Submitted:
02 April 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Area, Samples and Ethical Consideration
2.2. Bacterial Isolates Retrieval and Confirmation of IDs
2.3. Antimicrobial Susceptibility and Minimum Inhibitory Test
2.4. Detection and Confirmation of ESBLs
2.5. Statistical Analysis
3. Results
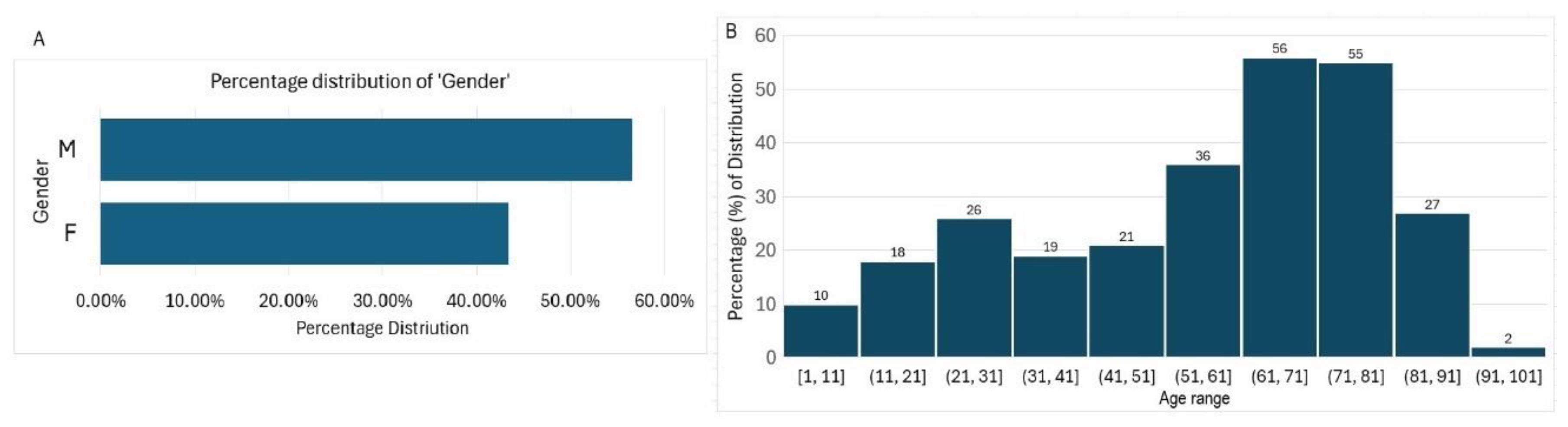
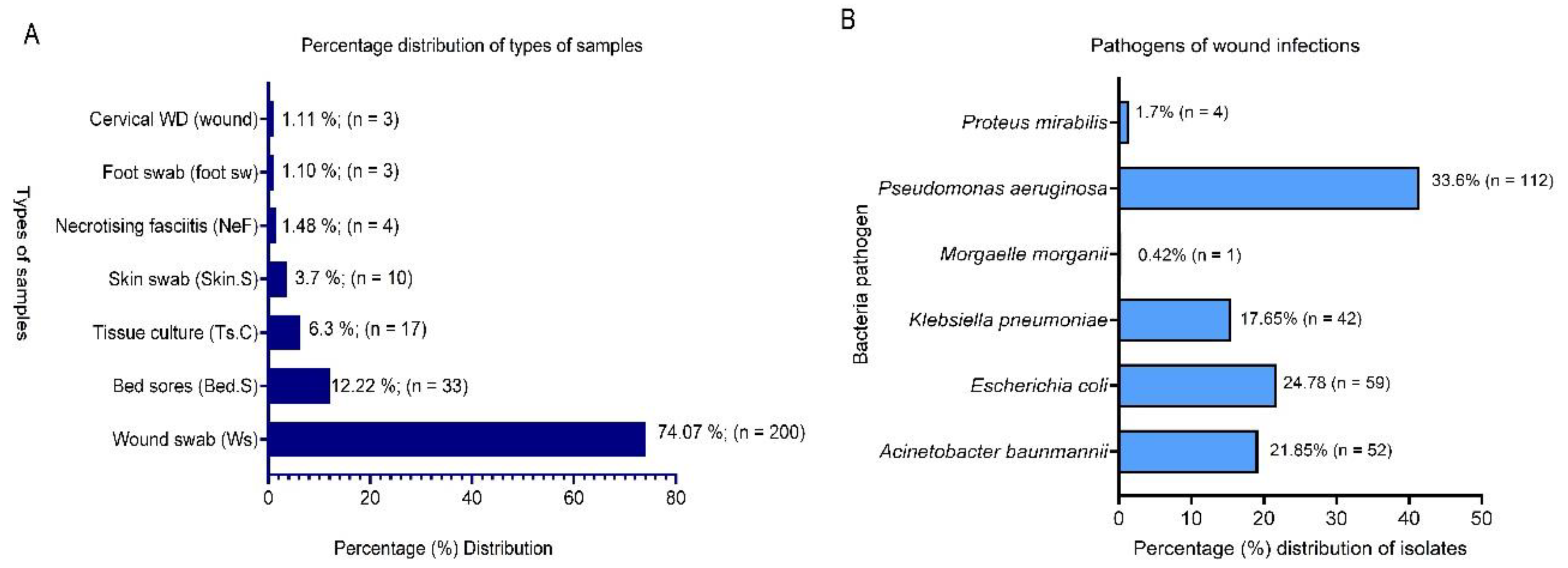
3.1. Demographics, Sample Types and Bacteria Pathogens
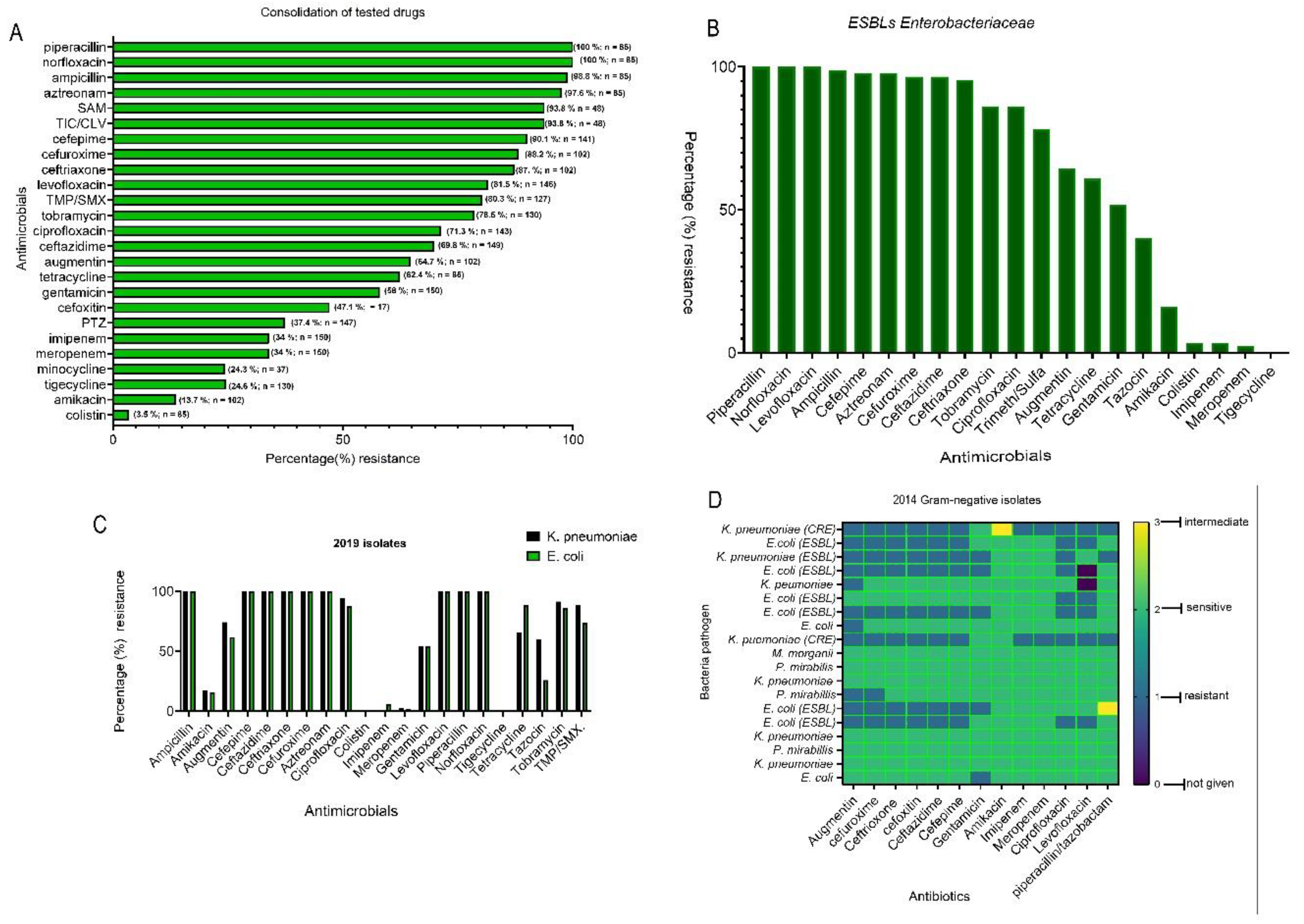
3.2. Consolidated Antimicrobial Susceptibility Profile of the Isolates and ESBLs
3.2.1. Subsubsection
3.3. Antimicrobial Assay of Bacteria Species and the Year of Isolation
3.3.1. Klebsiella Pneumoniae, Escherichia coli
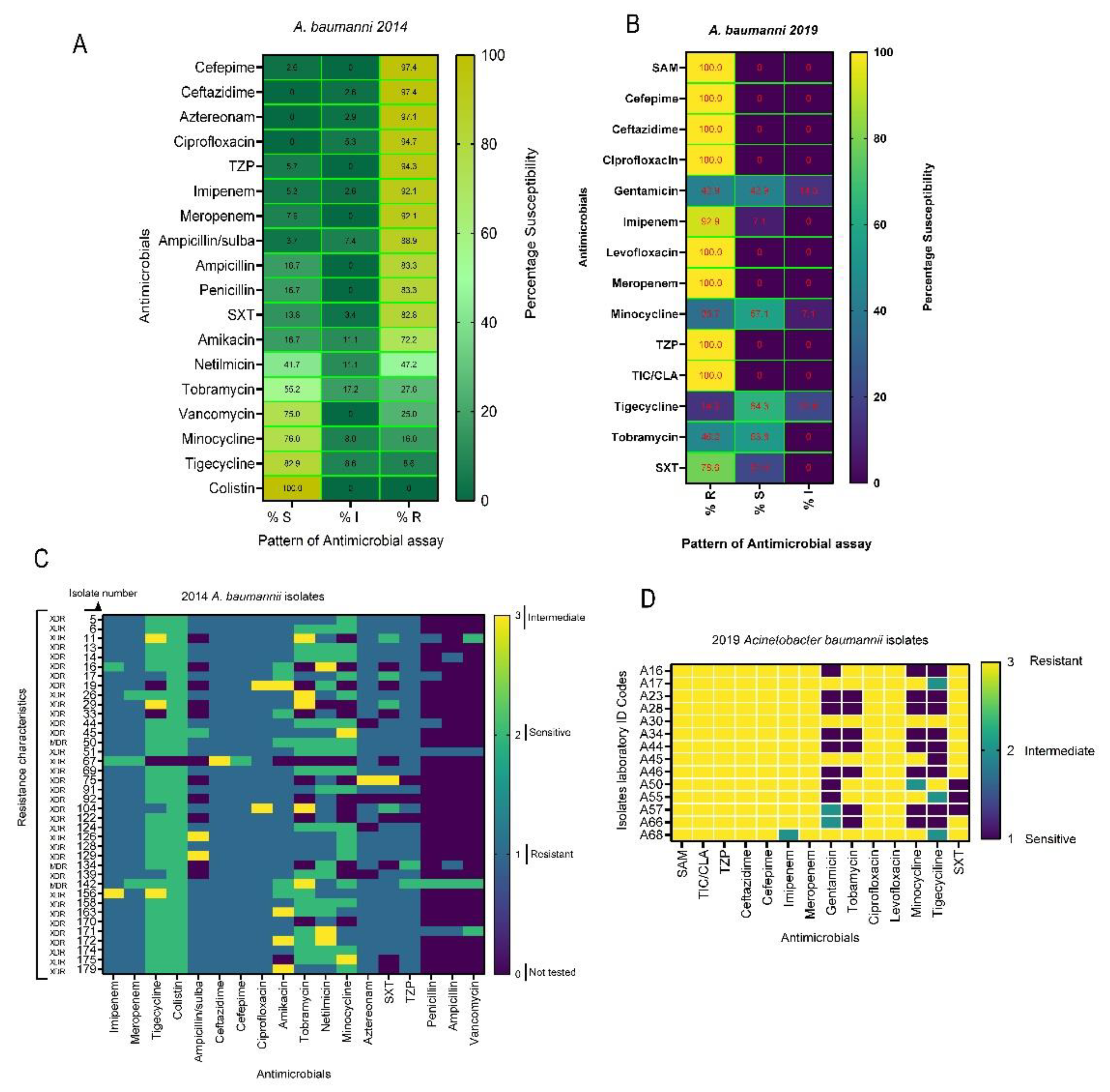
3.3.2. Acinetobacter baumannii
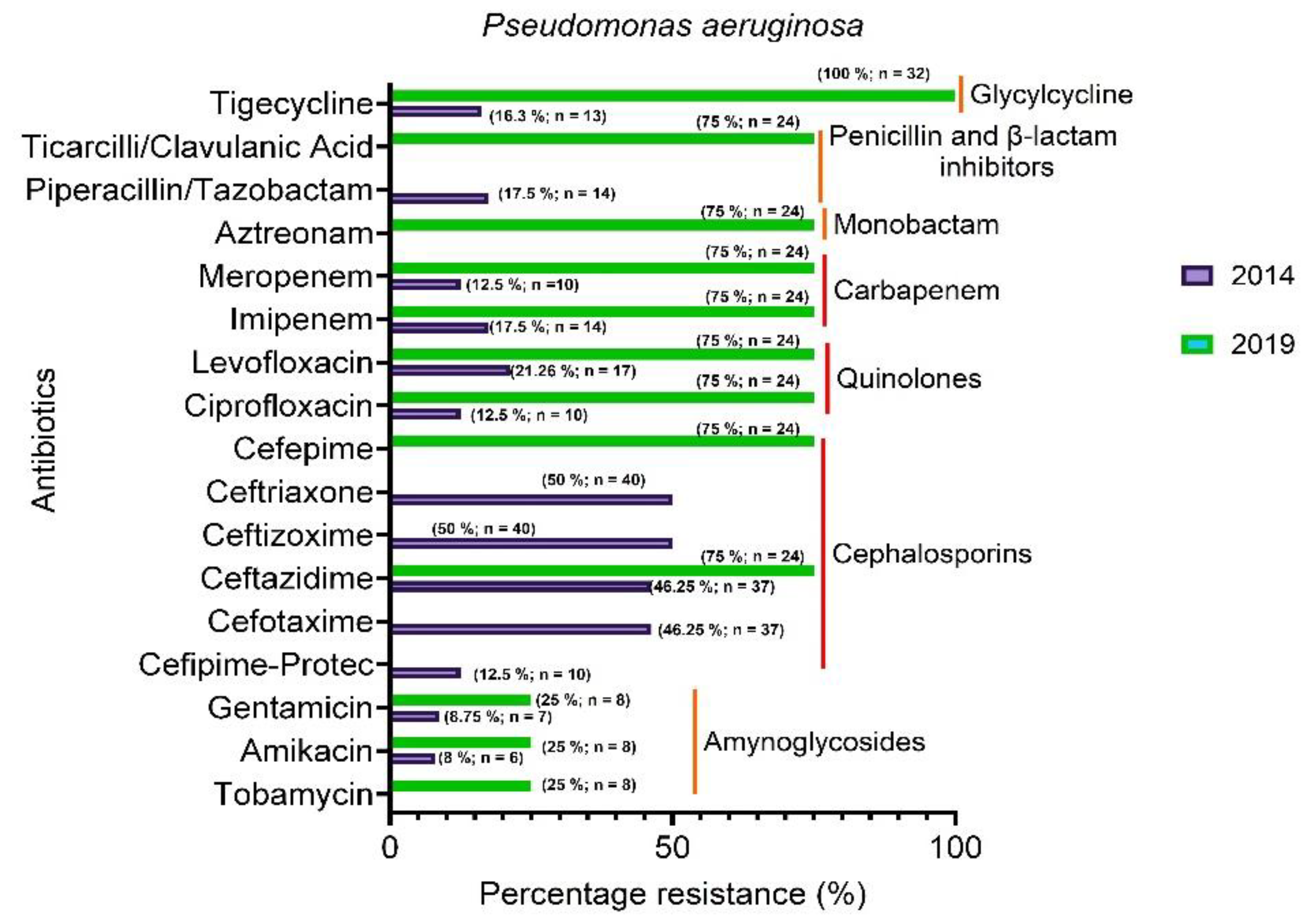
3.3.3. Pseudomonas aeruginosa
3.4. Minimum Inhibitory Concertation of Antibiotics and ESBL Gene Carriage
| Bacteria species (number) | No. positive (n / %) | No. negative (n / %) | p-value | |
|
E. coli (51) |
CTX/CLA |
47 (92) |
6 (8.0) |
0.000173* |
| CAZ/CLA | 43 (84.3) | 10 (15.7) | 0.00019* | |
|
K. pneumoniae (36) |
||||
| CTX/CLA | 32 (89) | 5 (11) | 0.00278* | |
| CAZ/CLA | 34 (94) | 3 (6) | 0.000595* |
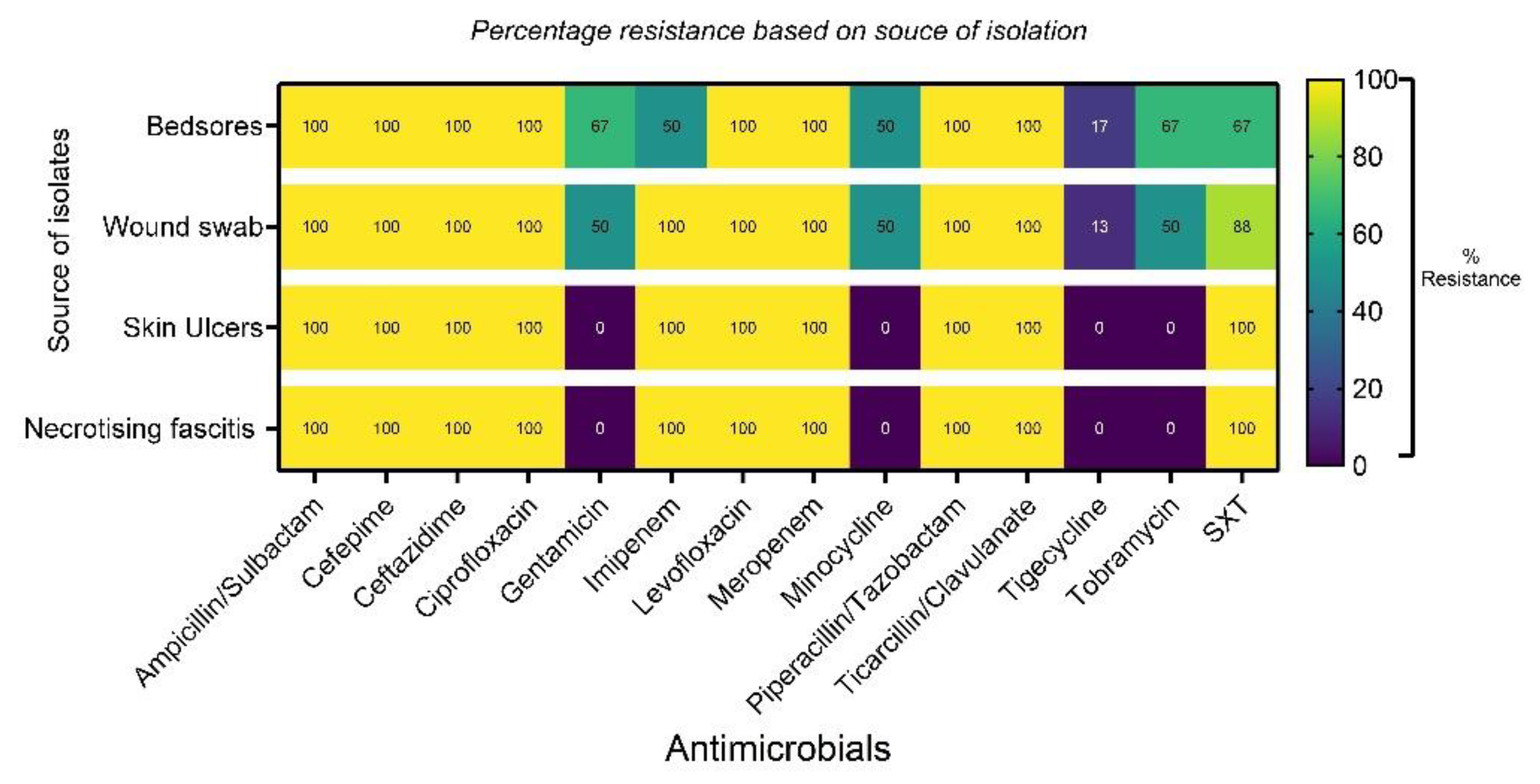
3.5. Antimicrobial Resistance Assay Based on Source of Bacteria Isolation
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MDR | Multidrug resistance |
| XDR | Extensive drug resistance |
| PDR | Pan drug resistance |
| AST | Antimicrobial susceptibility test |
| ESBLs | Extended-Spectrum -Lactamases |
| ESKAPE |
(Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter spp.) |
| (CRE) | Carbapenem-resistant Enterobacteriaceae. |
References
- Khalid, F.; Poulose, C.; Farah, D.F.M.; Mahmood, A.; Elsheikh, A.; Khojah, O.T. Prevalence and Antimicrobial Susceptibility Patterns of Wound and Pus Bacterial Pathogens at a Tertiary Care Hospital in Central Riyadh, Saudi Arabia. Microbiology Research 2024, 15, 2015–2034. [Google Scholar] [CrossRef]
- Zhou, L.; Zheng, H.; Liu, Z.; Wang, S.; Liu, Z.; Chen, F.; Zhang, H.; Kong, J.; Zhou, F.; Zhang, Q. Conductive Antibacterial Hemostatic Multifunctional Scaffolds Based on Ti3C2Tx MXene Nanosheets for Promoting Multidrug-Resistant Bacteria-Infected Wound Healing. ACS Nano 2021, 15, 2468–2480. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K. Human Wounds and Its Burden: An Updated Compendium of Estimates. Advances in Wound Care 2019, 8, 39–48. [Google Scholar] [CrossRef]
- Alharbi, A.S. Bacteriological Profile of Wound Swab and Their Antibiogram Pattern in a Tertiary Care Hospital, Saudi Arabia. Saudi Medical Journal 2022, 43, 1373–1382. [Google Scholar] [CrossRef]
- Ibrahim, M.E. Prevalence of Acinetobacter Baumannii in Saudi Arabia: Risk Factors, Antimicrobial Resistance Patterns and Mechanisms of Carbapenem Resistance. Annals of Clinical Microbiology and Antimicrobials 2019, 18. [Google Scholar] [CrossRef]
- Malaekah, H.M.; Alotaibi, A.E.; Alsebail, R.A.; Alelawi, G.T.; Alsarrani, R.H.; Banjar, W.M. Wound Care Knowledge and Perception of the Saudi General Population in Riyadh Region. Advances in Wound Care 2021, 10, 293–300. [Google Scholar] [CrossRef]
- Robert, A.; Al Dawish, M.; Braham, R.; Musallam, M.; Al Hayek, A.; Al Kahtany, N. Type 2 Diabetes Mellitus in Saudi Arabia: Major Challenges and Possible Solutions. Current Diabetes Reviews 2016, 13, 59–64. [Google Scholar] [CrossRef]
- Li, S.; Renick, P.; Senkowsky, J.; Nair, A.; Tang, L. Diagnostics for Wound Infections. Advances in Wound Care 2021, 10, 317–327. [Google Scholar] [CrossRef]
- Puca, V.; Marulli, R.Z.; Grande, R.; Vitale, I.; Niro, A.; Molinaro, G.; Prezioso, S.; Muraro, R.; Di Giovanni, P. Microbial Species Isolated from Infected Wounds and Antimicrobial Resistance Analysis: Data Emerging from a Three-Years Retrospective Study. Antibiotics 2021, 10, 1162. [Google Scholar] [CrossRef]
- Banawas, S.S.; Alobaidi, A.S.; Dawoud, T.M.; AlDehaimi, A.; Alsubaie, F.M.; Abdel-Hadi, A.; Manikandan, P. Prevalence of Multidrug-Resistant Bacteria in Healthcare-Associated Bloodstream Infections at Hospitals in Riyadh, Saudi Arabia. Pathogens 2023, 12, 1075. [Google Scholar] [CrossRef] [PubMed]
- Baraka, M.A.; Alboghdadly, A.; Alshawwa, S.; Elnour, A.A.; Alsultan, H.; Alsalman, T.; Alaithan, H.; Islam, M.A.; El-Fass, K.A.; Mohamed, Y.; et al. Perspectives of Healthcare Professionals Regarding Factors Associated with Antimicrobial Resistance (AMR) and Their Consequences: A Cross-Sectional Study in Eastern Province of Saudi Arabia. Antibiotics 2021, 10, 878. [Google Scholar] [CrossRef]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; CLSI supplement M100; Clinical and Laboratory Standards Institute: Pittsburgh, PA, USA, 2020; Available online: https://www.nih.org.pk/wp-content/uploads/2021/02/CLSI2020.pdf (accessed on 25th August 2024).
- Clinical Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Testing for Bacteria That Grows Aerobically, 9th ed.; (M07-A9); Clinical Laboratory Standards Institute: Wayne, PA, USA, 2012.
- Bajaj, V.; Gadi, N.; Spihlman, A.P.; Wu, S.C.; Choi, C.H.; Moulton, V.R. Aging, Immunity, and COVID-19: How Age Influences the Host Immune Response to Coronavirus Infections? Frontiers in Physiology 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Akhavizadegan, H.; Hosamirudsari, H.; Pirroti, H.; Akbarpour, S. Antibiotic Resistance: A Comparison between Inpatient and Outpatient Uropathogens, Islamic Republic of Iran. Eastern Mediterranean Health Journal 2020. [Google Scholar] [CrossRef]
- OECD Stemming the Superbug Tide; Just A Few Dollars More, OECD Health Policy Studies, OECD, 2018; ISBN 9789264307582. [CrossRef]
- Swami, S.K.; Banerjee, R. Comparison of Hospital-Wide and Age and Location - Stratified Antibiograms of S. Aureus, E. Coli, and S. Pneumoniae: Age and Location-Stratified Antibiograms. Springer Plus 2013, 2. [Google Scholar] [CrossRef]
- Moges, F.; Eshetie, S.; Abebe, W.; Mekonnen, F.; Dagnew, M.; Endale, A.; Amare, A.; Feleke, T.; Gizachew, M.; Tiruneh, M. High Prevalence of Extended-Spectrum Beta-Lactamase-Producing Gram-Negative Pathogens from Patients Attending Felege Hiwot Comprehensive Specialized Hospital, Bahir Dar, Amhara Region. PLOS ONE 2019, 14, e0215177. [Google Scholar] [CrossRef]
- Bandy, A.; A Wani, F.; H Mohammed, A.; F Dar, U.; R Dar, M.; A Tantry, B. Bacteriological Profile of Wound Infections and Antimicrobial Resistance in Selected Gram-Negative Bacteria. African Health Sciences 2022, 22, 576–586. [Google Scholar] [CrossRef] [PubMed]
- Shimekaw, M.; Tigabu, A.; Tessema, B. Bacterial Profile, Antimicrobial Susceptibility Pattern, and Associated Risk Factors among Patients with Wound Infections at Debre Markos Referral Hospital, Northwest, Ethiopia. The International Journal of Lower Extremity Wounds 2020, 153473462093373. [Google Scholar] [CrossRef]
- El-Saed, A.; Balkhy, H.H.; Alshamrani, M.M.; Aljohani, S.; Alsaedi, A.; Al Nasser, W.; El Gammal, A.; Almohrij, S.A.; Alyousef, Z.; Almunif, S.; et al. High Contribution and Impact of Resistant Gram-Negative Pathogens Causing Surgical Site Infections at a Multi-Hospital Healthcare System in Saudi Arabia, 2007. BMC Infectious Diseases 2020, 20. [Google Scholar] [CrossRef]
- Prastiyanto, M.E.; Darmawati, S.; Daryono, B.S.; Retnaningrum, E. Examining the Prevalence and Antimicrobial Resistance Profiles of Multidrug-Resistant Bacterial Isolates in Wound Infections from Indonesian Patients. Narra J 2024, 4, e980. [Google Scholar] [CrossRef]
- Peacock, S.J.; Parkhill, J.; Brown, N.M. Changing the Paradigm for Hospital Outbreak Detection by Leading with Genomic Surveillance of Nosocomial Pathogens. Microbiology 2018, 164, 1213–1219. [Google Scholar] [CrossRef]
- Eryilmaz-Eren, E.; Yalcin, S.; Ozan, F.; Saatci, E.; Suzuk-Yildiz, S.; Ture, Z.; Kilinc-Toker, A.; Celik, I. An Outbreak Analysis of Wound Infection due to Acinetobacter Baumannii in Earthquake-Trauma Patients. American Journal of Infection Control 2023, 52, 599–604. [Google Scholar] [CrossRef]
- El-Kholy, A.A.; Elanany, M.G.; Sherif, M.M.; Gad, M.A. High Prevalence of VIM, KPC, and NDM Expression among Surgical Site Infection Pathogens in Patients Having Emergency Surgery. Surgical Infections 2018, 19, 629–633. [Google Scholar] [CrossRef]
- Li, L.; Dai, J.; Xu, L.; Chen, Z.; Li, X.; Liu, M.; Wen, Y.; Chen, X. Antimicrobial Resistance and Pathogen Distribution in Hospitalized Burn Patients. Medicine 2018, 97, e11977. [Google Scholar] [CrossRef] [PubMed]
- Madu Emeka, P.; Ineta Badger, L.; Estrella, E.; Belgira An, G.; Ezzat Khal, H. Investigation of Colistin and Polymyxin B on Clinical Extreme Resistant Enterobacteriaceae Isolates for Surveillance Purposes. International Journal of Pharmacology 2022, 18, 699–713. [Google Scholar] [CrossRef]
- Momenah, A.M.; Bakri, R.A.; Jalal, N.A.; Ashgar, S.S.; Felemban, R.F.; Bantun, F.; Hariri, S.H.; Barhameen, A.A.; Faidah, H.; AL-Said, H.M. Antimicrobial Resistance Pattern of Pseudomonas Aeruginosa: An 11-Year Experience in a Tertiary Care Hospital in Makkah, Saudi Arabia. Infection and Drug Resistance 2023, Volume 16, 4113–4122. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Faiz, A. Antimicrobial Resistance Patterns of Pseudomonas Aeruginosa in Tertiary Care Hospitals of Makkah and Jeddah. Annals of Saudi Medicine 2016, 36, 23–28. [Google Scholar] [CrossRef]
| Bacteria pathogen | Wards | ICU | OPD | ER | Grand Total |
| P. aeruginosa | 90 | 12 | 0 | 10 | 112 |
| E.coli | 44 | 4 | 5 | 6 | 59 |
| K. pneumoniae | 25 | 13 | 2 | 2 | 42 |
| A. baumannii | 40 | 8 | 0 | 4 | 52 |
| M. morganii | 1 | 0 | 0 | 0 | 1 |
| P. mirabilis | 1 | 3 | 0 | 0 | 4 |
| Total n; (%) | 201; (74.44%) | 40; (14.81%) | 7; (2.6%) | 22; (8.15%) | 270; (100%) |
| Antimicrobials | Resistant | Sensitive | Intermediate |
| Amoxicillin | - | - | - |
| Ampicillin | 16, >= 32 | - | - |
| Amoxicillin/Clavulanic Acid | >= 32 | 2, 4, 8 | 16 |
| Ampicillin/Sulbactam | 4, 16, ≥ 32 | - | - |
| Piperacillin/Tazobactam | >= 128 | 4 | - |
| Cefalotin | >= 64 | 2, 8 | - |
| Cefoxitin | >= 64 | 4 | - |
| Ceftazidime | >= 64 | 1 | - |
| Ceftriaxone | >= 64 | 1 | - |
| Cefepime | >= 64 | 1 | - |
| Imipenem | >= 16 | <= 0.25 | 2 |
| Meropenem | >= 16 | <= 0.25 | - |
| Amikacin | >= 64 | <= 2 | 8 |
| Gentamicin | >= 16 | <= 1 | - |
| Ciprofloxacin | 2, >= 4 | <= 0.25, 1 | - |
| Tigecycline | >= 8 | <= 0.5, 2 | - |
| Nitrofurantoin | 128, 256 | <= 16 | 64 |
| Trimethroprim/Sulfamethoxazole | >= 320 | <= 20 | - |
| Ticarcillin/Clavulanic Acid | >= 128 | 16 | - |
| Aztreonam | 32, >= 64 | 4 | - |
| Tobramycin | >=16 | <= 1 | - |
| Levofloxacin | >= 8 | 1 | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




