Submitted:
05 April 2026
Posted:
08 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
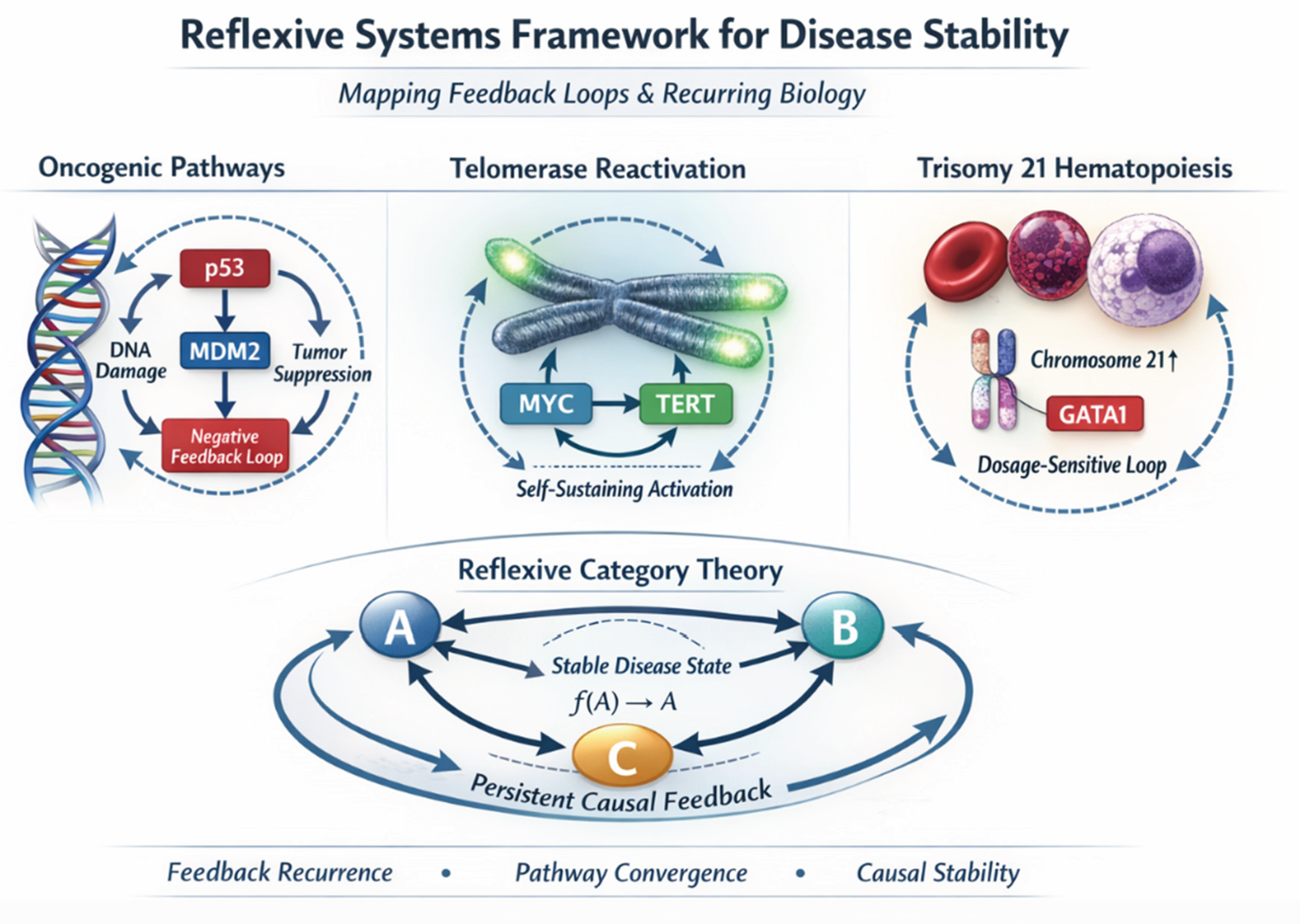
Biology-First Reflexive Systems Framework
Case Study I: Damage Surveillance as a Reflexive Tumor-Suppressor Program
Case Study II: Telomerase Reactivation as a Self-Sustaining Oncogenic State
Case Study III: Trisomy 21 as a Developmental Dosage-Rewiring Event
What RCT Adds Beyond Classical Experimental Readouts
Discussion
Conclusion
References
- Huang, S. Back to the biology in systems biology: what can we learn from biomolecular networks? Brief Funct Genomic Proteomic;PubMed 2004, 2(4), 279–97. [Google Scholar] [CrossRef] [PubMed]
- Sreekant Mudholkar, R; Virmani, A; Ranjeet Mudholkar, S. Reflexive Category Theory: A Unified Framework for Self-Reference in Mathematics and Computer Science.
- Lahav, G; Rosenfeld, N; Sigal, A; Geva-Zatorsky, N; Levine, AJ; Elowitz, MB; et al. Dynamics of the p53-Mdm2 feedback loop in individual cells. Nat Genet. 2004, 36(2), 147–50. [Google Scholar] [CrossRef] [PubMed]
- Venkitaraman, AR. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell;PubMed 2002, 108(2), 171–82. [Google Scholar] [CrossRef] [PubMed]
- Huang, R; Zhou, PK. DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduction and Targeted Therapy PubMed. 2021, 6(1), 1 254. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov.;PubMed 2022, 12(1), 31–46. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X; Larsson, C; Xu, D. Mechanisms underlying the activation of TERT transcription and telomerase activity in human cancer: old actors and new players. Oncogene PubMed. 2019, 38(34), 6172–83. [Google Scholar] [CrossRef] [PubMed]
- Dratwa, M; Wysoczańska, B; Łacina, P; Kubik, T; Bogunia-Kubik, K. TERT—Regulation and Roles in Cancer Formation. Front Immunol;PubMed 2020, 11, 589929. [Google Scholar] [CrossRef] [PubMed]
- Laurent, AP; Kotecha, RS; Malinge, S. Gain of chromosome 21 in hematological malignancies: lessons from studying leukemia in children with Down syndrome. Leukemia;PubMed 2020, 34(8), 1984–99. [Google Scholar] [CrossRef] [PubMed]
- Geva-Zatorsky, N; Rosenfeld, N; Itzkovitz, S; Milo, R; Sigal, A; Dekel, E; et al. Oscillations and variability in the p53 system. Mol Syst Biol. 2006, 2. [Google Scholar] [CrossRef] [PubMed]
- Marine, JC; Lozano, G. Mdm2-mediated ubiquitylation: p53 and beyond. Cell Death Differ;PubMed 2010, 17(1), 93–102. [Google Scholar] [CrossRef] [PubMed]
- Khattar, E; Tergaonkar, V. Transcriptional Regulation of Telomerase Reverse Transcriptase (TERT) by MYC. In Front Cell Dev Biol; PubMed, 26 Jan 2017; Volume 5(JAN). [Google Scholar] [CrossRef] [PubMed]
- Kato, S; Sato, T; Fujita, H; Kawatani, M; Yamada, Y. Effects of GLP-1 receptor agonist on changes in the gut bacterium and the underlying mechanisms. Sci Rep.;PubMed 2021, 11(1). [Google Scholar] [CrossRef] [PubMed]
- Liu, M; Zhang, Y; Jian, Y; Gu, L; Zhang, D; Zhou, H; et al. The regulations of telomerase reverse transcriptase (TERT) in cancer. Cell Death Dis. 2024, 15(1). [Google Scholar] [CrossRef] [PubMed]
- Chou, R; Fanciullo, GJ; Fine, PG; Adler, JA; Ballantyne, JC; Davies, P; et al. Clinical Guidelines for the Use of Chronic Opioid Therapy in Chronic Noncancer Pain. Journal of Pain 2009, 10(2). [Google Scholar] [CrossRef] [PubMed]
- de Castro, CPM; Cadefau, M; Cuartero, S. The Mutational Landscape of Myeloid Leukaemia in Down Syndrome. Cancers (Basel) PubMed. 2021, 13(16). [Google Scholar] [CrossRef] [PubMed]
- Gialesaki, S; Bräuer-Hartmann, D; Issa, H; Bhayadia, R; Alejo-Valle, O; Verboon, L; et al. RUNX1 isoform disequilibrium promotes the development of trisomy 21-associated myeloid leukemia. Blood 2023, 141(10), 1105–18. [Google Scholar] [CrossRef] [PubMed]
- Liu, X; Shin, N; Koblish, HK; Yang, G; Wang, Q; Wang, K; et al. Selective inhibition of IDO1 effectively regulates mediators of antitumor immunity. Blood PubMed. 2010, 115(17), 3520–30. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



