Submitted:
10 April 2026
Posted:
13 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Culture and Characterization
2.2. Evaluation of Cell Morphology with Epifluorescent Microspcope
2.3. Evaluation of Osteosarcoma Cell Lines Proliferation and Migration
2.3.1. MTS Assay
2.3.2. Cell Cycle Analysis
2.3.3. Colony Forming Unit (CFU) Assay
2.3.4. Wound Healing Assay (Scratch Test)
2.3.5. Transwell Migration Assay
2.4. Gene Expression Analysis
2.5. Protein Expression Analysis—Western Blot
2.6. Statistical Analysis
3. Results
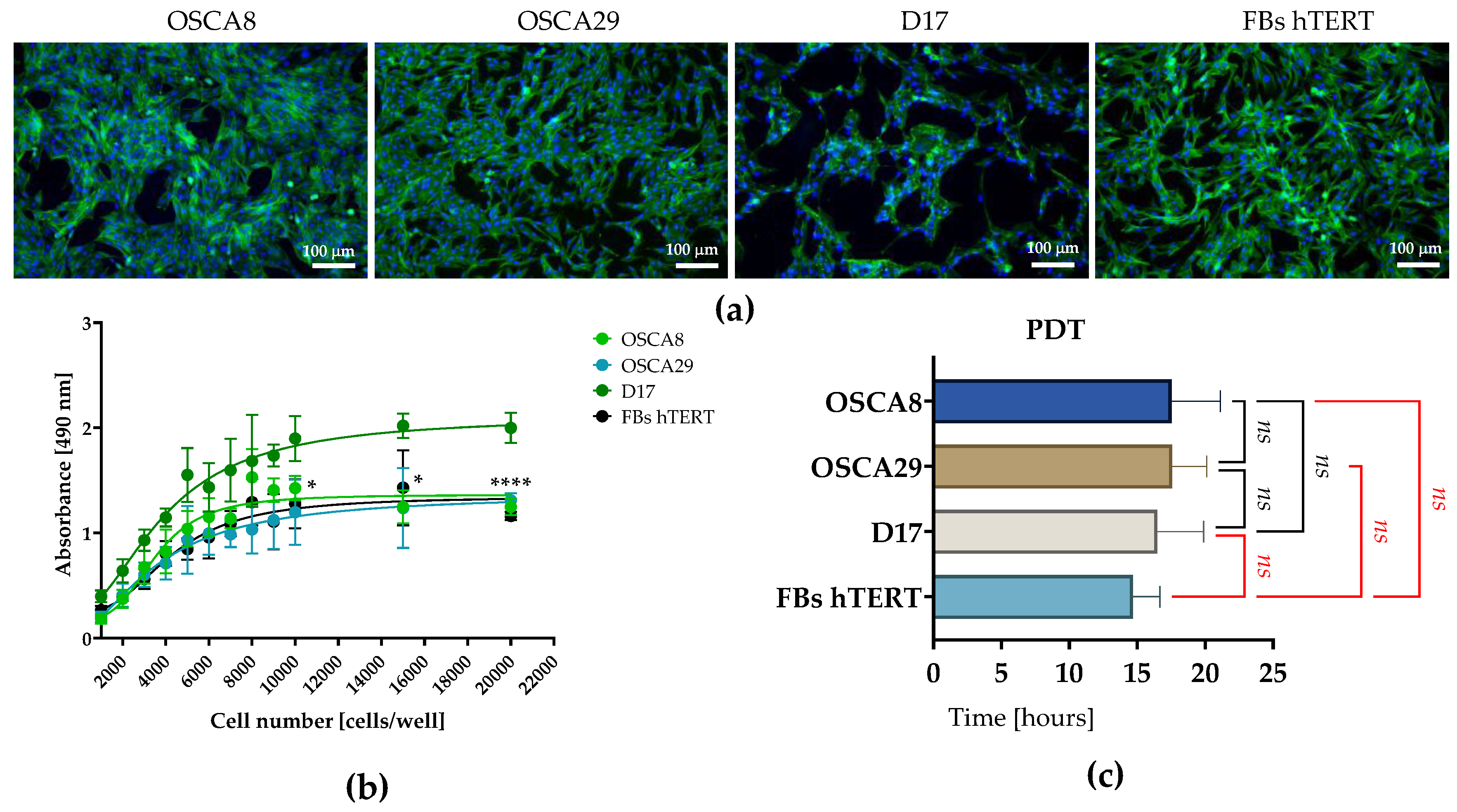
3.1. Primary Tumor- and Metastasis-Derived Canine Osteosarcoma Cell Lines Display Distinct Morphotypic Features In Vitro
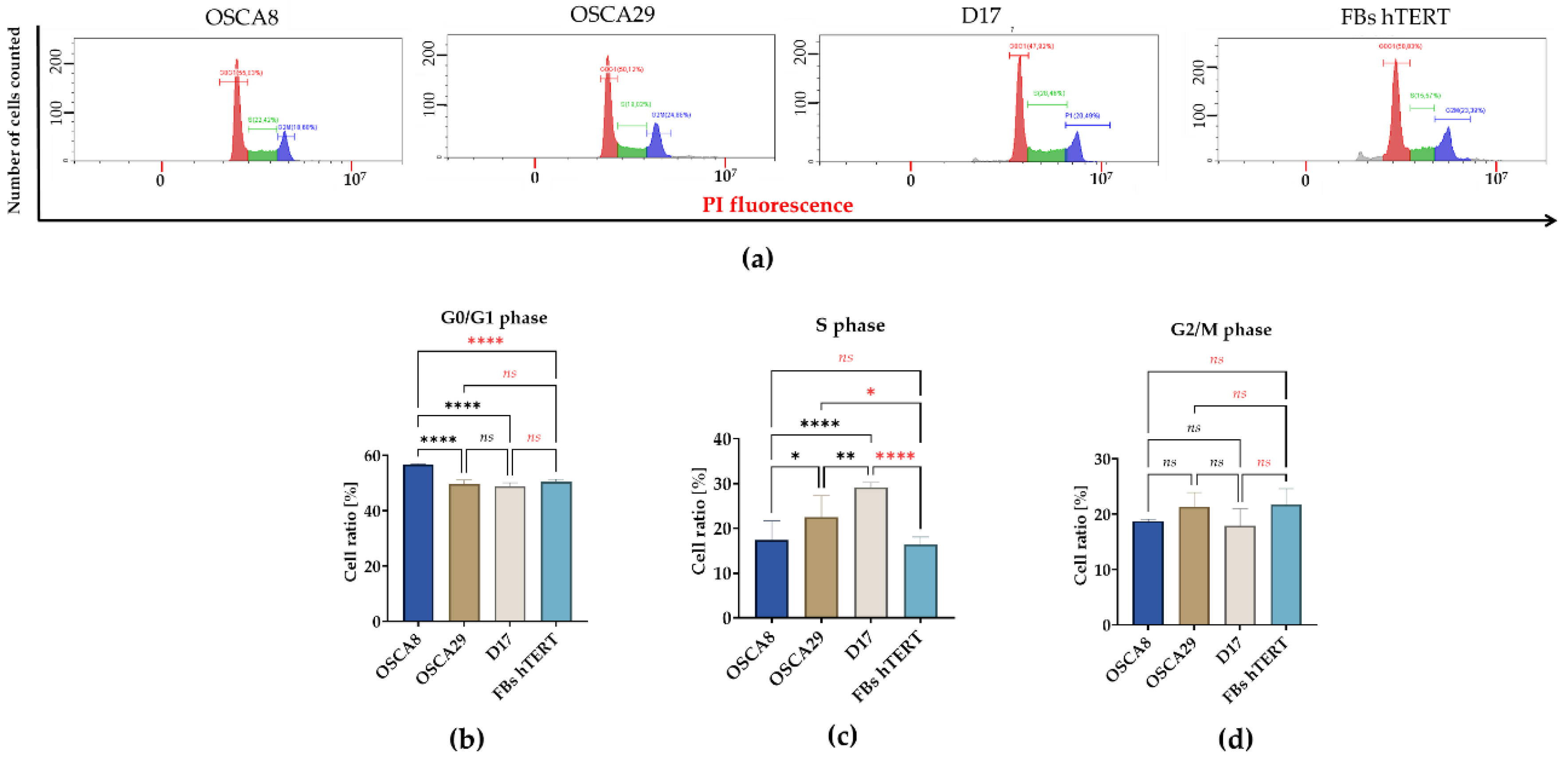
3.2. Canine Osteosarcoma Cell Lines Exhibit Distinct G0/G1 and S-Phase Distributions
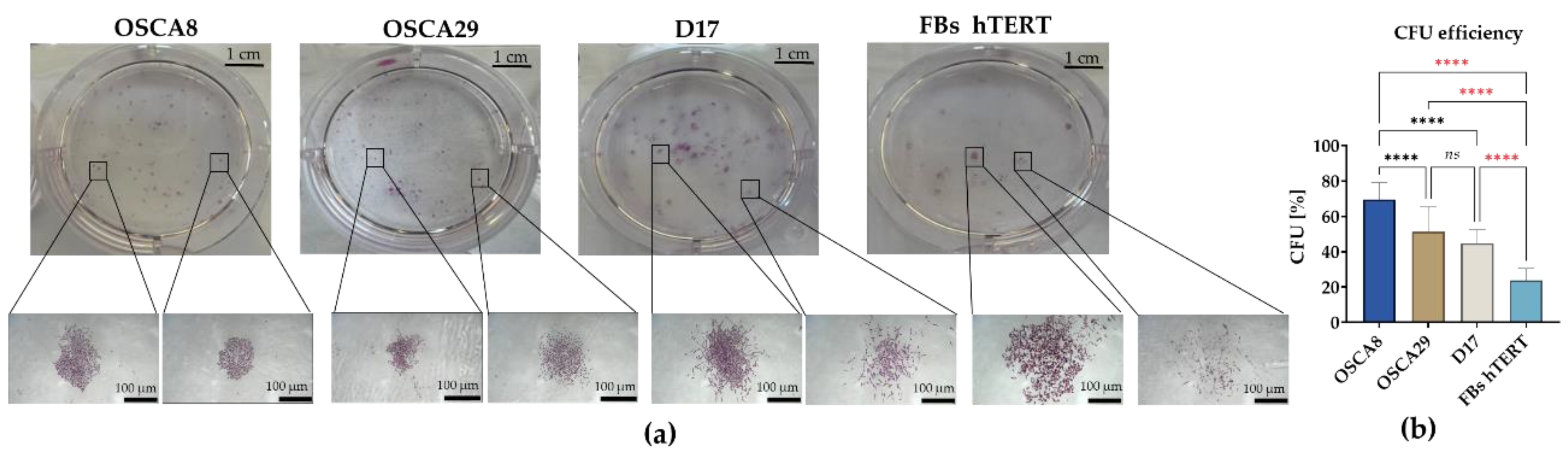
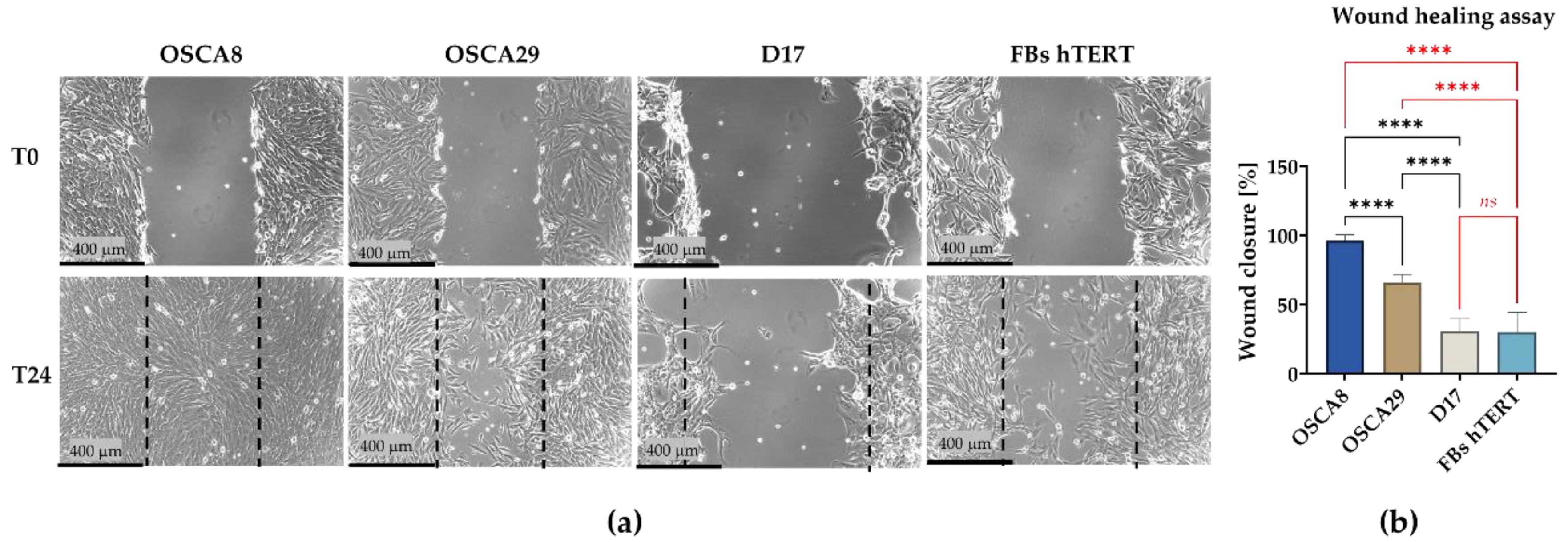
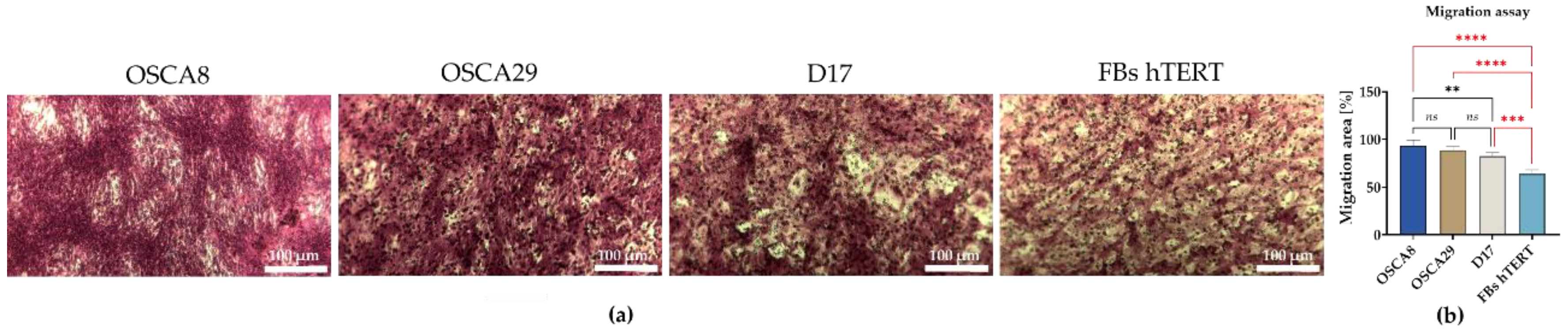
3.3. Canine Osteosarcoma Cell Lines Display Distinct Clonogenic and Migratory Phenotypes, with OSCA-8 Showing the Most Pronounced Aggressive Profile In Vitro
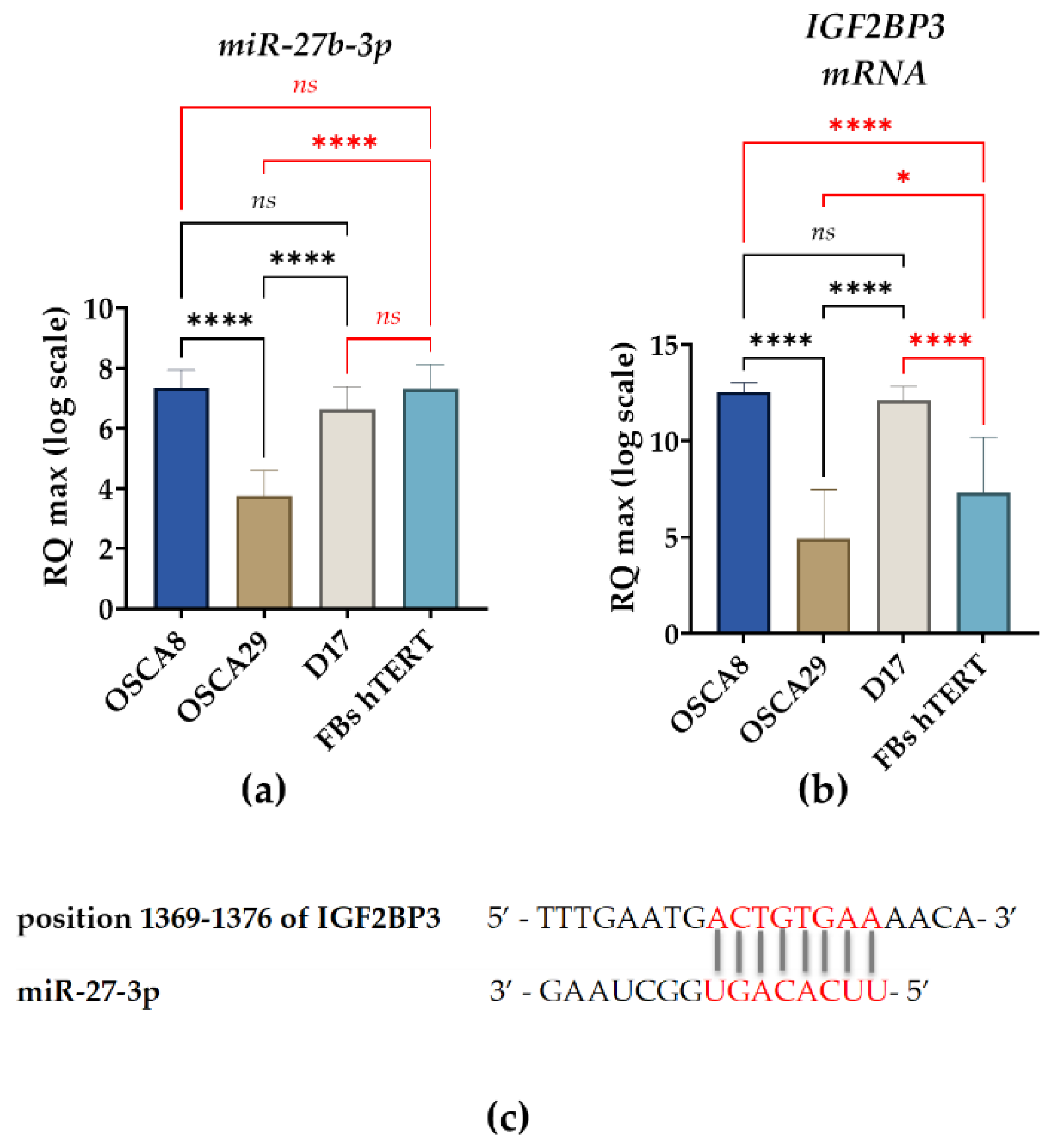
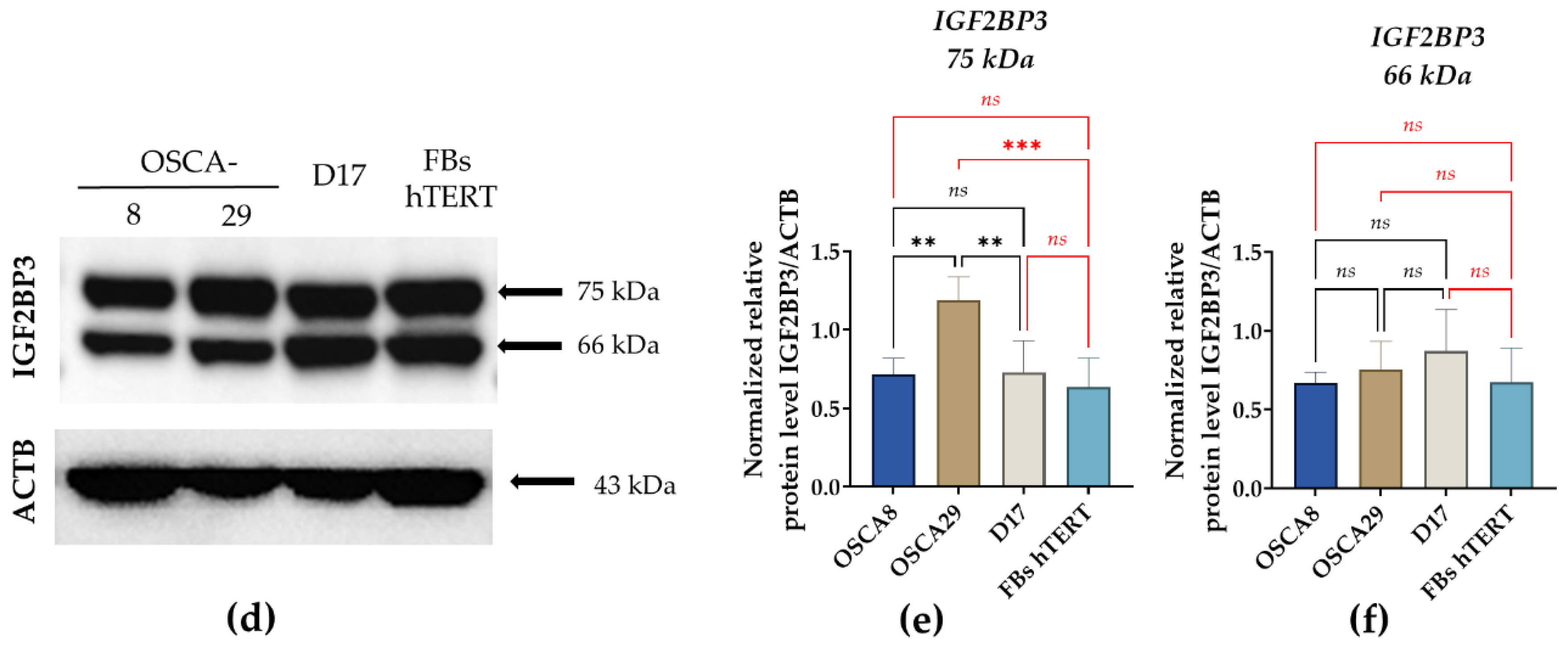
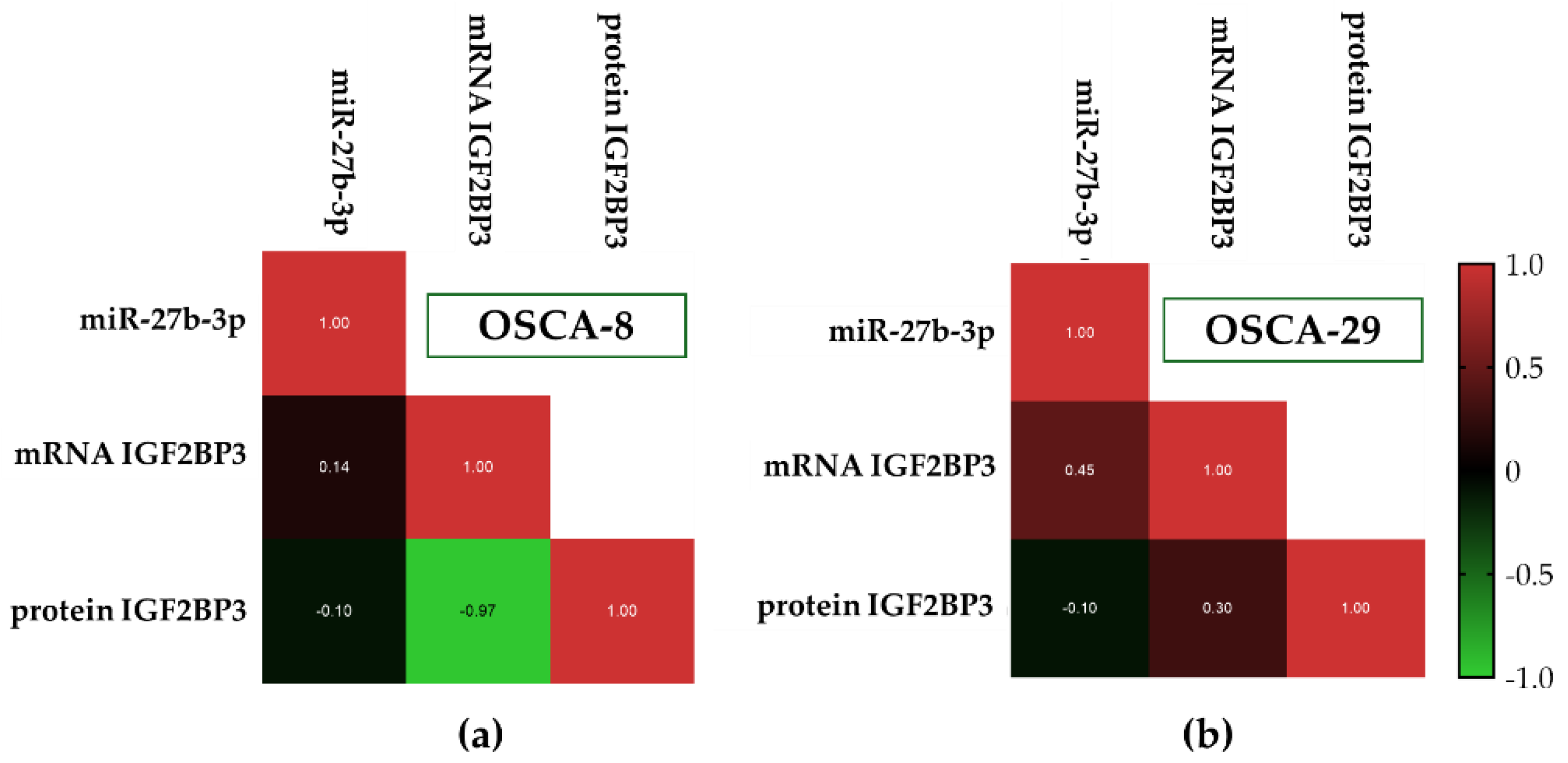
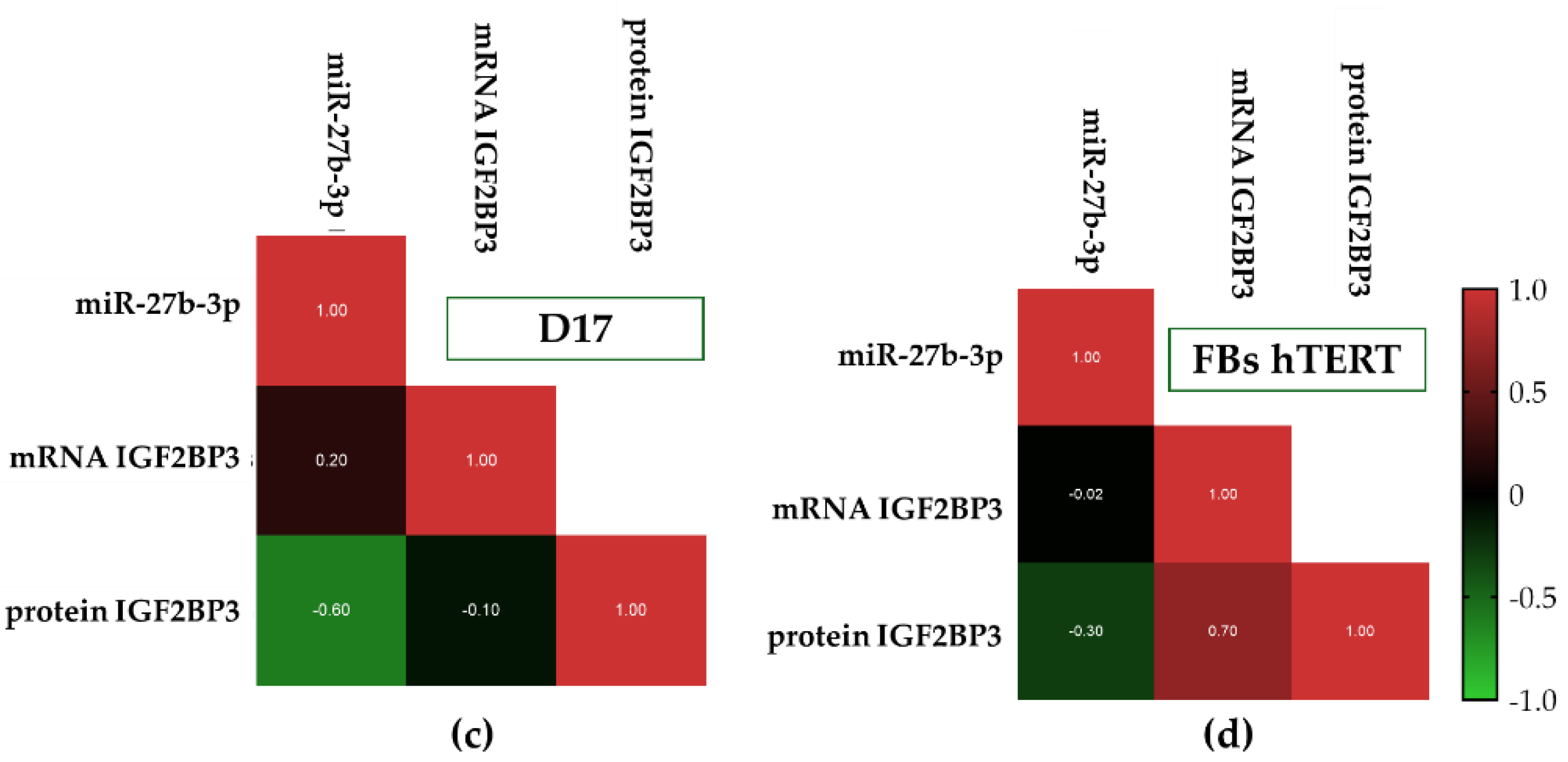
3.4. Distinct Post-Transcriptional Patterns of IGF2BP3 Expression Are Observed Across Canine Osteosarcoma Cell Lines Despite Partially Concordant miR-27b-3p and Transcript Profiles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ACTB | β-actin |
| AKT | Protein Kinase B |
| BCA | Bicinchoninic Acid Assay |
| BSA | Bovine Serum Albumin |
| CFU | Colony-Forming Unit |
| CGM | Complete Growth Medium |
| DAPI | 4′,6-diamidino-2-phenylindole dihydrochloride |
| DMEM | Dulbecco's Modified Eagle Medium |
| EMEM | Eagle's Minimum Essential Medium |
| FBS | Fetal Bovine Serum |
| GAPDH | Glyceraldehyde-3-Phosphate Dehydrogenase |
| HBSS | Hanks' Balanced Salt Solution |
| HRP | Chemiluminescent and Fluorescent Peroxidase |
| IGF-1R | Insulin-like Growth Factor 1 Receptor |
| IGF2BP3 | Insulin-like Growth Factor 2 mRNA-Binding Protein 3 |
| IGF-I/GH | Insulin-like growth factor 1/ Growth Hormone |
| MAPK | Mitogen-Activated Protein Kinases |
| OSA | Osteosarcoma |
| P/S | Penicillin-Streptomycin |
| PFA | paraformaldehyde |
| PVDF | Polyvinylidene Fluoride |
| TBS | Tris-Buffered Saline with Tween 20 |
Appendix A
| Parameter/Cell line | OSCA-8 | OSCA-29 | D17 |
|---|---|---|---|
| Morphology | spindle-shaped | spindle-shaped | epithelial-like |
| Tissue of origin | Bone | bone | bone |
| Tumor origin | primary tumor | primary tumor | lung metastasis |
| Age of donor dog (years) | 1 | 6 | 11 |
| Breed | Rottweiler | Rottweiler | Poodle |
| Sex | Male | Female | Female |
References
- Qin, S.; Hu, Y.; Deng, R.; Wang, Z. Exploring the Heterogeneity of Osteosarcoma Cell Characteristics and Metabolic States and Their Association with Clinical Prognosis. Front Immunol 2024, 15, 1507476. [Google Scholar] [CrossRef]
- Remiszewski, P.; Siedlecki, E.; Wełniak-Kamińska, M.; Mikula, M.; Czarnecka, A. Choosing the Right Animal Model for Sarcoma Research. Cell Mol Life Sci 2026, 83, 73. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Dong, X.; Chen, S. Advances in Osteosarcoma Research: Pathogenesis and Emerging Therapeutic Strategies. Crit Rev Oncol Hematol 2026, 222, 105252. [Google Scholar] [CrossRef] [PubMed]
- Dolnicka, A.; Fosse, V.; Raciborska, A.; Śmieszek, A. Building a Therapeutic Bridge Between Dogs and Humans: A Review of Potential Cross-Species Osteosarcoma Biomarkers. Int J Mol Sci 2025, 26, 5152. [Google Scholar] [CrossRef] [PubMed]
- Jackson-Oxley, J.; Alibhai, A.A.; Thompson, R.; Lothion-Roy, J.; de Brot, S.; Dunning, M.D.; Jeyapalan, J.N.; Mongan, N.P.; Rutland, C.S. Identification and Characterisation of Canine Osteosarcoma Biomarkers and Therapeutic Targets. Cancers (Basel) 2026, 18, 262. [Google Scholar] [CrossRef]
- Fan, T.M.; Khanna, C. Comparative Aspects of Osteosarcoma Pathogenesis in Humans and Dogs. Vet Sci 2015, 2, 210–230. [Google Scholar] [CrossRef]
- Edmunds, G.L.; Smalley, M.J.; Beck, S.; Errington, R.J.; Gould, S.; Winter, H.; Brodbelt, D.C.; O’Neill, D.G. Dog Breeds and Body Conformations with Predisposition to Osteosarcoma in the UK: A Case-Control Study. Canine Med Genet 2021, 8, 2. [Google Scholar] [CrossRef]
- Dixit, M.; Poudel, S.B.; Yakar, S. Effects of GH/IGF Axis on Bone and Cartilage. Mol Cell Endocrinol 2021, 519, 111052. [Google Scholar] [CrossRef]
- Maniscalco, L.; Iussich, S.; Morello, E.; Martano, M.; Gattino, F.; Miretti, S.; Biolatti, B.; Accornero, P.; Martignani, E.; Sánchez-Céspedes, R.; et al. Increased Expression of Insulin-like Growth Factor-1 Receptor Is Correlated with Worse Survival in Canine Appendicular Osteosarcoma. Vet J 2015, 205, 272–280. [Google Scholar] [CrossRef]
- Beck, J.; Ren, L.; Huang, S.; Berger, E.; Bardales, K.; Mannheimer, J.; Mazcko, C.; LeBlanc, A. Canine and Murine Models of Osteosarcoma. Vet Pathol 2022, 59, 399–414. [Google Scholar] [CrossRef]
- Leitner, N.; Ertl, R.; Gabner, S.; Fuchs-Baumgartinger, A.; Walter, I.; Hlavaty, J. Isolation and Characterization of Novel Canine Osteosarcoma Cell Lines from Chemotherapy-Naïve Patients. Cells 2023, 12, 1026. [Google Scholar] [CrossRef] [PubMed]
- Choong, P.F.; Teh, H.X.; Teoh, H.K.; Ong, H.K.; Choo, K.B.; Sugii, S.; Cheong, S.K.; Kamarul, T. Heterogeneity of Osteosarcoma Cell Lines Led to Variable Responses in Reprogramming. Int J Med Sci 2014, 11, 1154–1160. [Google Scholar] [CrossRef] [PubMed]
- Romero Rodríguez, M.I.; Vargas Pino, J.C.; Sierra-Ballén, E.L. Tumor Growth, Proliferation and Diffusion in Osteosarcoma. Acta Biotheor 2025, 73, 4. [Google Scholar] [CrossRef] [PubMed]
- Dutour, A.; Pasello, M.; Farrow, L.; Amer, M.H.; Entz-Werlé, N.; Nathrath, M.; Scotlandi, K.; Mittnacht, S.; Gomez-Mascard, A. Microenvironment Matters: Insights from the FOSTER Consortium on Microenvironment-Driven Approaches to Osteosarcoma Therapy. Cancer Metastasis Rev 2025, 44, 44. [Google Scholar] [CrossRef]
- Gola, C.; Giannuzzi, D.; Rinaldi, A.; Iussich, S.; Modesto, P.; Morello, E.; Buracco, P.; Aresu, L.; De Maria, R. Genomic and Transcriptomic Characterization of Canine Osteosarcoma Cell Lines: A Valuable Resource in Translational Medicine. Front Vet Sci 2021, 8, 666838. [Google Scholar] [CrossRef]
- Modesto, P.; Fernandez, J.L.C.; Martini, I.; Zoccola, R.; Pugliano, M.C.; Ciucis, C.G.D.; Goria, M.; Ferrari, A.; Razzuoli, E. Characterization of D-17 Canine Osteosarcoma Cell Line and Evaluation of Its Ability to Response to Infective Stressor Used as Alternative Anticancer Therapy. Animals (Basel) 2020, 10, 1981. [Google Scholar] [CrossRef]
- Scott, M.C.; Sarver, A.L.; Gavin, K.J.; Thayanithy, V.; Getzy, D.M.; Newman, R.A.; Cutter, G.R.; Lindblad-Toh, K.; Kisseberth, W.C.; Hunter, L.E.; et al. Molecular Subtypes of Osteosarcoma Identified by Reducing Tumor Heterogeneity through an Interspecies Comparative Approach. Bone 2011, 49, 356–367. [Google Scholar] [CrossRef]
- Gebhard, C.; Gabriel, C.; Walter, I. Morphological and Immunohistochemical Characterization of Canine Osteosarcoma Spheroid Cell Cultures. Anat Histol Embryol 2016, 45, 219–230. [Google Scholar] [CrossRef]
- Legare, M.E.; Bush, J.; Ashley, A.K.; Kato, T.; Hanneman, W.H. Cellular and Phenotypic Characterization of Canine Osteosarcoma Cell Lines. J Cancer 2011, 2, 262–270. [Google Scholar] [CrossRef]
- Do, S.-I.; Kim, Y.W.; Park, H.-R.; Park, Y.-K. Expression of Insulin-like Growth Factor-II mRNA Binding Protein 3 (IMP3) in Osteosarcoma. Oncol Res 2008, 17, 269–272. [Google Scholar] [CrossRef]
- Lederer, M.; Bley, N.; Schleifer, C.; Hüttelmaier, S. The Role of the Oncofetal IGF2 mRNA-Binding Protein 3 (IGF2BP3) in Cancer. Semin Cancer Biol 2014, 29, 3–12. [Google Scholar] [CrossRef]
- Mancarella, C.; Scotlandi, K. IGF2BP3 From Physiology to Cancer: Novel Discoveries, Unsolved Issues, and Future Perspectives. Front Cell Dev Biol 2019, 7, 363. [Google Scholar] [CrossRef] [PubMed]
- Shu, R.; Cheng, Q.; Yu, Z.; Zhou, H.; Lin, M.; Shi, M.; Chen, J.; Chen, J.; Zhao, M. The METTL3-IGF2BP3 Axis Drives Osteosarcoma Progression by Enhancing ID1 mRNA Stability. BMC Musculoskelet Disord 2026, 27, 179. [Google Scholar] [CrossRef]
- Liu, X.; Chen, J.; Chen, W.; Xu, Y.; Shen, Y.; Xu, X. Targeting IGF2BP3 in Cancer. Int J Mol Sci 2023, 24, 9423. [Google Scholar] [CrossRef]
- Chen, L.-J.; Liu, H.-Y.; Xiao, Z.-Y.; Qiu, T.; Zhang, D.; Zhang, L.-J.; Han, F.-Y.; Chen, G.-J.; Xu, X.-M.; Zhu, J.-H.; et al. IGF2BP3 Promotes the Progression of Colorectal Cancer and Mediates Cetuximab Resistance by Stabilizing EGFR mRNA in an m6A-Dependent Manner. Cell Death Dis 2023, 14, 581. [Google Scholar] [CrossRef]
- Zhu, T.; He, L.; Li, S.; Zhao, J. Identification of IGF2BP3 and CENPA as Key Regulators of Immunophenotypes in Renal Clear Cell Carcinoma. Front Genet 2025, 16, 1749780. [Google Scholar] [CrossRef]
- Yang, Y.; Gao, J.; Shi, H.; Sihto, H.; Kilpinen, S.; Vilcot, F.; Janská, L.; Jeschonneck, J.; Cvetanovic, T.; Höög, A.; et al. IGF2BP3 As a Prognostic Biomarker and Regulator of Metastasis in Merkel Cell Carcinoma. JID Innov 2025, 5, 100355. [Google Scholar] [CrossRef]
- Ye, P.; Ke, X.; Zang, X.; Sun, H.; Dong, Z.; Lin, J.; Wang, L.; Liu, W.; Miao, G.; Tan, Y.; et al. Up-Regulated MiR-27-3p Promotes the G1-S Phase Transition by Targeting Inhibitor of Growth Family Member 5 in Osteosarcoma. Biomed Pharmacother 2018, 101, 219–227. [Google Scholar] [CrossRef]
- Shen, S.-J.; Song, Y.; Ren, X.-Y.; Xu, Y.-L.; Zhou, Y.-D.; Liang, Z.-Y.; Sun, Q. MicroRNA-27b-3p Promotes Tumor Progression and Metastasis by Inhibiting Peroxisome Proliferator-Activated Receptor Gamma in Triple-Negative Breast Cancer. Front Oncol 2020, 10, 1371. [Google Scholar] [CrossRef]
- Zhu, L.; Long, C.; Tang, Y.; Wu, C.; Wei, L.; Ye, T.; Chen, J. miR-27b-3p Directly Targets PPARG to Regulate Pancreatic Cancer Progression. Discov Oncol 2026, 17, 398. [Google Scholar] [CrossRef]
- Amin, S.; Marcinkowska, K.; Wołoszyńska, M.; Opaliński, S.; Skrzypczak, D.; Wiercik, P.; Bobak, Ł; Śmieszek, A. Evaluation of the Cytotoxicity of Biochar Aqueous Extract in Caco-2 Cells: Time-Dependent Regulation of Apoptosis, Associated with miRNA Modulation. Molecules 2026, 31, 989. [Google Scholar] [CrossRef]
- Stein, C. A Two-Sample Test for a Linear Hypothesis Whose Power Is Independent of the Variance. The Annals of Mathematical Statistics 1945, 16, 243–258. [Google Scholar] [CrossRef]
- Megquier, K.; Turner-Maier, J.; Morrill, K.; Li, X.; Johnson, J.; Karlsson, E.K.; London, C.A.; Gardner, H.L. The Genomic Landscape of Canine Osteosarcoma Cell Lines Reveals Conserved Structural Complexity and Pathway Alterations. PLoS One 2022, 17, e0274383. [Google Scholar] [CrossRef]
- Cannon, C.M.; Pozniak, J.; Scott, M.C.; Ito, D.; Gorden, B.H.; Graef, A.J.; Modiano, J.F. Canine Osteosarcoma Cells Exhibit Resistance to Aurora Kinase Inhibitors. Vet Comp Oncol 2015, 13, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Małek, A.; Wojnicki, M.; Borkowska, A.; Wójcik, M.; Ziółek, G.; Lechowski, R.; Zabielska-Koczywąs, K. Gold Nanoparticles Inhibit Extravasation of Canine Osteosarcoma Cells in the Ex Ovo Chicken Embryo Chorioallantoic Membrane Model. Int J Mol Sci 2023, 24, 9858. [Google Scholar] [CrossRef]
- Wilk, S.S.; Kukier, K.I.; Michałowski, A.M.; Wojnicki, M.; Smereczyński, B.; Wójcik, M.; Zabielska-Koczywąs, K.A. The Anti-Metastatic Properties of Glutathione-Stabilized Gold Nanoparticles-A Preliminary Study on Canine Osteosarcoma Cell Lines. Int J Mol Sci 2025, 26, 6102. [Google Scholar] [CrossRef]
- McMahon, M.B.; Bear, M.D.; Kulp, S.K.; Pennell, M.L.; London, C.A. Biological Activity of Gemcitabine against Canine Osteosarcoma Cell Lines in Vitro. Am J Vet Res 2010, 71, 799–808. [Google Scholar] [CrossRef]
- Fowles, J.S.; Dailey, D.D.; Gustafson, D.L.; Thamm, D.H.; Duval, D.L. The Flint Animal Cancer Center (FACC) Canine Tumour Cell Line Panel: A Resource for Veterinary Drug Discovery, Comparative Oncology and Translational Medicine. Vet Comp Oncol 2017, 15, 481–492. [Google Scholar] [CrossRef]
- Lindström, H.J.G.; Friedman, R. Inferring Time-Dependent Population Growth Rates in Cell Cultures Undergoing Adaptation. BMC Bioinformatics 2020, 21, 583. [Google Scholar] [CrossRef]
- Musser, M.L.; Mahaffey, A.L.; Fath, M.A.; Buettner, G.R.; Wagner, B.A.; Schneider, B.K.; Seo, Y.-J.; Mochel, J.P.; Johannes, C.M. In Vitro Cytotoxicity and Pharmacokinetic Evaluation of Pharmacological Ascorbate in Dogs. Front Vet Sci 2019, 6, 385. [Google Scholar] [CrossRef]
- Pan, W.; Wang, H.; Jianwei, R.; Ye, Z. MicroRNA-27a Promotes Proliferation, Migration and Invasion by Targeting MAP2K4 in Human Osteosarcoma Cells. Cell Physiol Biochem 2014, 33, 402–412. [Google Scholar] [CrossRef]
- Li, Z.; Feng, L.; Wei, X.; Li, H.; Zhu, Y.; Wang, H.; Liu, J.; Luo, L.; Zheng, Z.; Song, B.; et al. IGF2BP3-Dependent Glutamine/BCAA Metabolic Rewiring Rejuvenates Aged Human Adipose-Derived Stem Cells for Enhanced Tissue Regeneration. Cell Discov 2026, 12, 5. [Google Scholar] [CrossRef]
- Martinelli, M.; Mancarella, C.; Scapoli, L.; Palmieri, A.; De Sanctis, P.; Ferrari, C.; Pasello, M.; Zucchini, C.; Scotlandi, K. Polymorphic Variants of IGF2BP3 and SENCR Have an Impact on Predisposition and/or Progression of Ewing Sarcoma. Front Oncol 2022, 12, 968884. [Google Scholar] [CrossRef]
- Mancarella, C.; Giusti, V.; Caldoni, G.; Laginestra, M.A.; Parra, A.; Toracchio, L.; Giordano, G.; Roncuzzi, L.; Piazzi, M.; Blalock, W.; et al. Extracellular Vesicle-Associated IGF2BP3 Tunes Ewing Sarcoma Cell Migration and Affects PI3K/Akt Pathway in Neighboring Cells. Cancer Gene Ther 2023, 30, 1285–1295. [Google Scholar] [CrossRef]
- Sánchez-Sendra, B.; Pérez-Debén, S.; González-Muñoz, J.F.; Murgui, A.; Monteagudo, C. Prognostic Value of IGF2 mRNA-Binding Protein 3 (IGF2BP3) Intratumoral Expression in Melanoma Patients at the Time of Diagnosis: Comparative Analysis of RT-qPCR Versus Immunohistochemistry. Cancers (Basel) 2022, 14, 2319. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




