Submitted:
06 April 2026
Posted:
08 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
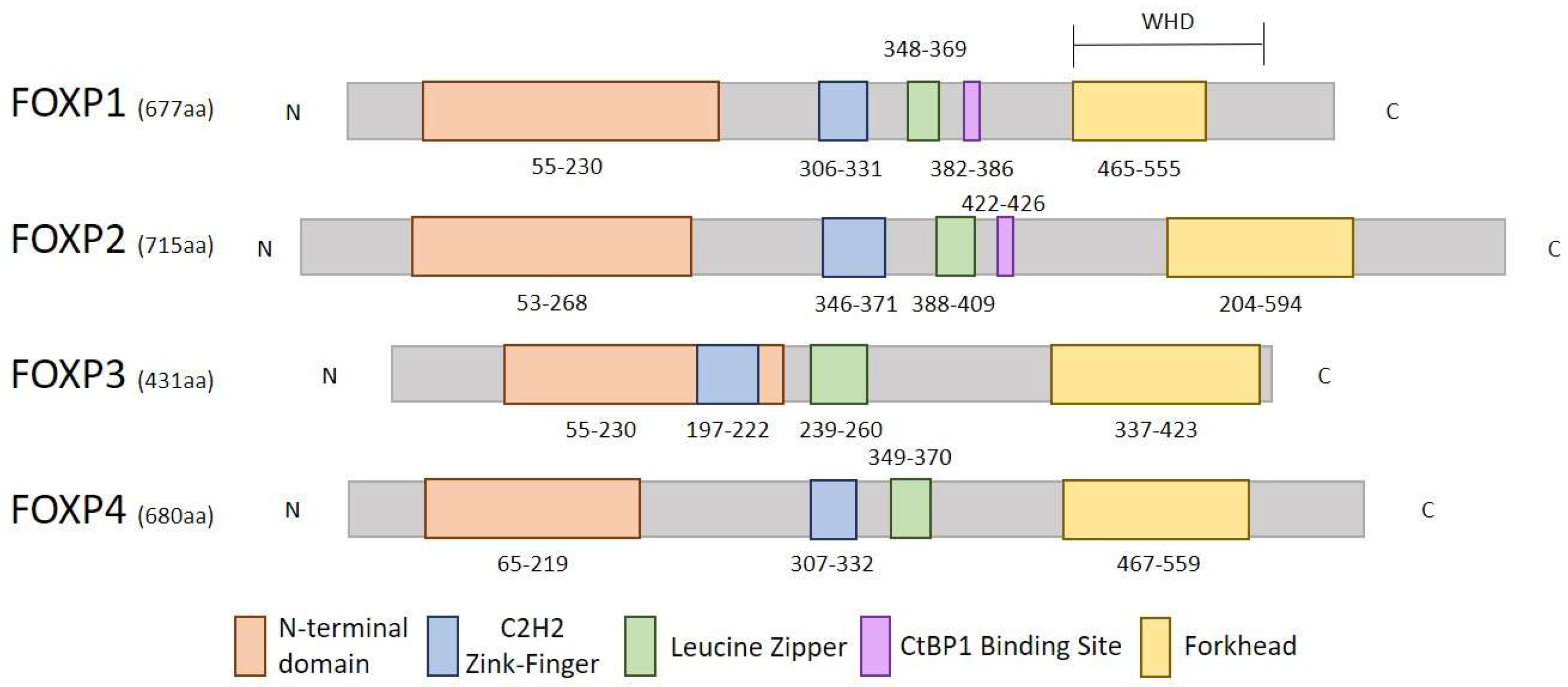
1.1. Structure and Function of FOXP Genes
1.2. FOXP Genes and Cancer
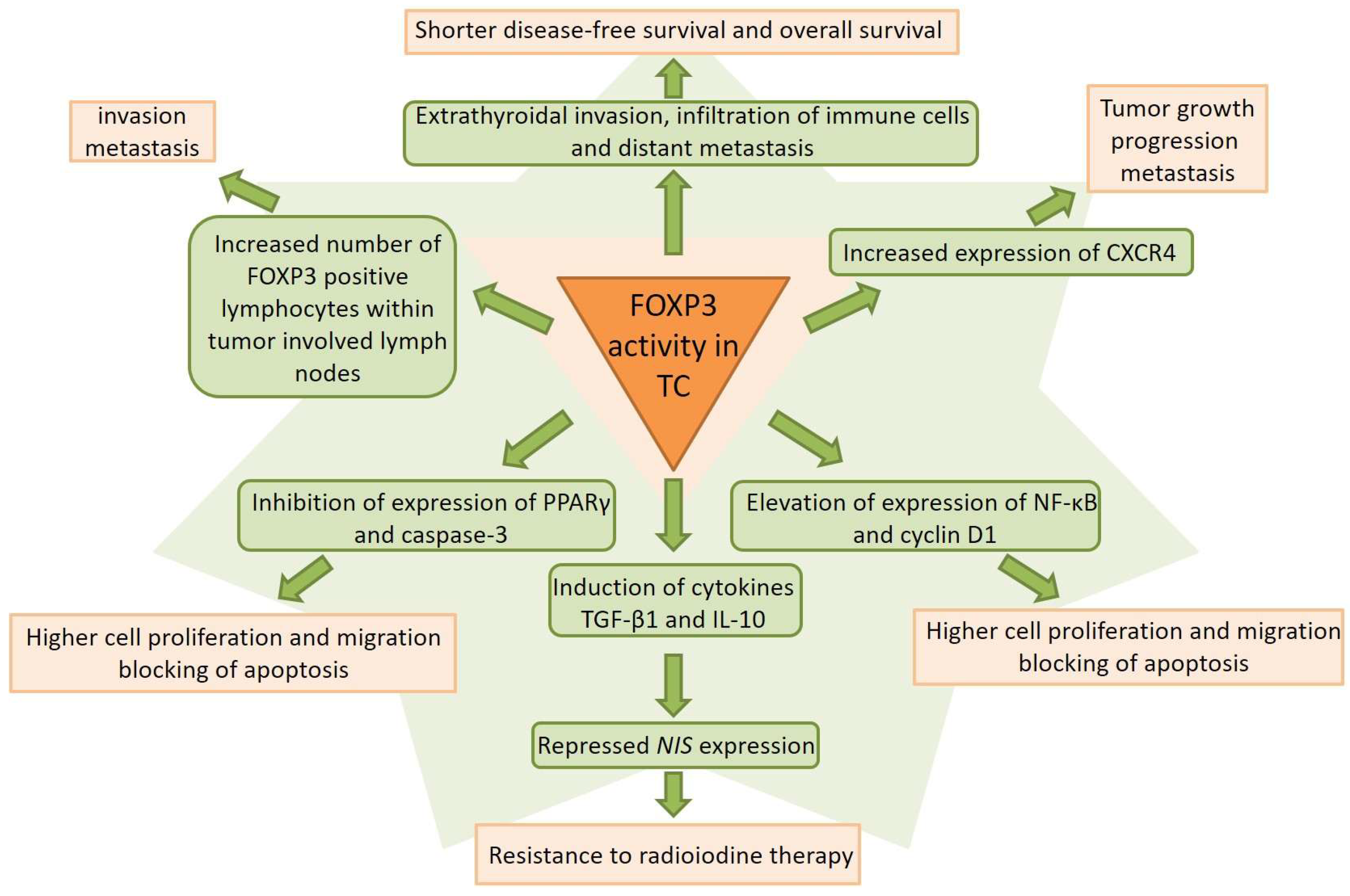
2. FOXP3 Gene in Thyroid Cancer
2.1. FOXP3 Expression And Its Localization
2.2. FOXP3 and Immune Evasion in Thyroid Cancer
2.3. Polymorphisms of FOXP3
2.4. Epigenetic Regulation of FOXP3 Expression
2.5. FOXP3 as a Therapeutic Target and PTC Biomarker
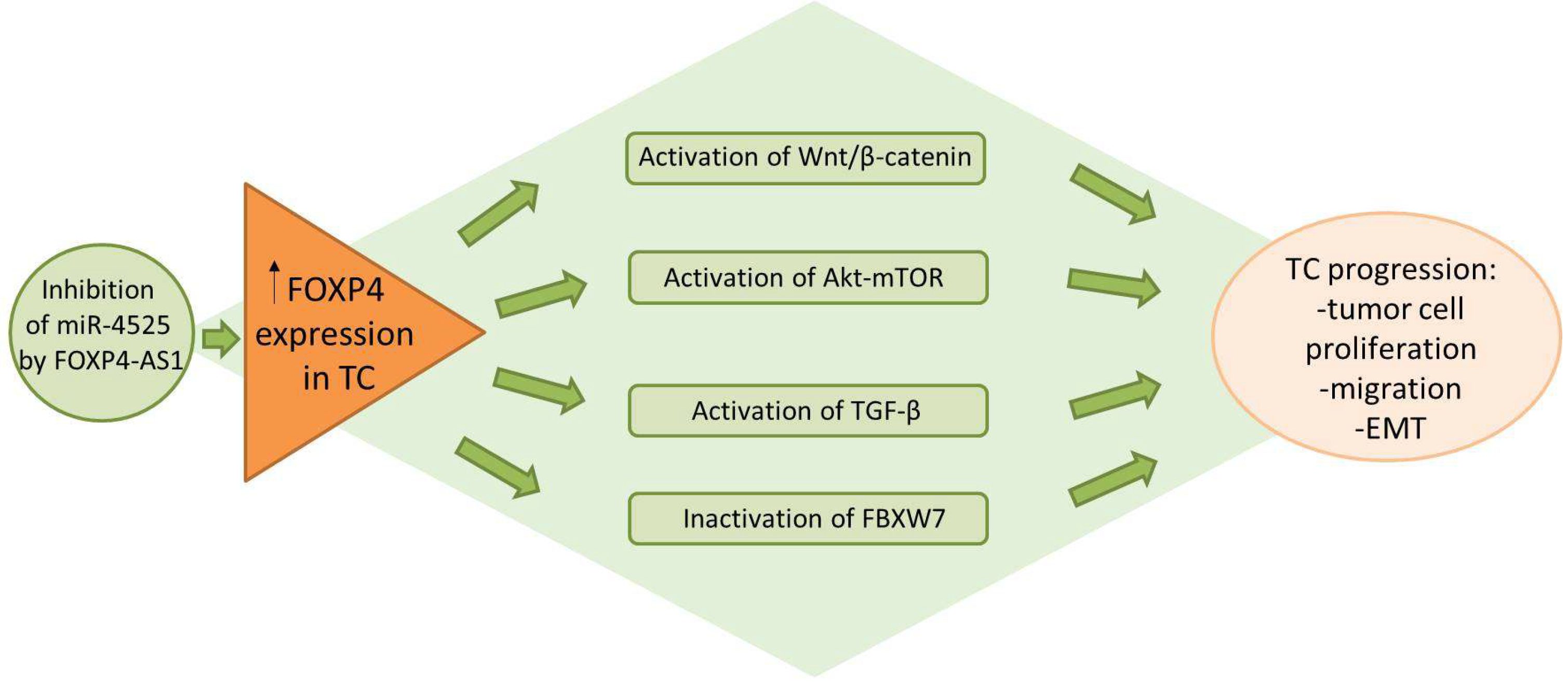
3. FOXP4 in Thyroid Cancer
3.1. FOXP4 Expression Patterns
3.2. FOXP4 Regulatory Networks
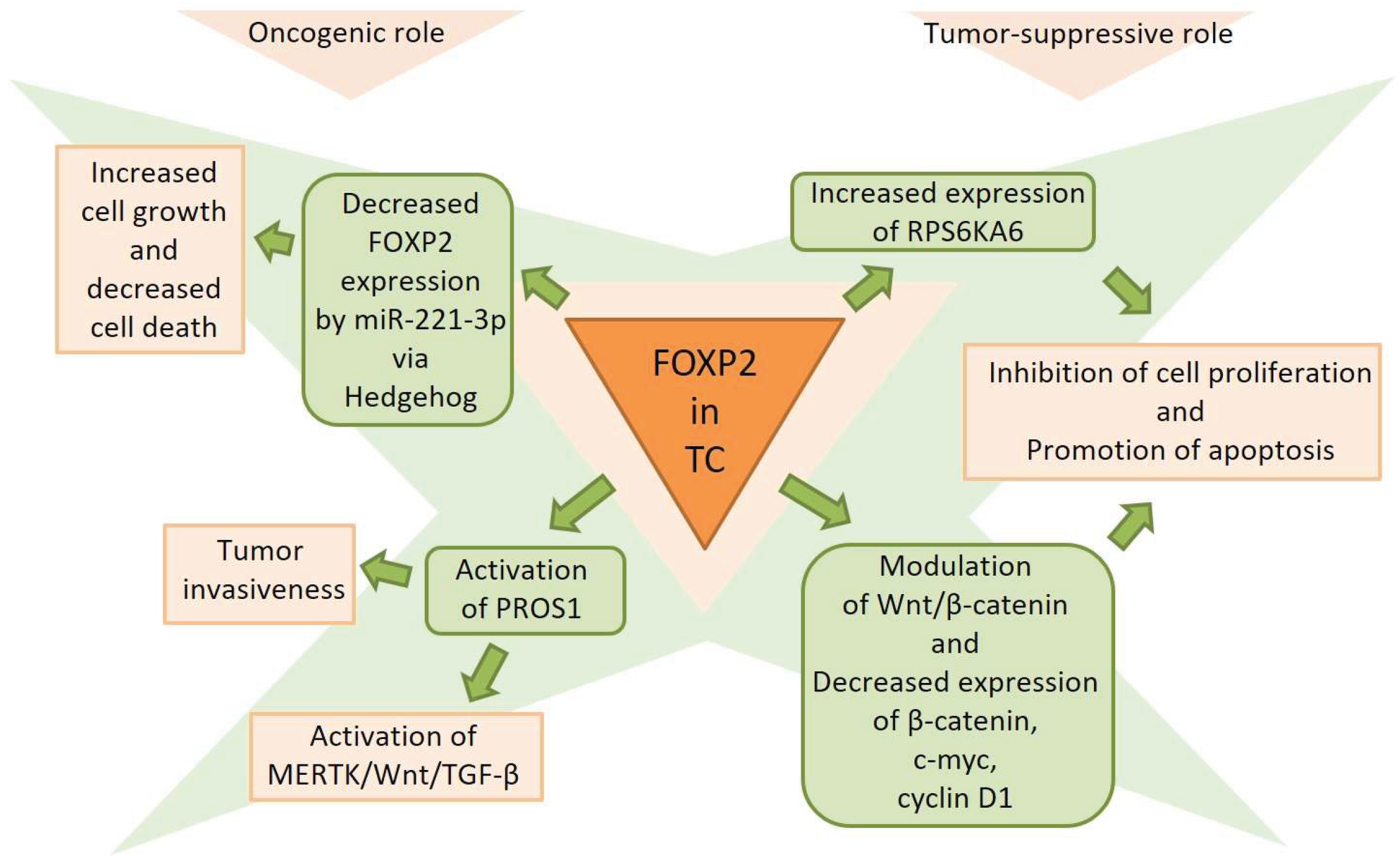
4. FOXP2 in Thyroid Cancer
4.1. FOXP2 Expression Patterns
4.2. FOXP2 Regulatory Networks
5. FOXP1 in Thyroid Cancer
6. Clinical Significance and Prognostic Implications
6.1. Associations with Tumor Aggressiveness
6.2. Lymph Node and Distant Metastasis
6.3. Treatment Resistance
6.4. Prognostic Value and Survival Outcomes
7. Conclusion and Future Perspectives
Conflicts of Interest
Abbreviations
| PCLBCL | Primary Cutaneous Large B-cell Lymphoma |
| TI-RADS ceRNA CtBP1 DHGTC DLBCL FBXW7 iTregs nTregs TGF-β1 AML-1 IGF-1 IL-10 NF-κB MALT PDTC Treg AML ATC |
Thyroid Imaging Reporting and Data System Competing Endogenous RNA C-terminal Binding site for C-terminal binding protein 1 Differentiated High-grade Thyroid Carcinoma Diffuse Large B-cell Lymphoma F-box and WD repeat domain containing 7 Induced Regulatory T cells NaturalRegulatory T cells Transforming Growth Factor-β1 Acute Myeloid Leukemia 1 Insulin-like Growth Factor 1 Interleukin-10 Nuclear factor kappa-light-chain-enhancer of activated B cells Gastric Mucosa Associated Lymphoid Tissue Lymphoma Poorly Differentiated Thyroid Carcinoma Regulatory T cells Acute Myeloid Leukemia Anaplastic Thyroid Carcinoma |
| CLT DTC EMT FTC HCC MTC NIS OTC PTC SNPs WHD WHO HT LT TC |
Chronic Lymphocytic Thyroiditis Differentiated Thyroid Carcinomas Epithelial–Mesenchymal Transition Follicular Thyroid Carcinoma Hepatocellular Carcinoma Medullary Thyroid Carcinoma Sodium Iodide Symporter OncocyticThyroid Carcinoma Papillary Thyroid Carcinoma Single Nucleotide Polymorphisms Winged Helix Domain World Health Organization Hashimoto’s Thyroiditis Lymphocytic Thyroiditis Thyroid Cancer |
References
- Filho, A.M.; Laversanne, M.; Ferlay, J.; Colombet, M.; Piñeros, M.; Znaor, A.; Parkin, D.M.; Soerjomataram, I.; Bray, F. The GLOBOCAN 2022 Cancer Estimates: Data Sources, Methods, and a Snapshot of the Cancer Burden Worldwide. Intl Journal of Cancer2025, 156, 1336–1346. [CrossRef]
- Kruger, E.; Toraih, E.A.; Hussein, M.H.; Shehata, S.A.; Waheed, A.; Fawzy, M.S.; Kandil, E. Thyroid Carcinoma: A Review for 25 Years of Environmental Risk Factors Studies. Cancers2022, 14, 6172. [CrossRef]
- Forma, A.; Kłodnicka, K.; Pająk, W.; Flieger, J.; Teresińska, B.; Januszewski, J.; Baj, J. Thyroid Cancer: Epidemiology, Classification, Risk Factors, Diagnostic and Prognostic Markers, and Current Treatment Strategies. Int J Mol Sci2025, 26, 5173. [CrossRef]
- Li, Y.-D.; Ye, Q.-Y.; Chen, Y.-X.; Hu, X.-R. Thyroid Cancer: Pathogenesis, Clinicopathology, Diagnosis, and Management. MedComm (2020)2025, 6, e70449. [CrossRef]
- Durante, C.; Hegedüs, L.; Czarniecka, A.; Paschke, R.; Russ, G.; Schmitt, F.; Soares, P.; Solymosi, T.; Papini, E. 2023 European Thyroid Association Clinical Practice Guidelines for Thyroid Nodule Management. European Thyroid Journal2023, 12, e230067. [CrossRef]
- Baloch, Z.W.; Asa, S.L.; Barletta, J.A.; Ghossein, R.A.; Juhlin, C.C.; Jung, C.K.; LiVolsi, V.A.; Papotti, M.G.; Sobrinho-Simões, M.; Tallini, G.; et al. Overview of the 2022 WHO Classification of Thyroid Neoplasms. EndocrPathol2022, 33, 27–63. [CrossRef]
- Badiu, C. Who Classification of Endocrine and Neuroendocrine Tumours, 5th Edition - Who Classification of Tumours, Volume 10. Acta Endo (Buc)2024, 20, 412–412. [CrossRef]
- Zhang, L.; Feng, Q.; Wang, J.; Tan, Z.; Li, Q.; Ge, M. Molecular Basis and Targeted Therapy in Thyroid Cancer: Progress and Opportunities. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer2023, 1878, 188928. [CrossRef]
- Nabhan, F.; Dedhia, P.H.; Ringel, M.D. Thyroid Cancer, Recent Advances in Diagnosis and Therapy. Intl Journal of Cancer2021, 149, 984–992. [CrossRef]
- Ringel, M.D.; Sosa, J.A.; Baloch, Z.; Bischoff, L.; Bloom, G.; Brent, G.A.; Brock, P.L.; Chou, R.; Flavell, R.R.; Goldner, W.; et al. 2025 American Thyroid Association Management Guidelines for Adult Patients with Differentiated Thyroid Cancer. Thyroid®2025, 35, 841–985. [CrossRef]
- Myatt, S.S.; Lam, E.W.-F. The Emerging Roles of Forkhead Box (Fox) Proteins in Cancer. Nat Rev Cancer2007, 7, 847–859. [CrossRef]
- Carlsson, P.; Mahlapuu, M. Forkhead Transcription Factors: Key Players in Development and Metabolism. Developmental Biology2002, 250, 1–23. [CrossRef]
- Fay, D.S. Cancer Metabolism: Feeding a Worm to Starve a Tumor. Current Biology2013, 23, R557–R559. [CrossRef]
- Katoh, M.; Katoh, M. Human FOX Gene Family (Review). Int J Oncol2004. [CrossRef]
- Kaestner, K.H.; Knöchel, W.; Martínez, D.E. Unified Nomenclature for the Winged Helix/Forkhead Transcription Factors. Genes Dev.2000, 14, 142–146. [CrossRef]
- Li, C.; Tucker, P.W. DNA-Binding Properties and Secondary Structural Model of the Hepatocyte Nuclear Factor 3/Fork Head Domain. Proc. Natl. Acad. Sci. U.S.A.1993, 90, 11583–11587. [CrossRef]
- Shu, W.; Yang, H.; Zhang, L.; Lu, M.M.; Morrisey, E.E. Characterization of a New Subfamily of Winged-Helix/Forkhead (Fox) Genes That Are Expressed in the Lung and Act as Transcriptional Repressors. Journal of Biological Chemistry2001, 276, 27488–27497. [CrossRef]
- Wang, B.; Lin, D.; Li, C.; Tucker, P. Multiple Domains Define the Expression and Regulatory Properties of Foxp1 Forkhead Transcriptional Repressors. Journal of Biological Chemistry2003, 278, 24259–24268. [CrossRef]
- Clark, K.L.; Halay, E.D.; Lait, E.; Burley, S.K. Co-Crystal Structure of the HNF-3/Fork Head DNA-Recognition Motif Resembles Histone HS. 1993, 364(6436), 412-20.
- Chen, Y.; Chen, C.; Zhang, Z.; Liu, C.-C.; Johnson, M.E.; Espinoza, C.A.; Edsall, L.E.; Ren, B.; Zhou, X.J.; Grant, S.F.A.; et al. DNA Binding by FOXP3 Domain-Swapped Dimer Suggests Mechanisms of Long-Range Chromosomal Interactions. Nucleic Acids Research2015, 43, 1268–1282. [CrossRef]
- Li, S.; Weidenfeld, J.; Morrisey, E.E. Transcriptional and DNA Binding Activity of the Foxp1/2/4 Family Is Modulated by Heterotypic and Homotypic Protein Interactions. Molecular and Cellular Biology2004, 24, 809–822. [CrossRef]
- Stroud, J.C.; Wu, Y.; Bates, D.L.; Han, A.; Nowick, K.; Paabo, S.; Tong, H.; Chen, L. Structure of the Forkhead Domain of FOXP2 Bound to DNA. Structure2006, 14, 159–166. [CrossRef]
- Perumal, K.; Dirr, H.W.; Fanucchi, S. A Single Amino Acid in the Hinge Loop Region of the FOXP Forkhead Domain Is Significant for Dimerisation. Protein J2015, 34, 111–121. [CrossRef]
- Chu, Y.; Chang, C.; Shiu, J.; Chang, Y.; Chen, C.; Chuang, W. Solution Structure and Backbone Dynamics of the DNA-binding Domain of FOXP1: Insight into Its Domain Swapping and DNA Binding. Protein Science2011, 20, 908–924. [CrossRef]
- Obsil, T.; Obsilova, V. Structure/Function Relationships Underlying Regulation of FOXO Transcription Factors. Oncogene2008, 27, 2263–2275. [CrossRef]
- Mendoza, E.; Scharff, C. Protein-Protein Interaction Among the FoxP Family Members and Their Regulation of Two Target Genes, VLDLR and CNTNAP2 in the Zebra Finch Song System. Front. Mol. Neurosci.2017, 10, 112. [CrossRef]
- Teufel, A.; Wong, E.A.; Mukhopadhyay, M.; Malik, N.; Westphal, H. FoxP4, a Novel Forkhead Transcription Factor. Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression2003, 1627, 147–152. [CrossRef]
- Lu, M.M.; Li, S.; Yang, H.; Morrisey, E.E. Foxp4: A Novel Member of the Foxp Subfamily of Winged-Helix Genes Co-Expressed with Foxp1 and Foxp2 in Pulmonary and Gut Tissues. Mechanisms of Development2002, 119, S197–S202. [CrossRef]
- Van Keimpema, M.; Grüneberg, L.J.; Mokry, M.; Van Boxtel, R.; Van Zelm, M.C.; Coffer, P.; Pals, S.T.; Spaargaren, M. The Forkhead Transcription Factor FOXP1 Represses Human Plasma Cell Differentiation. Blood2015, 126, 2098–2109. [CrossRef]
- Ren, J.; Han, L.; Tang, J.; Liu, Y.; Deng, X.; Liu, Q.; Hao, P.; Feng, X.; Li, B.; Hu, H.; et al. Foxp1 Is Critical for the Maintenance of Regulatory T-Cell Homeostasis and Suppressive Function. PLoS Biol2019, 17, e3000270. [CrossRef]
- Urreizti, R.; Damanti, S.; Esteve, C.; Franco-Valls, H.; Castilla-Vallmanya, L.; Tonda, R.; Cormand, B.; Vilageliu, L.; Opitz, J.M.; Neri, G.; et al. A De Novo FOXP1 Truncating Mutation in a Patient Originally Diagnosed as C Syndrome. Sci Rep2018, 8, 694. [CrossRef]
- Siper, P.M.; De Rubeis, S.; Trelles, M.D.P.; Durkin, A.; Di Marino, D.; Muratet, F.; Frank, Y.; Lozano, R.; Eichler, E.E.; Kelly, M.; et al. Prospective Investigation of FOXP1 Syndrome. Molecular Autism2017, 8, 57. [CrossRef]
- Meerschaut, I.; Rochefort, D.; Revençu, N.; Pètre, J.; Corsello, C.; Rouleau, G.A.; Hamdan, F.F.; Michaud, J.L.; Morton, J.; Radley, J.; et al. FOXP1 -Related Intellectual Disability Syndrome: A Recognisable Entity. J Med Genet2017, 54, 613–623. [CrossRef]
- Ferland, R.J.; Cherry, T.J.; Preware, P.O.; Morrisey, E.E.; Walsh, C.A. Characterization of Foxp2 and Foxp1 mRNA and Protein in the Developing and Mature Brain. J of Comparative Neurology2003, 460, 266–279. [CrossRef]
- Brown, P.; Marafioti, T.; Kusec, R.; Banham, A.H. The FOXP1 Transcription Factor Is Expressed in the Majority of Follicular Lymphomas but Is Rarely Expressed in Classical and Lymphocyte Predominant Hodgkin’s Lymphoma. J Mol Hist2005, 36, 249–256. [CrossRef]
- MacDermot, K.D.; Bonora, E.; Sykes, N.; Coupe, A.-M.; Lai, C.S.L.; Vernes, S.C.; Vargha-Khadem, F.; McKenzie, F.; Smith, R.L.; Monaco, A.P.; et al. Identification of FOXP2 Truncation as a Novel Cause of Developmental Speech and Language Deficits. The American Journal of Human Genetics2005, 76, 1074–1080. [CrossRef]
- Flores, S.V.; Figueroa-Barra, A.; Labraña-Palma, M.; Roco-Videla, A.; Caviedes-Olmos, M.; Perez-Jiménez, S.; Aguilera Eguía, R. Variability and Positive Selection in FOXP2, a Gene Associated with the Development of Language, Speech, and Cognition. Data and Metadata2024, 3, 439. [CrossRef]
- Lai, C.S.L.; Fisher, S.E.; Hurst, J.A.; Vargha-Khadem, F.; Monaco, A.P. A Forkhead-Domain Gene Is Mutated in a Severe Speech and Language Disorder. Nature2001, 413, 519–523. [CrossRef]
- Tsui, D.; Vessey, J.P.; Tomita, H.; Kaplan, D.R.; Miller, F.D. FoxP2 Regulates Neurogenesis during Embryonic Cortical Development. J. Neurosci.2013, 33, 244–258. [CrossRef]
- Spaeth, J.M.; Hunter, C.S.; Bonatakis, L.; Guo, M.; French, C.A.; Slack, I.; Hara, M.; Fisher, S.E.; Ferrer, J.; Morrisey, E.E.; et al. The FOXP1, FOXP2 and FOXP4 Transcription Factors Are Required for Islet Alpha Cell Proliferation and Function in Mice. Diabetologia2015, 58, 1836–1844. [CrossRef]
- METASTROKE Consortium, International Consortium for Blood Pressure GWAS; SpiroMeta Consortium; CHARGE Consortium Pulmonary Group, CHARGE Consortium Aging and Longevity Group; Hagenaars, S.P.; Harris, S.E.; Davies, G.; Hill, W.D.; Liewald, D.C.M.; Ritchie, S.J.; Marioni, R.E.; et al. Shared Genetic Aetiology between Cognitive Functions and Physical and Mental Health in UK Biobank (N=112 151) and 24 GWAS Consortia. Mol Psychiatry2016, 21, 1624–1632. [CrossRef]
- McCarthy-Jones, S.; Green, M.J.; Scott, R.J.; Tooney, P.A.; Cairns, M.J.; Wu, J.Q.; Oldmeadow, C.; Carr, V. Preliminary Evidence of an Interaction between the FOXP2 Gene and Childhood Emotional Abuse Predicting Likelihood of Auditory Verbal Hallucinations in Schizophrenia. Journal of Psychiatric Research2014, 50, 66–72. [CrossRef]
- Mozzi, A.; Forni, D.; Clerici, M.; Pozzoli, U.; Mascheretti, S.; Guerini, F.R.; Riva, S.; Bresolin, N.; Cagliani, R.; Sironi, M. The Evolutionary History of Genes Involved in Spoken and Written Language: Beyond FOXP2. Sci Rep2016, 6, 22157. [CrossRef]
- Jiang, L.-L.; Ruan, L.-W. Association between FOXP3 Promoter Polymorphisms and Cancer Risk: A Meta-Analysis. Oncology Letters2014, 8, 2795–2799. [CrossRef]
- Magg, T.; Mannert, J.; Ellwart, J.W.; Schmid, I.; Albert, M.H. Subcellular Localization of FOXP 3 in Human Regulatory and Nonregulatory T Cells. Eur J Immunol2012, 42, 1627–1638. [CrossRef]
- Müller, S.; Poehnert, D.; Müller, J.A.; Scheumann, G.W.F.; Koch, M.; Lück, R. Regulatory T Cells in Peripheral Blood, Lymph Node, and Thyroid Tissue in Patients with Medullary Thyroid Carcinoma. World j. surg.2010, 34, 1481–1487. [CrossRef]
- Grimmig, T.; Kim, M.; Germer, C.-T.; Gasser, M.; Maria Waaga-Gasser, A. The Role of FOXP3 in Disease Progression in Colorectal Cancer Patients. OncoImmunology2013, 2, e24521. [CrossRef]
- Waight, J.D.; Takai, S.; Marelli, B.; Qin, G.; Hance, K.W.; Zhang, D.; Tighe, R.; Lan, Y.; Lo, K.-M.; Sabzevari, H.; et al. Cutting Edge: Epigenetic Regulation of Foxp3 Defines a Stable Population of CD4+ Regulatory T Cells in Tumors from Mice and Humans. The Journal of Immunology2015, 194, 878–882. [CrossRef]
- Nik Tavakoli, N.; Hambly, B.D.; Sullivan, D.R.; Bao, S. Forkhead Box Protein 3: Essential Immune Regulatory Role. The International Journal of Biochemistry & Cell Biology2008, 40, 2369–2373. [CrossRef]
- Redpath, M.; Xu, B.; Van Kempen, L.C.; Spatz, A. The Dual Role of the X-linked FoxP3 Gene in Human Cancers. Molecular Oncology2011, 5, 156–163. [CrossRef]
- Martin, F.; Ladoire, S.; Mignot, G.; Apetoh, L.; Ghiringhelli, F. Human FOXP3 and Cancer. Oncogene2010, 29, 4121–4129. [CrossRef]
- Jia, H.; Qi, H.; Gong, Z.; Yang, S.; Ren, J.; Liu, Y.; Li, M.-Y.; Chen, G.G. The Expression of FOXP3 and Its Role in Human Cancers. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer2019, 1871, 170–178. [CrossRef]
- Jackson, B.C.; Carpenter, C.; Nebert, D.W.; Vasiliou, V. Update of Human and Mouse Forkhead Box (FOX) Gene Families. Hum Genomics2010, 4, 345. [CrossRef]
- Santos, M.E.; Athanasiadis, A.; Leitao, A.B.; DuPasquier, L.; Sucena, E. Alternative Splicing and Gene Duplication in the Evolution of the FoxP Gene Subfamily. Molecular Biology and Evolution2011, 28, 237–247. [CrossRef]
- Dong, D.; Sindhava, V.J.; Ganesan, A.; Naradikian, M.S.; Stephen, T.L.; Frisch, A.; Valentine, K.M.; Buza, E.; Wiehagen, K.R.; Cancro, M.P.; et al. Normal Treg Homeostasis and Suppressive Function Require Both FOXP1 and FOXP4.JCI insight. 2025, 10(18), e195981. [CrossRef]
- Tam, W.Y.; Leung, C.K.Y.; Tong, K.K.; Kwan, K.M. Foxp4 Is Essential in Maintenance of Purkinje Cell Dendritic Arborization in the Mouse Cerebellum. Neuroscience2011, 172, 562–571. [CrossRef]
- Co, M.; Anderson, A.G.; Konopka, G. FOXP Transcription Factors in Vertebrate Brain Development, Function, and Disorders. WIREs Developmental Biology2020, 9, e375. [CrossRef]
- Herrero, M.J.; Gitton, Y. The Untold Stories of the Speech Gene, the FOXP2 Cancer Gene. Genes Cancer2018, 9, 11–38. [CrossRef]
- Halacli, S.O.; Dogan, A.L. FOXP1 Regulation via the PI3K/Akt/p70S6K Signaling Pathway in Breast Cancer Cells. Oncology Letters2015, 9, 1482–1488. [CrossRef]
- Wlodarska, I.; Veyt, E.; De Paepe, P.; Vandenberghe, P.; Nooijen, P.; Theate, I.; Michaux, L.; Sagaert, X.; Marynen, P.; Hagemeijer, A.; et al. FOXP1, a Gene Highly Expressed in a Subset of Diffuse Large B-Cell Lymphoma, Is Recurrently Targeted by Genomic Aberrations. Leukemia2005, 19, 1299–1305. [CrossRef]
- Barrans, S.L.; Fenton, J.A.L.; Banham, A.; Owen, R.G.; Jack, A.S. Strong Expression of FOXP1 Identifies a Distinct Subset of Diffuse Large B-Cell Lymphoma (DLBCL) Patients with Poor Outcome. Blood2004, 104, 2933–2935. [CrossRef]
- Sagaert, X.; De Paepe, P.; Libbrecht, L.; Vanhentenrijk, V.; Verhoef, G.; Thomas, J.; Wlodarska, I.; De Wolf-Peeters, C. Forkhead Box Protein P1 Expression in Mucosa-Associated Lymphoid Tissue Lymphomas Predicts Poor Prognosis and Transformation to Diffuse Large B-Cell Lymphoma. JCO2006, 24, 2490–2497. [CrossRef]
- Hoefnagel, J.J.; Mulder, M.M.S.; Dreef, E.; Jansen, P.M.; Pals, S.T.; Meijer, C.J.L.M.; Willemze, R.; Vermeer, M.H. Expression of B-Cell Transcription Factors in Primary Cutaneous B-Cell Lymphoma. Modern Pathology2006, 19, 1270–1276. [CrossRef]
- Han, S.; Wu, X.; Wan, L.; Zeng, Q.; Li, J.; Liu, Z. FOXP1 Expression Predicts Polymorphic Histology and Poor Prognosis in Gastric Mucosa-Associated Lymphoid Tissue Lymphomas. Dig Surg2009, 26, 156–162. [CrossRef]
- Wang, X.; Sun, J.; Cui, M.; Zhao, F.; Ge, C.; Chen, T.; Yao, M.; Li, J. Downregulation of FOXP1 Inhibits Cell Proliferation in Hepatocellular Carcinoma by Inducing G1/S Phase Cell Cycle Arrest. IJMS2016, 17, 1501. [CrossRef]
- Fox, S.B.; Brown, P.; Han, C.; Ashe, S.; Leek, R.D.; Harris, A.L.; Banham, A.H. Expression of the Forkhead Transcription Factor FOXP1 Is Associated with Estrogen Receptor and Improved Survival in Primary Human Breast Carcinomas.Clinical Cancer Research2004, 10, 3521-7, doi.org/10.1158/1078-0432.CCR-03-0461.
- Bates, G.J.; Fox, S.B.; Han, C.; Launchbury, R.; Leek, R.D.; Harris, A.L.; Banham, A.H. Expression of the Forkhead Transcription Factor FOXP1 Is Associated with That of Estrogen Receptorβ in Primary Invasive Breast Carcinomas. Breast Cancer Res Treat2008, 111, 453–459. [CrossRef]
- Brown, P.J.; Wong, K.K.; Felce, S.L.; Lyne, L.; Spearman, H.; Soilleux, E.J.; Pedersen, L.M.; Møller, M.B.; Green, T.M.; Gascoyne, D.M.; et al. FOXP1 Suppresses Immune Response Signatures and MHC Class II Expression in Activated B-Cell-like Diffuse Large B-Cell Lymphomas. Leukemia2016, 30, 605–616. [CrossRef]
- Walker, M.P.; Stopford, C.M.; Cederlund, M.; Fang, F.; Jahn, C.; Rabinowitz, A.D.; Goldfarb, D.; Graham, D.M.; Yan, F.; Deal, A.M.; et al. FOXP1 Potentiates Wnt/b-Catenin Signaling in Diffuse Large B Cell Lymphoma.Science signaling,2015, 8(362), ra12-ra12. [CrossRef]
- Zhang, Y.; Zhang, S.; Wang, X.; Liu, J.; Yang, L.; He, S.; Chen, L.; Huang, J. Prognostic Significance of FOXP1 as an Oncogene in Hepatocellular Carcinoma. J Clin Pathol2012, 65, 528–533. [CrossRef]
- Takayama, K.; Suzuki, T.; Tsutsumi, S.; Fujimura, T.; Takahashi, S.; Homma, Y.; Urano, T.; Aburatani, H.; Inoue, S. Integrative Analysis of FOXP1 Function Reveals a Tumor-Suppressive Effect in Prostate Cancer.Molecular endocrinology2014, 28, 2012-2024, doi.org/10.1210/me.2014-1171.
- Koon, H.B.; Ippolito, G.C.; Banham, A.H.; Tucker, P.W. FOXP1: A Potential Therapeutic Target in Cancer. Expert Opinion on Therapeutic Targets2007, 11, 955–965. [CrossRef]
- Zhang, W.; Duan, N.; Song, T.; Li, Z.; Zhang, C.; Chen, X. The Emerging Roles of Forkhead Box (FOX) Proteins in Osteosarcoma. J. Cancer2017, 8, 1619–1628. [CrossRef]
- Sin, C.; Li, H.; Crawford, D.A. Transcriptional Regulation by FOXP1, FOXP2, and FOXP4 Dimerization. J Mol Neurosci2015, 55, 437–448. [CrossRef]
- Vernes, S.C.; Spiteri, E.; Nicod, J.; Groszer, M.; Taylor, J.M.; Davies, K.E.; Geschwind, D.H.; Fisher, S.E. High-Throughput Analysis of Promoter Occupancy Reveals Direct Neural Targets of FOXP2, a Gene Mutated in Speech and Language Disorders. The American Journal of Human Genetics2007, 81, 1232–1250. [CrossRef]
- DeLaughter, D.M.; Christodoulou, D.C.; Robinson, J.Y.; Seidman, C.E.; Baldwin, H.S.; Seidman, J.G.; Barnett, J.V. Spatial Transcriptional Profile of the Chick and Mouse Endocardial Cushions Identify Novel Regulators of Endocardial EMT in Vitro. Journal of Molecular and Cellular Cardiology2013, 59, 196–204. [CrossRef]
- Song, X.-L.; Tang, Y.; Lei, X.-H.; Zhao, S.-C.; Wu, Z.-Q. miR-618 Inhibits Prostate Cancer Migration and Invasion by Targeting FOXP2. J. Cancer2017, 8, 2501–2510. [CrossRef]
- Wong, K.K.; Gascoyne, D.M.; Soilleux, E.J.; Lyne, L.; Spearman, H.; Roncador, G.; Pedersen, L.M.; Møller, M.B.; Green, T.M.; Banham, A.H. FOXP2-Positive Diffuse Large B-Cell Lymphomas Exhibit a Poor Response to R-CHOP Therapy and Distinct Biological Signatures. Oncotarget2016, 7, 52940–52956. [CrossRef]
- Campbell, A.J.; Lyne, L.; Brown, P.J.; Launchbury, R.J.; Bignone, P.; Chi, J.; Roncador, G.; Lawrie, C.H.; Gatter, K.C.; Kusec, R.; et al. Aberrant Expression of the Neuronal Transcription Factor FOXP2 in Neoplastic Plasma Cells. Br J Haematol2010, 149, 221–230. [CrossRef]
- Cuiffo, B.G.; Karnoub, A.E. Silencing FOXP2 in Breast Cancer Cells Promotes Cancer Stem Cell Traits and Metastasis. Molecular & Cellular Oncology2016, 3, e1019022. [CrossRef]
- Yan, X.; Zhou, H.; Zhang, T.; Xu, P.; Zhang, S.; Huang, W.; Yang, L.; Gu, X.; Ni, R.; Zhang, T. Downregulation of FOXP2 Promoter Human Hepatocellular Carcinoma Cell Invasion. Tumor Biol.2015, 36, 9611–9619. [CrossRef]
- Liu, R.; Li, S.; Yang, W.-H.; Wang, L. IPEX Syndrome, FOXP3 and Cancer. Journal of syndromes2015, 1,7. [CrossRef]
- Szylberg, Ł.; Karbownik, D.; Marszałek, A. The Role of FOXP3 in Human Cancers. Anticancer Res2016, 36, 3789–3794.
- Li, X.; Gao, Y.; Li, J.; Zhang, K.; Han, J.; Li, W.; Hao, Q.; Zhang, W.; Wang, S.; Zeng, C.; et al. FOXP3 Inhibits Angiogenesis by Downregulating VEGF in Breast Cancer. Cell Death Dis2018, 9, 744. [CrossRef]
- He, Y.-Q.; Bo, Q.; Yong, W.; Qiu, Z.-X.; Li, Y.-L.; Li, W.-M. FoxP3 Genetic Variants and Risk of Non-Small Cell Lung Cancer in the Chinese Han Population. Gene2013, 531, 422–425. [CrossRef]
- Ma, G.-F.; Miao, Q.; Liu, Y.-M.; Gao, H.; Lian, J.-J.; Wang, Y.-N.; Zeng, X.-Q.; Luo, T.-C.; Ma, L.-L.; Shen, Z.-B.; et al. High FoxP3 Expression in Tumour Cells Predicts Better Survival in Gastric Cancer and Its Role in Tumour Microenvironment. Br J Cancer2014, 110, 1552–1560. [CrossRef]
- Hao, Q.; Zhang, C.; Gao, Y.; Wang, S.; Li, J.; Li, M.; Xue, X.; Li, W.; Zhang, W.; Zhang, Y. FOXP3 Inhibits NF-κB Activity and Hence COX2 Expression in Gastric Cancer Cells. Cellular Signalling2014, 26, 564–569. [CrossRef]
- Merlo, A.; Casalini, P.; Carcangiu, M.L.; Malventano, C.; Triulzi, T.; Mènard, S.; Tagliabue, E.; Balsari, A. FOXP3 Expression and Overall Survival in Breast Cancer. JCO2009, 27, 1746–1752. [CrossRef]
- Ladoire, S.; Arnould, L.; Mignot, G.; Coudert, B.; Rébé, C.; Chalmin, F.; Vincent, J.; Bruchard, M.; Chauffert, B.; Martin, F.; et al. Presence of Foxp3 Expression in Tumor Cells Predicts Better Survival in HER2-Overexpressing Breast Cancer Patients Treated with Neoadjuvant Chemotherapy. Breast Cancer Res Treat2011, 125, 65–72. [CrossRef]
- Zuo, T.; Wang, L.; Morrison, C.; Chang, X.; Zhang, H.; Li, W.; Liu, Y.; Wang, Y.; Liu, X.; Chan, M.W.Y.; et al. FOXP3 Is an X-Linked Breast Cancer Suppressor Gene and an Important Repressor of the HER-2/ErbB2 Oncogene. Cell2007, 129, 1275–1286. [CrossRef]
- Nakahira, K.; Morita, A.; Kim, N.-S.; Yanagihara, I. Phosphorylation of FOXP3 by LCK Downregulates MMP9 Expression and Represses Cell Invasion. PLoS ONE2013, 8, e77099. [CrossRef]
- Bettelli, E.; Dastrange, M.; Oukka, M. Foxp3 Interacts with Nuclear Factor of Activated T Cells and NF-κB to Repress Cytokine Gene Expression and Effector Functions of T Helper Cells. Proc. Natl. Acad. Sci. U.S.A.2005, 102, 5138–5143. [CrossRef]
- Ono, M.; Yaguchi, H.; Ohkura, N.; Kitabayashi, I.; Nagamura, Y.; Nomura, T.; Miyachi, Y.; Tsukada, T.; Sakaguchi, S. Foxp3 Controls Regulatory T-Cell Function by Interacting with AML1/Runx1. Nature2007, 446, 685–689. [CrossRef]
- Ma, T.; Zhang, J. Upregulation of FOXP4 in Breast Cancer Promotes Migration and Invasion through Facilitating EMT. CMAR2019, Volume 11, 2783–2793. [CrossRef]
- Yang, T.; Li, H.; Thakur, A.; Chen, T.; Xue, J.; Li, D.; Chen, M. FOXP4 Modulates Tumor Growth and Independently Associates with miR-138 in Non-Small Cell Lung Cancer Cells. Tumor Biol.2015, 36, 8185–8191. [CrossRef]
- Liu, M.; Shi, X.; Wang, J.; Xu, Y.; Wei, D.; Zhang, Y.; Yang, K.; Wang, X.; Liang, S.; Chen, X.; et al. Association of FOXP4 Gene with Prostate Cancer and the Cumulative Effects of Rs4714476 and 8q24 in Chinese Men. Clin. Lab.2015, 61. [CrossRef]
- Yang, L.; Ge, D.; Chen, X.; Qiu, J.; Yin, Z.; Zheng, S.; Jiang, C. FOXP4-AS1 Participates in the Development and Progression of Osteosarcoma by Downregulating LATS1 via Binding to LSD1 and EZH2. Biochemical and Biophysical Research Communications2018, 502, 493–500. [CrossRef]
- Niu, Y.; Wang, G.; Li, Y.; Guo, W.; Guo, Y.; Dong, Z. LncRNA FOXP4-AS1 Promotes the Progression of Esophageal Squamous Cell Carcinoma by Interacting With MLL2/H3K4me3 to Upregulate FOXP4. Front. Oncol.2021, 11, 773864. [CrossRef]
- Expression of Foxp3 and TLR4 in Human Papillary Thyroid Carcinoma and Its Clinical Significance. Histology and Histopathology2022, 38, 339–347. [CrossRef]
- Xu, L.; Yang, Z.; Zhao, Q.; Feng, H.; Kuang, J.; Liu, Z.; Chen, L.; Zhan, L.; Yan, J.; Cai, W.; et al. Effect of FOXP2 Transcription Factor on Immune Infiltration of Thyroid Cancer and Its Potential Clinical Value. Front. Immunol.2022, 13, 982812. [CrossRef]
- Szylberg, Ł.; Bodnar, M.; Harasymczuk, J.; Marszalek, A. Expression of FoxP3 Protein Plays a Key Role in Thyroid Tumors in Children. Fetal and Pediatric Pathology2014, 33, 84–91. [CrossRef]
- Gong, Z.; Jia, H.; Xue, L.; Li, D.; Zeng, X.; Wei, M.; Liu, Z.; Tong, M.C.F.; Chen, G.G. The Emerging Role of Transcription Factor FOXP3 in Thyroid Cancer. Rev EndocrMetabDisord2022, 23, 421–429. [CrossRef]
- Cunha, L.L.; Morari, E.C.; Nonogaki, S.; Soares, F.A.; Vassallo, J.; Ward, L.S. Foxp3 Expression Is Associated with Aggressiveness in Differentiated Thyroid Carcinomas. Clinics2012, 67, 483–488. [CrossRef]
- Lopes, J.E.; Torgerson, T.R.; Schubert, L.A.; Anover, S.D.; Ocheltree, E.L.; Ochs, H.D.; Ziegler, S.F. Analysis of FOXP3 Reveals Multiple Domains Required for Its Function as a Transcriptional Repressor. The Journal of Immunology2006, 177, 3133–3142. [CrossRef]
- Ugolini, C.; Elisei, R.; Proietti, A.; Pelliccioni, S.; Lupi, C.; Borrelli, N.; Viola, D.; Leocata, P.; Vitti, P.; Miccoli, P.; et al. FoxP3 Expression in Papillary Thyroid Carcinoma: A Possible Resistance Biomarker to Iodine 131 Treatment. Thyroid2014, 24, 339–346. [CrossRef]
- Hinz, S.; Pagerols-Raluy, L.; Oberg, H.-H.; Ammerpohl, O.; Grüssel, S.; Sipos, B.; Grützmann, R.; Pilarsky, C.; Ungefroren, H.; Saeger, H.-D.; et al. Foxp3 Expression in Pancreatic Carcinoma Cells as a Novel Mechanism of Immune Evasion in Cancer. Cancer Research2007, 67, 8344–8350. [CrossRef]
- Mohamed, S.Y.; Ibrahim, T.R.; Elbasateeny, S.S.; Abdelaziz, L.A.; Farouk, S.; Yassin, M.A.; Embaby, A. Clinicopathological Characterization and Prognostic Implication of FOXP3 and CK19 Expression in Papillary Thyroid Carcinoma and Concomitant Hashimoto’s Thyroiditis. Sci Rep2020, 10, 10651. [CrossRef]
- Farrag, M.S.; Emarah, Z.; Hamdy, O.; Hafez, M.T.; Abdelwahab, K.; Ros, M.H.; Farrag, N.S.; Ibrahiem, A.T. Relevance of CD4+ and CD8+ Tumor-Infiltrating Lymphocytes and Foxp3 as Prognostic Factors in Papillary Thyroid Carcinoma: New Mission for Immunotherapy. Egyptian Journal of Pathology2021, 41, 73–86. [CrossRef]
- French, J.D.; Kotnis, G.R.; Said, S.; Raeburn, C.D.; McIntyre, R.C.; Klopper, J.P.; Haugen, B.R. Programmed Death-1+ T Cells and Regulatory T Cells Are Enriched in Tumor-Involved Lymph Nodes and Associated with Aggressive Features in Papillary Thyroid Cancer. The Journal of Clinical Endocrinology & Metabolism2012, 97, E934–E943. [CrossRef]
- Zeng, R.; Lyu, Y.; Niu, H.; Yang, K.; Yan, X. FoxP3 Promotes Lymph Node Metastasis in Patients with Papillary Thyroid Carcinoma Complicated with Hashimoto’s Thyroiditis. Transl Cancer Res TCR2020, 9, 1337–1350. [CrossRef]
- Yang, X.; Lun, Y.; Jiang, H.; Liu, X.; Duan, Z.; Xin, S.; Zhang, J. SIRT1-Regulated Abnormal Acetylation of FOXP3 Induces Regulatory T-Cell Function Defect in Hashimoto’s Thyroiditis. Thyroid2018, 28, 246–256. [CrossRef]
- Neves Junior, M.P.; Carvalho Filho, E.M.D.; Camandaroba, M.P.G.; Mascarenhas, M.B.; Correia, T.C.; Neves, Y.C.S. FOXP3 Expression in Papillary Thyroid Carcinoma with and without Hashimoto’s Thyroiditis. J. Bras. Patol. Med. Lab.2013, 49, 283–287. [CrossRef]
- Zeng, R.; Lyu, Y.; Zhang, G.; Shou, T.; Wang, K.; Niu, H.; Yan, X. Positive Effect of RORγt on the Prognosis of Thyroid Papillary Carcinoma Patients Combined with Hashimoto’s Thyroiditis.American journal of translational research2018, 10, 3011.
- Chen, Z.; Ding, J.; Pang, N.; Du, R.; Meng, W.; Zhu, Y.; Zhang, Y.; Ma, C.; Ding, Y. The Th17/Treg Balance and the Expression of Related Cytokines in Uygur Cervical Cancer Patients. Diagn Pathol2013, 8, 61. [CrossRef]
- Torregrossa, L.; Giannini, R.; Borrelli, N.; Sensi, E.; Melillo, R.M.; Leocata, P.; Materazzi, G.; Miccoli, P.; Santoro, M.; Basolo, F. CXCR4 Expression Correlates with the Degree of Tumor Infiltration and BRAF Status in Papillary Thyroid Carcinomas. Modern Pathology2012, 25, 46–55. [CrossRef]
- Achilla, C.; Chorti, A.; Papavramidis, T.; Angelis, L.; Chatzikyriakidou, A. Genetic and Epigenetic Association of FOXP3 with Papillary Thyroid Cancer Predisposition. IJMS2024, 25, 7161. [CrossRef]
- Lei Zheng, W.J.; Yang Zhang, L.X. Association between FOXP3, FOXE1 Gene Polymorphisms and Risk of Differentiated Thyroid Cancer in Chinese Han Population. Mol Biol2015, 04. [CrossRef]
- Muayad, B.A.; Jassim, H.A. Role of FOXP3 (Rs3761548) Polymorphism in Modulating FOXP3 Protein Level in Iraqi Patients with Thyroid Disorder. Al-Rafidain J Med Sci2025, 8, 208–214. [CrossRef]
- Oda, J.M.M.; Hirata, B.K.B.; Guembarovski, R.L.; Watanabe, M.A.E. Genetic Polymorphism in FOXP3 Gene: Imbalance in Regulatory T-Cell Role and Development of Human Diseases. J Genet2013, 92, 163–171. [CrossRef]
- Yu, M.; Tan, X.; Huang, Y. Foxp-3 Variants Are Associated with Susceptibility to Graves’ Disease in Chinese Population. Eur J Inflamm2017, 15, 113–119. [CrossRef]
- Tan, G.; Wang, X.; Zheng, G.; Du, J.; Zhou, F.; Liang, Z.; Wei, W.; Yu, H. Meta-Analysis Reveals Significant Association between FOXP3 Polymorphisms and Susceptibility to Graves’ Disease. J Int Med Res2021, 49, 03000605211004199. [CrossRef]
- Wang, S.; Wu, J.; Ren, J.; Vlantis, A.C.; Li, M.; Liu, S.Y.W.; Ng, E.K.W.; Chan, A.B.W.; Luo, D.-C.; Liu, Z.; et al. MicroRNA-125b Interacts with Foxp3 to Induce Autophagy in Thyroid Cancer. Molecular Therapy2018, 26, 2295–2303. [CrossRef]
- Wang, J.; Gong, R.; Zhao, C.; Lei, K.; Sun, X.; Ren, H. Human FOXP3 and Tumour Microenvironment. Immunology2023, 168, 248–255. [CrossRef]
- Zhou, T.; Zhao, D.; Ma, N.; Zhu, X.; Chen, X.; Luo, X.; Chen, S.; Gao, Q. The Essential Role of Forkhead Box P4 (FOXP4) in Thyroid Cancer: A Study Related to The Cancer Genome Atlas and Experimental Data. Endocrine Connections2023, 12, e220390. [CrossRef]
- Zhou, T.; Ma, N.; Zhang, Y.; Chen, X.; Luo, X.; Zhang, M.; Gao, Q.; Zhao, D. Transcription Factor FOXP4 Inversely Governs Tumor Suppressor Genes and Contributes to Thyroid Cancer Progression. Heliyon2024, 10, e23875. [CrossRef]
- Luo, X.; Gao, Q.; Zhou, T.; Tang, R.; Zhao, Y.; Zhang, Q.; Wang, N.; Ye, H.; Chen, X.; Chen, S.; et al. FOXP4-AS1 Inhibits Papillary Thyroid Carcinoma Proliferation and Migration Through the AKT Signaling Pathway. Front. Oncol.2022, 12, 900836. [CrossRef]
- Yeh, C.-H.; Bellon, M.; Nicot, C. FBXW7: A Critical Tumor Suppressor of Human Cancers. Mol Cancer2018, 17, 115. [CrossRef]
- Wang, Z.; Inuzuka, H.; Zhong, J.; Wan, L.; Fukushima, H.; Sarkar, F.H.; Wei, W. Tumor Suppressor Functions of FBW7 in Cancer Development and Progression. FEBS Letters2012, 586, 1409–1418. [CrossRef]
- Rajabi, S.; Alix-Panabières, C.; Alaei, A.S.; Abooshahab, R.; Shakib, H.; Ashrafi, M.R. Looking at Thyroid Cancer from the Tumor-Suppressor Genes Point of View. Cancers2022, 14, 2461. [CrossRef]
- Chen, Y.; Li, X.; Li, Y.; Wu, Y.; Huang, G.; Wang, X.; Guo, S. Downregulation of microRNA-423-5p Suppresses TGF-β1-induced EMT by Targeting FOXP4 in Airway Fibrosis. Mol Med Rep2022, 26, 242. [CrossRef]
- E, C.; Yang, J.; Li, H.; Li, C. LncRNA LOC105372579 Promotes Proliferation and Epithelial-Mesenchymal Transition in Hepatocellular Carcinoma via Activating miR-4316/FOXP4 Signaling. CMAR2019, Volume 11, 2871–2879. [CrossRef]
- Zhang, G.; Wang, Y.; Han, X.; Lu, T.; Fu, L.; Jin, H.; Yang, K.; Cai, H. FOXP4-AS1 May Be a Potential Prognostic Biomarker in Human Cancers: A Meta−Analysis and Bioinformatics Analysis. Front. Oncol.2022, 12, 799265. [CrossRef]
- Yu, L.; Lu, J. Long Non-Coding RNA, FOXP4-AS1, Acts as a Novel Biomarker of Cancers. Oncologie2023, 25, 619–627. [CrossRef]
- Zhang, F.; Yu, X.; Lin, Z.; Wang, X.; Gao, T.; Teng, D.; Teng, W. Using Tumor-Infiltrating Immune Cells and a ceRNA Network Model to Construct a Prognostic Analysis Model of Thyroid Carcinoma. Front. Oncol.2021, 11, 658165. [CrossRef]
- Akhlaghipour, I.; Fanoodi, A.; Zangouei, A.S.; Taghehchian, N.; Khalili-Tanha, G.; Moghbeli, M. MicroRNAs as the Critical Regulators of Forkhead Box Protein Family in Pancreatic, Thyroid, and Liver Cancers. Biochem Genet2023, 61, 1645–1674. [CrossRef]
- Taghehchian, N.; Lotfi, M.; Zangouei, A.S.; Akhlaghipour, I.; Moghbeli, M. MicroRNAs as the Critical Regulators of Forkhead Box Protein Family during Gynecological and Breast Tumor Progression and Metastasis. Eur J Med Res2023, 28, 330. [CrossRef]
- Su, W.; Hu, S.; Zhou, L.; Bi, H.; Li, Z. FOXP2 Inhibits the Aggressiveness of Lung Cancer Cells by Blocking TGFβ Signaling. Oncol Lett2024, 27, 227. [CrossRef]
- Xiaoquan Zhu; Chao Chen; Wei Dong; Yong Xu; Siying Liang; Wenlong Jia; Jian Li; Ying Qu; JianpoZhai; Yaoguang Zhang; et al. FOXP2confers Oncogenic Effects in Prostate Cancer through Activating MET Signalling. 2022. [CrossRef]
- Yan, L.; Sun, H.; Chen, Y.; Yu, X.; Zhang, J.; Li, P. FOXP2 Suppresses the Proliferation, Invasion, and Aerobic Glycolysis of Hepatocellular Carcinoma Cells by Regulating the KDM5A / FBP1 Axis. Environmental Toxicology2024, 39, 341–356. [CrossRef]
- Yang, F.; Xiao, Z.; Zhang, S. FOXP2 Regulates Thyroid Cancer Cell Proliferation and Apoptosis via Transcriptional Activation of RPS6KA6. Exp Ther Med2022, 23, 434. [CrossRef]
- Chang, W.; Chang, Q.; Lu, H.; Li, Y.; Chen, C. MiR-221-3p Facilitates Thyroid Cancer Cell Proliferation and Inhibit Apoptosis by Targeting FOXP2 Through Hedgehog Pathway. Mol Biotechnol2022, 64, 919–927. [CrossRef]
- Fu, L.; Shi, Z.; Luo, G.; Tu, W.; Wang, X.; Fang, Z.; Li, X. Multiple microRNAs Regulate Human FOXP2 Gene Expression by Targeting Sequences in Its 3’ Untranslated Region. Mol Brain2014, 7, 71. [CrossRef]
- Tapia-Carrillo, D.; Zambada-Moreno, O.; Hernández-Lemus, E.; Tovar, H. Two Cohorts, One Network: Consensus Master Regulators Orchestrating Papillary Thyroid Carcinoma. IJMS2025, 26, 11231. [CrossRef]
- Zhang, Y.-L.; Sun, F.-T.; Zhang, Z.; Chen, X.-X.; Liu, A.-X.; Pan, J.-J.; Peng, F.; Zhou, S.; Sun, L.-J. Comprehensive Expression Analysis Suggests Functional Overlapping of Human FOX Transcription Factors in Cancer. Asian Pacific Journal of Cancer Prevention2015, 15, 10475–10481. [CrossRef]
- Kim, J.-H.; Hwang, J.; Jung, J.H.; Lee, H.-J.; Lee, D.Y.; Kim, S.-H. Molecular Networks of FOXP Family: Dual Biologic Functions, Interplay with Other Molecules and Clinical Implications in Cancer Progression. Mol Cancer2019, 18, 180. [CrossRef]
- Zou, H.; Tang, C.; Chen, H. FOXP2 Regulates the Proliferation, Migration, and Apoptosis of Thyroid Carcinoma Cells via Wnt/β-Catenin Signaling Pathway. Trop. J. Pharm Res2022, 20, 1609–1614. [CrossRef]
- Zhang, W.; Zhang, Y.; Liu, Z.; Wang, Z.; Wang, H.; Ji, X.; Su, H.; Yang, F.; Yan, L.; Xu, Y.; et al. PROS1-MERTK Axis Drives Tumor Microenvironment Crosstalk and Progression in Papillary Thyroid Microcarcinoma. Advanced Science2025, 12, e13474. [CrossRef]
- Agbektas, T.; Tas, A.; Sariakcali, B.; Hasbek, Z.; Silig, Y. Potential of the FOX Gene Family and LncRNAs as Biomarkers in Differentiated Thyroid Cancer Treated with I-131. Turkish Journal of Biochemistry2025, 50(6), 813-826. [CrossRef]
- Banham, A.H.; Beasley, N.; Campo, E.; Fernandez, P.L.; Fidler, C.; Gatter, K.; Jones, M.; Mason, D.Y.; Prime, J.E.; Trougouboff, P.; et al. The FOXP1 Winged Helix Transcription Factor Is a Novel Candidate Tumor Suppressor Gene on Chromosome 3p.Cancer research2001, 61(24), 8820-8829.
- Banham, A.H.; Connors, J.M.; Brown, P.J.; Cordell, J.L.; Ott, G.; Sreenivasan, G.; Farinha, P.; Horsman, D.E.; Gascoyne, R.D. Expression of the FOXP1 Transcription Factor Is Strongly Associated with Inferior Survival in Patients with Diffuse Large B-Cell Lymphoma.Clinical Cancer Research2005, 11(3), 1065-1072, doi.org/10.1158/1078-0432.1065.11.3.
- Jiang, W.; Li, L.; Tang, Y.; Zhang, W.; Liu, W.; Li, G. Expression of FOXP1 in Mucosa-Associated Lymphoid Tissue Lymphoma Suggests a Large Tumor Cell Transformation and Predicts a Poorer Prognosis in the Positive Thyroid Patients. Med Oncol2012, 29, 3352–3359. [CrossRef]
- Xiao, J.; He, B.; Zou, Y.; Chen, X.; Lu, X.; Xie, M.; Li, W.; He, S.; You, S.; Chen, Q. Prognostic Value of Decreased FOXP1 Protein Expression in Various Tumors: A Systematic Review and Meta-Analysis. Sci Rep2016, 6, 30437. [CrossRef]
- Sheng, H.; Li, X.; Xu, Y. Knockdown of FOXP1 Promotes the Development of Lung Adenocarcinoma. Cancer Biology & Therapy2019, 20, 537–545. [CrossRef]
- Wen, X.; Xu, Z.; Ni, H.; Liu, S.; Jin, Y.; Zhao, W.; Luo, S.; Fang, Y.; Mao, Z.; Lin, J.; et al. FOXP1 Is Associated with Oncogenesis and Clinical Outcomes in Hematologic Malignancies. Front. Immunol.2025, 16, 1569641. [CrossRef]
- Wang, L.; Luo, P.; Yang, Z.; Zhong, X.; Ji, C. FOXP1 Inhibits Pancreatic Cancer Growth by Transcriptionally Regulating IRF1 Expression. PLoS ONE2023, 18, e0280794. [CrossRef]
- Giatromanolaki, A.; Koukourakis, M.I.; Sivridis, E.; Gatter, K.C.; Harris, A.L.; Banham, A.H. Loss of Expression and Nuclear/Cytoplasmic Localization of the FOXP1 Forkhead Transcription Factor Are Common Events in Early Endometrial Cancer: Relationship with Estrogen Receptors and HIF-1α Expression. Modern Pathology2006, 19, 9–16. [CrossRef]
- Katoh, M.; Igarashi, M.; Fukuda, H.; Nakagama, H.; Katoh, M. Cancer Genetics and Genomics of Human FOX Family Genes. Cancer Letters2013, 328, 198–206. [CrossRef]
- Ginzberg, S.P.; Sharpe, J.; Passman, J.E.; Amjad, W.; Wirtalla, C.J.; SoegaardBallester, J.M.; Finn, C.B.; Mandel, S.J.; Kelz, R.R.; Wachtel, H. Revisiting the Relationship Between Tumor Size and Risk in Well-Differentiated Thyroid Cancer. Thyroid2024, 34, 980–989. [CrossRef]
- Kim, C.A.; Kim, H.I.; Kim, N.H.; Kim, T.Y.; Kim, W.B.; Chung, J.H.; Jeon, M.J.; Kim, T.H.; Kim, S.W.; Kim, W.G. Prognostic Impact of Primary Tumor Size in Papillary Thyroid Carcinoma without Lymph Node Metastasis. Endocrinol Metab2025, 40, 405–413. [CrossRef]
- Bouzehouane, N.; Roy, P.; Decaussin-Petrucci, M.; Bertholon-Grégoire, M.; Bully, C.; Perrin, A.; Lasolle, H.; Lifante, J.-C.; Borson-Chazot, F.; Bournaud, C. Prognostic Impact of Microscopic Extra-Thyroidal Extension (mETE) on Disease Free Survival in Patients with Papillary Thyroid Carcinoma (PTC). Cancers2022, 14, 2591. [CrossRef]
- Liu, P.; Yu, X. Lymphatic Metastasis of Papillary Thyroid Carcinoma: Mechanism and Clinicopathological Physiology. Front. Endocrinol.2026, 16, 1725077. [CrossRef]
- Zeng, R.; Lyu, Y.; Niu, H.; Yang, K.; Yan, X. FoxP3 Promotes Lymph Node Metastasis in Patients with Papillary Thyroid Carcinoma Complicated with Hashimoto’s Thyroiditis. Transl Cancer Res TCR2020, 9, 1337–1350. [CrossRef]
- Wang, S.; Luo, J.; Zhong, Z.; Qi, X.; Liu, T.; Qin, R.; Zhao, Z.; Xi, Y.; Sun, R.; Wu, Y.; et al. Treatment and Prognosis for Patients with Differentiated Thyroid Carcinoma Bone Metastases. Transl Cancer Res TCR2020, 9, 972–982. [CrossRef]
- Ma, S.; Wang, Q.; Ma, X.; Wu, L.; Guo, F.; Ji, H.; Liu, F.; Zhao, Y.; Qin, G. FoxP3 in Papillary Thyroid Carcinoma Induces NIS Repression through Activation of the TGF-Β1/Smad Signaling Pathway. Tumor Biol.2016, 37, 989–998. [CrossRef]
- Riesco-Eizaguirre, G.; Rodríguez, I.; De La Vieja, A.; Costamagna, E.; Carrasco, N.; Nistal, M.; Santisteban, P. The BRAFV600E Oncogene Induces Transforming Growth Factor β Secretion Leading to Sodium Iodide Symporter Repression and Increased Malignancy in Thyroid Cancer. Cancer Research2009, 69, 8317–8325. [CrossRef]
- He, Y.; Tang, Z.; Xu, M.; Huang, T. Dedifferentiation and Redifferentiation of Follicular-Cell-Derived Thyroid Carcinoma: Mechanisms and Therapeutic Implications. Biomedicines2025, 13, 2982. [CrossRef]
- Jiang, S.; Huang, Y.; Li, Y.; Gu, Q.; Jiang, C.; Tao, X.; Sun, J. Silencing FOXP2 Reverses Vemurafenib Resistance in BRAFV600E Mutant Papillary Thyroid Cancer and Melanoma Cells. Endocrine2022, 79, 86–97. [CrossRef]
- Li, S.; Chen, Z.; Liu, M.; Li, L.; Cai, W.; Lian, Z.-X.; Guan, H.; Xu, B. Immunophenotyping with High-Dimensional Flow Cytometry Identifies Treg Cell Subsets Associated with Recurrence in Papillary Thyroid Carcinoma. Endocrine-Related Cancer2024, 31, e230240. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



