1. Introduction
Cervical cancer continues to be a significant cause of morbidity and mortality among women globally, with a particularly pronounced impact in regions with limited access to systematic screening programs and specialized personnel [
1]. Cervical cytology, in either conventional or liquid-based modalities, is a consolidated tool for the early detection of precursor lesions and, by extension, the reduction of mortality associated with cervical cancer [
2]. However, cytological interpretation remains a predominantly manual, time-intensive process dependent on the experience of the cytotechnologist or cytopathologist, which introduces inter-observer variability and limits the operational scalability of screening programs [
3].
The progressive digitization of cytological slides through WSI technologies, coupled with the recent maturity of deep learning methods, has driven the development of automated medical image analysis systems aimed at supporting specialists in the detection, localization, and prioritization of findings [
4]. Recent studies have shown that artificial intelligence can achieve accuracy levels comparable to expert personnel in identifying cervical lesions [
5], reinforcing its potential as an assisted screening tool. In digital pathology, and more recently, in cervical cytology, these approaches address workflows based on high-resolution images, where cell pattern identification must be robust against staining variations, artifacts, blurring, and intra- and inter-patient heterogeneity [
6]. This automated analysis paradigm is especially relevant as a support mechanism for population-based screening, allowing for standardized criteria and reduced workload in environments with a shortage of qualified personnel [
7]. These capabilities make detection models promising tools that facilitate the integration of artificial intelligence into clinical screening workflows.
Among these, single-stage detectors, such as the YOLO family originally proposed by Redmon et al. [
8], stand out for their balance between accuracy and computational efficiency [
9]. Their ability to localize multiple small objects per image with reduced inference times makes them particularly suitable for cytological analysis, where numerous cells coexist, often partially overlapping, with high morphological variability and a complex visual context [
10]. Consequently, these detectors are positioned as practical candidates for localizing and characterizing abnormal cells in WSI-derived images, operating at the patch level as the unit of analysis, and serving as a component of evidence for clinical screening flows.
However, despite the reported progress, there is still a lack of consensus on methodological decisions that significantly affect the performance of detectors in realistic medical scenarios, particularly when working with limited and imbalanced datasets, which is a common situation in cervical cytology [
6]. Some studies have explored architectural variants or training improvements, but with heterogeneity in data partitioning, target variable definition, and metrics, which complicates direct comparisons between approaches [
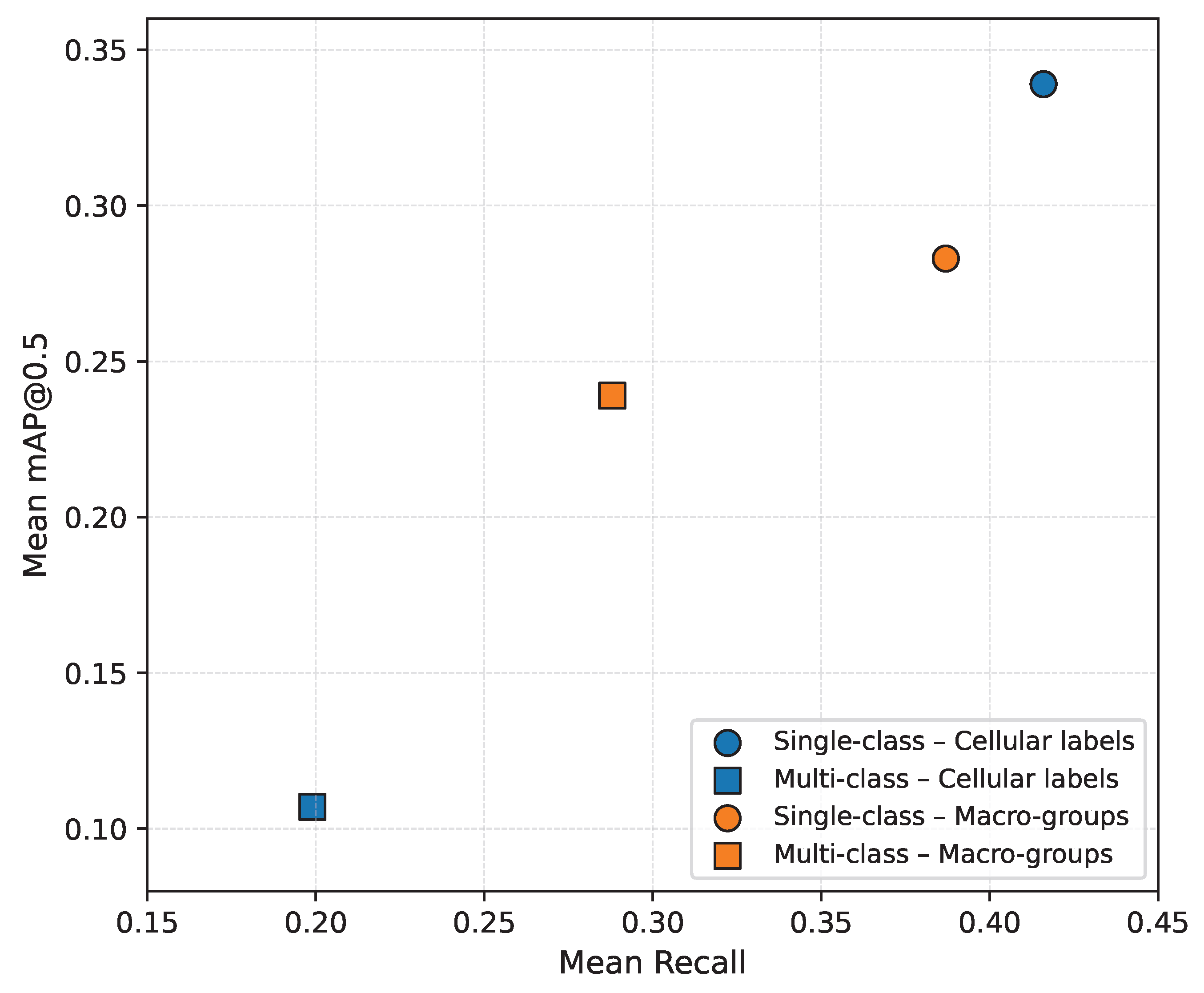
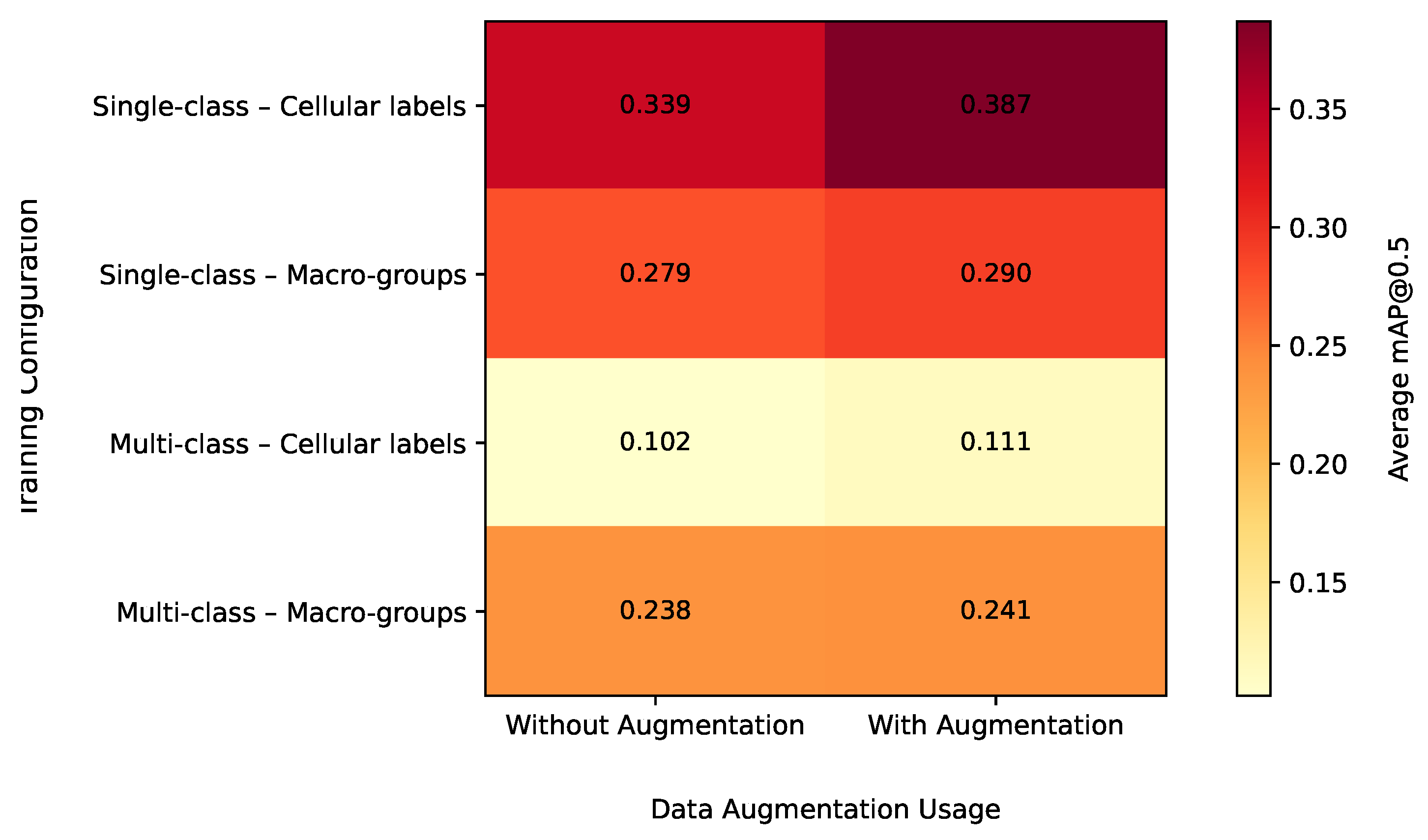
11]. Among the methodological decisions that seem to have the greatest impact are the following:
- (i)
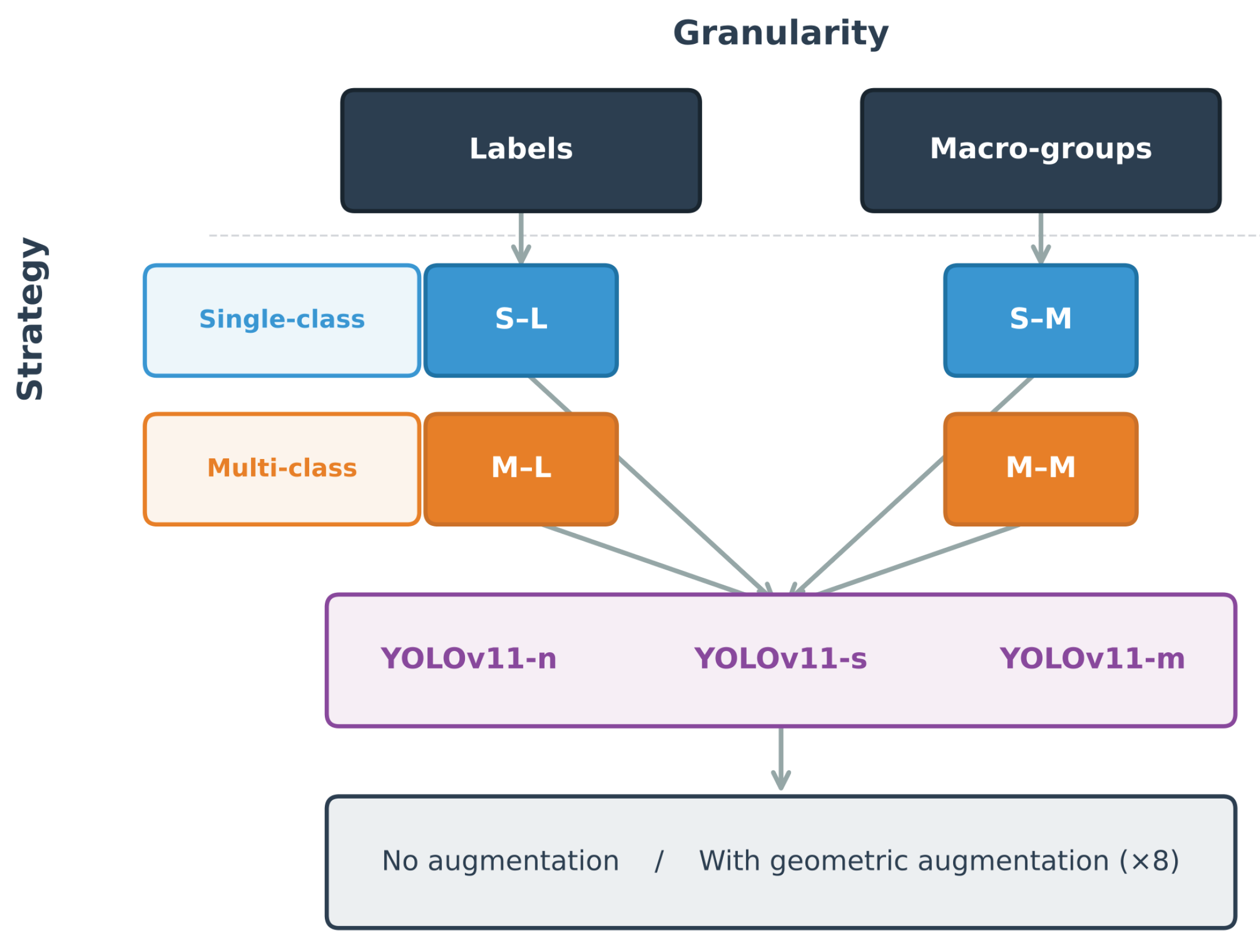
Single-class versus multi-class formulation; in this work, single-class formulation was defined as training a detector with a single target label using an independent model for each specific cell label or diagnostic macro-group, while multi-class formulation was defined as training a single detector capable of simultaneously discriminating all specific cell labels or diagnostic macro-groups.
- (ii)
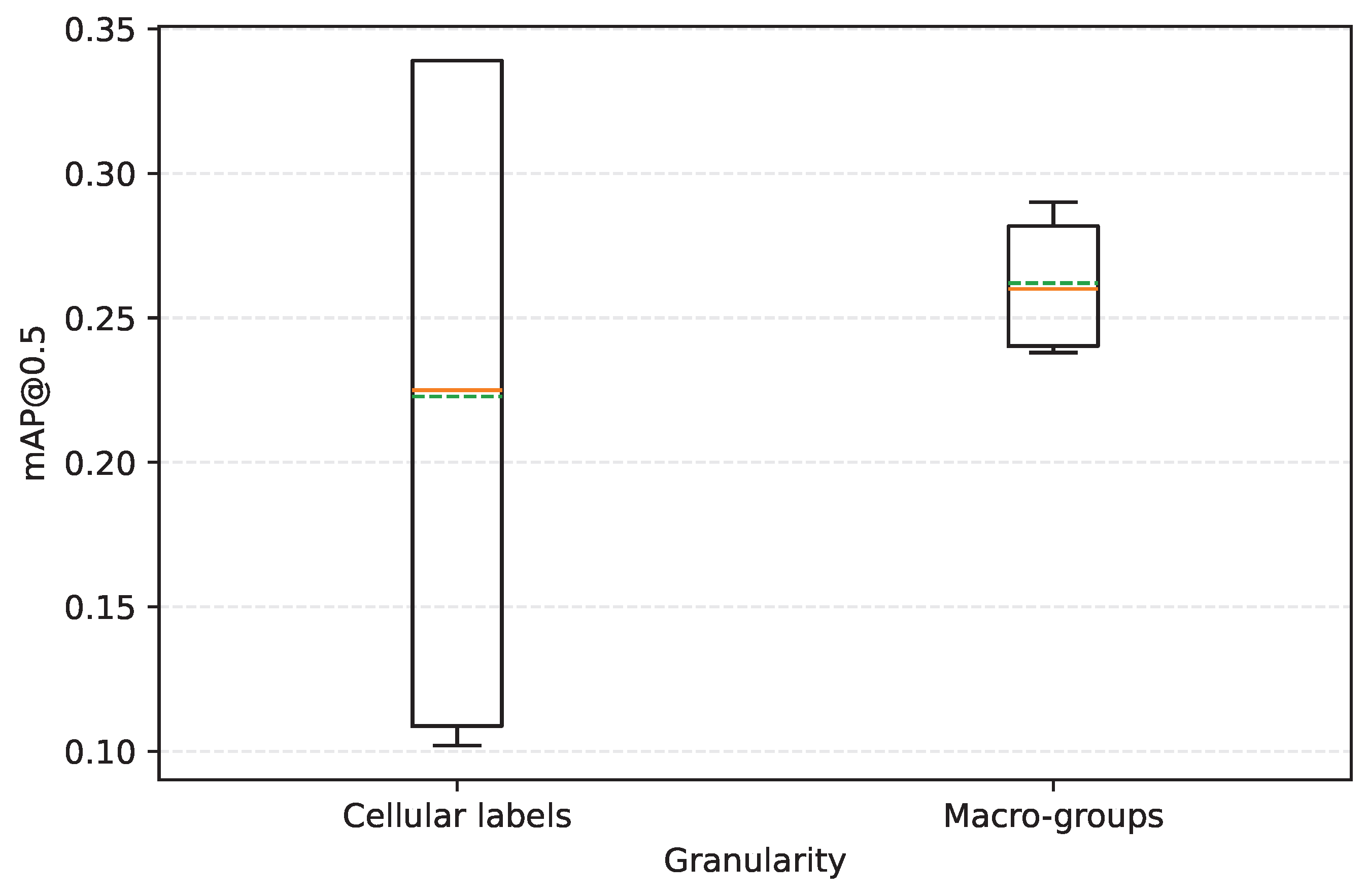
The granularity of the target variable, differentiating between a fine level based on specific cell labels and an aggregated level based on diagnostic macro-groups.
- (iii)
-
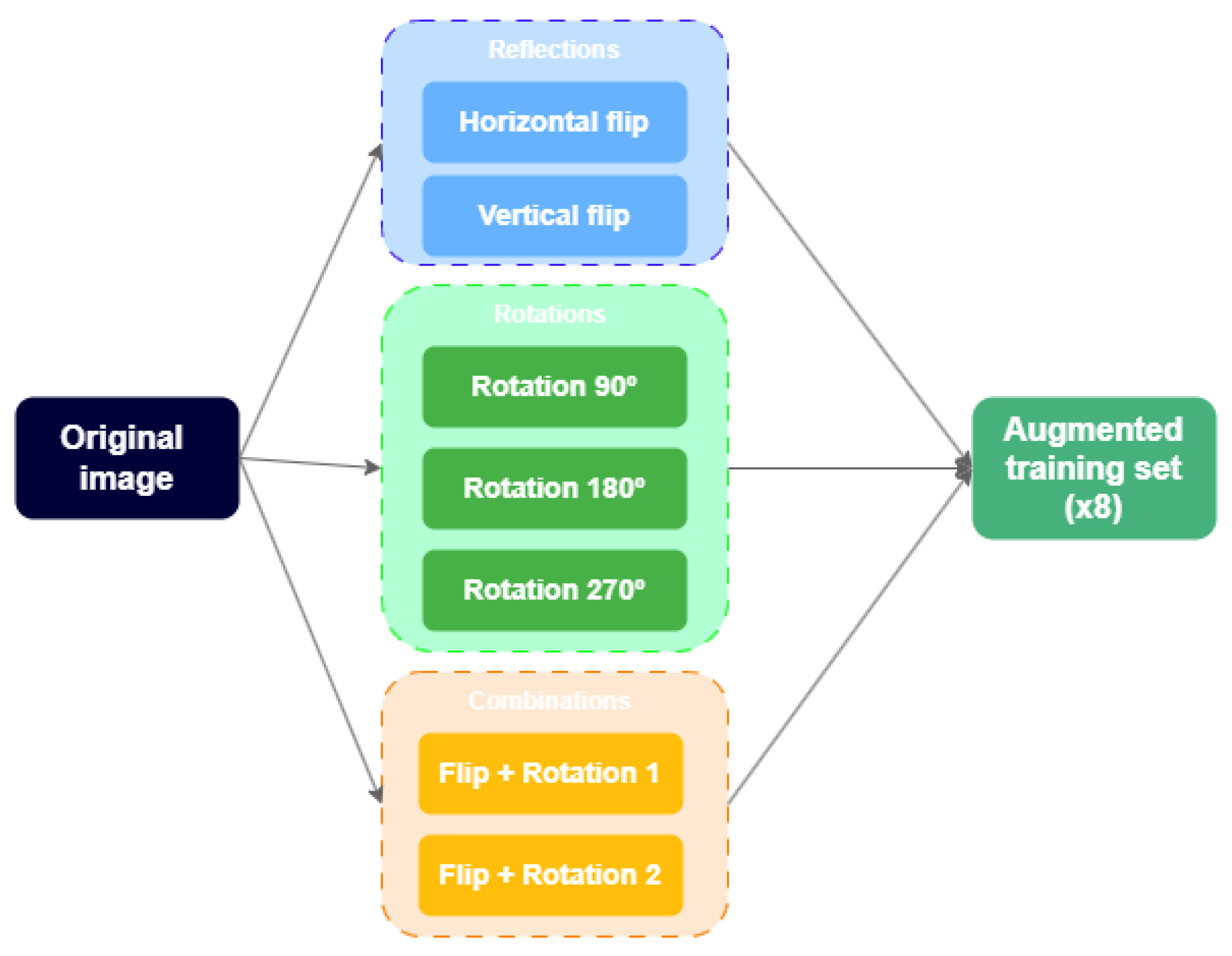
The use of geometric data augmentation applied only during training as a mechanism to improve generalization without biasing the evaluation.
Additionally, comparative evidence on how these choices interact with model size and capacity (variants with different trade-offs between precision and efficiency) remains limited, despite the availability of scalable architectures within the same family of models.
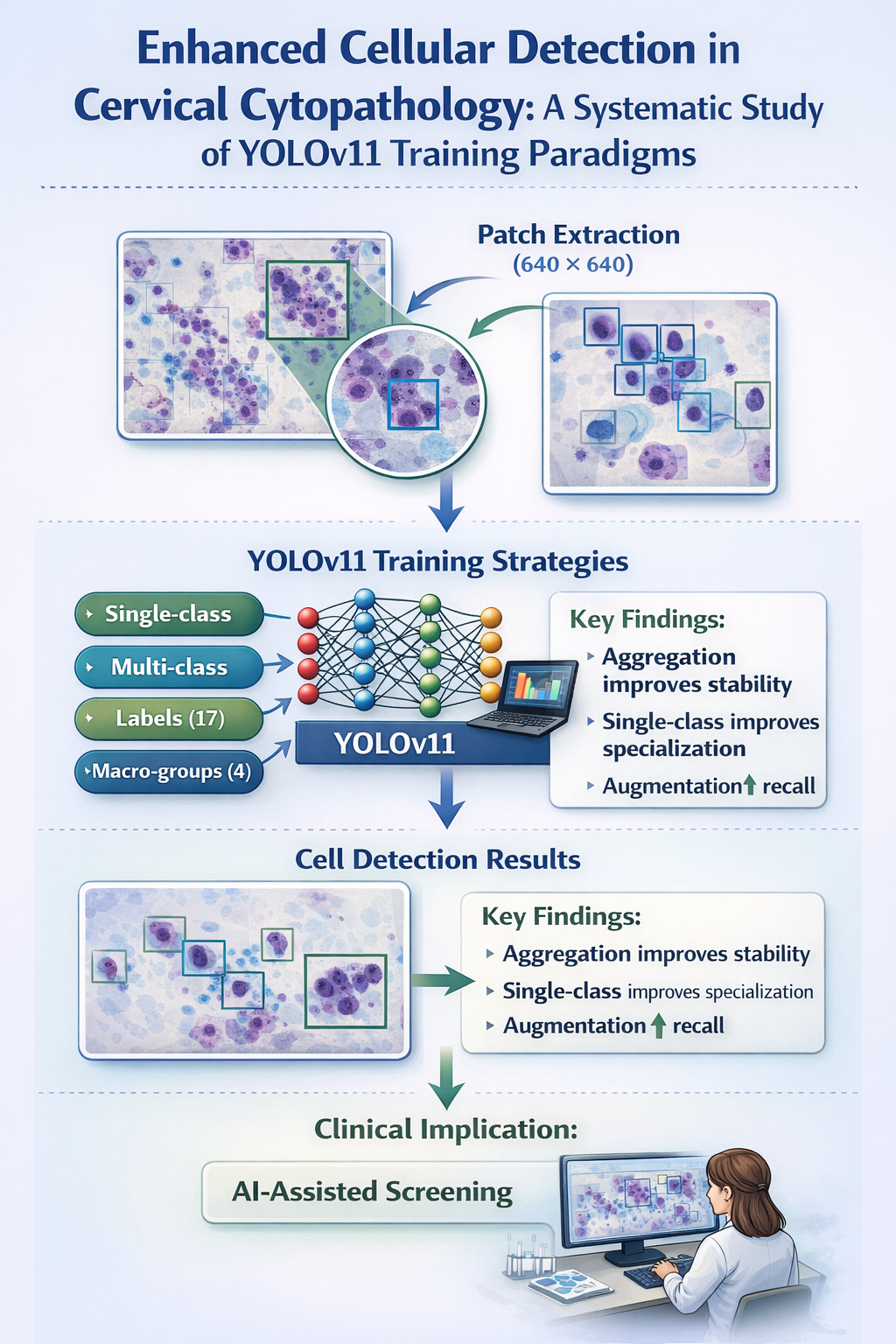
In this study, these variables were systematically studied through a controlled experimental evaluation of YOLOv11 for cell detection in digital cervical cytology. YOLOv11 was selected for its suitability for detecting small and numerous objects, as well as for offering scalable variants within the same architectural family, allowing for the comparison of models with different capacities under homogeneous conditions. Specifically, the
n (
nano),
s (
small), and
m (
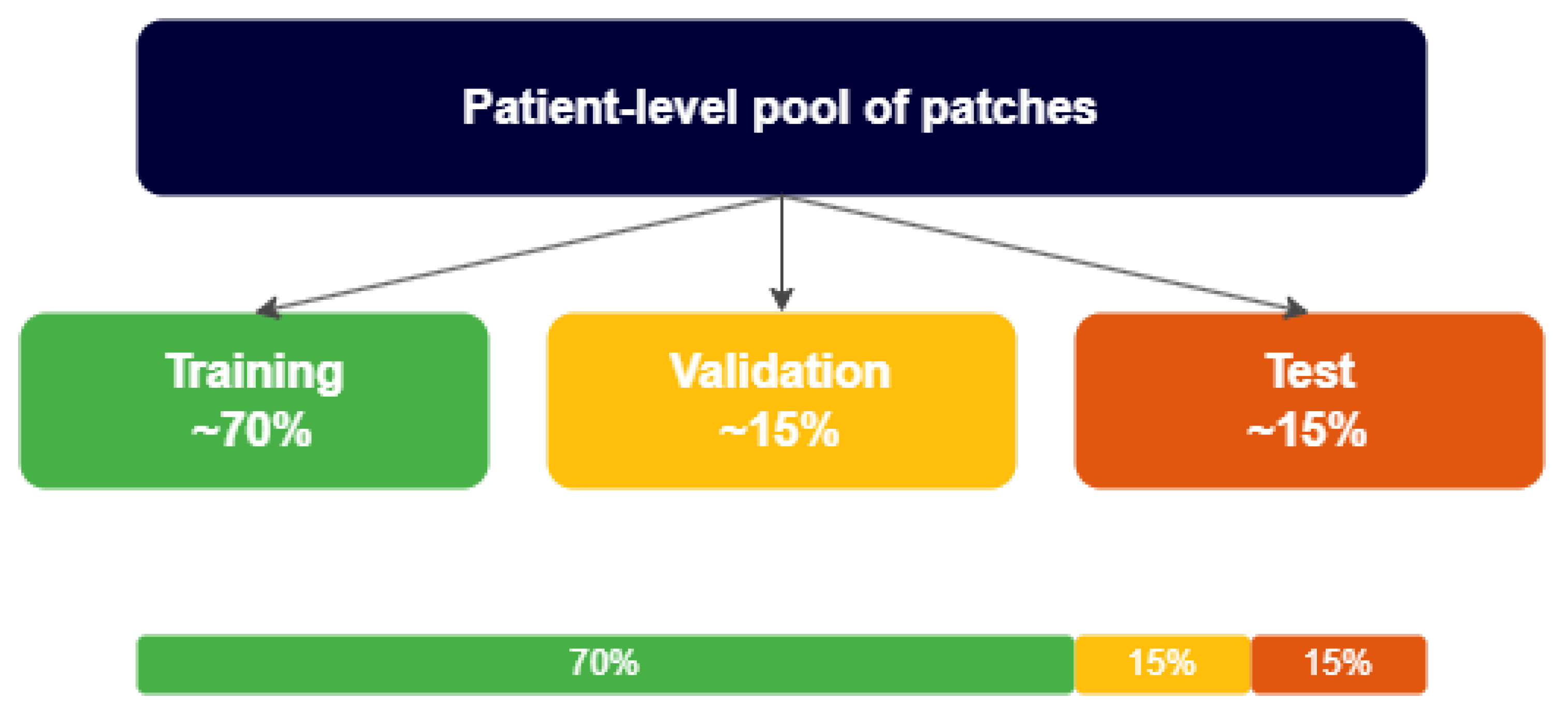
medium) variants, which represent configurations of increasing complexity, were analyzed. Training strategies that combined single-class/multi-class formulations with two levels of target variable granularity—fine (specific cell labels) and aggregated (diagnostic macro-groups)—were compared, considering the effect of data augmentation applied exclusively to the training set. The evaluation was designed with patient-level splitting to minimize information leakage between training, validation, and testing, following digital pathology best practices [
12] and approximating the measured performance to a real clinical use scenario.
The contribution of this study is to provide a reproducible comparative framework to understand the impact of (a) problem formulation (single-class vs. multi-class), (b) target variable granularity, (c) data augmentation, and (d) model size in a biomedical context with data constraints and class imbalance. In addition to informing the practical choice of training configurations for YOLOv11 detectors in cervical cytology, this analysis establishes a useful methodological foundation for integrating WSI-level information and exploring global aggregation schemes, including approaches such as Multiple Instance Learning (MIL).
Based on the issues presented and the need to establish robust methodological criteria for using state-of-the-art detection models in digital cytology, this paper is structured as follows:
Section 2 reviews the state of the art, addressing the challenges of AI in cervical cytology, the influence of the Bethesda taxonomy on model design, and the positioning of the YOLO family in this field.
Section 3 details the experimental methodology, including the data source, factorial design of the experiments (single-class vs. multi-class strategies and target variable granularity), and strict patient-level partitioning protocol to ensure clinical validity. Subsequently,
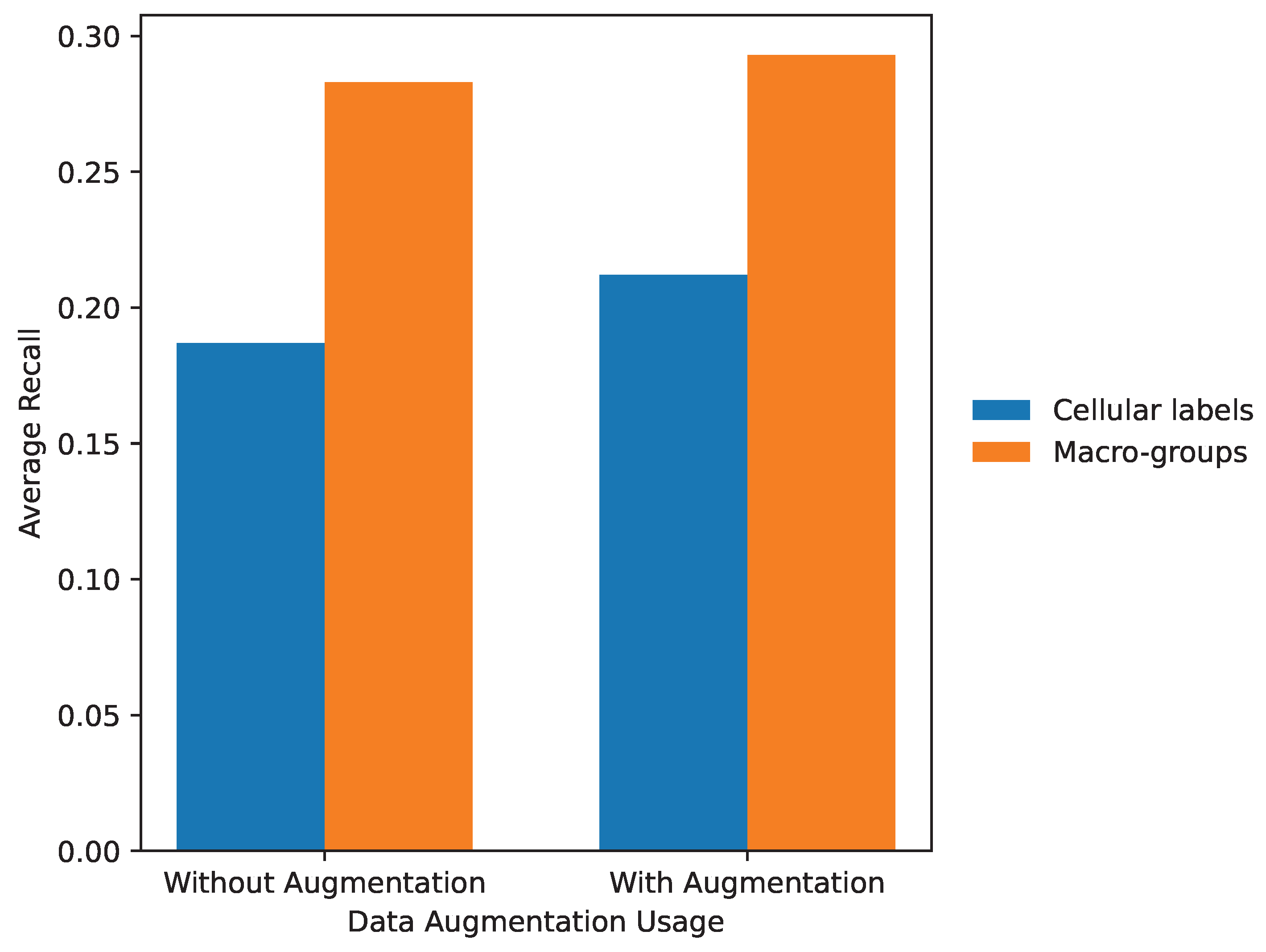
Section 4 presents the quantitative results, analyzing the effects of data augmentation, model capacity, and the interaction between the evaluated factors using precision, recall, and mAP metrics. Finally,
Section 5 discusses the implications of these findings.
Section 6 summarizes the study’s conclusions, main limitations, and future lines of work.