Submitted:
08 April 2026
Posted:
09 April 2026
You are already at the latest version
Abstract
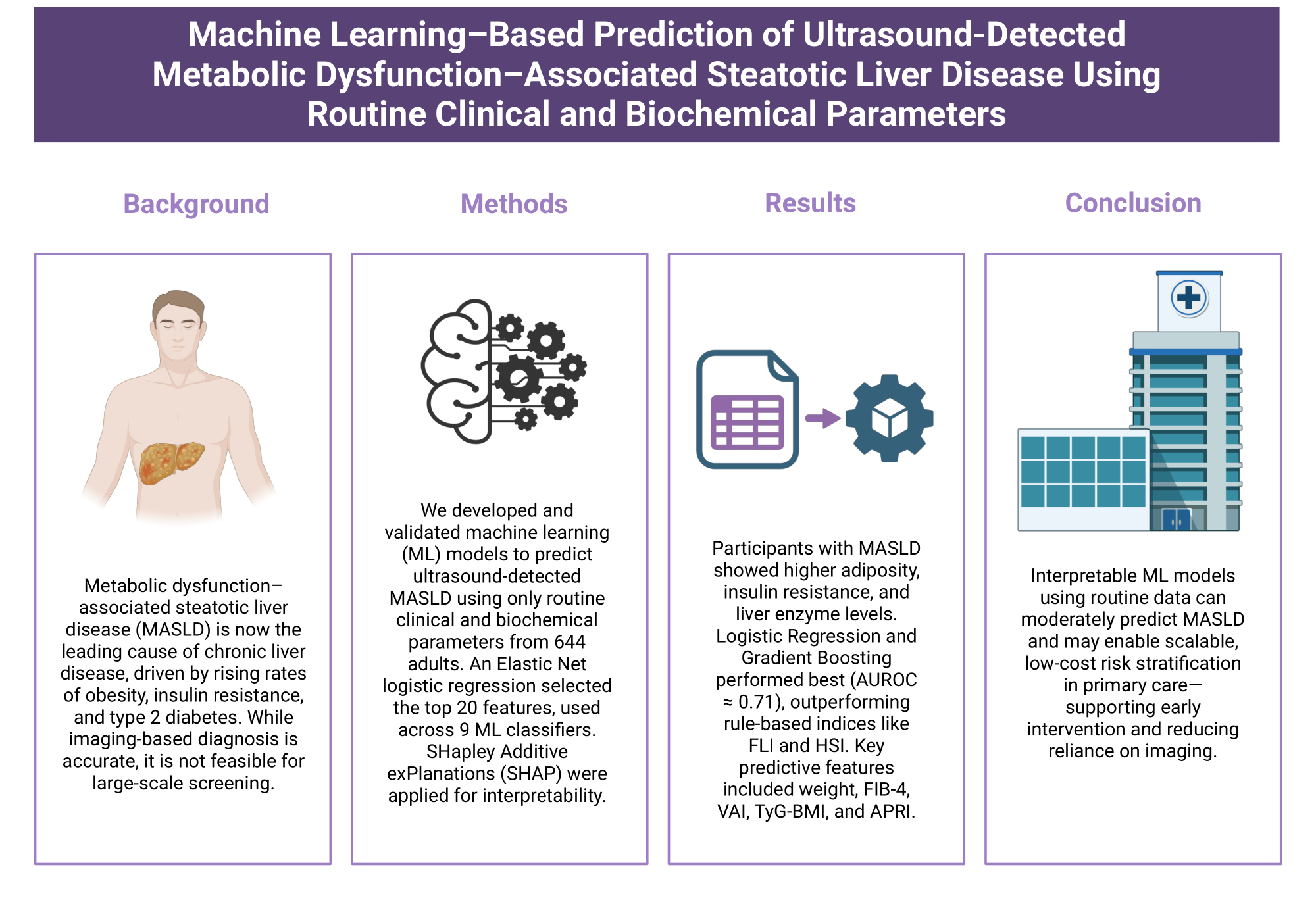
Keywords:
1. Introduction
2. Methods
2.1. Dataset and Ultrasonography Outcome Definition
2.2. Study Design and Modeling
2.3. Preprocessing and Feature Selection (Single Pipeline)
2.4. Data Splitting and Resampling
2.5. Classifiers
2.6. Evaluation Metrics and Interpretability
2.7. Comparative Evaluation of Simple Steatosis Scores and ML Models
3. Results
3.1. Baseline Characteristics
3.2. Model Development and Evaluation
3.3. Elastic Net–Based Variable Selection
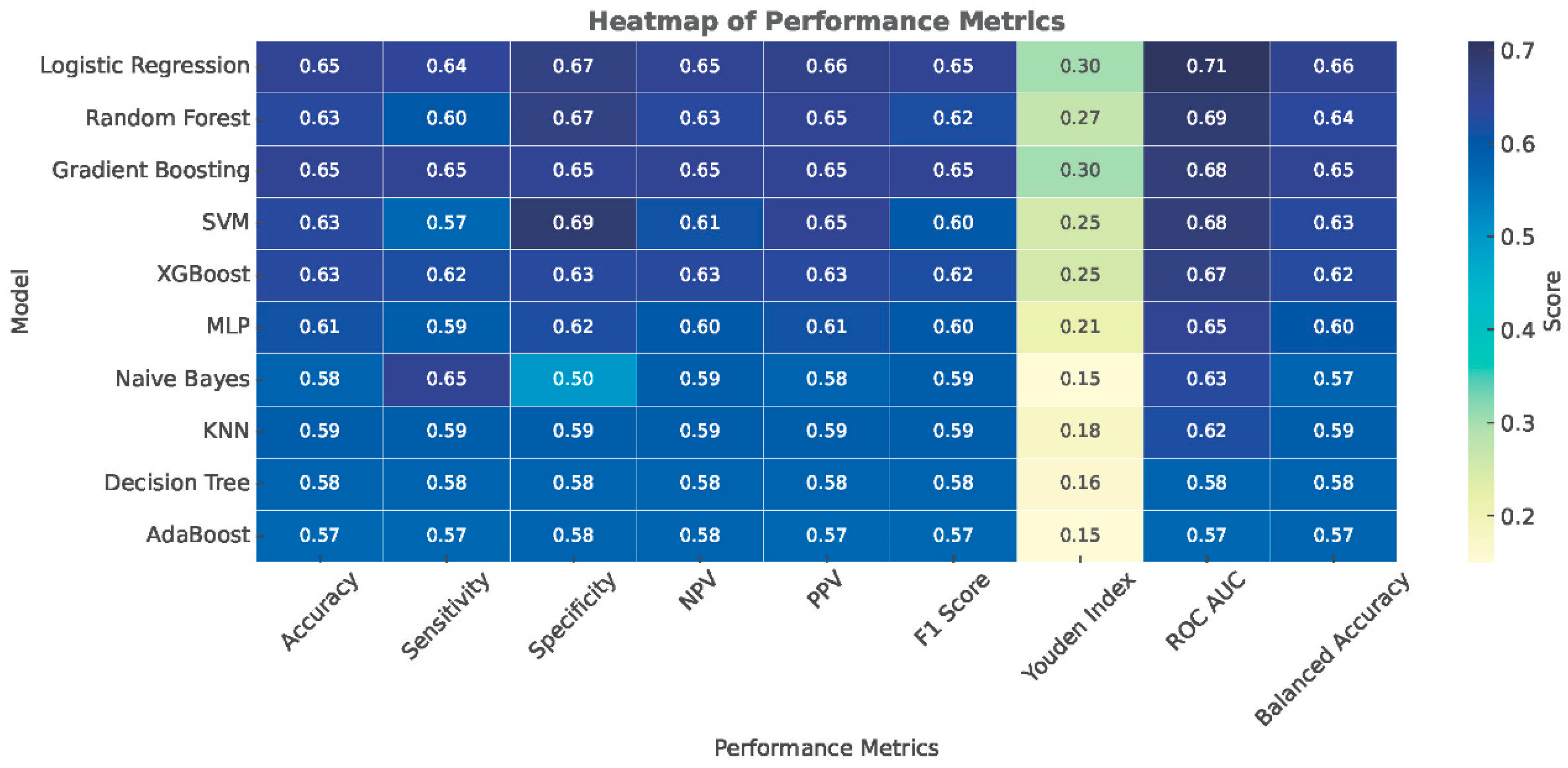
3.4. Accuracy
3.5. F1 Score
3.6. Sensitivity
3.7. Specificity
3.8. Youden’s J
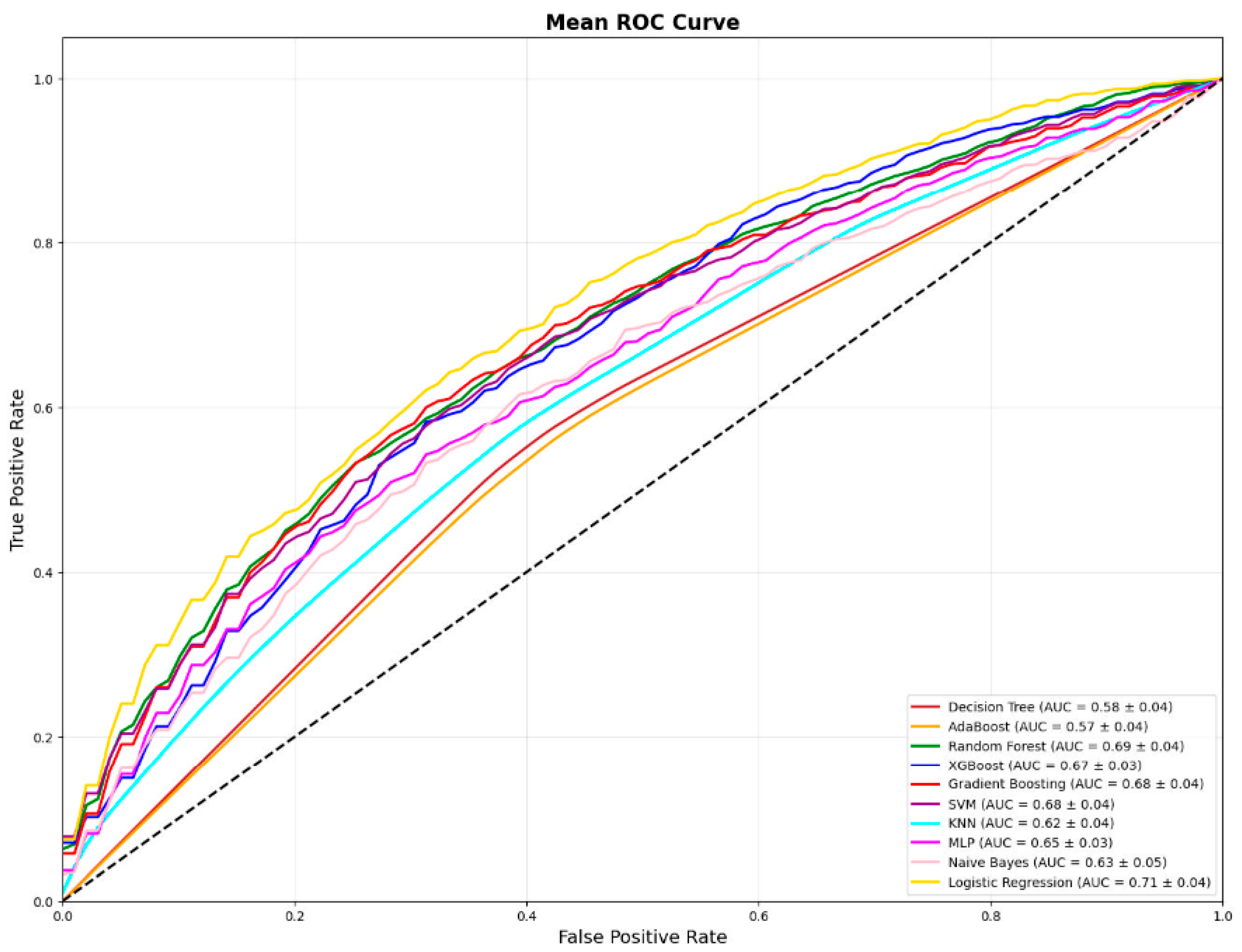
3.9. AUROC
3.10. Balanced Accuracy
3.11. ROC Visualization
3.12. PPV/NPV
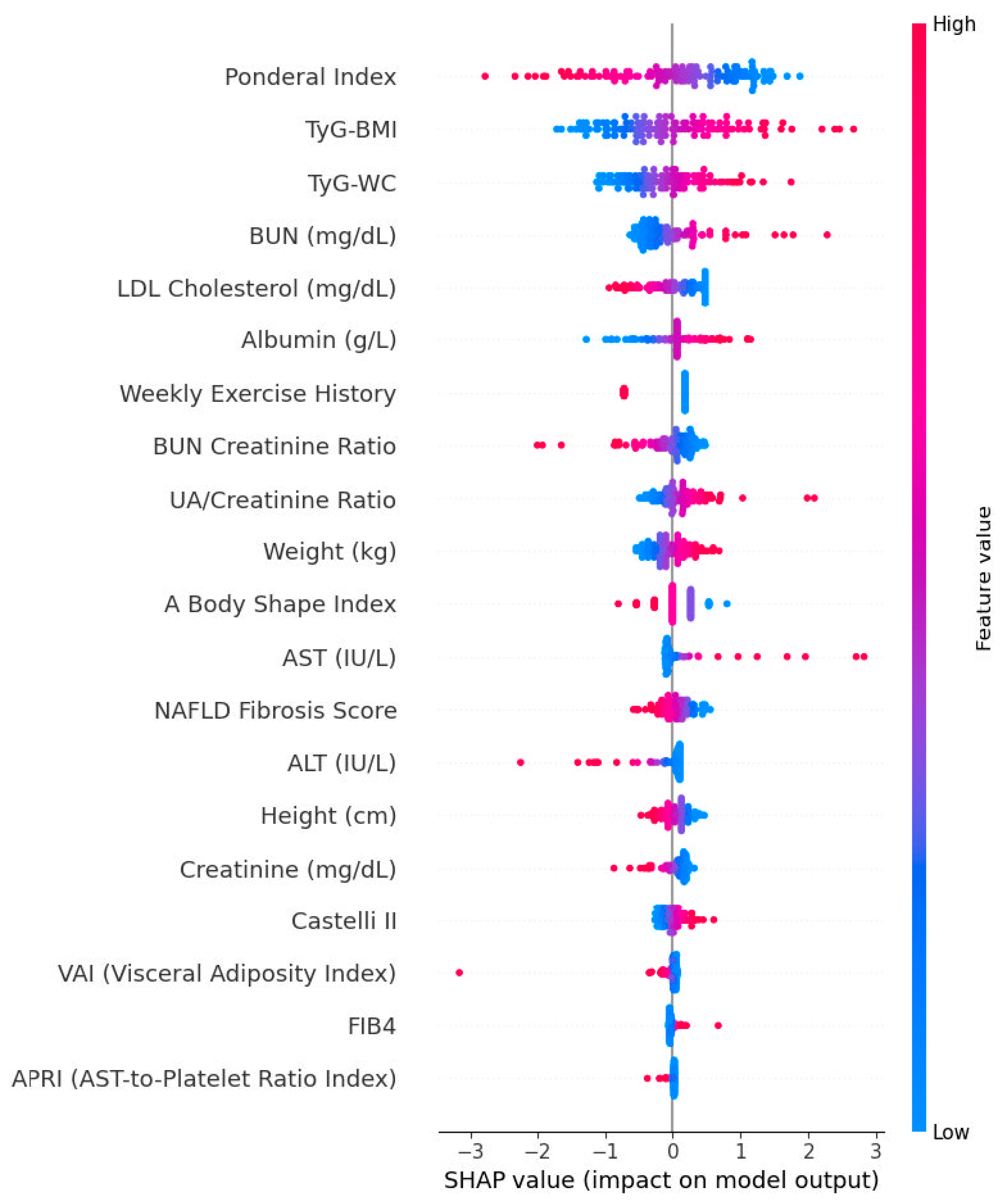
3.13. SHAP Summaries and Directionality
3.14. Comparison Between ML Models and Simple Clinic Scores for Ultrasound-Detected Hepatic Steatosis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Chan, W., et al., Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD): A State-of-the-Art Review. JOURNAL OF OBESITY & METABOLIC SYNDROME, 2023. 32(3): p. 197-213.
- Li, Y., et al., Updated mechanisms of MASLD pathogenesis. LIPIDS IN HEALTH AND DISEASE, 2024. 23(1). [CrossRef]
- Hong, S., et al., From NAFLD to MASLD: When metabolic comorbidity matters. ANNALS OF HEPATOLOGY, 2024. 29(2). [CrossRef]
- Zazueta, A., et al., Alteration of Gut Microbiota Composition in the Progression of Liver Damage in Patients with Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD). INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES, 2024. 25(8). [CrossRef]
- Fan, J., et al., Guideline for the Prevention and Treatment of Metabolic Dysfunction-associated Fatty Liver Disease (Version 2024). JOURNAL OF CLINICAL AND TRANSLATIONAL HEPATOLOGY, 2024. 12(11): p. 955-974. [CrossRef]
- Lu, H., et al., Identification of hub gene for the pathogenic mechanism and diagnosis of MASLD by enhanced bioinformatics analysis and machine learning. PLOS ONE, 2025. 20(5). [CrossRef]
- DiBattista, J., et al., Accuracy of Non-invasive Indices for Diagnosing Hepatic Steatosis Compared to Imaging in a Real-World Cohort. DIGESTIVE DISEASES AND SCIENCES, 2022. 67(11): p. 5300-5308. [CrossRef] [PubMed]
- Wu, J., et al., Population-specific cut-off points of fatty liver index for the diagnosis of hepatic steatosis. JOURNAL OF HEPATOLOGY, 2021. 75(3): p. 726-728. [CrossRef]
- Su, P., et al., Comparison of Machine Learning Models and the Fatty Liver Index in Predicting Lean Fatty Liver. DIAGNOSTICS, 2023. 13(8). [CrossRef] [PubMed]
- Frey, L., et al., Use of machine learning for early prediction of short-term mortality in veterans with metabolic dysfunction-associated steatotic liver disease. PLOS ONE, 2025. 20(10). [CrossRef] [PubMed]
- Chen, H., et al., Development and validation of machine learning models for MASLD: based on multiple potential screening indicators. FRONTIERS IN ENDOCRINOLOGY, 2025. 15. [CrossRef]
- Soliman, R., A. Helmy, and G. Shiha, Precision in Diagnosis of Liver Fibrosis in MASLD: Machine Learning-Based Scores May Be More Accurate Than Conventional NITs. LIVER INTERNATIONAL, 2025. 45(4). [CrossRef]
- Weng, S., et al., Prediction of fatty liver disease in a Chinese population using machine-learning algorithms. Diagnostics, 2023. 13(6): p. 1168. [CrossRef] [PubMed]
- Marques, R., et al., Ultrasound versus elastography in the diagnosis of hepatic steatosis: evaluation of traditional machine learning versus deep learning. Sensors, 2024. 24(23): p. 7568. [CrossRef]
- Mahzari, A., Artificial intelligence in nonalcoholic fatty liver disease. Egyptian Liver Journal, 2022. 12(1): p. 69. [CrossRef]
- Kirik, A., et al., Prevalence of MASLD and fibrosis risk in Turkish adults with cardiometabolic risk factors: a nationwide multicenter study (DAHUDER MASLD study). Journal of Clinical Medicine, 2025. 14(19): p. 7098. [CrossRef]
- Bedogni, G., et al., The Fatty Liver Index: a simple and accurate predictor of hepatic steatosis in the general population. BMC GASTROENTEROLOGY, 2006. 6. [CrossRef]
- Lee, J., et al., Hepatic steatosis index: A simple screening tool reflecting nonalcoholic fatty liver disease. DIGESTIVE AND LIVER DISEASE, 2010. 42(7): p. 503-508. [CrossRef] [PubMed]
- Huang, D., H. El-Serag, and R. Loomba, Global epidemiology of NAFLD-related HCC: trends, predictions, risk factors and prevention. NATURE REVIEWS GASTROENTEROLOGY & HEPATOLOGY, 2021. 18(4): p. 223-238.
- Lazarus, J., et al., A global action agenda for turning the tide on fatty liver disease. HEPATOLOGY, 2024. 79(2): p. 502-523. [CrossRef]
- Crudele, L., et al., Fatty Liver Index (FLI) is the best score to predict MASLD with 50% lower cut-off value in women than in men. Biology of sex Differences, 2024. 15(1): p. 43. [CrossRef]
- McTeer, M., et al., Machine learning approaches to enhance diagnosis and staging of patients with MASLD using routinely available clinical information. Plos one, 2024. 19(2): p. e0299487. [CrossRef]
- Cubillos, G., et al., Development of a novel deep learning method that transforms tabular input variables into images for the prediction of SLD. SCIENTIFIC REPORTS, 2025. 15(1). [CrossRef]
- Lim, D., et al., Use of Machine Learning to Predict Onset of NAFLD in an All-Comers Cohort-Development and Validation in 2 Large Asian Cohorts. GASTRO HEP ADVANCES, 2024. 3(7): p. 1005-1011. [CrossRef] [PubMed]
- Fabbrini, E., S. Sullivan, and S. Klein, Obesity and Nonalcoholic Fatty Liver Disease: Biochemical, Metabolic, and Clinical Implications. HEPATOLOGY, 2010. 51(2): p. 679-689. [CrossRef]
- Demirci, S. and S. Sezer, Fatty Liver Index vs. Biochemical-Anthropometric Indices: Diagnosing Metabolic Dysfunction-Associated Steatotic Liver Disease with Non-Invasive Tools. DIAGNOSTICS, 2025. 15(5). [CrossRef]
- Qian, X., et al., Value of triglyceride glucose-body mass index in predicting nonalcoholic fatty liver disease in individuals with type 2 diabetes mellitus. FRONTIERS IN ENDOCRINOLOGY, 2025. 15. [CrossRef]
- Sheng, G., et al., The usefulness of obesity and lipid-related indices to predict the presence of Non-alcoholic fatty liver disease. LIPIDS IN HEALTH AND DISEASE, 2021. 20(1). [CrossRef] [PubMed]
- Xuan, Y., et al., Elevated ALT/AST ratio as a marker for NAFLD risk and severity: insights from a cross-sectional analysis in the United States. FRONTIERS IN ENDOCRINOLOGY, 2024. 15. [CrossRef] [PubMed]
- Rigor, J., et al., Noninvasive fibrosis tools in NAFLD: validation of APRI, BARD, FIB-4, NAFLD fibrosis score, and Hepamet fibrosis score in a Portuguese population. POSTGRADUATE MEDICINE, 2022. 134(4): p. 435-440. [CrossRef]
- Ouzan, D., et al., Using the FIB-4, automatically calculated, followed by the ELF test in second line to screen primary care patients for liver disease. SCIENTIFIC REPORTS, 2024. 14(1). [CrossRef]
- Yang, C., et al., A Bidirectional Relationship Between Hyperuricemia and Metabolic Dysfunction-Associated Fatty Liver Disease. FRONTIERS IN ENDOCRINOLOGY, 2022. 13.
- Francoz, C., et al., Hepatorenal Syndrome. CLINICAL JOURNAL OF THE AMERICAN SOCIETY OF NEPHROLOGY, 2019. 14(5): p. 774-781. [CrossRef]
- Bucurica, S., et al., Exploring the Relationship between Lipid Profile, Inflammatory State and 25-OH Vitamin D Serum Levels in Hospitalized Patients. BIOMEDICINES, 2024. 12(8). [CrossRef]
- Belalcazar, S., et al., CONVENTIONAL BIOMARKERS FOR CARDIOVASCULAR RISKS AND THEIR CORRELATION WITH THE CASTELLI RISK INDEX-INDICES AND TG/HDL-C. ARCHIVOS DE MEDICINA, 2020. 20(1): p. 11-22.
- Xiao, L., et al., Development and Validation of Machine Learning-Based Marker for Early Detection and Prognosis Stratification of Nonalcoholic Fatty Liver Disease. ADVANCED SCIENCE, 2025. 12(33). [CrossRef]
- Verschuren, L., et al., Development of a novel non-invasive biomarker panel for hepatic fibrosis in MASLD. NATURE COMMUNICATIONS, 2024. 15(1). [CrossRef] [PubMed]
- Yu, Y., et al., Predicting metabolic dysfunction associated steatotic liver disease using explainable machine learning methods. SCIENTIFIC REPORTS, 2025. 15(1). [CrossRef] [PubMed]
- Koliaki, C., et al., Metabolically Healthy Obesity and Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD): Navigating the Controversies in Disease Development and Progression. CURRENT OBESITY REPORTS, 2025. 14(1). [CrossRef]
- Weng, S., et al., Prediction of Fatty Liver Disease in a Chinese Population Using Machine-Learning Algorithms. DIAGNOSTICS, 2023. 13(6). [CrossRef]
- Collins, G., et al., TRIPOD plus AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ-BRITISH MEDICAL JOURNAL, 2024. 385.
| Features | Normal USG | Hepatic steatosis on USG | p-value |
| n (%) | 322 (50) | 322 (50) | - |
| Demographic and Anthropometric Characteristics | |||
| Age | 62.36 ± 19.12 [60.26 – 64.46] | 59.81 ± 15.95 [58.06 – 61.56] | 0.004* |
| Sex (Female) | 179 (55.59) | 170 (52.80) | 0.477† |
| Educational Status | 57 (17.70) | 57 (17.70) | 1.000† |
| Smoking | 131 (40.68) | 144 (44.72) | 0.300† |
| Height (cm) | 164.19 ± 10.18 [163.08 – 165.31] | 165.91 ± 9.98 [164.82 – 167.01] | 0.043* |
| Weight (kg) | 70.43 ± 14.58 [68.83 – 72.03] | 79.82 ± 18.24 [77.82 – 81.82] | <0.001* |
| WC (cm) | 87.71 ± 17.17 [85.83 – 89.59] | 95.65 ± 18.24 [93.65 – 97.65] | <0.001* |
| BMI | 26.24 ± 5.77 [25.61 – 26.87] | 29.02 ± 6.57 [28.30 – 29.74] | <0.001* |
| Weekly Exercise History | 71 (22.05) | 44 (13.66) | 0.005† |
| Hemodynamic Parameters | |||
| SBP (mmHg) | 124.29 ± 16.14 [122.52 – 126.06] | 126.64 ± 17.28 [124.75 – 128.54] | 0.026* |
| DBP (mmHg) | 72.63 ± 9.73 [71.56 – 73.69] | 75.15 ± 11.10 [73.93 – 76.37] | <0.001* |
| HR (beats/min) | 80.35 ± 12.87 [78.94 – 81.76] | 82.67 ± 13.16 [81.23 – 84.11] | 0.031* |
| Clinical Comorbidities and Medication Use | |||
| DM | 116 (36.02) | 165 (51.24) | <0.001† |
| HTN | 157 (48.76) | 175 (54.35) | 0.156† |
| DLP | 59 (18.32) | 103 (31.99) | <0.001† |
| ASCVD | 73 (22.67) | 82 (25.47) | 0.407† |
| CVD | 14 (4.35) | 17 (5.28) | 0.581† |
| PCOS (in females) | 1 (0.56) | 2 (1.18) | 0.563† |
| OSA Syndrome | 0 (0) | 5 (1.55) | 0.025† |
| Metformin Use | 40 (12.42) | 83 (25.78) | <0.001† |
| Pioglitazone Use | 3 (0.93) | 4 (1.24) | 0.704† |
| SGLT-2i Use | 26 (8.07) | 45 (13.98) | 0.017† |
| Statin Use | 39 (12.11) | 65 (20.19) | 0.005† |
| Hematologic Parameters | |||
| Hb (g/dL) | 11.12 ± 2.56 [10.84 – 11.40] | 11.81 ± 2.61 [11.52 – 12.09] | <0.001‡ |
| WBC (10³/µL) | 8.41 ± 3.6 [8.01 – 8.80] | 8.82 ± 3.52 [8.43 – 9.20] | 0.110* |
| Lymphocyte Count (10³/µL) | 2.16 ± 3.04 [1.83 – 2.50] | 2.10 ± 1.52 [1.93 – 2.27] | 0.004* |
| Neutrophil Count (10³/µL) | 5.83 ± 3.34 [5.46 – 6.19] | 5.96 ± 3.35 [5.59 – 6.33] | 0.630* |
| Monocyte Count (10³/µL) | 0.74 ± 0.78 [0.66 – 0.83] | 0.79 ± 0.84 [0.69 – 0.88] | 0.339* |
| Platelet Count (10³/µL) | 253.55 ± 113.88 [241.07 – 266.04] | 262.55 ± 103.84 [251.16 – 273.93] | 0.290* |
| Biochemical Parameters | |||
| Fasting Plasma Glucose (mg/dL) | 126.06 ± 63.27 [119.12 – 133] | 144.47 ± 81.20 [135.57 – 153.37] | <0.001* |
| BUN (mg/dL) | 28.06 ± 23.42 [25.49 – 30.63] | 27.79 ± 23.97 [25.16 – 30.42] | 0.710* |
| Cr (mg/dL) | 1.28 ± 1.08 [1.16 – 1.40] | 1.19 ± 0.80 [1.10 – 1.28] | 0.658* |
| eGFR (mL/min/1.73 m²) | 70.74 ± 34.64 [66.94 – 74.54] | 73.12 ± 33.29 [69.47 – 76.77] | 0.287* |
| Total Cholesterol (mg/dL) | 152.09 ± 52.52 [146.34 – 157.85] | 159.67 ± 54.58 [153.69 – 165.66] | 0.039* |
| LDL-C (mg/dL) | 91.07 ± 39.71 [86.72 – 95.42] | 94.37 ± 40.61 [89.92 – 98.82] | 0.271* |
| HDL-C (mg/dL) | 38.64 ± 14.27 [37.07 – 40.20] | 37.32 ± 13.71 [35.81 – 38.82] | 0.202* |
| TG (mg/dL) | 134.36 ± 153.57 [117.53 – 151.20] | 170.91 ± 116.66 [158.12 – 183.70] | <0.001* |
| AST (U/L) | 38.76 ± 87.87 [29.13 – 48.40] | 43.94 ± 93.24 [33.71 – 54.16] | 0.198* |
| ALT (U/L) | 40.60 ± 98.78 [29.77 – 51.43] | 39.87 ± 78.22 [31.30 – 48.45] | <0.001* |
| GGT (U/L) | 66.91 ± 106.53 [55.23 – 78.59] | 88.30 ± 175.22 [69.09 – 107.51] | 0.167* |
| HbA1c (%) | 6.51 ± 2.34 [6.26 – 6.77] | 7.11 ± 2.78 [6.81 – 7.41] | <0.001* |
| Albumin (g/L) | 36.03 ± 5.05 [35.47 – 36.58] | 37.69 ± 4.56 [37.19 – 38.19] | <0.001* |
| Direct Bilirubin (mg/dL) | 0.30 ± 0.64 [0.23 – 0.37] | 0.31 ± 0.56 [0.25 – 0.37] | 0.714* |
| Indirect Bilirubin (mg/dL) | 0.35 ± 0.27 [0.32 – 0.38] | 0.40 ± 0.45 [0.35 – 0.45] | 0.488* |
| TSH (mIU/L) | 2.02 ± 3.10 [1.68 – 2.36] | 2.65 ± 6.72 [1.91 – 3.38] | 0.120* |
| FT4 (ng/dL) | 1.26 ± 0.27 [1.23 – 1.29] | 1.24 ± 0.23 [1.21 – 1.26] | 0.490* |
| UA (mg/dL) | 5.53 ± 2.06 [5.30 – 5.75] | 5.75 ± 1.79 [5.55 – 5.94] | 0.096* |
| Ferritin (μg/L) | 221.65 ± 251.39 [194.09 – 249.21] | 241.63 ± 305.89 [208.10 – 275.17] | 0.227* |
| Vitamin B12 (ng/L) | 448.49 ± 294.90 [416.16 – 480.82] | 445.07 ± 269.07 [415.57 – 474.57] | 0.199* |
| ALP (U/L) | 111.11 ± 96.62 [100.51 – 121.70] | 107.58 ± 92.81 [97.41 – 117.76] | 0.561* |
| Features | Normal USG | Hepatic steatosis on USG | p-value |
| n (%) | 322 (50) | 322 (50) | - |
| Body Composition Indices | |||
| WtHR | 0.54 ± 0.11 [0.52 – 0.55] | 0.58 ± 0.11 [0.57 – 0.59] | <0.001* |
| ABSI | 0.08 ± 0.01 [0.08 – 0.08] | 0.08 ± 0.01 [0.08 – 0.08] | 0.074* |
| Body Fat Percentage | 35.63 ± 10.98 [34.43 – 36.84] | 38.08 ± 11.74 [36.80 – 39.37] | 0.006‡ |
| PI | 16.13 ± 4.43 [15.65 – 16.62] | 17.59 ± 4.39 [17.11 – 18.07] | <0.001* |
| CI | 1.23 ± 0.17 [1.21 – 1.25] | 1.27 ± 0.16 [1.25 – 1.28] | 0.004‡ |
| RFM | 31.84 ± 10.38 [30.70 – 32.97] | 34.40 ± 10.04 [33.30 – 35.50] | 0.002‡ |
| Metabolic Indices | |||
| TyG | 8.76 ± 0.75 [8.68 – 8.84] | 9.14 ± 0.81 [9.05 – 9.22] | <0.001‡ |
| TyG/HDL Ratio | 4.73 ± 11.37 [3.49 – 5.98] | 5.60 ± 5.85 [4.96 – 6.24] | <0.001* |
| AIP | 0.12 ± 0.34 [0.09 – 0.16] | 0.25 ± 0.33 [0.22 – 0.29] | <0.001‡ |
| LAP | 43.90 ± 84.95 [34.59 – 53.22] | 68.88 ± 62.30 [62.05 – 75.71] | <0.001* |
| VAI | 3.31 ± 7.41 [2.50 – 4.12] | 3.95 ± 4.20 [3.49 – 4.41] | <0.001* |
| TyG-BMI | 230.16 ± 55.79 [224.05 – 236.28] | 266.41 ± 71.26 [258.60 – 274.22] | <0.001* |
| TyG-WC | 769.58 ± 171.00 [750.83 – 788.33] | 876.31 ± 195.00 [854.93 – 897.69] | <0.001‡ |
| TyG-WHtR | 4.70 ± 1.08 [4.58 – 4.82] | 5.29 ± 1.20 [5.16 – 5.42] | <0.001* |
| Cardiovascular Indices | |||
| Castelli I | 4.41 ± 2.68 [4.12 – 4.70] | 4.70 ± 2.26 [4.46 – 4.95] | 0.004* |
| Castelli II | 2.55 ± 1.38 [2.40 – 2.70] | 2.75 ± 1.33 [2.60 – 2.89] | 0.047* |
| Non-HDL-C | 113.46 ± 49.47 [108.03 – 118.88] | 122.36 ± 52.14 [116.64 – 128.07] | 0.020* |
| RC | 22.39 ± 30.81 [19.01 – 25.77] | 27.99 ± 26.75 [25.06 – 30.92] | <0.001* |
| PP | 51.66 ± 12.98 [50.24 – 53.09] | 51.49 ± 14.05 [49.95 – 53.03] | 0.783* |
| RPP | 9977.79 ± 2025.54 [9755.71 – 10199.87] | 10463.94 ± 2157.03 [10227.45 – 10700.43] | <0.001* |
| Liver Indices | |||
| De Ritis Ratio | 1.25 ± 0.56 [1.19 – 1.32] | 1.19 ± 0.63 [1.12 – 1.26] | 0.006* |
| APRI | 0.77 ± 5.61 [0.16 – 1.39] | 0.87 ± 5.69 [0.25 – 1.50] | 0.893* |
| FIB-4 | 2.23 ± 5.71 [1.60 – 2.85] | 2.43 ± 8.33 [1.52 – 3.35] | 0.088* |
| HSI | 36.08 ± 8.07 [35.19 – 36.96] | 39.41 ± 7.95 [38.53 – 40.28] | <0.001‡ |
| NFS | -0.93 ± 1.99 [-1.15 – -0.71] | -0.88 ± 2.06 [-1.11 – -0.65] | 0.768‡ |
| ALBI | -2.41 ± 0.56 [-2.47 – -2.35] | -2.51 ± 0.59 [-2.57 – -2.44] | 0.003* |
| HALP | 43.02 ± 54.53 [37.04 – 49.00] | 44.43 ± 57.84 [38.09 – 50.77] | 0.024* |
| Immune/Hematologic Scores | |||
| NLR | 3.79 ± 2.90 [3.47 – 4.11] | 3.64 ± 3.09 [3.30 – 3.98] | 0.160* |
| PLR | 158.85 ± 96.20 [148.30 – 169.40] | 148.72 ± 79.59 [139.99 – 157.44] | 0.401* |
| MLR | 0.43 ± 0.27 [0.40 – 0.46] | 0.45 ± 0.53 [0.39 – 0.50] | 0.279* |
| SII | 985.99 ± 936.22 [883.35 – 1088.64] | 962.46 ± 1118.44 [839.83 – 1085.08] | 0.682* |
| SIRI | 2.80 ± 2.92 [2.48 – 3.12] | 2.94 ± 3.67 [2.54 – 3.35] | 0.644* |
| PNI | 46.84 ± 16.49 [45.04 – 48.65] | 48.19 ± 9.37 [47.16 – 49.22] | <0.001* |
| Renal Indices | |||
| BUN/Cr Ratio | 33.74 ± 71.53 [25.89 – 41.58] | 29.68 ± 54.69 [23.68 – 35.67] | 0.359* |
| UHR | 0.17 ± 0.12 [0.16 – 0.18] | 0.18 ± 0.10 [0.17 – 0.19] | 0.031* |
| UA/Cr Ratio | 6.50 ± 7.65 [5.66 – 7.34] | 6.48 ± 6.05 [5.81 – 7.14] | 0.055* |
| Model | Accuracy | Sensitivity | Specificity | NPV | PPV | F1 Score | Youden Index | ROC AUC |
| Decision Tree | 0.5823 ± 0.0413 95% CI: 0.5652–0.5994 |
0.5852 ± 0.0736 95% CI: 0.5549–0.6156 |
0.5795 ± 0.0417 95% CI: 0.5623–0.5968 |
0.5851 ± 0.0461 95% CI: 0.5661–0.6041 |
0.5806 ± 0.0381 95% CI: 0.5649–0.5964 |
0.5819 ± 0.0533 95% CI: 0.5599–0.6039 |
0.1648 ± 0.0826 95% CI: 0.1307–0.1989 |
0.5824 ± 0.0413 95% CI: 0.5653–0.5994 |
| AdaBoost | 0.5764 ± 0.0420 95% CI: 0.5591–0.5938 |
0.5735 ± 0.0674 95% CI: 0.5457–0.6013 |
0.5796 ± 0.0457 95% CI: 0.5607–0.5984 |
0.5777 ± 0.0463 95% CI: 0.5586–0.5968 |
0.5763 ± 0.0397 95% CI: 0.5599–0.5927 |
0.5740 ± 0.0501 95% CI: 0.5533–0.5946 |
0.1531 ± 0.0838 95% CI: 0.1185–0.1877 |
0.5765 ± 0.0419 95% CI: 0.5592–0.5938 |
| Random Forest | 0.6336 ± 0.0417 95% CI: 0.6164–0.6508 |
0.6013 ± 0.0494 95% CI: 0.5809–0.6217 |
0.6659 ± 0.0702 95% CI: 0.6369–0.6949 |
0.6253 ± 0.0388 95% CI: 0.6093–0.6413 |
0.6457 ± 0.0518 95% CI: 0.6243–0.6670 |
0.6213 ± 0.0400 95% CI: 0.6047–0.6378 |
0.2672 ± 0.0834 95% CI: 0.2328–0.3017 |
0.6846 ± 0.0394 95% CI: 0.6683–0.7008 |
| XGBoost | 0.6264 ± 0.0303 95% CI: 0.6139–0.6389 |
0.6204 ± 0.0512 95% CI: 0.5993–0.6415 |
0.6324 ± 0.0554 95% CI: 0.6096–0.6553 |
0.6256 ± 0.0303 95% CI: 0.6131–0.6381 |
0.6291 ± 0.0351 95% CI: 0.6147–0.6436 |
0.6235 ± 0.0343 95% CI: 0.6094–0.6377 |
0.2528 ± 0.0605 95% CI: 0.2279–0.2778 |
0.6740 ± 0.0269 95% CI: 0.6629–0.6851 |
| Gradient Boosting | 0.6479 ± 0.0340 95% CI: 0.6338–0.6619 |
0.6443 ± 0.0559 95% CI: 0.6213–0.6674 |
0.6517 ± 0.0591 95% CI: 0.6273–0.6761 |
0.6479 ± 0.0368 95% CI: 0.6327–0.6631 |
0.6506 ± 0.0382 95% CI: 0.6348–0.6664 |
0.6460 ± 0.0366 95% CI: 0.6309–0.6611 |
0.2960 ± 0.0678 95% CI: 0.2681–0.3240 |
0.6824 ± 0.0383 95% CI: 0.6666–0.6983 |
| SVM | 0.6267 ± 0.0356 95% CI: 0.6120–0.6414 |
0.5665 ± 0.0436 95% CI: 0.5485–0.5844 |
0.6870 ± 0.0565 95% CI: 0.6637–0.7103 |
0.6130 ± 0.0312 95% CI: 0.6001–0.6259 |
0.6459 ± 0.0453 95% CI: 0.6272–0.6646 |
0.6026 ± 0.0370 95% CI: 0.5873–0.6179 |
0.2535 ± 0.0713 95% CI: 0.2240–0.2829 |
0.6783 ± 0.0427 95% CI: 0.6607–0.6959 |
| KNN | 0.5916 ± 0.0347 95% CI: 0.5773–0.6059 |
0.5939 ± 0.0617 95% CI: 0.5685–0.6194 |
0.5895 ± 0.0551 95% CI: 0.5668–0.6123 |
0.5931 ± 0.0376 95% CI: 0.5776–0.6086 |
0.5916 ± 0.0343 95% CI: 0.5774–0.6057 |
0.5915 ± 0.0421 95% CI: 0.5741–0.6089 |
0.1835 ± 0.0693 95% CI: 0.1549–0.2121 |
0.6207 ± 0.0369 95% CI: 0.6055–0.6360 |
| MLP | 0.6137 ± 0.0367 95% CI: 0.5985–0.6288 |
0.5994 ± 0.0582 95% CI: 0.5754–0.6234 |
0.6280 ± 0.0512 95% CI: 0.6068–0.6491 |
0.6115 ± 0.0373 95% CI: 0.5962–0.6269 |
0.6174 ± 0.0392 95% CI: 0.6013–0.6336 |
0.6072 ± 0.0429 95% CI: 0.5895–0.6249 |
0.2274 ± 0.0731 95% CI: 0.1972–0.2575 |
0.6558 ± 0.0389 95% CI: 0.6397–0.6718 |
| Naive Bayes | 0.5767 ± 0.0583 95% CI: 0.5527–0.6008 |
0.6510 ± 0.2014 95% CI: 0.5678–0.7341 |
0.5032 ± 0.2168 95% CI: 0.4137–0.5927 |
0.5929 ± 0.0866 95% CI: 0.5571–0.6286 |
0.5769 ± 0.0568 95% CI: 0.5534–0.6003 |
0.5905 ± 0.1088 95% CI: 0.5456–0.6354 |
0.1541 ± 0.1159 95% CI: 0.1063–0.2020 |
0.6326 ± 0.0550 95% CI: 0.6098–0.6553 |
| Logistic Regression | 0.6516 ± 0.0347 95% CI: 0.6373–0.6659 |
0.6379 ± 0.0577 95% CI: 0.6141–0.6617 |
0.6653 ± 0.0641 95% CI: 0.6389–0.6918 |
0.6485 ± 0.0340 95% CI: 0.6345–0.6626 |
0.6578 ± 0.0419 95% CI: 0.6406–0.6751 |
0.6460 ± 0.0387 95% CI: 0.6301–0.6620 |
0.3033 ± 0.0694 95% CI: 0.2746–0.3319 |
0.7148 ± 0.0399 95% CI: 0.6983–0.7312 |
| Approach | Output type | Coverage of cohort | Sensitivity | Specificity | Accuracy | ROC AUC | Key clinical characteristics |
| Logistic Regression (ML) | Continuous probability | 100% (644/644) Indeterminate cases: 0% |
0.64 | 0.67 | 0.65 | 0.71 | Balanced discrimination; interpretable coefficients; no indeterminate zone; suitable for automated screening |
| Gradient Boosting (ML) | Continuous probability | 100% (644/644) Indeterminate cases: 0% |
0.65 | 0.65 | 0.65 | 0.68 | Nonlinear modeling; stable performance; full cohort applicability |
|
HSI (rule-based) |
Binary decision rule (rule-in / rule-out) |
71.30% (459/644) Indeterminate cases: 28.70% |
0.84 | 0.42 | 0.63 | 0.63 | Designed for rule-out; high sensitivity but low specificity; large indeterminate group |
|
FLI (rule-based) |
Binary decision rule (rule-in / rule-out) |
76.60% (493/644) Indeterminate cases: 23.40% |
0.71 | 0.55 | 0.63 | 0.63 | Balanced rule-in/rule-out tool; moderate discrimination; indeterminate zone remains |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




