballeria L, Roulot D, Wong GLH, et al. Low Accuracy of FIB-4 and NAFLD Fibrosis Scores for Screening for Liver Fibrosis in the Population. Clinical Gastroenterology and Hepatology. 2022 Nov;20(11):2567-2576.e6. [CrossRef]
- Archer AJ, Belfield KJ, Orr JG, Gordon FH, Abeysekera KW. EASL clinical practice guidelines: non-invasive liver tests for evaluation of liver disease severity and prognosis. Frontline Gastroenterol. 2022 Sep;13(5):436–9. [CrossRef]
- Neureiter D, Kiesslich T, Ocker M. Translating Fibrosis to Malignancy: Biomarkers and Therapeutic Opportunities in Liver Fibrosis and Hepatocellular Carcinoma. Medical Sciences. 2026 Feb 25;14(1):110. [CrossRef]
- Sangro B, Argemi J, Ronot M, Paradis V, Meyer T, Mazzaferro V, et al. EASL Clinical Practice Guidelines on the management of hepatocellular carcinoma. Journal of Hepatology. 2025 Feb;82(2):315–74. [CrossRef]
- Jee YM, Lee JY, Ryu T. Chronic Inflammation and Immune Dysregulation in Metabolic-Dysfunction-Associated Steatotic Liver Disease Progression: From Steatosis to Hepatocellular Carcinoma. Biomedicines. 2025 May 21;13(5):1260. [CrossRef]
- Du D, Liu C, Qin M, Zhang X, Xi T, Yuan S, et al. Metabolic dysregulation and emerging therapeutical targets for hepatocellular carcinoma. Acta Pharmaceutica Sinica B. 2022 Feb;12(2):558–80. [CrossRef]
- Chen TH, Lin SH, Lee MY, Wang HC, Tsai KF, Chou CK. Mitochondrial alterations and signatures in hepatocellular carcinoma. Cancer Metastasis Rev. 2025 Mar;44(1):34. [CrossRef]
- Addison T. Observations on fatty degeneration of the liver. In: Guy’s Hospital reports. London: S. Highley. p. 476–85.
- Véronique Terrasse. Alcohol: a major preventable cause of cancer [Pressemitteilung] [Internet]. [cited 2025 Oct 18]. (IARC Evidence Summary Brief No. 6). Report. Available from: https://www.iarc.who.int/wp-content/uploads/2025/10/IARC_Evidence_Summary_Brief_6.pdf.
- Rumgay H, Ferlay J, De Martel C, Georges D, Ibrahim AS, Zheng R, et al. Global, regional and national burden of primary liver cancer by subtype. European Journal of Cancer. 2022 Jan;161:108–18. [CrossRef]
- WHO. European Health Alliance on Alcohol launched to reduce the unsustainable toll of alcohol harms in Europe [Internet]. 2025 May 8 [cited 2025 Oct 18]. Available from: https://www.who.int/europe/news/item/08-05-2025-european-health-alliance-on-alcohol-launched-to-reduce-the-unsustainable-toll-of-alcohol-harms-in-europe.
- Rinella ME, Lazarus JV, Ratziu V, Francque SM, Sanyal AJ, Kanwal F, et al. A multisociety Delphi consensus statement on new fatty liver disease nomenclature. Journal of Hepatology. 2023 Dec;79(6):1542–56. [CrossRef]
- Younossi ZM, Kalligeros M, Henry L. Epidemiology of metabolic dysfunction-associated steatotic liver disease. Clin Mol Hepatol. 2025 Feb 28;31(Suppl):S32–50. [CrossRef]
- Swarup S, Ahmed I, Grigorova Y, Zeltser R. Metabolic Syndrome. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 [cited 2025 Oct 18]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK459248/ PubMed PMID: 29083742.
- Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J Clinicians. 2024 May;74(3):229–63. [CrossRef]
- McNeil CA, Pramfalk C, Humphreys SM, Hodson L. The storage stability and concentration of acetoacetate differs between blood fractions. Clinica Chimica Acta. 2014 Jun;433:278–83. [CrossRef]
- Yamaoka K, Kanayama M, Tajiri K, Yamane M, Marumo F, Sato C. Clinical Significance of Arterial Ketone Body Ratio in Chronic Liver Disease. Digestion. 1998;59(4):360–3. [CrossRef]
- Kemperman RH, Ganetzky RD, Master SR. B-169 Beta-hydroxybutyrate/acetoacetate Ratio as Indicator for Mitochondrial Diseases Utilizing a Novel LC-MS/MS Based Ketone Body Panel. Clinical Chemistry. 2023 Oct 1;69(Supplement_1):hvad097.501. [CrossRef]
- Yassen KA, Galley HF, Lee A, Webster NR. Mitochondrial redox state in the critically ill. Br J Anaesth. 1999 Aug;83(2):325–7. PubMed PMID: 10618950. [CrossRef]
- Dishart MK, Schlichtig R, Tønnessen TI, Rozenfeld RA, Simplaceanu E, Williams D, et al. Mitochondrial redox state as a potential detector of liver dysoxia in vivo. Journal of Applied Physiology. 1998 Mar;84(3):791–7. [CrossRef]
- Mey JT, Erickson ML, Axelrod CL, King WT, Flask CA, McCullough AJ, et al. β-Hydroxybutyrate is reduced in humans with obesity-related NAFLD and displays a dose-dependent effect on skeletal muscle mitochondrial respiration in vitro. American Journal of Physiology-Endocrinology and Metabolism. 2020 Jul 1;319(1):E187–95. [CrossRef]
- Liu Y, Hong Z, Tan G, Dong X, Yang G, Zhao L, et al. NMR and LC/MS-based global metabolomics to identify serum biomarkers differentiating hepatocellular carcinoma from liver cirrhosis: Serum Biomarkers in Hepatocellular Carcinoma. Int J Cancer. 2014 Aug 1;135(3):658–68. [CrossRef]
- Liu Z, Zheng Y, Li Y, West China School of Medicine, Sichuan University, Chengdu, China, College of Computer Science, Sichuan University, Chengdu, China, Department of General Surgery, the First Affiliated Hospital of Wenzhou Medical University, China, et al. Association between ketone body levels and chronic liver disease: Epidemiological studies and potential mechanisms. AIMSPH. 2026;13(1):82–104. [CrossRef]
- Laffel L. Ketone bodies: a review of physiology, pathophysiology and application of monitoring to diabetes. Diabetes/Metabolism Research and Reviews. 1999;15(6):412–26. [CrossRef]
- Wishart DS. Emerging applications of metabolomics in drug discovery and precision medicine. Nat Rev Drug Discov. 2016 Jul;15(7):7. [CrossRef]
- Kim HY. Recent advances in nonalcoholic fatty liver disease metabolomics. Clin Mol Hepatol. 2021 Oct;27(4):553–9. PubMed PMID: 34098712; PubMed Central PMCID: PMC8524071. [CrossRef]
- Mudgal P, Kumar Gupta S, Malik S, Nith RB, Kumar S, K R, et al. Biomarker discovery in NAFLD: insights from metabolomics and vote counting meta-analysis. Metabolomics. 2025 Aug 12;21(5):116. [CrossRef]
- Kano N, Want EJ, McPhail MJW. Metabolomics in Advanced Liver Disease. Curr Treat Options Gastro. 2021 Jun 1;19(2):380–97. [CrossRef]
- Daniels NJ, Hershberger CE, Kerosky M, Wehrle CJ, Raj R, Aykun N, et al. Biomarker Discovery in Liver Disease Using Untargeted Metabolomics in Plasma and Saliva. IJMS. 2024 Sep 21;25(18):10144. [CrossRef]
- Semenza GL. HIF-1 and tumor progression: pathophysiology and therapeutics. Trends in Molecular Medicine. 2002 Apr;8(4):S62–7. [CrossRef]
- Galle PR, Foerster F, Kudo M, Chan SL, Llovet JM, Qin S, et al. Biology and significance of alpha-fetoprotein in hepatocellular carcinoma. Liver Int. 2019 Dec;39(12):2214–29. [CrossRef]
- Kim D, Cho E, Yu KS, Jang IJ, Yoon JH, Park T, et al. Comprehensive Metabolomic Search for Biomarkers to Differentiate Early Stage Hepatocellular Carcinoma from Cirrhosis. Cancers. 2019 Oct 6;11(10):1497. [CrossRef]
- Ishimoto T, Nagano O, Yae T, Tamada M, Motohara T, Oshima H, et al. CD44 Variant Regulates Redox Status in Cancer Cells by Stabilizing the xCT Subunit of System xc− and Thereby Promotes Tumor Growth. Cancer Cell. 2011 Mar;19(3):387–400. [CrossRef]
- Lee TKW, Guan XY, Ma S. Cancer stem cells in hepatocellular carcinoma — from origin to clinical implications. Nat Rev Gastroenterol Hepatol. 2022 Jan;19(1):26–44. [CrossRef]
- Yamashita T, Kaneko S. Orchestration of hepatocellular carcinoma development by diverse liver cancer stem cells. J Gastroenterol. 2014 Jul;49(7):1105–10. [CrossRef]
- Magee JA, Piskounova E, Morrison SJ. Cancer Stem Cells: Impact, Heterogeneity, and Uncertainty. Cancer Cell. 2012 Mar;21(3):283–96. [CrossRef]
- Zhang K, Che S, Su Z, Zheng S, Zhang H, Yang S, et al. CD90 promotes cell migration, viability and sphere-forming ability of hepatocellular carcinoma cells. Int J Mol Med. 2017 Dec 8. [CrossRef]
- Kinsey E, Lee HM. Management of Hepatocellular Carcinoma in 2024: The Multidisciplinary Paradigm in an Evolving Treatment Landscape. Cancers. 2024 Feb 4;16(3):666. [CrossRef]
- Rodriguez LA, Schmittdiel JA, Liu L, Macdonald BA, Balasubramanian S, Chai KP, et al. Hepatocellular Carcinoma in Metabolic Dysfunction-Associated Steatotic Liver Disease. JAMA Netw Open. 2024 Jul 11;7(7):e2421019. [CrossRef]
- Tan DJH, Ng CH, Lin SY, Pan XH, Tay P, Lim WH, et al. Clinical characteristics, surveillance, treatment allocation, and outcomes of non-alcoholic fatty liver disease-related hepatocellular carcinoma: a systematic review and meta-analysis. The Lancet Oncology. 2022 Apr;23(4):521–30. [CrossRef]
- Tsochatzis EA, Bosch J, Burroughs AK. Liver cirrhosis. The Lancet. 2014 May;383(9930):1749–61. [CrossRef]
- Sarich P, Canfell K, Egger S, Banks E, Joshy G, Grogan P, et al. Alcohol consumption, drinking patterns and cancer incidence in an Australian cohort of 226,162 participants aged 45 years and over. Br J Cancer. 2021 Jan 19;124(2):513–23. [CrossRef]
- Esser MB, Sherk A, Liu Y, Henley SJ, Naimi TS. Reducing Alcohol Use to Prevent Cancer Deaths: Estimated Effects Among U.S. Adults. American Journal of Preventive Medicine. 2024 Apr;66(4):725–9. [CrossRef]
- Leal-Lassalle H, Estévez-Vázquez O, Cubero FJ, Nevzorova YA. Metabolic and alcohol-associated liver disease (MetALD): a representation of duality. npj Gut Liver. 2025 Jan 10;2(1):1. [CrossRef]
- Bosque-Prous M, Espelt A, Borrell C, Bartroli M, Guitart AM, Villalbí JR, et al. Gender differences in hazardous drinking among middle-aged in Europe: the role of social context and women’s empowerment. Eur J Public Health. 2015 Aug;25(4):698–705. [CrossRef]
- Chen Y, Zhou J, Li J, Feng J, Chen Z, Wang X. Plasma metabolomic analysis of human hepatocellular carcinoma: Diagnostic and therapeutic study. Oncotarget. 2016 Jul 26;7(30):47332–42. [CrossRef]
- Huang F, Jiang H, Shen M, Zhang C, Chen Y, Pan B, et al. Plasma metabolomic profiling for hepatocellular carcinoma diagnosis and microvascular invasion prediction. Intl Journal of Cancer. 2025 Nov 15;157(10):2187–96. [CrossRef]
- Fotakis C, Kalafati IP, Amanatidou AI, Andreou V, Matzapetakis M, Kafyra M, et al. Serum metabolomic profiling unveils distinct sex-related metabolic patterns in NAFLD. Front Endocrinol. 2023 Oct 3;14:1230457. [CrossRef]
- Balcar L, Scheiner B, Fulgenzi CAM, D’Alessio A, Pomej K, Roig MB, et al. A meta-analysis and real-world cohort study on the sex-related differences in efficacy and safety of immunotherapy for hepatocellular carcinoma. JHEP Reports. 2024 Feb;6(2):100982. [CrossRef]
- Erlandsen EJ, Randers E. Reference intervals for plasma cystatin C and plasma creatinine in adults using methods traceable to international calibrators and reference methods. Clinical Laboratory Analysis. 2018 Jul;32(6):e22433. [CrossRef]
- Krishnan A, Woreta TA, Vaidya D, Liu Y, Hamilton JP, Hong K, et al. MELD or MELD-Na as a Predictive Model for Mortality Following Transjugular Intrahepatic Portosystemic Shunt Placement. J Clin Transl Hepatol. 2022 Jul 13;000(000):000–000. [CrossRef]
- Varnaccia G, Manz K, Zeiher J, Rattay P, Lange C. Risky Alcohol Consumption and Heavy Episodic Drinking among Parents in Germany: Results of a Nationwide Cross-Sectional Study. Journal of Environmental and Public Health. 2019 Dec 5;2019:1–13. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




