Submitted:
08 April 2026
Posted:
10 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Results
3.1. The Genomic Periodicity Index Defines a Universal Structural Rhythm of the Human Genome
3.2. Dinucleotide Composition Resolves into Five Universal Grammar Classes—The Panchamahābhūta States
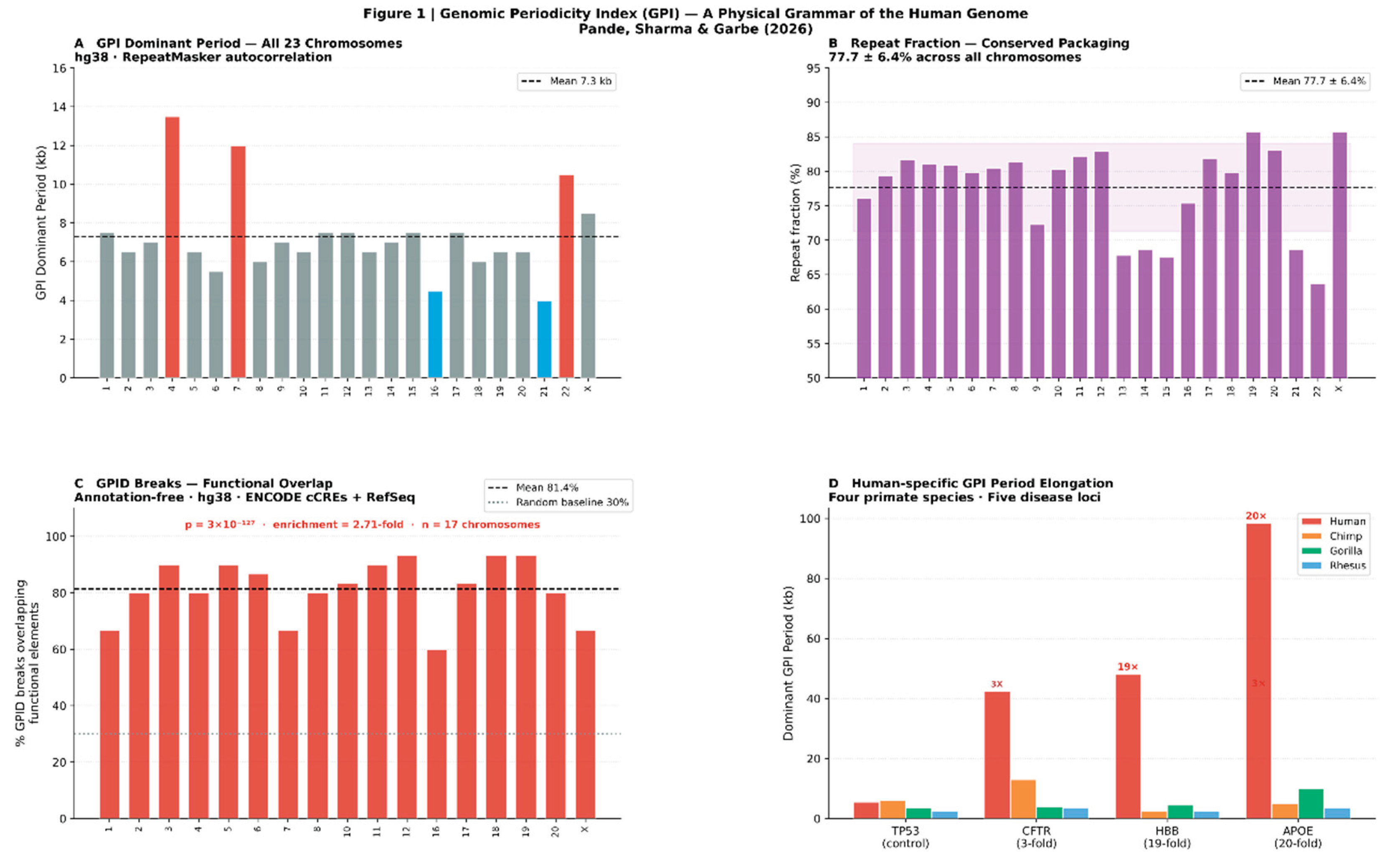
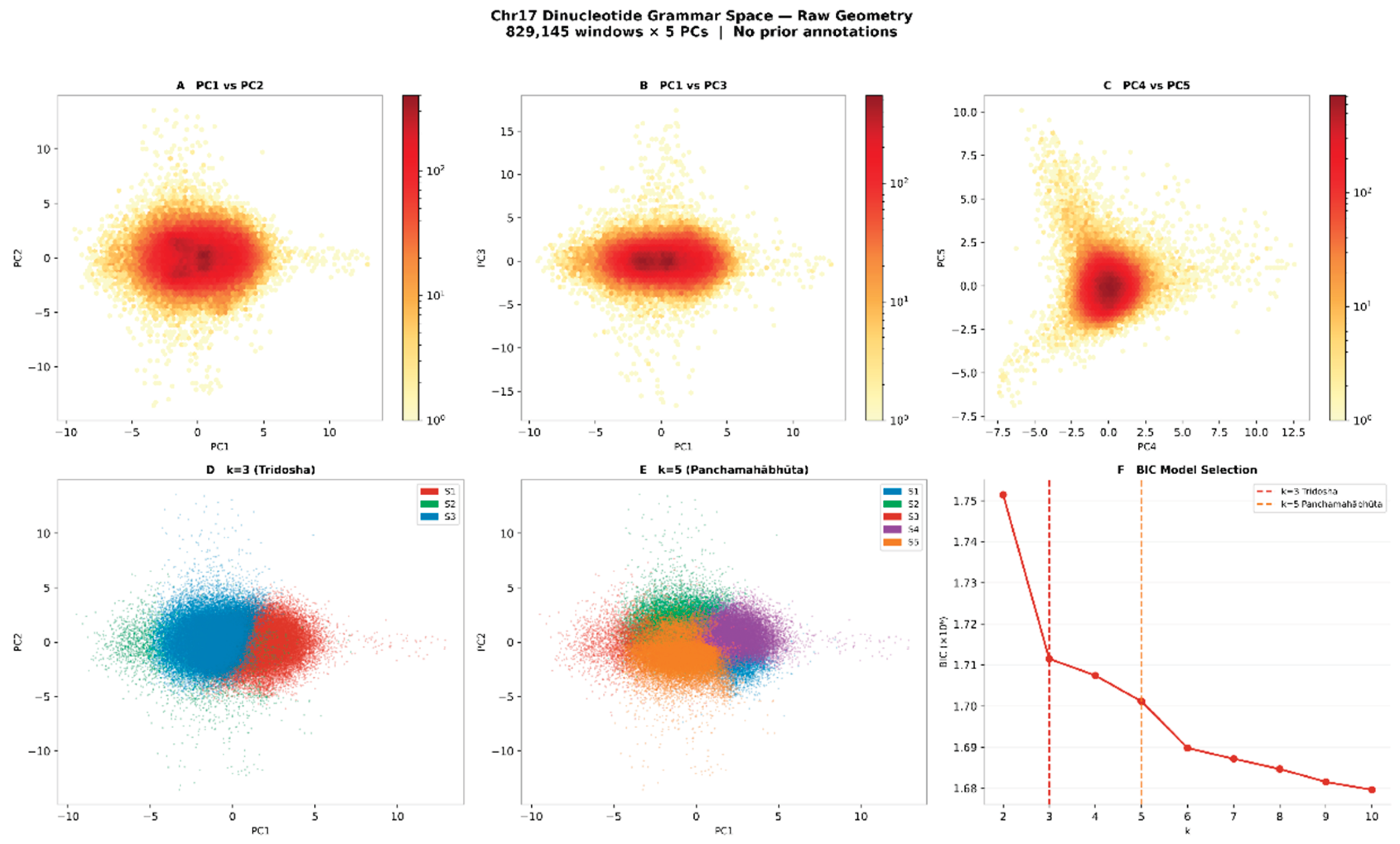
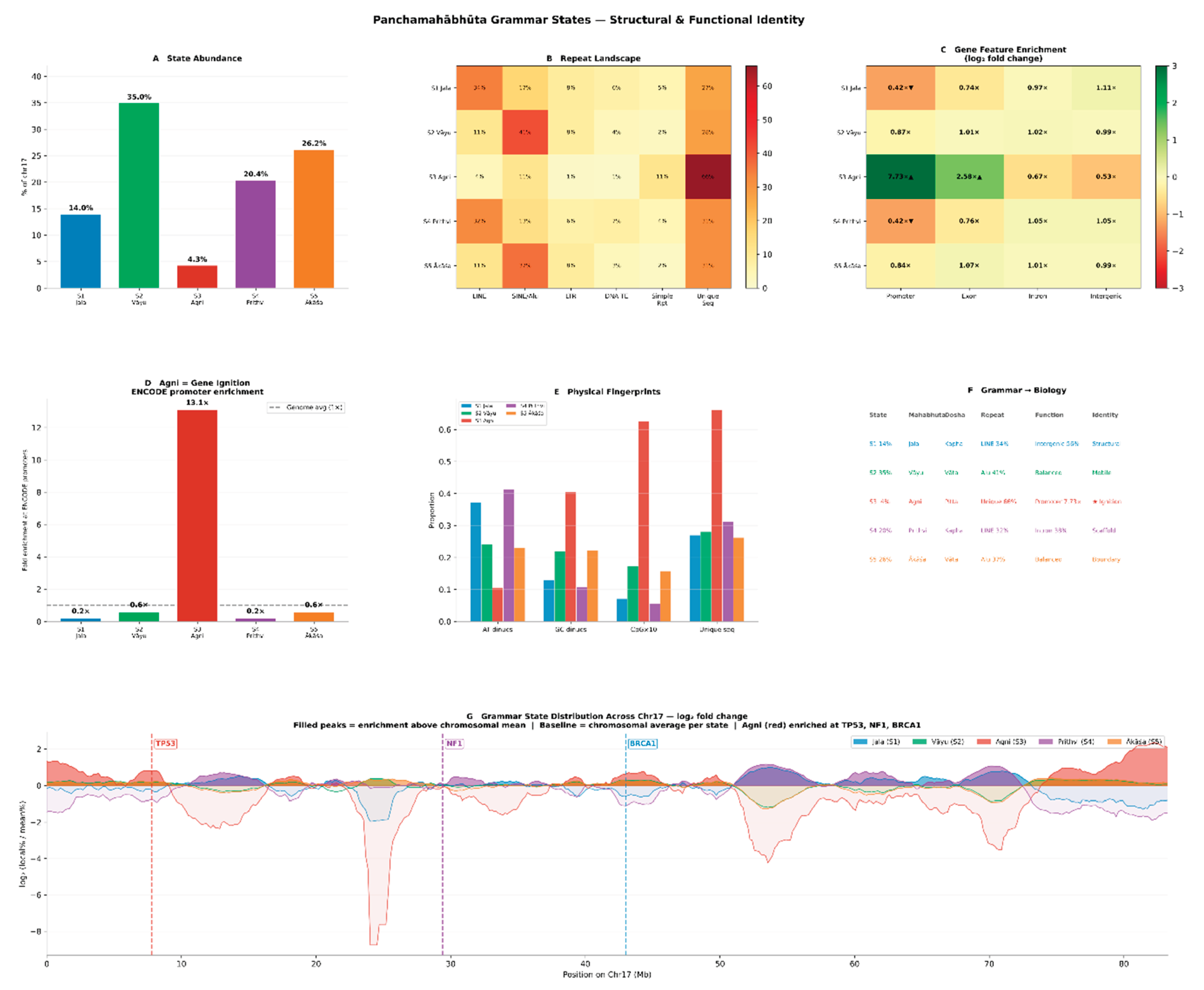
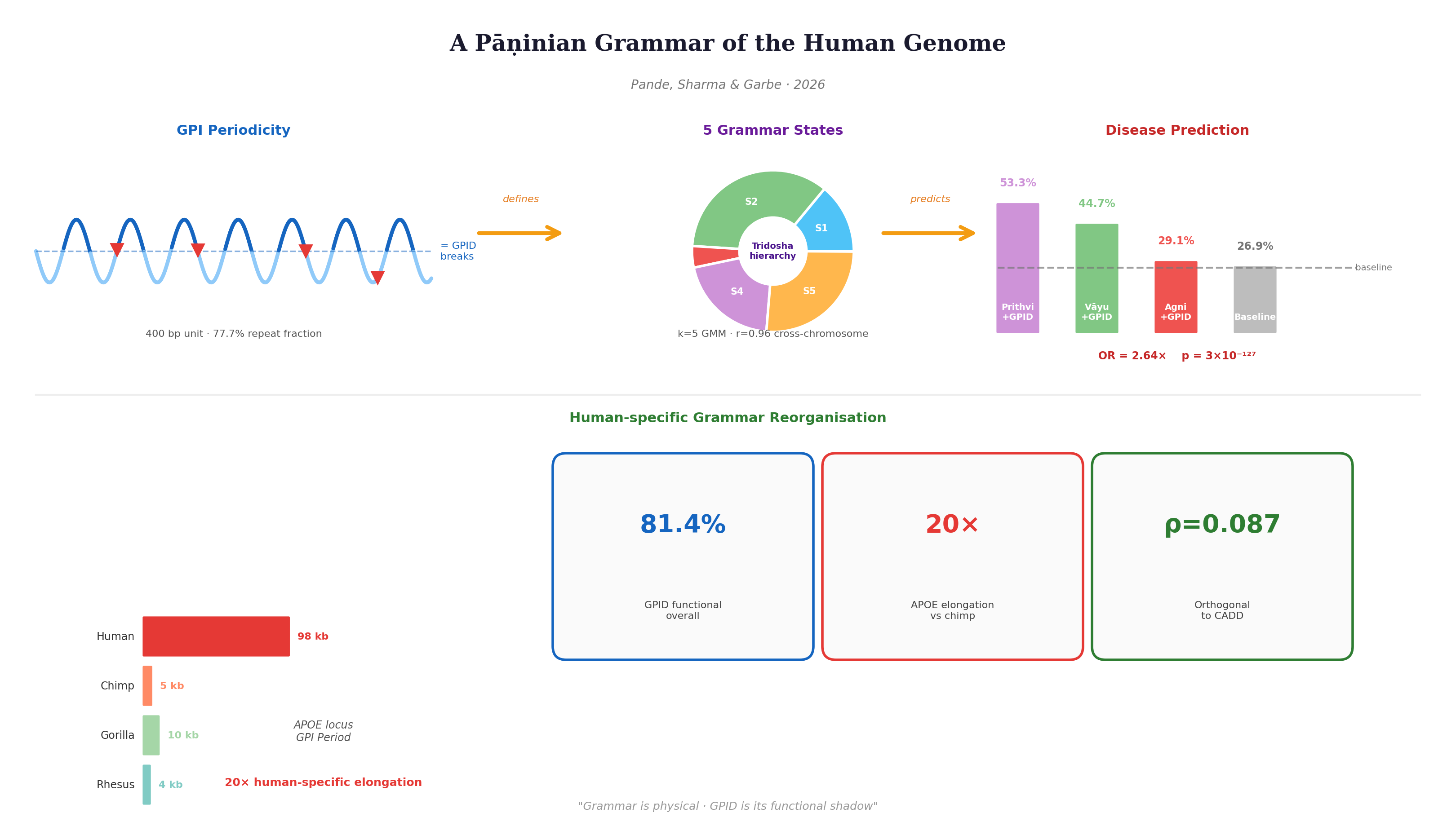
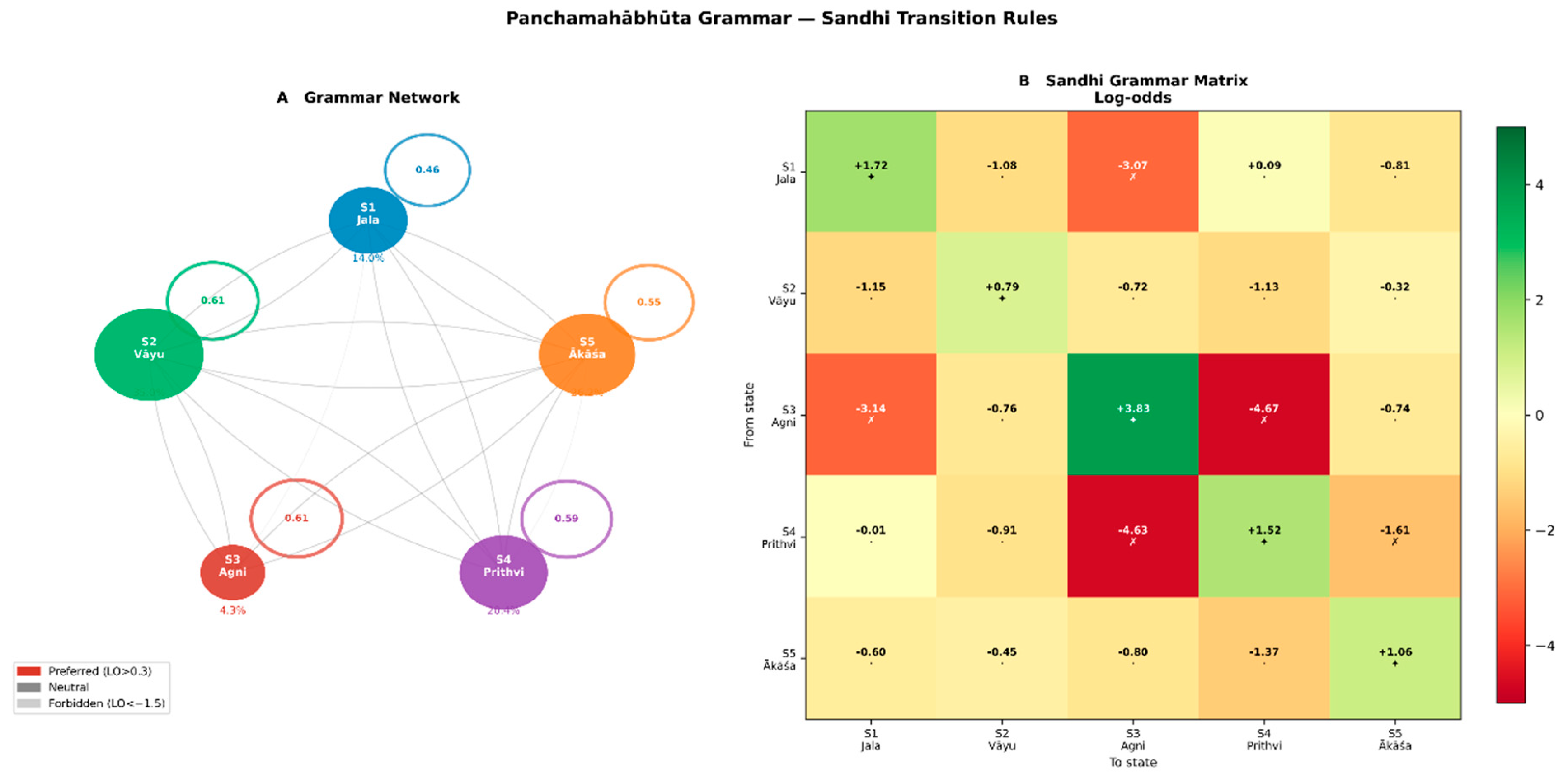
3.3. Sandhi Transition Rules Define a Formal Positional Grammar
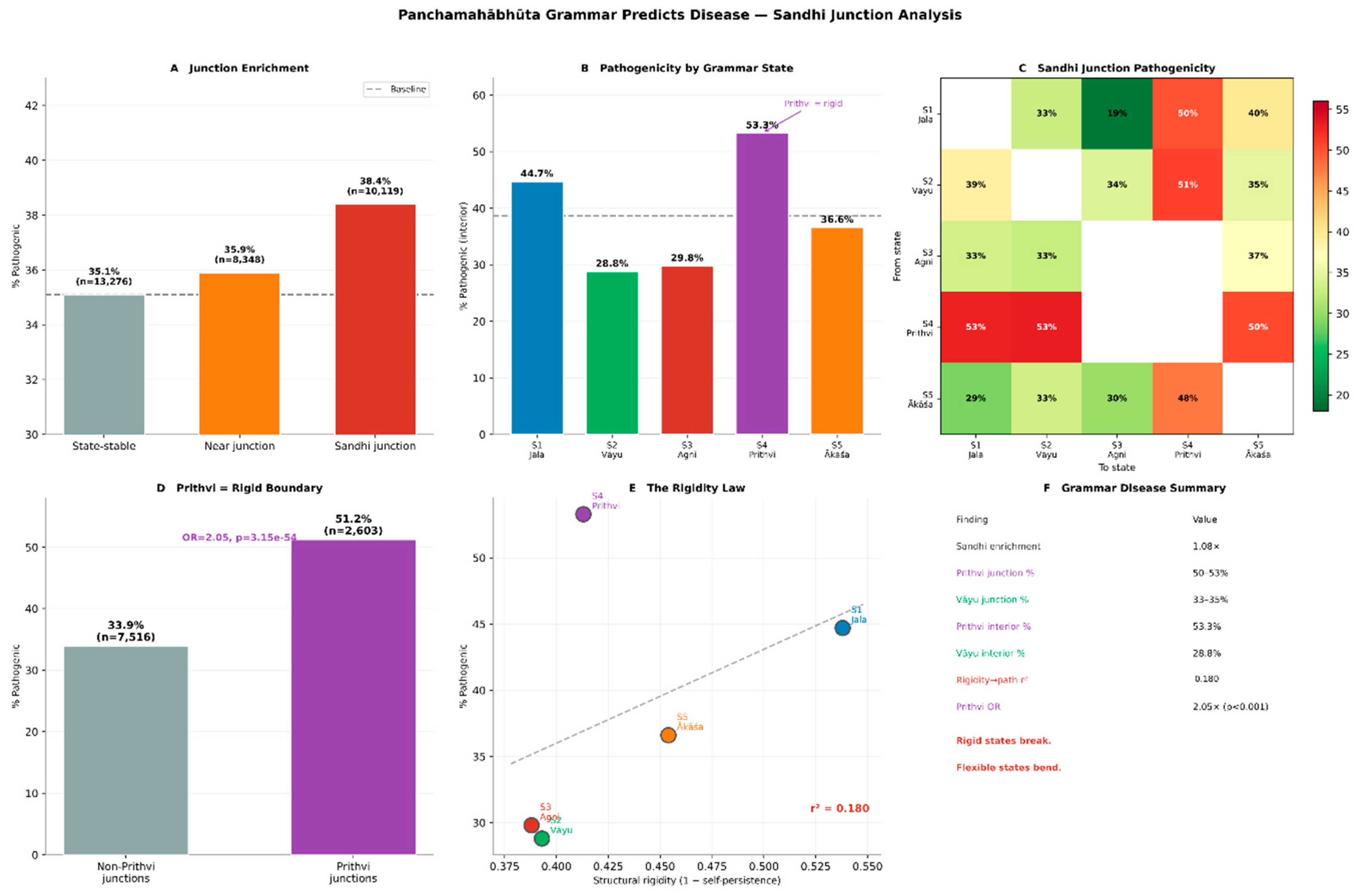
3.4. Grammatically Constrained Positions Are Enriched for Pathogenic Variants
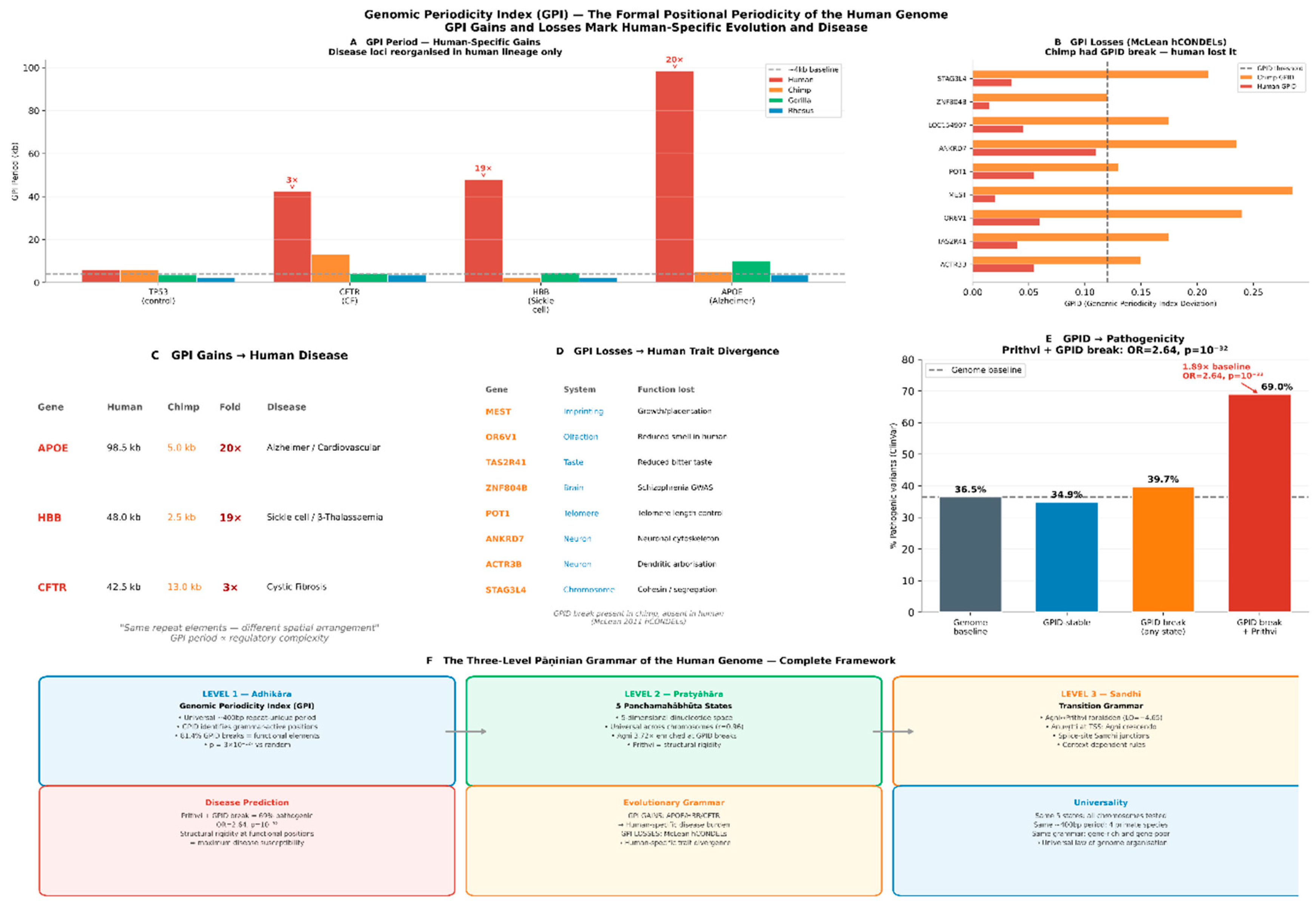
3.5. Human-Specific GPI Reorganization Marks Evolutionary Innovation at Disease and Trait-Divergence Loci
3.6. Grammar Predicts cis-Regulatory Element Identity
3.7. Tridosha Segmentation of the Genome and the Prakriti of the Regulome
3.8. Evolutionary Conservation Amplifies the Positional Constraint Signal
3.9. Genome-Wide Grammar Validation: Tridosha Biochemical Hierarchy and Disease Associations
Discussion
The GPI as a Dissipative Structure
The Metric (Anuṣṭubh Chandas) and the Scale of Genomic Grammar
The Panchamahābhūta States and Biological Function
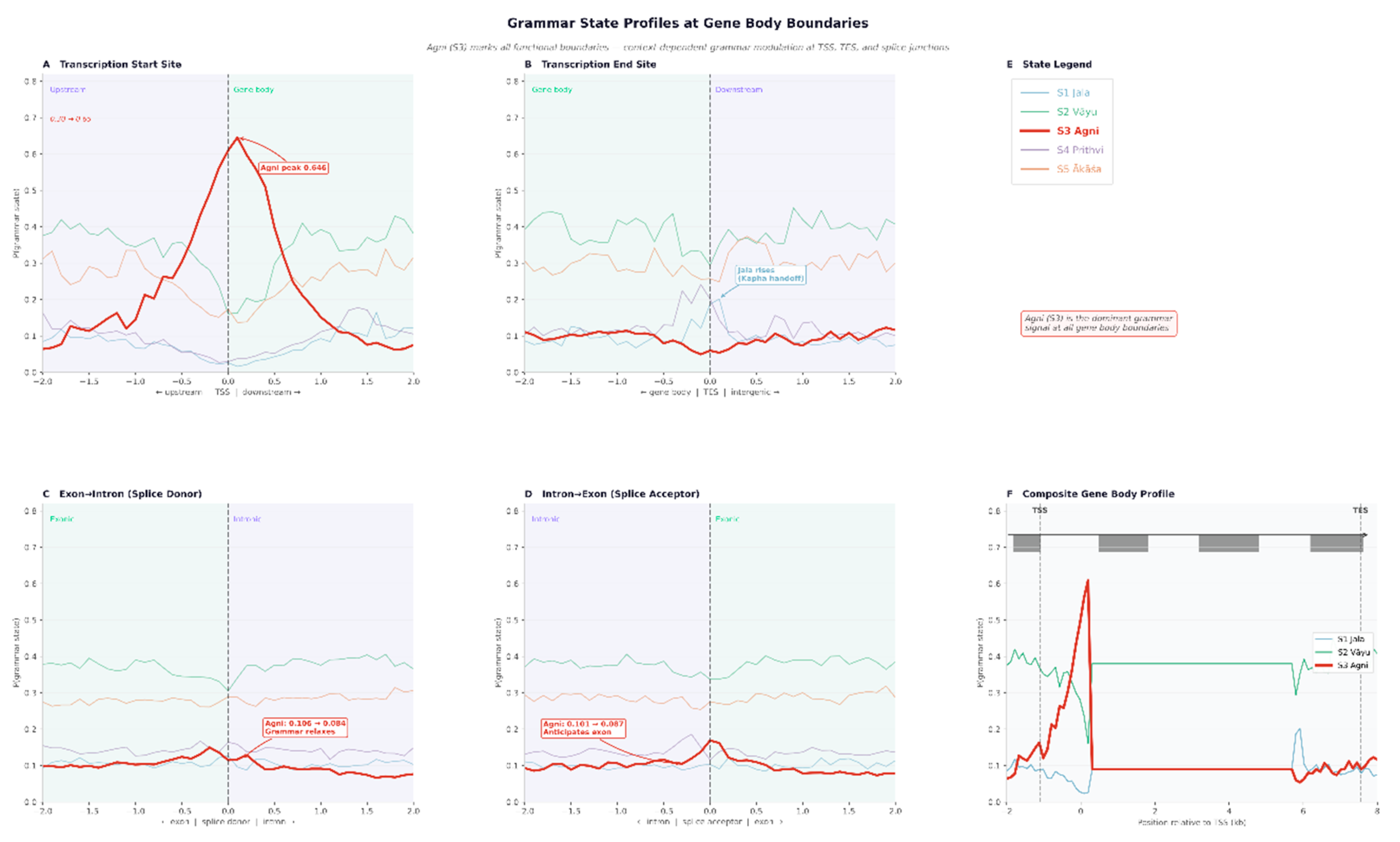
Sandhi Rules and Splice Site Recognition
Human-Specific Grammar Reorganization and Disease
Limitations and Future Directions
Conclusions
Methods
Genomic Periodicity Index (GPI) Computation
Functional Annotation of GPID Breaks
Panchamahābhūta Grammar State Inference
Sandhi Transition Matrix
Metagene Grammar Profiles
ClinVar Variant Analysis
Non-B DNA Structure Detection
Primate GPI Period Comparison
McLean hCONDEL Analysis
Tridosha Segmentation
Human-Specific Transposable Element Analysis
Software and Data Availability
Supplementary Materials
References
- Sharma; Pāṇini, R.N.; Ashtadhyayi. 1987.
- Kadvany, J. Pāṇini’s Grammar and Modern Computation. Hist. Philos. Log. 2016, 37, 325–346. [Google Scholar] [CrossRef]
- Lowe, J. J. Modern Linguistics in Ancient India; Cambridge University Press, 2024. [Google Scholar] [CrossRef]
- Ramkrishnamacharyulu, K. V. Annotating Sanskrit Texts Based on Śābdabodha Systems. In Sanskrit Computational Linguistics; Kulkarni, A., Huet, G., Eds.; Springer Berlin Heidelberg, Berlin, Heidelberg, 2008; vol. 5406 26–39. [Google Scholar]
- Caraka Samhita; Sharma, P. V., Ed.; vols 1, Sharira Sthana 6/4.
- Cardona, G. Some principles of Pānini’s grammargrammar. J. Indian Philos. 1970, 1, 40–74. [Google Scholar] [CrossRef]
- Huet, G.; Kulkarni, A.; Scharf, P. M. Sanskrit Computational Linguistics: First and Second International Symposia Rocquencourt, France, October 29-31, 2007 Providence, RI, USA, May 15-17, 2008 Revised Selected and Invited Papers; Springer Berlin Heidelberg Springer e-books, Berlin, Heidelberg, 2009. [Google Scholar]
- Prasher, B.; et al. Whole genome expression and biochemical correlates of extreme constitutional types defined in Ayurveda. J. Transl. Med. 2008, 6, 48. [Google Scholar] [CrossRef]
- Mukerji, M. Ayurgenomics-based frameworks in precision and integrative medicine: Translational opportunities. Camb. Prisms Precis. Med. 2023, 1, e29. [Google Scholar] [CrossRef]
- Govindaraj, P.; et al. Genome-wide analysis correlates Ayurveda Prakriti. Sci. Rep. 2015, 5, 15786. [Google Scholar] [CrossRef]
- Knudsen, B.; Hein, J. RNA secondary structure prediction using stochastic context-free grammars and evolutionary history. Bioinformatics 1999, 15, 446–454. [Google Scholar] [CrossRef]
- Eddy, S. R. Profile hidden Markov models. Bioinformatics 1998, 14, 755–763. [Google Scholar] [CrossRef]
- Prigogine; Stengers. Order Out of Chaos; 1984. [Google Scholar]
- Smit; Hubley; Green. RepeatMasker. 2013. [Google Scholar]
- et al.; The ENCODE Project Consortium Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 2020, 583, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Lyon, M. F. Gene Action in the X-chromosome of the Mouse (Mus musculus L.). Nature 1961, 190, 372–373. [Google Scholar] [CrossRef] [PubMed]
- Korenberg, J. R.; Rykowski, M. C. Human genome organization: Alu, LINES, and the molecular structure of metaphase chromosome bands. Cell 1988, 53, 391–400. [Google Scholar] [CrossRef]
- Khurana, S.; et al. A Macrohistone Variant Links Dynamic Chromatin Compaction to BRCA1-Dependent Genome Maintenance. Cell Rep. 2014, 8, 1049–1062. [Google Scholar] [CrossRef]
- et al.; Roadmap Epigenomics Consortium Integrative analysis of 111 reference human epigenomes. Nature 2015, 518, 317–330. [Google Scholar] [CrossRef]
- Thurman, R. E.; et al. The accessible chromatin landscape of the human genome. Nature 2012, 489, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; et al. Genome-wide Chromatin State Transitions Associated with Developmental and Environmental Cues. Cell 2013, 152, 642–654. [Google Scholar] [CrossRef]
- Landrum, M. J.; et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018, 46, D1062–D1067. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Martin-Pintado, N.; Post, H.; Altelaar, M.; Knipscheer, P. Multistep mechanism of G-quadruplex resolution during DNA replication. Sci. Adv. 2021, 7, eabf8653. [Google Scholar] [CrossRef] [PubMed]
- Horton, C. A.; et al. Short tandem repeats bind transcription factors to tune eukaryotic gene expression. Science 2023, 381, eadd1250. [Google Scholar] [CrossRef]
- Noll, A; et al. GPAC—Genome Presence/Absence Compiler. Mol Biol Evol 2015, 32, 275–286. [Google Scholar] [CrossRef]
- Chen, S.; et al. A genomic mutational constraint map using variation in 76,156 human genomes. Nature 2024, 625, 92–100. [Google Scholar] [CrossRef]
- Smit, A.F.A.; Hubley, R.; Green, P. RepeatMasker Open-4.0 2013–2015. Available online: http://www.repeatmasker.org.
- Kent, W.J.; et al. The human genome browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef] [PubMed]
- The ENCODE Project Consortium. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 2020, 583, 699–710. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018, 46, D1062–D1067. [Google Scholar] [CrossRef] [PubMed]
- McLean, C.Y.; et al. Human-specific loss of regulatory DNA and the evolution of human-specific traits. Science 2011, 334, 1573–1577. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



