Submitted:
13 April 2026
Posted:
15 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. The Tolerogenic Default: The Complete Mechanism
2.1. The Upstream Cause
2.2. The Silencing Event
2.3. The Seven Locks on IL-12
2.4. Bilateral Disarmament
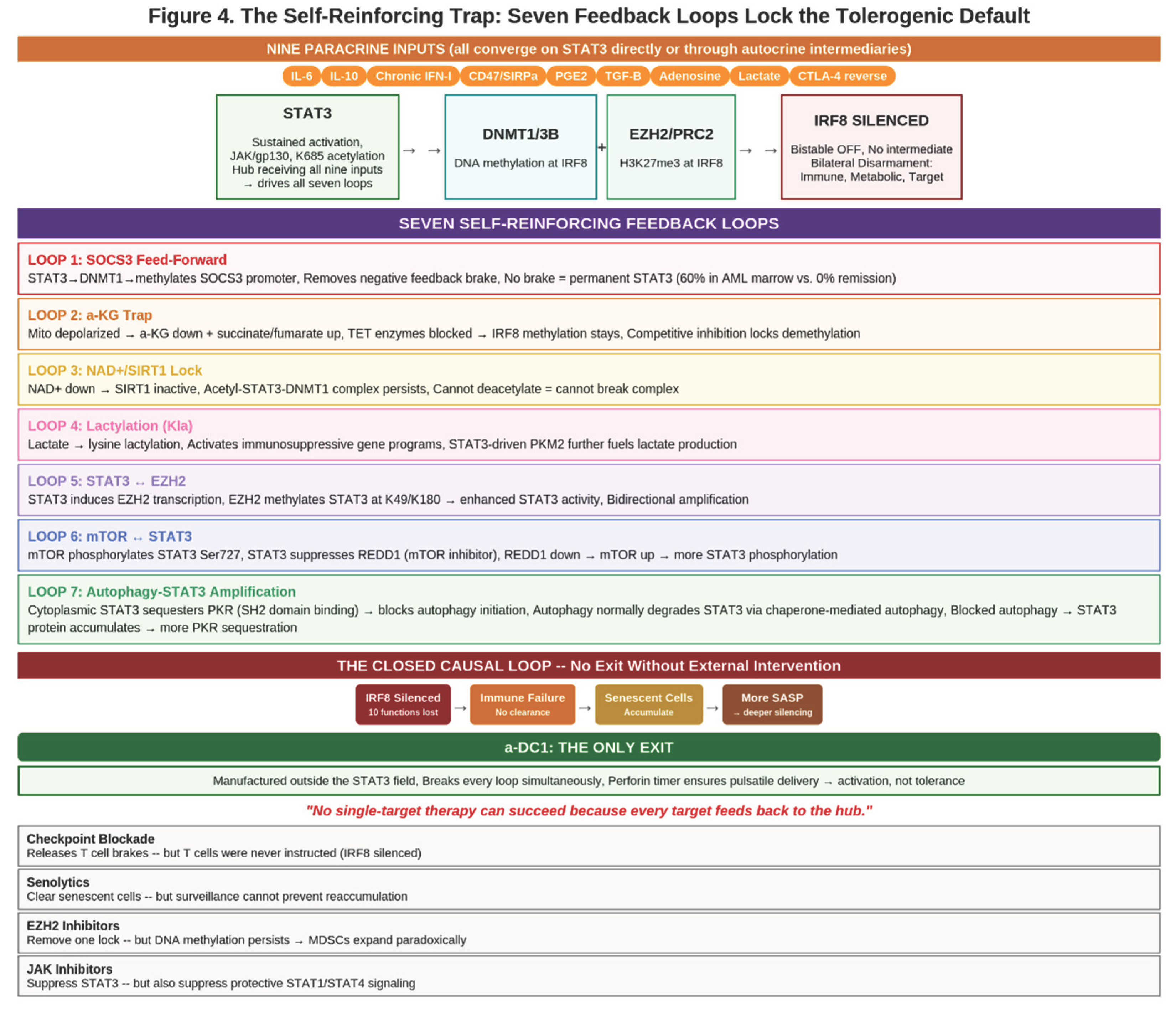
2.5. The Self-Reinforcing Architecture
2.6. Six Lines of Convergent Proof
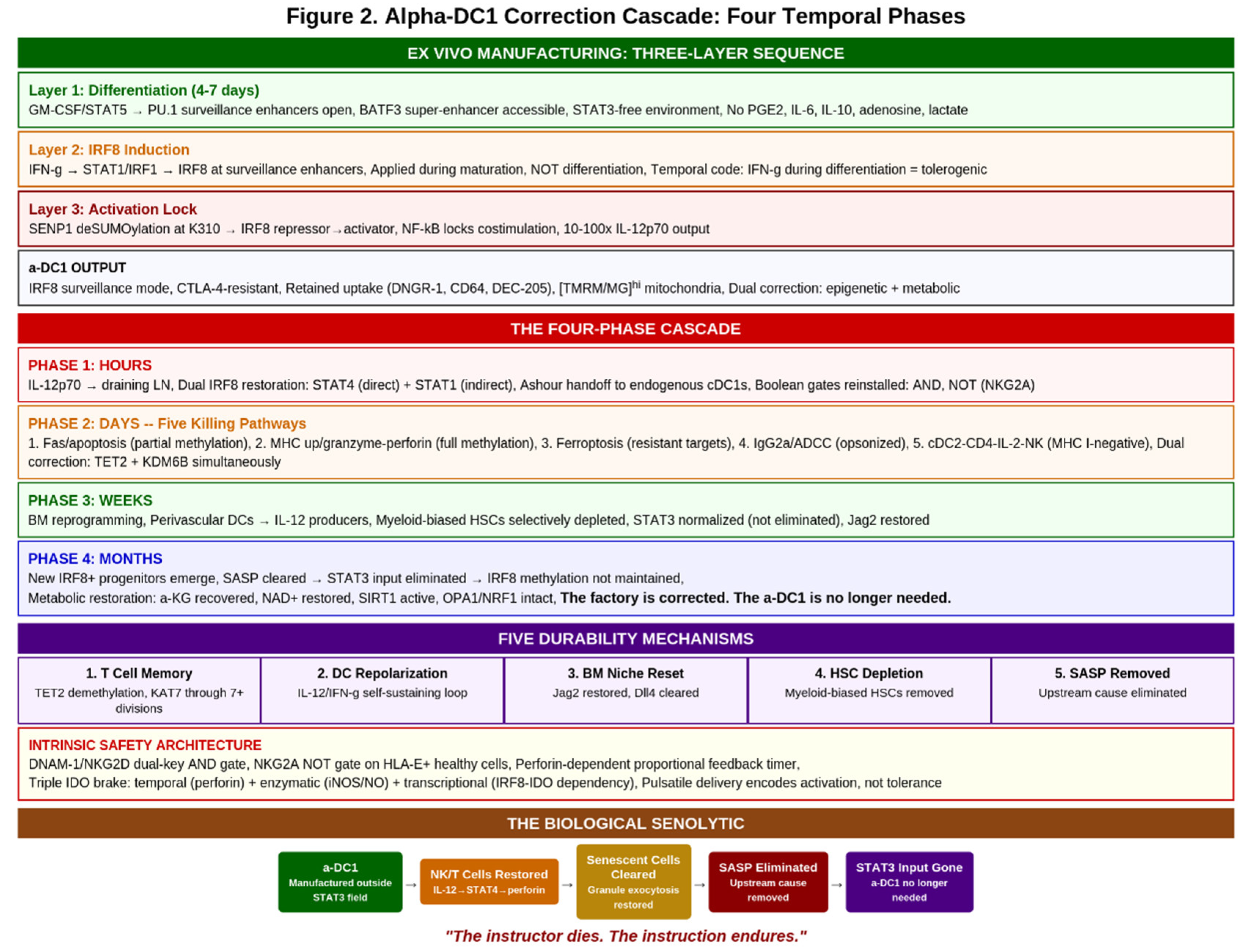
3. The Alpha-DC1 as Restored Instructor
3.1. Manufacturing as Chromatin Programming
3.2. Bidirectional Discrimination and Breaking the Tolerogenic Loop
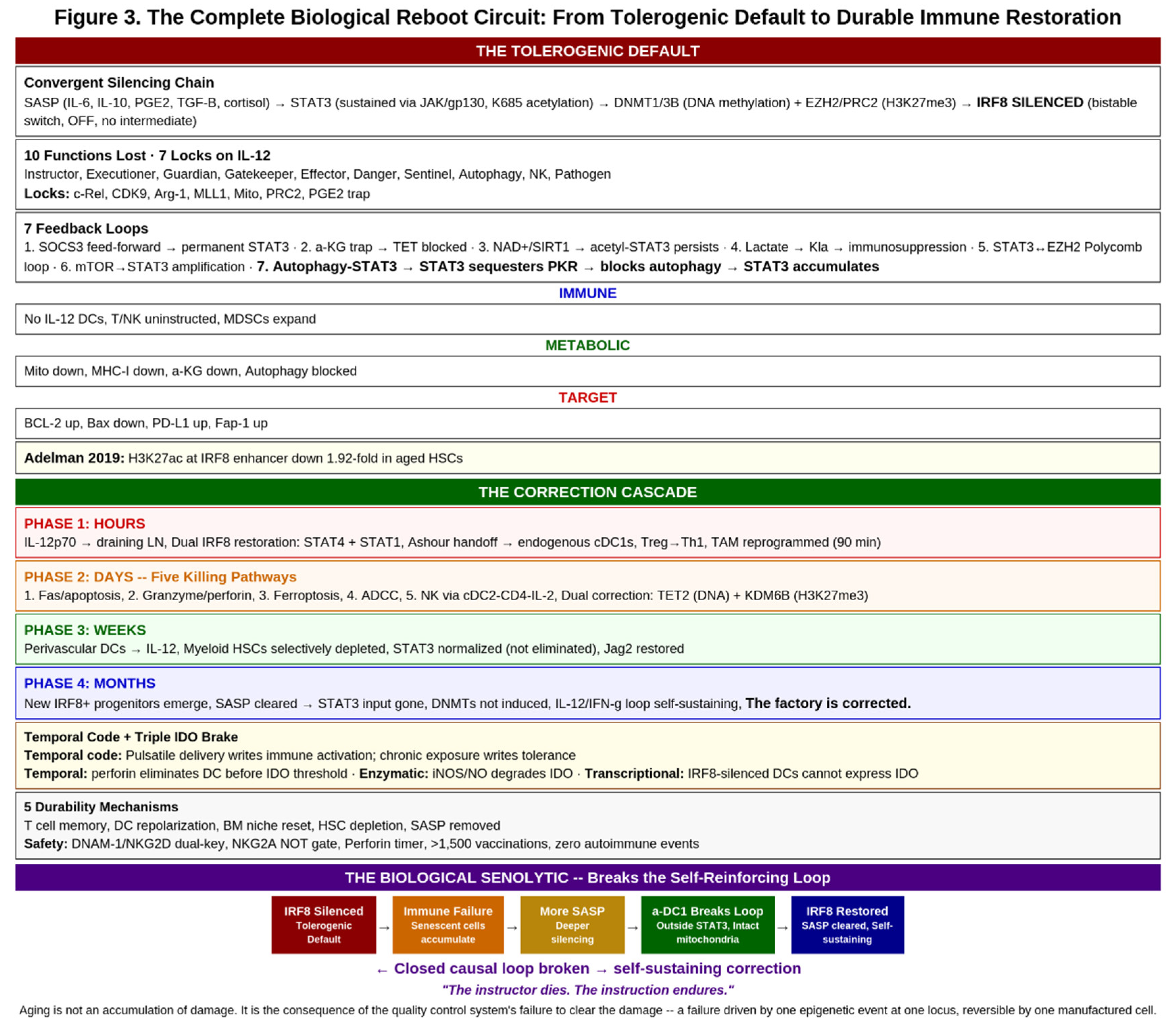
4. The Correction Cascade
4.1. Four Temporal Phases
4.2. Resolution, Durability, and Epigenetic Persistence
5. Platform Architecture and Clinical Evidence
6. Discussion
6.1. Summary of the Framework
6.2. Limitations
6.3. Relationship to Existing Paradigms
6.4. Testable Predictions
6.5. Broader Implications
6.6. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest Statement
Abbreviations
References
- Steinman, R.M.; Cohn, Z.A. Identification of a novel cell type in peripheral lymphoid organs of mice. J Exp Med 1973, 137, 1142–1162. [Google Scholar] [CrossRef] [PubMed]
- Banchereau, J.; Steinman, R.M. Dendritic cells and the control of immunity. Nature 1998, 392, 245–252. [Google Scholar] [CrossRef]
- Hawiger, D.; Inaba, K.; Dorsett, Y.; et al. Dendritic cells induce peripheral T cell unresponsiveness under steady state conditions in vivo. J Exp Med 2001, 194, 769–779. [Google Scholar] [CrossRef] [PubMed]
- Krummel, M.F.; Allison, J.P. CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. J Exp Med 1995, 182, 459–465. [Google Scholar] [CrossRef] [PubMed]
- Leach, D.R.; Krummel, M.F.; Allison, J.P. Enhancement of antitumor immunity by CTLA-4 blockade. Science 1996, 271, 1734–1736. [Google Scholar] [CrossRef]
- Sakaguchi, S.; Sakaguchi, N.; Asano, M.; et al. Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). J Immunol 1995, 155, 1151–1164. [Google Scholar] [CrossRef]
- Hori, S.; Nomura, T.; Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 2003, 299, 1057–1061. [Google Scholar] [CrossRef]
- Wing, K.; Onishi, Y.; Prieto-Martin, P.; et al. CTLA-4 control over Foxp3+ regulatory T cell function. Science 2008, 322, 271–275. [Google Scholar] [CrossRef]
- Qureshi, O.S.; Zheng, Y.; Nakamura, K.; et al. Trans-endocytosis of CD80 and CD86: A molecular basis for the cell-extrinsic function of CTLA-4. Science 2011, 333, 261–263. [Google Scholar] [CrossRef]
- Kennedy, A.; Waters, E.; Rowshanravan, B.; et al. Differences in CD80 and CD86 transendocytosis reveal CD86 as a key target for CTLA-4 immune regulation. Nat Immunol 2022, 23, 1365–1378. [Google Scholar] [CrossRef]
- Mailliard, R.B.; Wieckowski, E.; Muthuswamy, R.; et al. Alpha-type-1 polarized dendritic cells: A novel immunization tool with optimized CTL-inducing activity. Cancer Res 2004, 64, 5934–5937. [Google Scholar] [CrossRef] [PubMed]
- Hansen, M.; Hjorto, G.M.; Donia, M.; et al. Comparison of clinical grade type 1-polarized and standard matured dendritic cells for cancer immunotherapy. Vaccine 2013, 31, 639–646. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Obermajer, N.; Tsuji, T.; et al. Contributions of DNAM-1 and NKG2D to functional activation of human CTLs. Cancer Immunol Res 2024, 12, 339–353. [Google Scholar] [CrossRef]
- Feng, T.; Cao, A.T.; Weaver, C.T.; et al. Interleukin-12 converts Foxp3+ regulatory T cells to interferon-gamma-producing Foxp3+ T cells that inhibit colitis. Gastroenterology (2011) 140, 2031–2043. [CrossRef]
- Wilson, C.B.; Makar, K.W.; Shnyreva, M.; et al. DNA methylation and the expanding epigenetics of T cell lineage commitment. Semin Immunol 2005, 17, 105–119. [Google Scholar] [CrossRef]
- Qiao, Y.; Giannopoulou, E.G.; Chan, C.H.; et al. Synergistic activation of inflammatory cytokine genes by interferon-gamma-induced chromatin remodeling and toll-like receptor signaling. Immunity 2013, 39, 454–469. [Google Scholar] [CrossRef]
- Ostuni, R.; Piccolo, V.; Barozzi, I.; et al. Latent enhancers activated by stimulation in differentiated cells. Cell 2013, 152, 157–171. [Google Scholar] [CrossRef]
- Mikulski, P.; Schubert, S.; Giesselmann, S.; et al. IFN-gamma-induced chromatin memory in human cells is maintained through mitotic divisions by the histone acetyltransferase KAT7. Nat Struct Mol Biol 2025, 32, 512–523. [Google Scholar] [CrossRef]
- Cowley, C.J.; Sajjath, S.M.; Soto-Ugaldi, L.F.; et al. Distinctive DNA sequence features define epigenetic longevity of inflammatory memory. Science 2026, 391(6792), 1347–1354. [Google Scholar] [CrossRef]
- Subbiah, V.; Murthy, R.; Hong, D.S.; et al. Cytokines produced by dendritic cells administered intratumorally correlate with clinical outcome in patients with diverse cancers. Clin Cancer Res 2018, 24, 3845–3856. [Google Scholar] [CrossRef]
- Liau, L.M.; Ashkan, K.; Brem, S.; et al. Association of autologous tumor lysate-loaded dendritic cell vaccination with extension of survival among patients with newly diagnosed and recurrent glioblastoma. JAMA Oncol 2023, 9, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Jackson, A.L.; Matsumoto, H.; Janszen, M.; et al. Restricted expression of p55 interleukin 2 receptor (CD25) on normal T cells. Clin Immunol Immunopathol 1990, 54, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Coppe, J.P.; Patil, C.K.; Rodier, F.; et al. Senescence-associated secretory phenotypes reveal cell nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol 2008, 6, e301. [Google Scholar] [CrossRef] [PubMed]
- Manz, M.G.; Boettcher, S. Emergency granulopoiesis. Nat Rev Immunol 2014, 14, 302–314. [Google Scholar] [CrossRef]
- Park, S.J.; Nakagawa, T.; Kitamura, H.; et al. IL-6 regulates in vivo dendritic cell differentiation through STAT3 activation. J Immunol 2004, 173, 3844–3854. [Google Scholar] [CrossRef]
- Kalinski, P.; Hilkens, C.M.; Snijders, A.; et al. IL-12-deficient dendritic cells, generated in the presence of prostaglandin E2, promote type 2 cytokine production in maturing human naive T helper cells. J Immunol 1997, 159, 28–35. [Google Scholar] [CrossRef]
- Gabrilovich, D.I.; Nagaraj, S. Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol 2009, 9, 162–174. [Google Scholar] [CrossRef]
- Spranger, S.; Bao, R.; Gajewski, T.F. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature 2015, 523, 231–235. [Google Scholar] [CrossRef]
- Goriely, S.; Demonte, D.; Ziegler-Heitbrock, L.; et al. Human IL-12(p35) gene activation involves selective remodeling of a single nucleosome within a region of the promoter containing critical Sp1-binding sites. Blood 2003, 101, 4894–4902. [Google Scholar] [CrossRef]
- Tamura, T.; Yanai, H.; Savitsky, D.; et al. The IRF family transcription factors in immunity and oncogenesis. Annu Rev Immunol 2008, 26, 535–584. [Google Scholar] [CrossRef]
- Salem, S.; Salem, D.; Gros, P. Role of IRF8 in immune cells functions, protection against infections, and susceptibility to inflammatory diseases. Hum Genet 2020, 139, 707–721. [Google Scholar] [CrossRef]
- Langlais, D.; Barreiro, L.B.; Gros, P. The macrophage IRF8/IRF1 regulome is required for protection against infections and is associated with chronic inflammation. J Exp Med 2016, 213, 585–603. [Google Scholar] [CrossRef]
- Ibrahim, M.L.; Klement, J.D.; Lu, C.; et al. Myeloid-derived suppressor cells produce IL-10 to elicit DNMT3b-dependent IRF8 silencing to promote colitis-associated colon tumorigenesis. Cell Rep 2018, 25, 3036–3046.e6. [Google Scholar] [CrossRef] [PubMed]
- McGough, J.M.; Yang, D.; Huang, S.; et al. DNA methylation represses IFN-gamma-induced and signal transducer and activator of transcription 1-mediated IFN regulatory factor 8 activation in colon carcinoma cells. Mol Cancer Res 2008, 6, 1841–1851. [Google Scholar] [CrossRef] [PubMed]
- Ohno, Y.; Kitamura, H.; Takahashi, N.; et al. IL-6 down-regulates HLA class II expression and IL-12 production of human dendritic cells to impair activation of antigen-specific CD4+ T cells. Cancer Immunol Immunother 2016, 65, 193–204. [Google Scholar] [CrossRef] [PubMed]
- Pathak, S.; Ma, S.; Shukla, V.; et al. A role for IRF8 in B cell anergy. J Immunol 2013, 191, 6222–6230. [Google Scholar] [CrossRef]
- Zhao, B.; Takami, M.; Yamada, A.; et al. Interferon regulatory factor-8 regulates bone metabolism by suppressing osteoclastogenesis. Nat Med 2009, 15, 1066–1071. [Google Scholar] [CrossRef]
- Lung, T.L.; Saurwein-Teissl, M.; Parson, W.; et al. Unimpaired dendritic cells can be derived from monocytes in old age and can mobilize residual function in senescent T cells. Vaccine 2000, 18, 1606–1612. [Google Scholar] [CrossRef]
- Zhou, J.; Tison, K.; Zhou, H.; et al. STAT5 and STAT3 balance shapes dendritic cell function and tumour immunity. Nature 2025, 643, 519–528. [Google Scholar] [CrossRef]
- Zhivaki, D.; Kennedy, S.N.; Park, J.; et al. Correction of age-associated defects in dendritic cells enables CD4+ T cells to eradicate tumors. Cell 2024, 187, 3888–3903.e18. [Google Scholar] [CrossRef]
- Tsuji, S.; Matsumoto, M.; Takeuchi, O.; et al. Maturation of human dendritic cells by cell wall skeleton of Mycobacterium bovis bacillus Calmette-Guerin: Involvement of toll-like receptors. Infect Immun 2000, 68, 6883–6890. [Google Scholar] [CrossRef] [PubMed]
- Shankar, G.; Pestano, L.A.; Bosch, M.L. Interferon-gamma added during bacillus Calmette-Guerin induced dendritic cell maturation stimulates potent Th1 immune responses. J Transl Med 2003, 1, 7. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.H.; Xu, S.; Tailor, P.; et al. The small ubiquitin-like modifier-deconjugating enzyme sentrin-specific peptidase 1 switches IFN regulatory factor 8 from a repressor to an activator during macrophage activation. J Immunol 2012, 189, 3548–3556. [Google Scholar] [CrossRef] [PubMed]
- Fesneau, O.; Theate, I.; Caux, C.; et al. IL-12 drives NKG2A expression on tumor-reactive CD8+ T cells. Immunity 2024, 57, 1614–1631. [Google Scholar]
- Ashour, D.; Arampatzi, P.; Pavlovic, V.; et al. IL-12 from endogenous cDC1, and not vaccine DC, is required for Th1 induction. J Clin Invest 2020, 130, 3051–3060. [Google Scholar] [CrossRef]
- Matatall, K.A.; Jeong, M.; Chen, S.; et al. Chronic infection depletes hematopoietic stem cells through stress-induced terminal differentiation. Cell Rep 2016, 17, 2584–2595. [Google Scholar] [CrossRef]
- Panda, A.; Qian, F.; Mohanty, S.; et al. Age-associated decrease in TLR function in primary human dendritic cells predicts influenza vaccine response. J Immunol 2010, 184, 2518–2527. [Google Scholar] [CrossRef]
- Compte, N.; Zouaoui Boudjeltia, K.; Vanhaeverbeek, M.; et al. Frailty in old age is associated with decreased interleukin-12/23 production in response to toll-like receptor ligation. PLoS One 2013, 8, e65325. [Google Scholar] [CrossRef]
- Guo, Z.; Tilburgs, T.; Wong, B.; et al. Dysfunction of dendritic cells in aged C57BL/6 mice leads to failure of natural killer cell activation and of tumor eradication. Proc Natl Acad Sci U S A 2014, 111, 14199–14204. [Google Scholar] [CrossRef]
- Wagner, A.H.; Conzelmann, M.; Fitzer, F.; et al. JAK1/STAT3 activation directly inhibits IL-12 production in dendritic cells by preventing CDK9/P-TEFb recruitment to the p35 promoter. Biochem Pharmacol 2015, 96, 52–64. [Google Scholar] [CrossRef]
- Kim, D.; Park, G.; Huuhtanen, J.; et al. STAT3 activation in large granular lymphocyte leukemia is associated with cytokine signaling and DNA hypermethylation. Leukemia 2021, 35, 3430–3443. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Canales, D.; Krishnan, R.; Jessup, C.F.; et al. Early exposure of interferon-gamma inhibits signal transducer and activator of transcription-6 signalling and nuclear factor kappaB activation in a short-term monocyte-derived dendritic cell culture promoting regulatory dendritic cells. Clin Exp Immunol 2012, 167, 447–458. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.M.; Qualls, J.E.; O’Brien, K.; et al. A distal enhancer in Il12b is the target of transcriptional repression by the STAT3 pathway and requires the basic leucine zipper (B-ZIP) protein NFIL3. J Biol Chem 2011, 286, 23582–23590. [Google Scholar] [CrossRef] [PubMed]
- Hoentjen, F.; Sartor, R.B.; Ozaki, M.; et al. STAT3 regulates NF-kappaB recruitment to the IL-12p40 promoter in dendritic cells. Blood 2005, 105, 689–696. [Google Scholar] [CrossRef]
- Molony, R.D.; Nguyen, J.T.; Kong, Y.; et al. Aging impairs both primary and secondary RIG-I signaling for interferon induction in human monocytes. Sci Signal 2017, 10(509), eaan2392. [Google Scholar] [CrossRef]
- Zhang, Z.; Huang, X.; Wang, E.; et al. Suppression of Mll1-complex by Stat3/Cebpbeta-induced miR-21a/21b/181b maintains the accumulation, homeostasis, and immunosuppressive function of polymorphonuclear myeloid-derived suppressor cells. J Immunol 2020, 204, 3400–3415. [Google Scholar] [CrossRef]
- Zaccard, C.R.; Watkins, S.C.; Kalinski, P.; et al. CD40L induces functional tunneling nanotube networks exclusively in dendritic cells programmed by mediators of type 1 immunity. J Immunol 2015, 194, 1047–1056. [Google Scholar] [CrossRef]
- Wang, W.; Green, M.; Choi, J.E.; et al. CD8+ T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 2019, 569, 270–274. [Google Scholar] [CrossRef]
- Ferris, S.T.; Ohara, R.A.; Ou, F.; et al. cDC1 vaccines drive tumor rejection by direct presentation independently of host cDC1. Cancer Immunol Res 2022, 10, 920–931. [Google Scholar] [CrossRef]
- Khateb, M.; Azriel, A.; Levi, B.Z. The third intron of IRF8 is a cell-type-specific chromatin priming element during mouse embryonal stem cell differentiation. J Mol Biol 2019, 431, 210–222. [Google Scholar] [CrossRef]
- Politis, A.D.; Sivo, J.; Driggers, P.H.; et al. Modulation of interferon consensus sequence binding protein mRNA in murine peritoneal macrophages. J Immunol 1992, 148, 801–22429. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Guan, X.; Tamura, T.; Ozato, K.; Ma, X. Synergistic activation of interleukin-12 p35 gene transcription by interferon regulatory factor-1 and interferon consensus sequence-binding protein. J Biol Chem 2004, 279, 55609–55617. [Google Scholar] [CrossRef] [PubMed]
- Lanca, T.; Ungerback, J.; Da Silva, C.; et al. IRF8 deficiency induces the transcriptional, functional, and epigenetic reprogramming of cDC1 into the cDC2 lineage. Immunity 2022, 55, 1431–1447.e11. [Google Scholar] [CrossRef] [PubMed]
- Burchert, A.; Cai, D.; Hofbauer, L.C.; et al. Interferon consensus sequence binding protein (ICSBP; IRF-8) antagonizes BCR/ABL and down-regulates bcl-2. Blood 2004, 103, 3480–3489. [Google Scholar] [CrossRef]
- Yang, D.; Thangaraju, M.; Green, K.; et al. IFN regulatory factor 8 sensitizes soft tissue sarcoma cells to death receptor-initiated apoptosis via repression of FLICE-like protein. Cancer Res 2009, 69, 1080–1088. [Google Scholar] [CrossRef]
- Hu, X.; Yang, D.; Zimmerman, M.; et al. IRF8 regulates acid ceramidase expression to mediate apoptosis and suppresses tumourigenesis. Nat Commun 2011, 2, 1–12. [Google Scholar] [CrossRef]
- Yang, D.; Wang, S.; Brooks, C.; et al. IFN regulatory factor 8 mediates apoptosis in nonhemopoietic tumor cells via regulation of Fas expression. J Immunol 2007, 179, 4775–4782. [Google Scholar] [CrossRef]
- Huang, W.; Zhu, C.; Wang, H.; et al. IRF8 represses PTPN13/FAP-1 expression and sensitizes tumor cells to FAS-mediated apoptosis. Cancer Immunol Res 2015, 3, 73–83. [Google Scholar] [CrossRef]
- Kurotaki, D.; Yamamoto, M.; Nishiyama, A.; et al. IRF8 inhibits C/EBP-alpha activity to restrain mononuclear phagocyte progenitors from differentiating into neutrophils. Nat Commun 2014, 5, 4978. [Google Scholar] [CrossRef]
- Lee, J.; Zhou, Y.J.; Ma, W.; et al. Lineage specification of human dendritic cells is marked by IRF8 expression in hematopoietic stem cells and multipotent progenitors. Nat Immunol 2017, 18, 877–888. [Google Scholar] [CrossRef]
- Meyer, M.A.; Baer, J.M.; Knolhoff, B.L.; et al. Breast and pancreatic cancer interrupt IRF8-dependent dendritic cell development to overcome immune surveillance. Nat Commun 2018, 9, 1250. [Google Scholar] [CrossRef] [PubMed]
- Miyagawa, F.; Zhang, H.; Terunuma, A.; et al. Interferon regulatory factor 8 integrates T-cell receptor and cytokine-signaling pathways and drives effector differentiation of CD8 T cells. Proc Natl Acad Sci U S A 2012, 109, 12123–12128. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Wei, K.; Sun, Q.; et al. Analysis of the three-dimensional genome of exhausted CD8+ T cells reveals a critical role of IRF8 in their differentiation and functions in cancer. Nat Immunol 2025, 26, 2280–2295. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Zou, Z.; Tao, S.; et al. IRF8 is associated with STING to modulate DNA damage-induced cellular senescence. Nat Commun 2023, 14, 5504. [Google Scholar] [CrossRef]
- Zhao, J.; Kong, H.J.; Li, H.; et al. IRF-8/interferon (IFN) consensus sequence-binding protein is involved in Toll-like receptor (TLR) signaling and contributes to the cross-talk between TLR and IFN-gamma signaling pathways. J Biol Chem 2006, 281, 10073–10080. [Google Scholar] [CrossRef]
- Saeki, K.; Shichino, S.; Ueha, S.; et al. IRF8 configures enhancer landscape in postnatal microglia and directs microglia specific transcriptional programs. Nat Immunol 2024, 25, 1928–1942. [Google Scholar] [CrossRef]
- Yan, J.; Chen, G.; Tong, S.; et al. IRF8 contributes to mucosal immunity by promoting IFN-gamma expression in gastric epithelial cells. J Cell Biochem 2018, 119, 1274–1281. [Google Scholar]
- Gupta, M.; Shin, D.M.; Ramakrishna, L.; et al. IRF8 directs stress-induced autophagy in macrophages and promotes clearance of Listeria monocytogenes. Nat Commun 2015, 6, 6379. [Google Scholar] [CrossRef]
- Adams, N.M.; Lau, C.M.; Fan, X.; et al. Transcription factor IRF8 orchestrates the adaptive natural killer cell response. Immunity 2018, 48, 1172–1182.e6. [Google Scholar] [CrossRef]
- Mace, E.M.; Bigley, V.; Gunesch, J.T.; et al. Biallelic mutations in IRF8 impair human NK cell maturation and function. J Clin Invest 2017, 127, 306–320. [Google Scholar] [CrossRef]
- Alter-Koltunoff, M.; Goren, S.; Nousbeck, J.; et al. Innate immunity to intraphagosomal pathogens is mediated by interferon regulatory factor 8 (IRF-8) that stimulates the expression of macrophage-specific Nramp1 through antagonizing repression by c-Myc. J Biol Chem 2008, 283, 2724–2733. [Google Scholar] [CrossRef] [PubMed]
- Marquis, J.F.; Vachon, V.; Bhatt, D.L.; et al. The IRF-8 regulome includes genome-wide enrichment for GBP and IRG family antimicrobial effectors. Eur J Immunol 2011, 41, 3270–3282. [Google Scholar] [CrossRef]
- Murphy, T.L.; Murphy, K.M. The making of a cDC1: Precision programming of progenitor potential. Immunol Rev 2026, 337, e70081. [Google Scholar] [CrossRef] [PubMed]
- Lehtonen, A.; Lund, R.; Lahesmaa, R.; Julkunen, I.; Sareneva, T.; Matikainen, S. IFN-alpha and IL-12 activate IFN regulatory factor 1 (IRF-1), IRF-4, and IRF-8 gene expression in human NK and T cells. Cytokine 2003, 24, 81–90. [Google Scholar] [CrossRef]
- Zebley, C.C.; Abdelsamed, H.A.; Ghoneim, H.E.; et al. Proinflammatory cytokines promote TET2-mediated DNA demethylation during CD8 T cell effector differentiation. Cell Rep 2021, 37, 109796. [Google Scholar] [CrossRef]
- Stirewalt, D.L.; Choi, Y.E.; Sharpless, N.E.; et al. Decreased IRF8 expression found in aging hematopoietic progenitor/stem cells. Leukemia 2009, 23, 391–393. [Google Scholar] [CrossRef]
- Elsner, R.A.; Smita, S.; Engiles, J.B.; Shlomchik, M.J. Coordinated regulation of extrafollicular B cell responses by IL-12 and IFN-gamma. Immunol Rev 2025, 329, e70027. [Google Scholar] [CrossRef]
- Watkins, S.K.; Egilmez, N.K.; Suttles, J.; et al. IL-12 rapidly alters the functional profile of tumor-associated and tumor-infiltrating macrophages in vitro and in vivo. J Immunol 2007, 178, 1357–1362. [Google Scholar] [CrossRef]
- Ewen, E.M.; Pahl, J.H.W.; Miller, M.; et al. KIR downregulation by IL-12/15/18 unleashes human NK cells from KIR/HLA-I inhibition and enhances killing of tumor cells. Eur J Immunol 2018, 48, 355–365. [Google Scholar] [CrossRef]
- Hambleton, S.; Salem, S.; Bustamante, J.; et al. IRF8 mutations and human dendritic-cell immunodeficiency. N Engl J Med 2011, 365, 127–138. [Google Scholar] [CrossRef]
- Basrai, S.; Nofech-Mozes, I.; Detroja, R.; et al. Blood-based epigenetic instability linked to human aging and disease. Nat Commun 2026, 17, 2754. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Wang, H.Y.; Woetmann, A.; et al. STAT3 induces transcription of the DNA methyltransferase 1 gene (DNMT1) in malignant T lymphocytes. Blood 2006, 108, 1058–1064. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Harden, J.L.; Anderson, C.D.; Egilmez, N.K. Tolerogenic phenotype of IFN-gamma-induced IDO+ dendritic cells is maintained via an autocrine IDO-kynurenine/AhR-IDO loop. J Immunol 2016, 197, 962–970. [Google Scholar] [CrossRef] [PubMed]
- Edelstein, M.P.; Ozaki, Y.; Duch, D.S. Synergistic effects of phorbol ester and IFN-gamma on the induction of indoleamine 2,3-dioxygenase in THP-1 monocytic leukemia cells. J Immunol 1989, 143, 2969–2973. [Google Scholar] [CrossRef]
- Kowalczyk, A.; D’Souza, C.A.; Zhang, L. Cell-extrinsic CTLA4-mediated regulation of dendritic cell maturation depends on STAT3. Eur J Immunol 2014, 44, 1143–1155. [Google Scholar] [CrossRef]
- Yang, J.; Huck, S.P.; McHugh, R.S.; Hermans, I.F.; Ronchese, F. Perforin-dependent elimination of dendritic cells regulates the expansion of antigen-specific CD8+ T cells in vivo. Proc Natl Acad Sci U S A 2006, 103, 147–152. [Google Scholar] [CrossRef]
- Chen, M.; Felix, K.; Wang, J. Critical role for perforin and Fas-dependent killing of dendritic cells in the control of inflammation. Blood 2012, 119, 127–136. [Google Scholar] [CrossRef]
- Harden, J.L.; Gu, T.; Kilinc, M.O.; et al. Dichotomous effects of IFN-gamma on dendritic cell function determine the extent of IL-12-driven antitumor T cell immunity. J Immunol 2011, 187, 126–132. [Google Scholar] [CrossRef]
- Greeneltch, K.M.; Schneider, M.; Steinberg, S.M.; et al. Host immunosurveillance controls tumor growth via IFN regulatory factor-8 dependent mechanisms. Cancer Res 2007, 67, 10406–10416. [Google Scholar] [CrossRef]
- Hu, X.; Bardhan, K.; Paschall, A.V.; et al. Deregulation of apoptotic factors Bcl-xL and Bax confers apoptotic resistance to myeloid-derived suppressor cells and contributes to their persistence in cancer. J Biol Chem 2013, 288, 19103–19115. [Google Scholar] [CrossRef]
- Mattei, F.; Schiavoni, G.; Sestili, P.; et al. IRF-8 controls melanoma progression by regulating the cross talk between cancer and immune cells within the tumor microenvironment. Neoplasia 2012, 14, 1223–1235. [Google Scholar] [CrossRef]
- Montoya, M.; Collins, S.A.; Chuntova, P.; et al. Interferon regulatory factor 8-driven reprogramming of the immune microenvironment enhances antitumor adaptive immunity and reduces immunosuppression in murine glioblastoma. Neuro-Oncology 2024, 26, 2272–2287. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Thangaraju, M.; Greeneltch, K.; et al. Repression of IFN regulatory factor 8 by DNA methylation is a molecular determinant of apoptotic resistance and metastatic phenotype in metastatic tumor cells. Cancer Res 2007, 67, 3301–3309. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.W.; Tong, Z.; Cao, P.; et al. Transcription-independent regulation of STING activation and innate immune responses by IRF8 in monocytes. Nat Commun 2022, 13, 4822. [Google Scholar] [CrossRef] [PubMed]
- Braun, D.A.; Fribourg, M.; Sealfon, S.C. Cytokine response is determined by duration of receptor and signal transducers and activators of transcription 3 (STAT3) activation. J Biol Chem 2013, 288, 2986–2993. [Google Scholar] [CrossRef]
- Qing, Y.; Stark, G.R. Alternative activation of STAT1 and STAT3 in response to interferon-gamma. J Biol Chem 2004, 279, 41679–41685. [Google Scholar] [CrossRef]
- Dees, C.; Pötter, S.; Zhang, Y.; et al. TGF-beta-induced epigenetic deregulation of SOCS3 facilitates STAT3 signaling to promote fibrosis. J Clin Invest 2020, 130, 2347–2363. [Google Scholar] [CrossRef]
- Zhao, B.; Takami, M.; Yamada, A.; et al. Interferon regulatory factor-8 regulates bone metabolism by suppressing osteoclastogenesis. Nat Med 2009, 15, 1066–1071. [Google Scholar] [CrossRef]
- Li, C.C.; Munitic, I.; Mittelstadt, P.R.; et al. Suppression of dendritic cell-derived IL-12 by endogenous glucocorticoids is protective in LPS-induced sepsis. PLoS Biol 2015, 13, e1002269. [Google Scholar] [CrossRef]
- Soulier, A.; Blois, S.M.; Sivakumaran, S.; et al. Cell-intrinsic regulation of murine dendritic cell function and survival by prereceptor amplification of glucocorticoid. Blood 2013, 122, 3288–3297. [Google Scholar] [CrossRef]
- Murphy, T.L.; Murphy, K.M. The making of a cDC1: Precision programming of progenitor potential. Immunol Rev 2026, 337, e70081. [Google Scholar] [CrossRef]
- Liu, T.T.; Kim, S.; Desai, P.; et al. Ablation of cDC2 development by triple mutations within the Zeb2 enhancer. Nature 2022, 607, 142–148. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Zhang, P.; Herrmann, A.; et al. Acetylated STAT3 is crucial for methylation of tumor-suppressor gene promoters and inhibition by resveratrol results in demethylation. Proc Natl Acad Sci U S A 2012, 109, 7765–7769. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Deuring, J.; Peppelenbosch, M.P.; et al. IL-6-induced DNMT1 activity mediates SOCS3 promoter hypermethylation in ulcerative colitis-related colorectal cancer. Carcinogenesis 2012, 33, 1889–1896. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, K.; Zhang, J.; et al. DNMTs-mediated SOCS3 methylation promotes the occurrence and development of AML. Eur J Haematol 2024, 112, 439–449. [Google Scholar] [CrossRef]
- Thomas, S.R.; Mohr, D.; Stocker, R. Nitric oxide inhibits indoleamine 2,3-dioxygenase activity in interferon-gamma primed mononuclear phagocytes. J Biol Chem 1994, 269, 14457–14464. [Google Scholar] [CrossRef]
- Hucke, C.; MacKenzie, C.R.; Adjogble, K.D.; et al. Nitric oxide-mediated regulation of gamma interferon-induced bacteriostasis: Inhibition and degradation of human indoleamine 2,3-dioxygenase. Infect Immun 2004, 72, 2723–2730. [Google Scholar] [CrossRef]
- López, A.S.; Alegre, E.; Díaz, A.; et al. Bimodal effect of nitric oxide in the enzymatic activity of indoleamine 2,3-dioxygenase in human monocytic cells. Immunol Lett 2006, 106, 163–171. [Google Scholar] [CrossRef]
- Egilmez, N.K.; Harden, J.L.; Virtuoso, L.P.; et al. Nitric oxide short-circuits interleukin-12-mediated tumor regression. Cancer Immunol Immunother 2011, 60, 839–845. [Google Scholar] [CrossRef]
- Zhu, J.; Huang, X.; Yang, G. Autocrine IL6-mediated activation of the STAT3–DNMT axis silences the TNF-alpha–RIP1 necroptosis pathway to sustain survival and accumulation of myeloid-derived suppressor cells. Cancer Res 2020, 80, 3145–3156. [Google Scholar] [CrossRef]
- Matatall, K.A.; Shen, C.C.; Challen, G.A.; King, K.Y. Type II interferon promotes differentiation of myeloid-biased hematopoietic stem cells. Stem Cells 2014, 32, 3023–3030. [Google Scholar] [CrossRef]
- Patel, B.; Zhou, Y.; Babcock, R.L.; et al. STAT3 protects hematopoietic stem cells by preventing activation of a deleterious autocrine type-I interferon response. Leukemia 2024, 38, 1143–1155. [Google Scholar] [CrossRef]
- Mantel, C.; Messina-Graham, S.; Moh, A.; et al. Mouse hematopoietic cell-targeted STAT3 deletion: Stem/progenitor cell defects, mitochondrial dysfunction, ROS overproduction, and a rapid aging-like phenotype. Blood 2012, 120, 2589–2599. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, B.J.; Roberts, A.W.; Najdovska, M.; Grail, D.; Ernst, M. The threshold of gp130-dependent STAT3 signaling is critical for normal regulation of hematopoiesis. Blood 2005, 105, 3512–3520. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Iwama, A.; Tadokoro, Y.; et al. Selective activation of STAT5 unveils its role in stem cell self-renewal in normal and leukemic hematopoiesis. J Exp Med 2005, 202, 169–179. [Google Scholar] [CrossRef] [PubMed]
- Liao, P.; Wang, W.; Wang, W.; et al. CD8+ T cells and fatty acids orchestrate tumor ferroptosis and immunity via ACSL4. Cancer Cell 2022, 40, 365–378.e6. [Google Scholar] [CrossRef]
- Lin, Z.; Zou, S.; Wen, K. The crosstalk of CD8+ T cells and ferroptosis in cancer. Front Immunol 2023, 14, 1255443. [Google Scholar] [CrossRef]
- Flem-Karlsen, K.; Talty, R.; McGeary, M.K.; et al. Cytotoxic CD8+ T cells downregulate GPX4 to promote ferroptosis in melanoma that drives antitumor immunity. Cancer Res 2025. [Google Scholar] [CrossRef]
- Onodera, A.; González-Avalos, E.; Lio, C.J.; et al. Roles of TET and TDG in DNA demethylation in proliferating and non-proliferating immune cells. Genome Biol 2021, 22, 186. [Google Scholar] [CrossRef]
- Blander, J.M.; Medzhitov, R. Toll-dependent selection of microbial antigens for presentation by dendritic cells. Nature 2006, 440, 808–812. [Google Scholar] [CrossRef]
- Leonard, J.P.; Sherman, M.L.; Fisher, G.L.; et al. Effects of single-dose interleukin-12 exposure on interleukin-12-associated toxicity and interferon-gamma production. Blood 1997, 90, 2541–2548. [Google Scholar] [PubMed]
- Yoshida, Y.; Yoshimi, R.; Yoshii, H.; et al. The transcription factor IRF8 activates integrin-mediated TGF-beta signaling and promotes neuroinflammation. Immunity 2014, 40, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Khan, O.; Giles, J.R.; McDonald, S.; et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 2019, 571, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Tucker, C.G.; Mitchell, J.S.; Martinov, T.; et al. Adoptive T cell therapy with IL-12-preconditioned low-avidity T cells prevents exhaustion and results in enhanced T cell activation, enhanced tumor clearance, and decreased risk for autoimmunity. J Immunol 2020, 205, 1449–1460. [Google Scholar] [CrossRef]
- Grassmann, S.; Santosa, E.K.; Kim, H.; et al. Stepwise epigenetic signal integration drives adaptive programming of cytotoxic lymphocytes. Immunity 2026. [Google Scholar] [CrossRef]
- Hajishengallis, G.; Chavakis, T. Inflammageing and clonal haematopoiesis interplay and their impact on human disease. Nat Rev Mol Cell Biol 2026. [Google Scholar] [CrossRef]
- Mohammadnia, N.; Xue, L.; Vestjens, L.T.W.; et al. Colchicine and longitudinal dynamics of clonal hematopoiesis: An exploratory substudy of the LoDoCo2 trial. J Am Coll Cardiol (2025) 86, 1983–1996. [CrossRef]
- Wang, D.; Kaniowski, D.; Jacek, K.; et al. Bi-functional CpG-STAT3 decoy oligonucleotide triggers multilineage differentiation of acute myeloid leukemia in mice. Mol Ther Nucleic Acids 2024, 35, 102268. [Google Scholar] [CrossRef]
- Soyano, A.; Lee, M.C.; Han, H.S.; et al. Intratumoral dendritic cell immunotherapy controls dissemination of metastasis-initiating cancer cells, even in patients with metastatic breast cancer. J Immunother Cancer 2026, 14, e013590. [Google Scholar] [CrossRef]
- Daman, A.W.; Antonelli, A.C.; Redelman-Sidi, G.; et al. Microbial cancer immunotherapy reprograms hematopoiesis to enhance myeloid-driven anti-tumor immunity. Cancer Cell 2025. [Google Scholar] [CrossRef]
- Grajales-Reyes, G.E.; Iwata, A.; Albring, J.; et al. Batf3 maintains autoactivation of Irf8 for commitment of a CD8α+ conventional DC clonogenic progenitor. Nat Immunol 2015, 16, 708–717. [Google Scholar] [CrossRef]
- Matteini, F.; Thambyrajah, R.; Montserrat-Vazquez, S.; et al. A Notch trans-activation to cis-inhibition switch underlies hematopoietic stem cell aging. Blood 2025. [Google Scholar] [CrossRef]
- You, Z.; Chi, Y.; Wang, L.; et al. Mitochondrial fitness controls cDC1-mediated antitumor immunity. Science 2026, 392, eadv6582. [Google Scholar] [CrossRef]
- Molina, I.S.; Haldar, M. Mitochondria power immunity against cancer. Science 2026, 392, 24–25. [Google Scholar] [CrossRef] [PubMed]
- Du, X.; Wen, J.; Wang, Y.; et al. Hippo/Mst signalling couples metabolic state and immune function of CD8-alpha-positive dendritic cells. Nature 2018, 558, 141–145. [Google Scholar] [CrossRef]
- Krawczyk, C.M.; Holowka, T.; Sun, J.; et al. Toll-like receptor-induced changes in glycolytic metabolism regulate dendritic cell activation. Blood 2010, 115, 4742–4749. [Google Scholar] [CrossRef] [PubMed]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Yang, H.; Xu, W.; et al. Inhibition of alpha-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev 2012, 26, 1326–1338. [Google Scholar] [CrossRef]
- Liu, P.S.; Wang, H.; Li, X.; et al. Alpha-ketoglutarate orchestrates macrophage activation through metabolic and epigenetic reprogramming. Nat Immunol 2017, 18, 985–994. [Google Scholar] [CrossRef]
- Zhou, B.; Magana, L.; Hong, Z.; et al. The angiocrine Rspondin3 instructs interstitial macrophage transition via metabolic-epigenetic reprogramming and resolves inflammatory injury. Nat Immunol 2020, 21, 1430–1443. [Google Scholar] [CrossRef]
- Nie, Y.; Erion, D.M.; Yuan, Z.; et al. STAT3 inhibition of gluconeogenesis is downregulated by SirT1. Nat Cell Biol 2009, 11, 492–500. [Google Scholar] [CrossRef] [PubMed]
- Limagne, E.; Thibaudin, M.; Euvrard, R.; et al. Sirtuin-1 activation controls tumor growth by impeding Th17 differentiation via STAT3 deacetylation. Cell Rep 2017, 19, 746–759. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.P.; Price, N.L.; Ling, A.J.Y.; et al. Declining NAD+ induces a pseudohypoxic state disrupting nuclear-mitochondrial communication during aging. Cell 2013, 155, 1624–1638. [Google Scholar] [CrossRef] [PubMed]
- Everts, B.; Amiel, E.; Huang, S.C.; et al. TLR-driven early glycolytic reprogramming via the kinases TBK1-IKK-epsilon supports the anabolic demands of dendritic cell activation. Nat Immunol 2014, 15, 323–332. [Google Scholar] [CrossRef]
- Ugele, I.; Cardenas-Conejo, Z.E.; Gessner, A.; et al. D-2-hydroxyglutarate and L-2-hydroxyglutarate inhibit IL-12 secretion by human monocyte-derived dendritic cells. Int J Mol Sci 2019, 20, 742. [Google Scholar] [CrossRef]
- Hammon, K.; Renner, K.; Kreutz, M.; et al. D-2-hydroxyglutarate supports a tolerogenic phenotype with lowered major histocompatibility class II expression in non-malignant dendritic cells and acute myeloid leukemia cells. Haematologica 2024, 109, 2500–2514. [Google Scholar] [CrossRef]
- Tian, Y.; Machado, H.B.; Bhatt, D.L.; et al. IFN-gamma regulates NAD+ metabolism to promote the respiratory burst in human monocytes. Blood Adv 2022, 6, 3821–3834. [Google Scholar] [CrossRef]
- Wölfle, S.J.; Strebovsky, J.; Bartz, H.; et al. PD-L1 expression on tolerogenic APCs is controlled by STAT-3. Eur J Immunol 2011, 41, 413–424. [Google Scholar] [CrossRef]
- Zerdes, I.; Wallerius, M.; Sifakis, E.G.; et al. STAT3 activity promotes programmed-death ligand 1 expression and suppresses immune responses in breast cancer. Cancers 2019, 11, E1479. [Google Scholar] [CrossRef]
- Bu, L.L.; Yu, G.T.; Wu, L.; et al. STAT3 induces immunosuppression by upregulating PD-1/PD-L1 in HNSCC. J Dent Res 2017, 96, 1027–1034. [Google Scholar] [CrossRef]
- Zhang, B.; Xu, A.; Wang, H.; et al. MPC-mediated lactate production drives histone lactylation in dendritic cells to affect tumor progression and immunotherapy. Cell Mol Life Sci 2025, 82, 371. [Google Scholar] [CrossRef] [PubMed]
- Sun, K.; Zhang, X.; Shi, J.; et al. Elevated protein lactylation promotes immunosuppressive microenvironment and therapeutic resistance in pancreatic ductal adenocarcinoma. J Clin Invest 2025, 135, e187024. [Google Scholar] [CrossRef] [PubMed]
- Doolittle, M.L.; et al. Aged murine bone marrow myeloid and mesenchymal cells develop unique senescence phenotypes. J Clin Invest 2026, 136, e195772. [Google Scholar] [CrossRef] [PubMed]
- Hofbauer, L.C.; Rauner, M. Deconstructing senescence phenotypes in cells of the bone and bone marrow. J Clin Invest 2026, 136, e204645. [Google Scholar] [CrossRef]
- Sladitschek-Martens, H.L.; et al. YAP/TAZ activity in stromal cells prevents ageing by controlling cGAS-STING. Nature 2022, 607, 790–798. [Google Scholar] [CrossRef]
- Ambrosi, T.H.; et al. Aged skeletal stem cells generate an inflammatory degenerative niche. Nature 2021, 597, 256–262. [Google Scholar] [CrossRef]
- Mitchell, C.A.; et al. Stromal niche inflammation mediated by IL-1 signaling is a targetable driver of hematopoietic aging. Cell Stem Cell (2023) 30, 820–837. [CrossRef]
- Molina, C.M.; Haldar, M. Mitochondrial metabolism and signaling direct dendritic cell function in antitumor immunity. Science 2026, 387, eadv6582. [Google Scholar] [CrossRef]
- Kim, T.W.; et al. IL-6 selectively suppresses cDC1 specification via C/EBP-beta. J Exp Med 2023, 220, e20230235. [Google Scholar] [CrossRef]
- Luo, Y.; et al. Senolytic drugs dasatinib and quercetin promote skin papilloma progression by eliminating senescent cells and impairing immune surveillance. One Health Adv 2025, 3, 5. [Google Scholar] [CrossRef]
- Patel, S.B.; et al. STAT3 protects hematopoietic stem cells by preventing activation of a deleterious autocrine type-I interferon response. Leukemia 2024, 38, 1707–1721. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, M.J.; et al. Transient hepatic reconstitution of trophic factors enhances aged immunity. Nature 2025, 639, 756–764. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.B.; et al. Depleting myeloid-biased haematopoietic stem cells rejuvenates aged immunity. Nature 2024, 628, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Gadecka, A.; et al. The senolytic cocktail, dasatinib and quercetin, impacts the chromatin structure of both young and senescent vascular smooth muscle cells. GeroScience 2025, 47, 1299–1315. [Google Scholar] [CrossRef]
- Fang, C.; Qiao, Y.; Mun, S.H.; et al. Cutting edge: EZH2 promotes osteoclastogenesis by epigenetic silencing of the negative regulator IRF8. J Immunol 2016, 196, 4452–4456. [Google Scholar] [CrossRef]
- Pan, Y.M.; Wang, C.G.; Zhu, M.; et al. STAT3 signaling drives EZH2 transcriptional activation and mediates poor prognosis in gastric cancer. Mol Cancer 2016, 15, 79. [Google Scholar] [CrossRef]
- Kim, D.; Park, G.; Huuhtanen, J.; et al. STAT3 activation in large granular lymphocyte leukemia is associated with cytokine signaling and DNA hypermethylation. Leukemia 2021, 35, 3430–3443. [Google Scholar] [CrossRef]
- Dasgupta, M.; Dermawan, J.K.; Willard, B.; Stark, G.R. STAT3-driven transcription depends upon the dimethylation of K49 by EZH2. Proc Natl Acad Sci USA 2015, 112, 3985–3990. [Google Scholar] [CrossRef]
- Kim, E.; Kim, M.; Woo, D.H.; et al. Phosphorylation of EZH2 activates STAT3 signaling via STAT3 methylation and promotes tumorigenicity of glioblastoma stem-like cells. Cancer Cell 2013, 23, 839–852. [Google Scholar] [CrossRef]
- Beerman, I.; Bock, C.; Garrison, B.S.; et al. Proliferation-dependent alterations of the DNA methylation landscape underlie hematopoietic stem cell aging. Cell Stem Cell 2013, 12, 413–425. [Google Scholar] [CrossRef]
- Wen, H.; Dou, Y.; Hogaboam, C.M.; Kunkel, S.L. Epigenetic regulation of dendritic cell-derived interleukin-12 facilitates immunosuppression after a severe innate immune response. Blood 2008, 111, 1797–1804. [Google Scholar] [CrossRef] [PubMed]
- DiNardo, A.R.; Netea, M.G.; Musher, D.M. Postinfectious epigenetic immune modifications—a double-edged sword. N Engl J Med 2021, 384, 261–270. [Google Scholar] [CrossRef] [PubMed]
- Zhan, Y.; Zhang, Y.; Zhang, S.; et al. Differential requirement for the polycomb repressor complex 2 in dendritic cell and tissue-resident myeloid cell homeostasis. Sci Immunol 2021, 6, eabf7268. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Wang, Z.; Zhou, J.; et al. EZH2 inhibitor GSK126 suppresses antitumor immunity by driving production of myeloid-derived suppressor cells. Cancer Res (2019) 79, 2009–2020. [CrossRef]
- Qiang, N.; Ao, J.; Nakamura, M.; et al. Alteration of the tumor microenvironment by pharmacological inhibition of EZH2 in hepatocellular carcinoma. Int Immunopharmacol 2023, 118, 110068. [Google Scholar] [CrossRef]
- Chibaya, L.; Murphy, K.C.; DeMarco, K.D.; et al. EZH2 inhibition remodels the inflammatory senescence-associated secretory phenotype to potentiate pancreatic cancer immune surveillance. Nat Cancer 2023, 4, 872–892. [Google Scholar] [CrossRef]
- Sun, D.; Luo, M.; Jeong, M.; et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 2014, 14, 673–688. [Google Scholar] [CrossRef]
- Kuribayashi, W.; Tanaka-Yano, M.; Park, B.; et al. Age-like methylation changes of HSCs in GADD45B knockout mice define methylation sites associated with loss of function. Aging Cell 2026, 25, e70453. [Google Scholar] [CrossRef]
- Adelman, E.R.; Huang, H.T.; Roisman, A.; et al. Aging human hematopoietic stem cells manifest profound epigenetic reprogramming of enhancers that may predispose to leukemia. Cancer Discov 2019, 9, 1080–1101. [Google Scholar] [CrossRef]
- Flohr Svendsen, A.; Yang, D.; Kim, K.; et al. A comprehensive transcriptome signature of murine hematopoietic stem cell aging. Blood 2021, 138, 439–451. [Google Scholar] [CrossRef]
- Williams, O.; Hu, L.; Huang, W.; et al. Nore1 inhibits age-associated myeloid lineage skewing and clonal hematopoiesis but facilitates termination of emergency (stress) granulopoiesis. J Biol Chem (2023) 299, 104867. [CrossRef] [PubMed]
- De Santa, F.; Totaro, M.G.; Prosperini, E.; et al. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell 2007, 130, 1083–1094. [Google Scholar] [CrossRef] [PubMed]
- Yıldırım-Buharalıoğlu, G.; Bond, M.; Sala-Newby, G.B.; et al. Regulation of epigenetic modifiers, including KDM6B, by interferon-gamma and interleukin-4 in human macrophages. Front Immunol 2017, 8, 92. [Google Scholar] [CrossRef] [PubMed]
- Mikulski, P.; Tehrani, S.S.H.; Kogan, A.; et al. Heritable maintenance of chromatin modifications confers transcriptional memory of interferon-gamma signaling. Nat Struct Mol Biol 2025. [Google Scholar] [CrossRef]
- Ovadya, Y.; Landsberger, T.; Leins, H.; et al. Impaired immune surveillance accelerates accumulation of senescent cells and aging. Nat Commun 2018, 9, 5435. [Google Scholar] [CrossRef]
- Antonangeli, F.; Zingoni, A.; Soriani, A.; Santoni, A. Senescent cells: Living or dying is a matter of NK cells. J Leukoc Biol 2019, 105, 1275–1283. [Google Scholar] [CrossRef]
- Waight, J.D.; Netherby, C.; Hensen, M.L.; et al. Myeloid-derived suppressor cell development is regulated by a STAT/IRF-8 axis. J Clin Invest 2013, 123, 4464–4478. [Google Scholar] [CrossRef]
- Harroch, S.; Gothelf, Y.; Watanabe, N.; Revel, M.; Chebath, J. Interleukin-6 activates and regulates transcription factors of the interferon regulatory factor family in M1 cells. J Biol Chem 1993, 268, 9092–9097. [Google Scholar] [CrossRef]
- Orabona, C.; Puccetti, P.; Vacca, C.; et al. Toward the identification of a tolerogenic signature in IDO-competent dendritic cells. Blood (2006) 107, 2846–2854. [CrossRef]
- Gargaro, M.; Scalisi, G.; Manni, G.; et al. Indoleamine 2,3-dioxygenase 1 activation in mature cDC1 promotes tolerogenic education of inflammatory cDC2 via metabolic communication. Immunity 2022, 55, 1032–1050.e14. [Google Scholar] [CrossRef]
- Ye, Z.H.; Jiang, X.M.; Huang, M.Y.; et al. Regulation of CD47 expression by interferon-gamma in cancer cells. Transl Oncol 2021, 14, 101162. [Google Scholar] [CrossRef]
- Hendriks, M.A.J.M.; Britsch, I.; Ke, X.; et al. Cancer cells under immune attack acquire CD47-mediated adaptive immune resistance independent of the myeloid CD47-SIRPα axis. Oncoimmunology 2021, 10, 2005344. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.M.; Pu, Y.; Han, D.; et al. Dendritic cells but not macrophages sense tumor mitochondrial DNA for cross-priming through signal regulatory protein α signaling. Immunity 2017, 47, 363–373.e5. [Google Scholar] [CrossRef] [PubMed]
- Santarelli, R.; Gonnella, R.; Di Giovenale, G.; et al. STAT3 activation by KSHV correlates with IL-10, IL-6 and IL-23 release and an autophagic block in dendritic cells. Sci Rep 2014, 4, 4241. [Google Scholar] [CrossRef] [PubMed]
- Masztalerz, A.; Van Rooijen, N.; Den Otter, W.; Everse, L.A. Mechanisms of macrophage cytotoxicity in IL-2 and IL-12 mediated tumour regression. Cancer Immunol Immunother 2003, 52, 235–242. [Google Scholar] [CrossRef]
- Yokogami, K.; Wakisaka, S.; Avruch, J.; Reeves, S.A. Serine phosphorylation and maximal activation of STAT3 during CNTF signaling is mediated by the rapamycin target mTOR. Curr Biol 2000, 10, 47–50. [Google Scholar] [CrossRef]
- Pinno, J.; Bongartz, H.; Klepsch, O.; et al. Interleukin-6 influences stress-signalling by reducing the expression of the mTOR-inhibitor REDD1 in a STAT3-dependent manner. Cell Signal 2016, 28, 907–916. [Google Scholar] [CrossRef]
- Weichhart, T.; Costantino, G.; Poglitsch, M.; et al. The TSC-mTOR signaling pathway regulates the innate inflammatory response. Immunity 2008, 29, 565–577. [Google Scholar] [CrossRef]
- Braun, M.; Ress, M.L.; Yoo, Y.E.; et al. IL12-mediated sensitizing of T-cell receptor-dependent and -independent tumor cell killing. Oncoimmunology 2016, 5, e1188245. [Google Scholar] [CrossRef]
- Derré, L.; Corvaisier, M.; Pandolfino, M.C.; et al. Expression of CD94/NKG2-A on human T lymphocytes is induced by IL-12: Implications for adoptive immunotherapy. J Immunol 2002, 168, 4864–4870. [Google Scholar] [CrossRef]
- Vom Berg, J.; Prokop, S.; Miller, K.R.; et al. Inhibition of IL-12/IL-23 signaling reduces Alzheimer's disease-like pathology and cognitive decline. Nat Med 2012, 18, 1812–1819. [Google Scholar] [CrossRef]
- Schneeberger, S.; Kim, S.J.; Geesdorf, M.N.; et al. Interleukin-12 signaling drives Alzheimer's disease pathology through disrupting neuronal and oligodendrocyte homeostasis. Nat Aging 2025, 5, 622–641. [Google Scholar] [CrossRef]
- Li, H.; Bullock, K.; Gurber, C.; et al. Metabolomic adaptations and correlates of survival to immune checkpoint blockade. Nat Commun 2019, 10, 4346. [Google Scholar] [CrossRef]
- Zhang, C.; Chien, C.; Jurgaitytė, E.; et al. Intratumoral Treg cell ablation elicits NK cell-mediated control of CD8 T cell-resistant tumors. Sci Immunol 2026. [Google Scholar] [CrossRef]
- Tsui, K.H.; Cheng, S.H.; Wang, B.; et al. Mitochondrial quality control as a central pharmacological target in aging. Pharmacol Res 2026, 227, 108188. [Google Scholar] [CrossRef]
- Wang, W.; Hu, Y.; Yang, C.; et al. Decreased NAD activates STAT3 and integrin pathways to drive epithelial-mesenchymal transition. Mol Cell Proteomics 2018, 17, 2005–2017. [Google Scholar] [CrossRef]
- Minhas, P.S.; Liu, L.; Moon, P.K.; et al. Macrophage de novo NAD synthesis specifies immune function in aging and inflammation. Nat Immunol 2019, 20, 50–63. [Google Scholar] [CrossRef] [PubMed]
- O'Hagan, H.M.; Mohammad, H.P.; Baylin, S.B. Double strand breaks can initiate gene silencing and SIRT1-dependent onset of DNA methylation in an exogenous promoter CpG island. PLoS Genet 2008, 4, e1000155. [Google Scholar] [CrossRef] [PubMed]
- Pruitt, K.; Zinn, R.L.; Ohm, J.E.; et al. Inhibition of SIRT1 reactivates silenced cancer genes without loss of promoter DNA hypermethylation. PLoS Genet 2006, 2, e40. [Google Scholar] [CrossRef] [PubMed]
- Petrocelli, J.J.; McKenzie, A.I.; de Hart, N.M.M.P.; et al. Disuse-induced muscle fibrosis, cellular senescence, and senescence-associated secretory phenotype in older adults are alleviated during re-ambulation with metformin pre-treatment. Aging Cell 2023, 22, e13936. [Google Scholar] [CrossRef]
- Vasamsetti, S.B.; Karnewar, S.; Kanugula, A.K.; et al. Metformin inhibits monocyte-to-macrophage differentiation via AMPK-mediated inhibition of STAT3 activation. Diabetes 2015, 64, 2028–2041. [Google Scholar] [CrossRef]
- Khan, J.; Pernicova, I.; Nisar, K.; Korbonits, M. Mechanisms of ageing: Growth hormone, dietary restriction, and metformin. Lancet Diabetes Endocrinol 2023, 11, 261–281. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yu, P.; Xu, Y.; et al. Metformin induces tolerogenicity of dendritic cells by promoting metabolic reprogramming. Cell Mol Life Sci 2023, 80, 283. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.J.; Zhu, B.T. Inhibition of DNA methylation by caffeic acid and chlorogenic acid, two common catechol-containing coffee polyphenols. Carcinogenesis 2006, 27, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Legaz, A.; Moguilner, S.; Barttfeld, P.; et al. The exposome of brain aging across 34 countries. Nat Med 2026. [Google Scholar] [CrossRef]
- Tang, Q.; Fang, J.; Lai, W.; et al. Hippo pathway monomerizes STAT3 to regulate prostate cancer growth. Cancer Sci 2022, 113, 2753–2762. [Google Scholar] [CrossRef]
- Hopwood, T.W.; Hall, S.; Begley, N.; et al. The circadian regulator BMAL1 programmes responses to parasitic worm infection via a dendritic cell clock. Sci Rep 2018, 8, 3782. [Google Scholar] [CrossRef]
- Cervantes-Silva, M.P.; Carroll, R.G.; Wilk, M.M.; et al. The circadian clock influences T cell responses to vaccination by regulating dendritic cell antigen processing. Nat Commun 2022, 13, 7217. [Google Scholar] [CrossRef]
- Manicassamy, S.; Reizis, B.; Ravindran, R.; et al. Activation of beta-catenin in dendritic cells regulates immunity versus tolerance in the intestine. Science 2010, 329, 849–853. [Google Scholar] [CrossRef]
- Sheen, J.H.; Strainic, M.G.; Liu, J.; et al. TLR-induced murine dendritic cell (DC) activation requires DC-intrinsic complement. J Immunol 2017, 199, 278–291. [Google Scholar] [CrossRef]
- Gargaro, M.; Vacca, C.; Massari, S.; et al. Engagement of nuclear coactivator 7 by 3-hydroxyanthranilic acid enhances activation of aryl hydrocarbon receptor in immunoregulatory dendritic cells. Front Immunol 2019, 10, 1973. [Google Scholar] [CrossRef]
- Zhao, Y.; Shen, X.; Na, N.; et al. mTOR masters monocyte development in bone marrow by decreasing the inhibition of STAT5 on IRF8. Blood 2018, 131, 1587–1599. [Google Scholar] [CrossRef] [PubMed]
- Wiernicki, B.; Maschalidi, S.; Pinney, J.; et al. Cancer cells dying from ferroptosis impede dendritic cell-mediated anti-tumor immunity. Nat Commun 2022, 13, 3676. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Bai, L.; Qu, C.; et al. PPARG-mediated ferroptosis in dendritic cells limits antitumor immunity. Biochem Biophys Res Commun 2021, 576, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Schaupp, L.; Muth, S.; Rogell, L.; et al. Microbiota-induced type I interferons instruct a poised basal state of dendritic cells. Cell 2020, 181, 1080–1096. [Google Scholar] [CrossRef]
- Ocaña-Morgner, C. Neutral sphingomyelinase and ceramide in dendritic cell activation and IL-12 production during immune responses. J Immunol 2007, 178, 6741–6751. [Google Scholar]
- Novitskiy, S.V.; Ryzhov, S.; Zaynagetdinov, R.; et al. Adenosine receptors in regulation of dendritic cell differentiation and function. Blood 2008, 112, 1822–1831. [Google Scholar] [CrossRef]
- Kalinski, P.; Vieira, P.L.; Schuitemaker, J.H.; de Jong, E.C.; Kapsenberg, M.L. Prostaglandin E(2) is a selective inducer of interleukin-12 p40 (IL-12p40) production and an inhibitor of bioactive IL-12p70 heterodimer. Blood 2001, 97, 3466–3469. [Google Scholar] [CrossRef]
- Kalinski, P.; Schuitemaker, J.H.; Hilkens, C.M.; Kapsenberg, M.L. Prostaglandin E2 induces the final maturation of IL-12-deficient CD1a+CD83+ dendritic cells: the levels of IL-12 are determined during the final dendritic cell maturation and are resistant to further modulation. J Immunol 1998, 161, 2804–2809. [Google Scholar] [CrossRef]
- Khayrullina, T.; Yen, J.H.; Jing, H.; Ganea, D. In vitro differentiation of dendritic cells in the presence of prostaglandin E2 alters the IL-12/IL-23 balance and promotes differentiation of Th17 cells. J Immunol 2008, 181, 721–735. [Google Scholar] [CrossRef]
- Van Elssen, C.H.; Vanderlocht, J.; Oth, T.; et al. Inflammation-restraining effects of prostaglandin E2 on natural killer-dendritic cell (NK-DC) interaction are imprinted during DC maturation. Blood 2011, 118, 2473–2482. [Google Scholar] [CrossRef]
- Bayerl, F.; Meiser, P.; Donakonda, S.; et al. Tumor-derived prostaglandin E2 programs cDC1 dysfunction to impair intratumoral orchestration of anti-cancer T cell responses. Immunity 2023, 56, 1341–1358.e11. [Google Scholar] [CrossRef] [PubMed]
- Anguille, S.; Smits, E.L.; Lion, E.; van Tendeloo, V.F.; Berneman, Z.N. Clinical use of dendritic cells for cancer therapy. Lancet Oncol 2014, 15, e257–e267. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Jia, X.; Liu, M.; et al. Impact of NSAID type, initiation timing, duration and dose on clinical outcomes of immunotherapy in NSCLC: A multicenter two-cohort study. J Immunother Cancer 2026, 14, e014360. [Google Scholar] [CrossRef] [PubMed]
- Punyawatthananukool, S.; Matsuura, R.; Wongchang, T.; et al. Prostaglandin E2-Ep2/Ep4 signaling induces immunosuppression in human cancer by impairing bioenergetics and ribosome biogenesis in immune cells. Nat Commun 2024, 15, 9464. [Google Scholar] [CrossRef]
- Jang, C.; Chen, L.; Rabinowitz, J.D. Metabolomics and isotope tracing. Cell 2018, 173, 822–837. [Google Scholar] [CrossRef]
- Kaya-Okur, H.S.; Wu, S.J.; Codomo, C.A.; et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun 2019, 10, 1930. [Google Scholar] [CrossRef]
- Fahy, G.M.; Brooke, R.T.; Watson, J.P.; et al. Reversal of epigenetic aging and immunosenescent trends in humans. Aging Cell 2019, 18, e13028. [Google Scholar] [CrossRef]
- Li, B.; Peng, X.; Li, H.; et al. Single-cell immune aging clocks reveal inter-individual heterogeneity during infection and vaccination. Nat Aging 2025, 5, 635–651. [Google Scholar] [CrossRef]
- Kantoff, P.W.; Higano, C.S.; Shore, N.D.; et al. Sipuleucel-T immunotherapy for castration-resistant prostate cancer. N Engl J Med 2010, 363, 411–422. [Google Scholar] [CrossRef]
- Yamamoto, K.; Shibata, F.; Miyasaka, N.; Miura, O. The human perforin gene is a direct target of STAT4 activated by IL-12 in NK cells. Biochem Biophys Res Commun 2002, 297, 1245–1252. [Google Scholar] [CrossRef]
- Hazeldine, J.; Hampson, P.; Lord, J.M. Reduced release and binding of perforin at the immunological synapse underlies the age-related decline in natural killer cell cytotoxicity. Aging Cell 2012, 11, 751–759. [Google Scholar] [CrossRef]
- Koroma, A.K.; Singh, K.; Wang, Y.; et al. Overactivation of Cdc42 GTPase impairs the cytotoxic function of NK cells from old individuals towards senescent fibroblasts. Aging Cell 2026, 25, e70398. [Google Scholar] [CrossRef] [PubMed]
- Sagiv, A.; Biran, A.; Yon, M.; et al. Granule exocytosis mediates immune surveillance of senescent cells. Oncogene 2013, 32, 1971–1977. [Google Scholar] [CrossRef] [PubMed]
- Shehata, H.M.; Hoebe, K.; Chougnet, C.A. The aged nonhematopoietic environment impairs natural killer cell maturation and function. Aging Cell 2015, 14, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Chiu, B.C.; Martin, B.E.; Stolberg, V.R.; Chensue, S.W. The host environment is responsible for aging-related functional NK cell deficiency. J Immunol 2013, 191, 4688–4698. [Google Scholar] [CrossRef]
- Bonnema, J.D.; Rivlin, K.A.; Ting, A.T.; et al. Cytokine-enhanced NK cell-mediated cytotoxicity: Positive modulatory effects of IL-2 and IL-12 on stimulus-dependent granule exocytosis. J Immunol 1994, 152, 2098–2104. [Google Scholar] [CrossRef]
- Nogusa, S.; Murasko, D.M.; Gardner, E.M. Differential effects of stimulatory factors on natural killer cell activities of young and aged mice. J Gerontol A Biol Sci Med Sci 2012, 67, 947–954. [Google Scholar] [CrossRef]
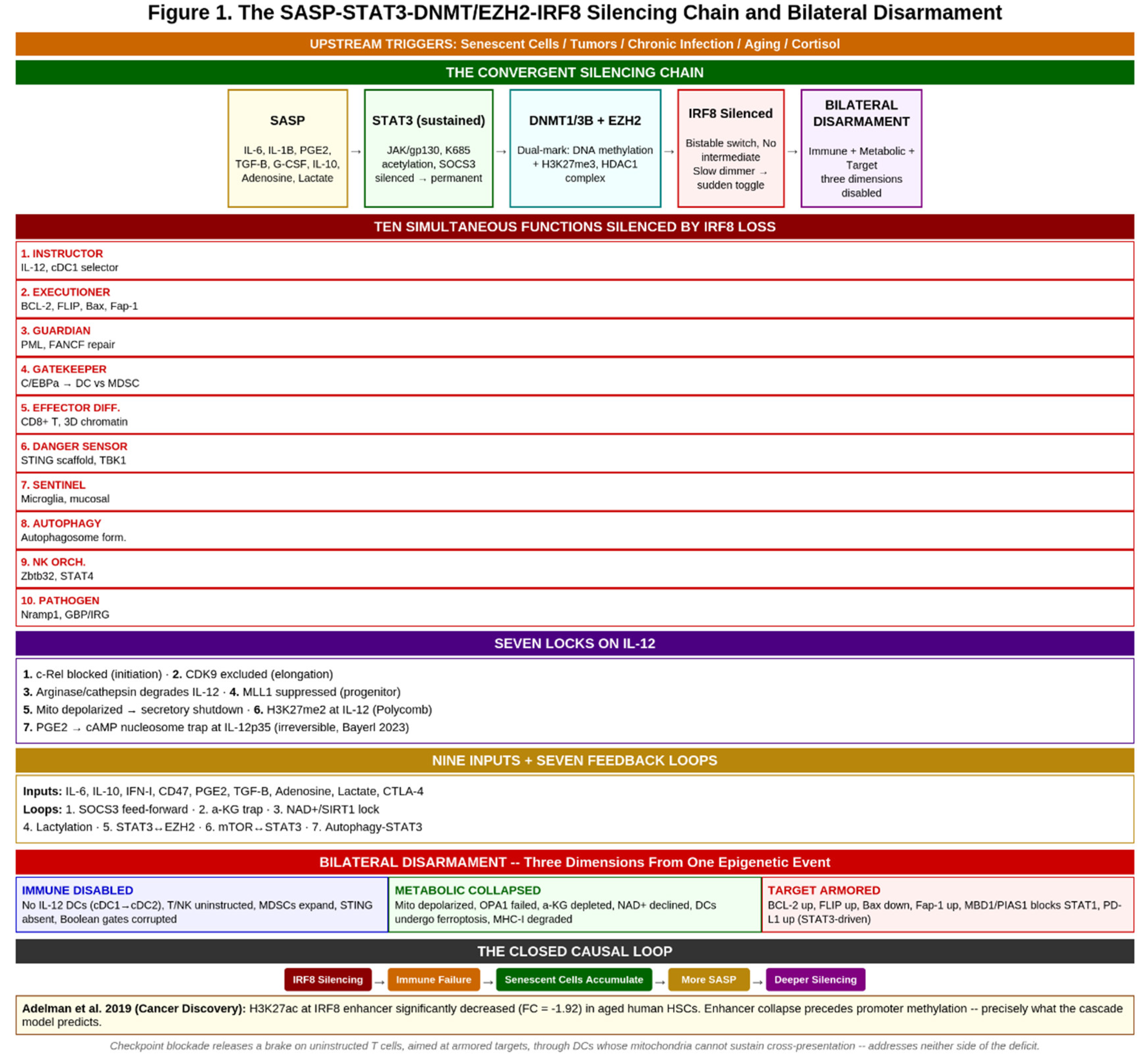

| Prediction | Biomarker | Assay | Expected (α-DC1) | Comparator (CPI) | Falsification |
|---|---|---|---|---|---|
| SASP–STAT3–IRF8 chain | IRF8 promoter CpG methylation (CD14+ monocytes) | Bisulfite-seq / ATAC-seq | Declining with each cycle | Not measured | Methylation unchanged after Phase 4 |
| IL-12 restoration | Serum IL-12p70 | ELISA | Pulsatile spikes (hours post-injection) | Absent | No detectable IL-12p70 |
| IDO suppression (bimodal NO) | Serum K/T ratio | LC-MS/MS | Declining from ~60–80 toward ~30–35 μmol/mmol | 37% increase at wk 4 (Li 2019) | K/T rises or unchanged |
| iNOS threshold crossed | Serum NOx (nitrite/nitrate) | Griess assay | Rising above ~60 μM | Not measured | NOx unchanged from baseline |
| Inverse NO–IDO correlation | NOx↑ with K/T↓ (paired draws) | Paired Griess + LC-MS/MS | Inverse correlation | Never documented | No correlation |
| STING competence | IRF8 methylation as companion Dx | Bisulfite-seq | Predicts STING agonist response | Not available | IRF8-silenced respond equally |
| Senescent cell clearance | p16/p21; SASP (IL-6, GDF-15) | IHC / Luminex | Declining (Phase 3–4) | Not measured | SASP unchanged at 6 mo |
| STAT3 hub activation | pSTAT3 (Y705) in CD14+ monocytes | Phospho-flow cytometry | Declining toward baseline | Elevated; predicts CPI failure | pSTAT3 unchanged |
| Loop 1 engagement | SOCS3 promoter methylation (CD14+) | Bisulfite-seq | Declining (brake restored) | 60% in AML vs 0% remission | SOCS3 methylation persists |
| Three failure modes | IRF8 meth + K/T + IL-12p70 combination | Composite panel | Distinguishes epigenetic / metabolic / adaptive | Not measured as combination | Combination not predictive |
| Mitochondrial restoration | Circulating cell-free mtDNA | qPCR / ddPCR | Declining (Phase 4) | Not measured | ccf-mtDNA unchanged |
| Metabolic fitness | PBMC mitochondrial respiration | Seahorse XF / Oroboros | OXPHOS capacity increasing | Not measured | Respiration unchanged |
| Loop 2 metabolic trap | α-KG/succinate ratio (CD14+ monocytes or plasma) | LC-MS/MS or colorimetric kit | Ratio rising toward 1.5–2.0 (Phase 4) | Not measured | Ratio unchanged or declining |
| Loop 3 acetylation lock | Whole blood NAD+ | NADMED enzymatic cycling (CE-marked) or LC-MS/MS | Rising from ~20 toward ~33 μmol/L (Phase 4) | Not measured | NAD+ unchanged from baseline |
| Dual-mark reversal | H3K27me3 at IRF8 locus (CD14+ monocytes) | CUT&Tag-qPCR or CUT&RUN | H3K27me3 declining at IRF8 enhancer (Phase 2–4) | Not measured | H3K27me3 persists at IRF8 |
| Systemic epigenetic reversal | Epigenetic age (DunedinPACE / GrimAge2) | CLIA-certified methylation array | Rate of aging decelerating; biological age decreasing | Not measured in immunotherapy | DunedinPACE unchanged or accelerating |
| Safety: autoimmune surveillance | Anti-dsDNA Ab; ANA titer | ELISA / IIF (HEp-2) | Stable or declining (anergy restored) | ANA rises in 15–25% of CPI patients | New anti-dsDNA or ANA ≥1:160 post-vaccination |
| Adaptive resistance | CD47 MFI on CTCs or sSIRPα | Flow cytometry (B6H12) / ELISA | Rising CD47 signals need for anti-CD47 combination | Not monitored | CD47 unchanged despite progression |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




