Submitted:
07 April 2026
Posted:
09 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Mendelian Randomization
3. MR Assumption Examples
-
Relevance: The instrument must be associated with the exposure.
- Valid example: Genetic variants in the FTO locus are strongly associated with body mass index (BMI), making them suitable instruments for studying the causal effect of BMI on health outcomes.
- Invalid example: A genetic variant that shows no measurable association with the exposure (e.g., a SNP not associated with cholesterol levels) would violate the relevance assumption because it provides no useful variation in the exposure.
-
Independence: The instrument must not be associated with confounders.
- Valid example: A SNP that influences circulating LDL cholesterol but is not associated with socioeconomic status, diet, smoking, or other lifestyle factors satisfies the independence assumption.
- Invalid example: A SNP whose allele frequency differs systematically across ancestral groups, where ancestry is also related to the outcome, violates independence due to population stratification acting as a confounder.
-
Exclusion restriction: The instrument must affect the outcome only through the exposure.
- Valid example: A variant in the HMGCR gene affects cardiovascular disease risk solely through its impact on LDL cholesterol levels, consistent with exclusion restriction.
- Invalid example: A pleiotropic SNP that influences both BMI and blood pressure through separate biological pathways violates the exclusion restriction assumption because it affects the outcome through mechanisms other than the exposure.
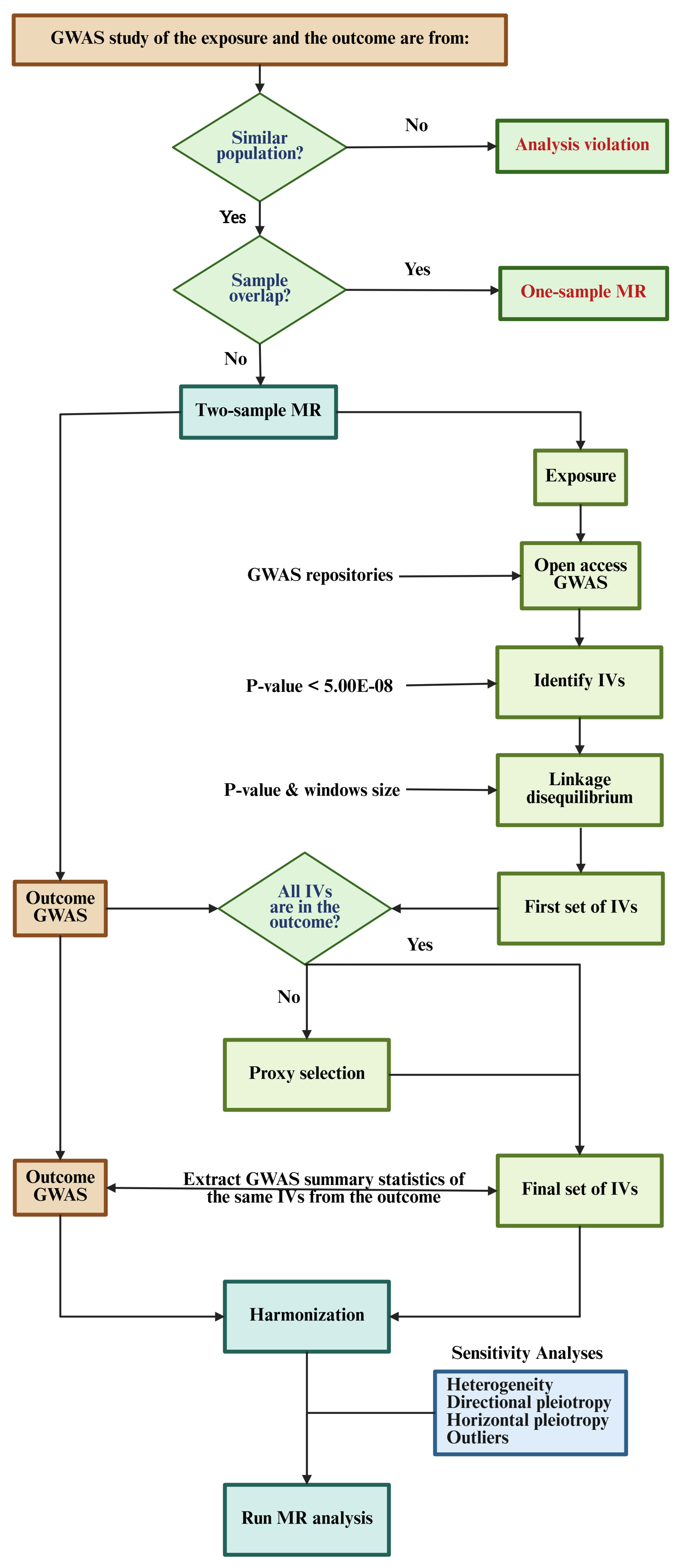
4. Workflow
4.1. Exposure and Outcome Population
4.2. Exposure and Outcome Selection
5. Illustrative Case Study
5.1. Power Calculation

5.2. Reading GWAS File (Exposure)
| Listing 1: Read the GWAS file into the TwoSampleMR package |
 |
5.3. Instrumental Variables Selection
| Listing 2: Select significant IVs based on the standard GWAS p-value threshold |
 |
5.4. Linkage Disequilibrium Clumping
| Listing 3: Local Linkage disequilibrium clumping |
 |
5.5. Instrument Strength

5.6. Reading GWAS File (Outcome)
5.7. Proxy Selection
5.8. Data Harmonization
| Listing 4: Data harmonization and save the results in a text file |
 |
5.9. Linear Methods
| Listing 5: Run and save the results of the MR analysis |
 |
-
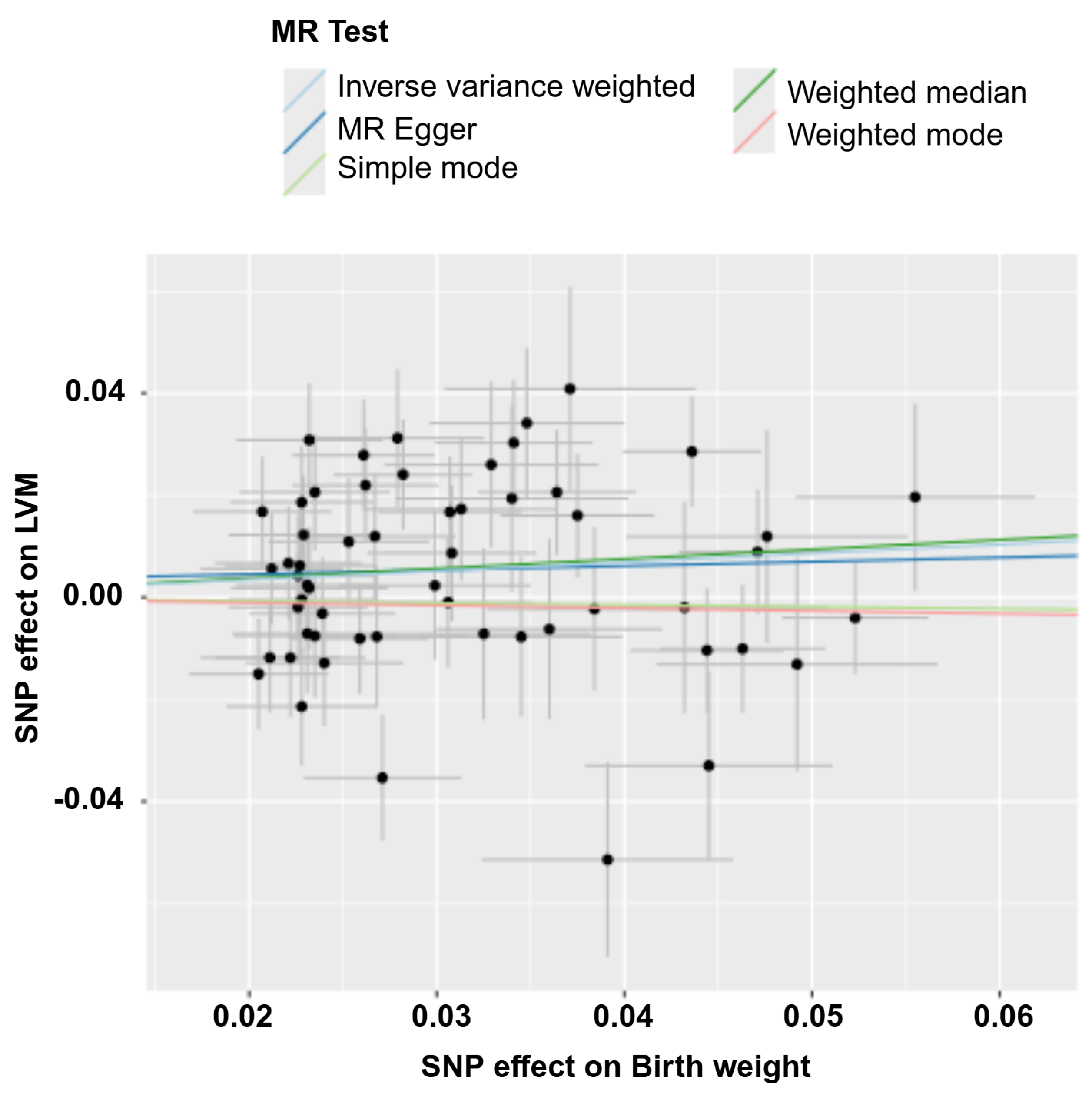
Inverse variance weighted: This is the primary estimate. With pval < 0.05, it suggests a 1-SD increase in genetically predicted birth weight is associated with a 0.1724-SD increase in left ventricular mass.

- Weighted Median: With pval < 0.05, this confirms the significant positive causal effect. This method is robust to up to 50% invalid instruments.
- MR Egger: This estimate is non-significant (pval > 0.05). Its significance here usually confirms the IVW conclusion unless heterogeneity or pleiotropy is detected.
- Weighted Mode: Non-significant (pval > 0.05). This method is highly robust to invalid instruments but has very low power. Its significance here usually confirms the IVW conclusion.
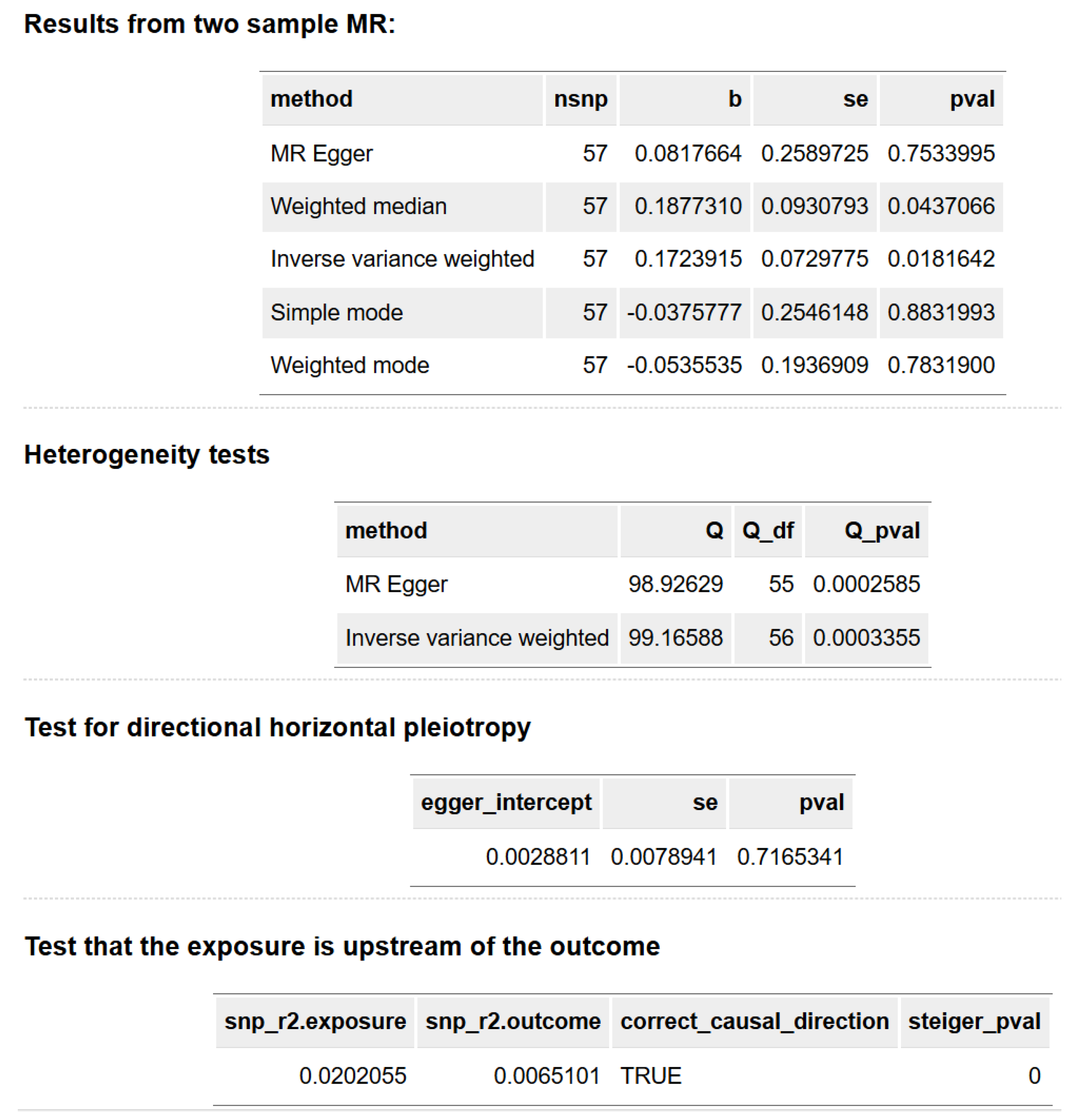

- MR Egger: With pval < 0.05, the result is highly significant. This indicates strong heterogeneity across the 57 instruments.
- IVW: Similarly, this is highly significant (pval < 0.05), confirming that there is substantial inconsistency in the causal estimates derived from the different SNPs.

| Listing 6: Run and save MR_PRESSO analysis in a text file |
 |
 |
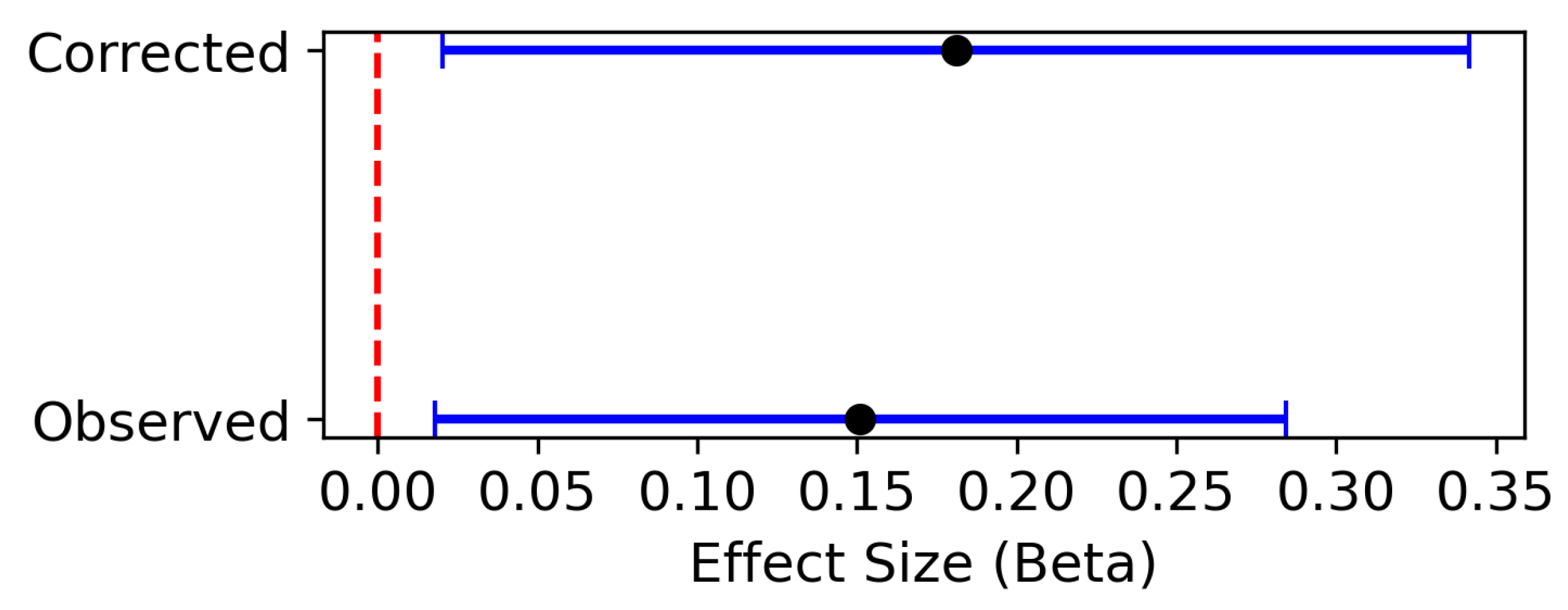
6. Possible Overlap

| Listing 7: MRlap analysis with the specified parameters |
 |
7. Multiple Variables MR
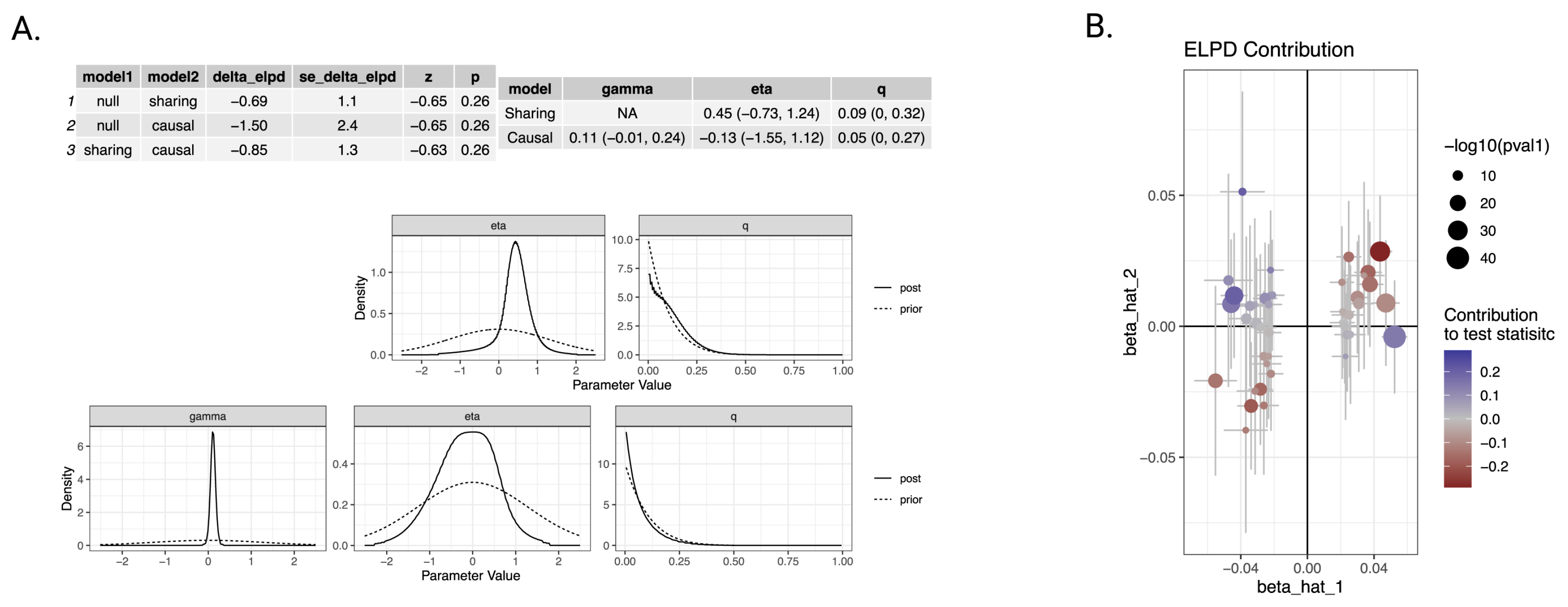
8. Causal Analysis Using CAUSE
- Causal model: the exposure has a direct causal effect on the outcome.
- Sharing model: both traits share common genetic influences, but there is no direct causal relationship.
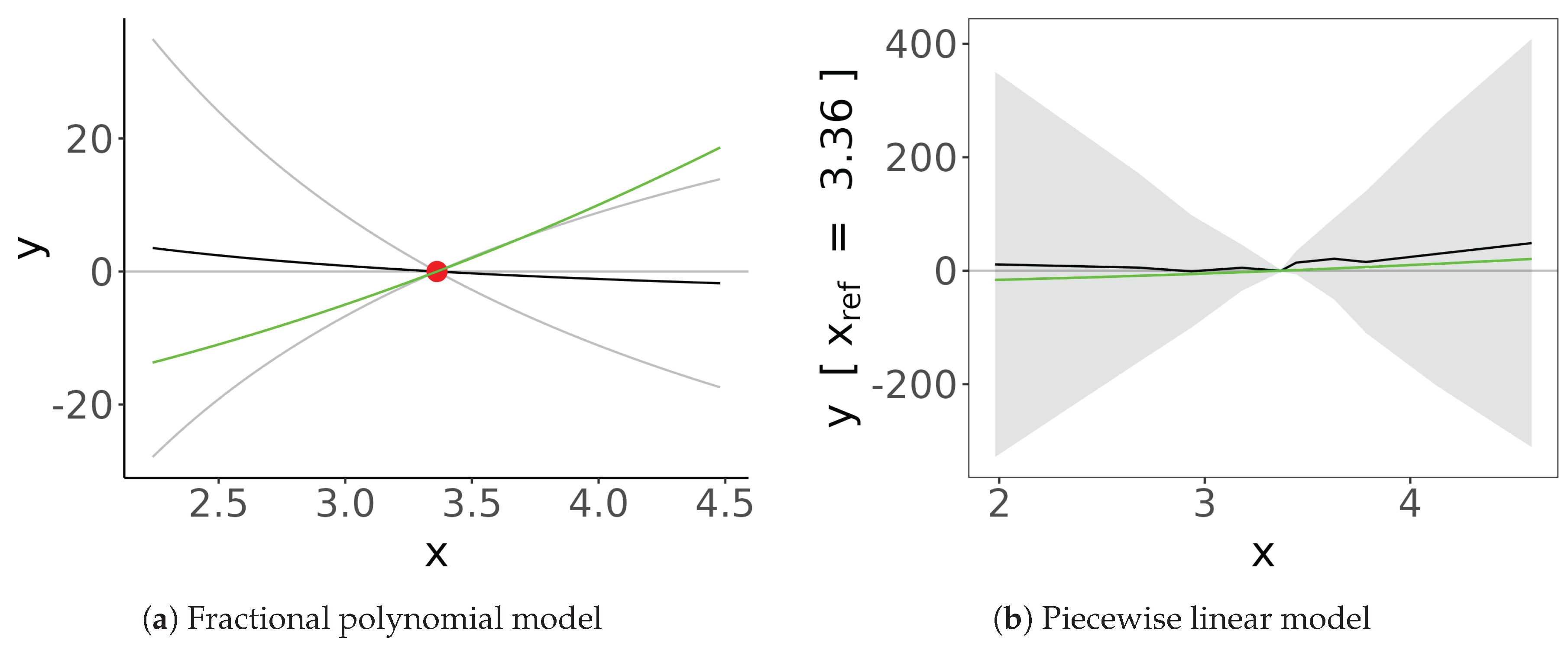
9. Non-Linear Methods

10. Multi-Traits Analysis
- #SNPs: The 3.8 million variants that passed quality control in all three files and were included in the analysis.
- or : The input sample sizes from the original GWAS summary statistics.
- GWAS mean : The original statistic, which measures the average association strength from the single-trait GWAS results.
- MTAG mean : The updated statistic, which measures the average association strength from the MTAG results after borrowing information across traits.
- GWAS equiv. : The effective sample size estimated from the increase in expected which quantifies the gain in statistical power.
11. Challenges and Limitations
- Sensitivity to GWAS Discovery Parameters. The reliability of MR estimates is heavily contingent on the quality of the underlying GWAS. Smaller sample sizes often lack the statistical power to identify robust instruments, leading to “weak instrument bias” which can pull results toward the observational association. Furthermore, as sample sizes increase, more genetic variants reach significance; however, including too many variants especially those with marginal effects can introduce noise or include pleiotropic variants that violate core MR assumptions and shift the final causal estimate.
- Lack of Ancestral Diversity in Phenotypic Data. A major bottleneck in MR is the “Eurocentric bias” within genomic databases. Most large-scale GWAS for complex traits, particularly resource-intensive imaging phenotypes like brain morphology or cardiac volumes, are conducted in populations of European descent. This lack of diversity limits the generalizability of findings. Genetic architecture and LD patterns vary across ethnicities; applying instruments discovered in one population to another can lead to biased estimates or a total loss of statistical power.
- Assumption of Linearity in Causal Effects. Most used standard MR frameworks, such as IVW, assume a linear relationship between the exposure and the outcome. However, biological systems often exhibit non-linear associations, such as U-shaped or threshold effects (e.g., the impact of alcohol consumption or BMI on mortality). When these methods are applied to non-linear data, they provide a “population-average” causal estimate that may mask critical nuances, potentially leading to misleading conclusions about the nature of the risk factor.
- Constraints of Individual-Level Data Access. While methods exist to address non-linearity, they typically require individual-level data rather than summary statistics. Obtaining such data is often hindered by stringent data transfer agreements, privacy regulations (like GDPR), or the sheer logistical challenge of harmonizing datasets across different biobanks. Consequently, researchers are frequently restricted to two-sample MR using public summary data, which, despite its convenience, is mathematically limited to estimating linear effects and cannot easily explore complex dose-response relationships.
- Influence of Linkage Disequilibrium Settings. The selection of parameters for clumping and pruning is a critical yet often subjective step that can significantly alter MR outcomes. These methods are used to ensure that genetic instruments are independent by removing variants in LD. If the r2 threshold is too relaxed, redundant variants may be included, artificially inflating the precision of the estimate. Conversely, overly stringent pruning can discard valid instruments, reducing statistical power and making the study susceptible to “winner’s curse” bias.
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bycroft, C.; Freeman, C.; Petkova, D.; Band, G.; Elliott, L.T.; Sharp, K.; Motyer, A.; Vukcevic, D.; Delaneau, O.; O’Connell, J.; et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 2018, 562, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Kurki, M.I.; Karjalainen, J.; Palta, P.; Sipila, T.P.; Kristiansson, K.; Donner, K.M.; Reeve, M.P.; Laivuori, H.; Aavikko, M.; Kaunisto, M.A.; et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 2023, 613, 508–518. [Google Scholar] [CrossRef]
- Sanderson, E.; Glymour, M.M.; Holmes, M.V.; Kang, H.; Morrison, J.; Munafò, M.R.; Palmer, T.; Schooling, C.M.; Wallace, C.; Zhao, Q.; et al. Mendelian randomization. Nature reviews Methods primers 2022, 2, 6. [Google Scholar] [CrossRef]
- Burgess, S.; Smith, G.D.; Davies, N.M.; Dudbridge, F.; Gill, D.; Glymour, M.M.; Hartwig, F.P.; Kutalik, Z.; Holmes, M.V.; Minelli, C.; et al. Guidelines for performing Mendelian randomization investigations: update for summer 2023. Wellcome open research 2023, 4, 186. [Google Scholar] [CrossRef]
- Davies, N.M.; Holmes, M.V.; Smith, G.D. Reading Mendelian randomisation studies: a guide, glossary, and checklist for clinicians. bmj 2018, 362. [Google Scholar] [CrossRef]
- Li, J.; Morrison, J. Mind the gap: characterizing bias due to population mismatch in two-sample Mendelian randomization. medRxiv 2025, 2025–07. [Google Scholar] [CrossRef]
- Sanderson, E.; Richardson, T.G.; Hemani, G.; Davey Smith, G. The use of negative control outcomes in Mendelian randomization to detect potential population stratification. International journal of epidemiology 2021, 50, 1350–1361. [Google Scholar] [CrossRef]
- Horikoshi, M.; Beaumont, R.N.; Day, F.R.; Warrington, N.M.; Kooijman, M.N.; Fernandez-Tajes, J.; Feenstra, B.; Van Zuydam, N.R.; Gaulton, K.J.; Grarup, N.; et al. Genome-wide associations for birth weight and correlations with adult disease. Nature 2016, 538, 248–252. [Google Scholar] [CrossRef] [PubMed]
- Aung, N.; Vargas, J.D.; Yang, C.; Cabrera, C.P.; Warren, H.R.; Fung, K.; Tzanis, E.; Barnes, M.R.; Rotter, J.I.; Taylor, K.D.; et al. Genome-wide analysis of left ventricular image-derived phenotypes identifies fourteen loci associated with cardiac morphogenesis and heart failure development. Circulation 2019, 140, 1318–1330. [Google Scholar] [CrossRef] [PubMed]
- Hemani, G.; Zheng, J.; Elsworth, B.; Wade, K.H.; Haberland, V.; Baird, D.; Laurin, C.; Burgess, S.; Bowden, J.; Langdon, R.; et al. The MR-Base platform supports systematic causal inference across the human phenome. elife 2018, 7, e34408. [Google Scholar] [CrossRef] [PubMed]
- Jones, S.R.; Carley, S.; Harrison, M. An introduction to power and sample size estimation. Emergency Medicine Journal 2003, 20, 453–458. [Google Scholar] [CrossRef]
- Deng, L.; Zhang, H.; Yu, K. Power calculation for the general two-sample Mendelian randomization analysis. Genetic Epidemiology 2020, 44, 290–299. [Google Scholar] [CrossRef]
- Marees, A.T.; De Kluiver, H.; Stringer, S.; Vorspan, F.; Curis, E.; Marie-Claire, C.; Derks, E.M. A tutorial on conducting genome-wide association studies: Quality control and statistical analysis. International journal of methods in psychiatric research 2018, 27, e1608. [Google Scholar] [CrossRef] [PubMed]
- Burgess, S.; Thompson, S.G.; Collaboration, C.C.G. Avoiding bias from weak instruments in Mendelian randomization studies. International journal of epidemiology 2011, 40, 755–764. [Google Scholar] [CrossRef]
- Pierce, B.L.; Burgess, S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. American journal of epidemiology 2013, 178, 1177–1184. [Google Scholar] [CrossRef] [PubMed]
- Verbanck, M.; Chen, C.Y.; Neale, B.; Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nature genetics 2018, 50, 693–698. [Google Scholar] [CrossRef]
- Burgess, S.; Davies, N.M.; Thompson, S.G. Bias due to participant overlap in two-sample Mendelian randomization. Genetic epidemiology 2016, 40, 597–608. [Google Scholar] [CrossRef] [PubMed]
- Mounier, N.; Kutalik, Z. Bias correction for inverse variance weighting Mendelian randomization. Genetic epidemiology 2023, 47, 314–331. [Google Scholar] [CrossRef]
- Huang, J.; Kleman, N.; Basu, S.; Shriver, M.D.; Zaidi, A.A. Interpreting SNP heritability in admixed populations. Genetics 2025, iyaf100. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Helenius, D.; Skotte, L.; Beaumont, R.N.; Wielscher, M.; Geller, F.; Juodakis, J.; Mahajan, A.; Bradfield, J.P.; Lin, F.T.; et al. Variants in the fetal genome near pro-inflammatory cytokine genes on 2q13 associate with gestational duration. Nature communications 2019, 10, 3927. [Google Scholar] [CrossRef]
- Morrison, J.; Knoblauch, N.; Marcus, J.H.; Stephens, M.; He, X. Mendelian randomization accounting for correlated and uncorrelated pleiotropic effects using genome-wide summary statistics. Nature genetics 2020, 52, 740–747. [Google Scholar] [CrossRef] [PubMed]
- Staley, J.R.; Burgess, S. Semiparametric methods for estimation of a nonlinear exposure-outcome relationship using instrumental variables with application to Mendelian randomization. Genetic epidemiology 2017, 41, 341–352. [Google Scholar] [CrossRef] [PubMed]
- Korte, A.; Farlow, A. The advantages and limitations of trait analysis with GWAS: a review. Plant methods 2013, 9, 29. [Google Scholar] [CrossRef]
- Turley, P.; Walters, R.K.; Maghzian, O.; Okbay, A.; Lee, J.J.; Fontana, M.A.; Nguyen-Viet, T.A.; Wedow, R.; Zacher, M.; Furlotte, N.A.; et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nature genetics 2018, 50, 229–237. [Google Scholar] [CrossRef] [PubMed]
| Repository | Phenotypes | Ethnicity | Link |
|---|---|---|---|
| China Kadoorie Biobank PheWeb | Multi-phenotype | Chinese | https://pheweb.ckbiobank.org/ CKB |
| BioBank Japan PheWeb | Multi-phenotype | Japanese | https://pheweb.jp/ BJP |
| GWAS Catalog | Multi-phenotype | Multi-ethnicity from different studies | https://www.ebi.ac.uk/gwas/ GC |
| GWAS Atlas | Multi-phenotype | Multi-ethnicity from different studies | https://atlas.ctglab.nl/ GA |
| GWAS UKBB (nealelab) | Multi-phenotype | White European | https://www.nealelab.is/uk-biobank GUKBB |
| FinnGen | Multi-phenotype | White European | https://www.finngen.fi/en/access_results FinnGen |
| OpenGWAS | Multi-phenotype | Multi-ethnicity from different studies | https://opengwas.io/ OG |
| Broad Institute Cardiovascular Disease Knowledge Portal | CVDs and CMR IDPs | Multi-ethnicity from different studies | https://hugeamp.org/downloads.html BICDKP |
| Michigan Genomics Initiative PheWeb | Multi-phenotype | Multi-ethnicity | https://pheweb.org/MGI/ MGIP |
| NIAGADS Data Sharing Service | Multi-phenotype | Multi-ethnicity from different studies | https://dss.niagads.org/ NIAGADS |
| UKBB Brain MRI GWAS | Brain MRI IDPs | White European | https://open.win.ox.ac.uk/ukbiobank/big40/pheweb33k/ UKBBMG |
| UKBB Plasma Proteomic GWAS | 2,923 proteins | White European | https://metabolomips.org/ukbbpgwas/ UKBPPG |
| Genetic factors for osteoporosis consortium | Osteoporosis | Multi-ethnicity from different studies | http://www.gefos.org/ GEFOS |
| Early Growth Genetics Consortium | Early growth factors | Multi-ethnicity from different studies | https://egg-consortium.org/ GCEG |
| UKB imputed data | Multi-phenotype | White European | https://yanglab.westlake.edu.cn/data/ukb_fastgwa/imp/ fastGWA |
| Centre For Cancer Genetic Epidemiology | Cancer | Multi-ethnicity from different studies | https://www.ccge.medschl.cam.ac.uk/breast-cancer-association-consortium-bcac/data-data-access/summary-results CCGE |
| National Cancer Institute (NIH) | Cancer | Multi-ethnicity | https://exploregwas.cancer.gov/#/downloads NCI |
| Genetic Investigation of ANthropometric Traits | Anthropometric Traits | Multi-ethnicity from different studies | https://giant-consortium.web.broadinstitute.org/index.php/GIANT_consortium_data_files GIANT |
| Common Metabolic Diseases | Metabolic Diseases | Multi-ethnicity from different studies | https://hugeamp.org/datasets.html CMD |
| Disease-centric Mendelian randomization database | Multi-phenotype | Multi-ethnicity from different studies | http://www.inbirg.com/DMRdb/#/base DMRdb |
| University of Bristol | Multi-phenotype | Multi-ethnicity from different studies | https://data.bris.ac.uk/data/dataset?level=top UoB |
| Dryad | Multi-phenotype | Multi-ethnicity from different studies | https://datadryad.org/ DRYAD |
| The Global Lipids Genetics Consortium | Lipid traits | Multi-ethnicity from different studies | https://csg.sph.umich.edu/willer/public/glgc-lipids2021/ GLGC |
| Exposure | Outcome | nsnps | Beta | SE | P_value |
|---|---|---|---|---|---|
| Gestational duration | LVM | 2 | 0.0983 | 0.211 | 6.55E-01 |
| Birth weight | LVM | 44 | 0.176 | 0.083 | 3.29E-02 |
| model1 | model2 | delta_elpd | se_delta_elpd | p |
|---|---|---|---|---|
| sharing | causal | -0.852 | 1.343 | 0.26 |
| Trait | #SNPs | GWAS mean | MTAG mean | GWAS equivalent | ||
|---|---|---|---|---|---|---|
| Birth weight | 3829459 | 143677 | 138095 | 1.276 | 1.279 | 145127 |
| LVM | 3829459 | 16923 | 16923 | 1.089 | 1.089 | 16993 |
| Gestational duration | 3829459 | 84689 | 84609 | 1.142 | 1.148 | 87897 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




