Submitted:
02 April 2026
Posted:
03 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
| Parameter | Description |
| BMI—body mass index [kg/m2] | A value derived from body mass divided by the square of the body height, traditionally used to group individuals as underweight, normal, overweight or obese. |
| FFM—fat free mass [kg], relative to weight [%] | Calculated by subtracting body fat weight from total body weight; also referred to as “lean body mass”. |
| FFMI—fat free mass index [kg/m2] | Describes the amount of fat-free mass (“lean body mass”) in relation to height and weight. Similar to BMI. |
| FM—fat mass [kg], relative to weight [%] | Total amount of fat; percentage of total bodyweight that is fat. |
| FMI—fat mass index [kg/m2] | Describes the amount of fat mass in relation to height and weight. Similar to BMI. |
| TBW—total body water [l], relative to weight [%] | The sum of intracellular water and extracellular water volume; approx. 60% of body weight of a normovolemic individual. |
| Phase angle φ [°] | Calculated by reactance/resistance ratio during bioelectrical impedance measurement. Used as an indicator of cell wall stability. Helpful in health risk assessment. |
| VAT—visceral adipose tissue [l] | Also known as abdominal fat, describes adipose tissue that surrounds the organs in the abdominal cavity. Overdeposition of visceral fat in the abdomen is known as visceral obesity. |
- Body Mass Index (BMI): categorized into starvation (< 16.0 kg/m²), emaciation (16.0–16.99 kg/m²), underweight (17.0–18.49 kg/m²), normal weight (18.5–24.99 kg/m²), overweight (25.0–29.99 kg/m²), obesity class I (30.0–34.99 kg/m²), obesity class II (35.0–39.99 kg/m²), and obesity class III (> 40.0 kg/m²).
- Fat Mass Index (FMI): divided into four levels: low, normal, elevated, and high, as specified in the database.
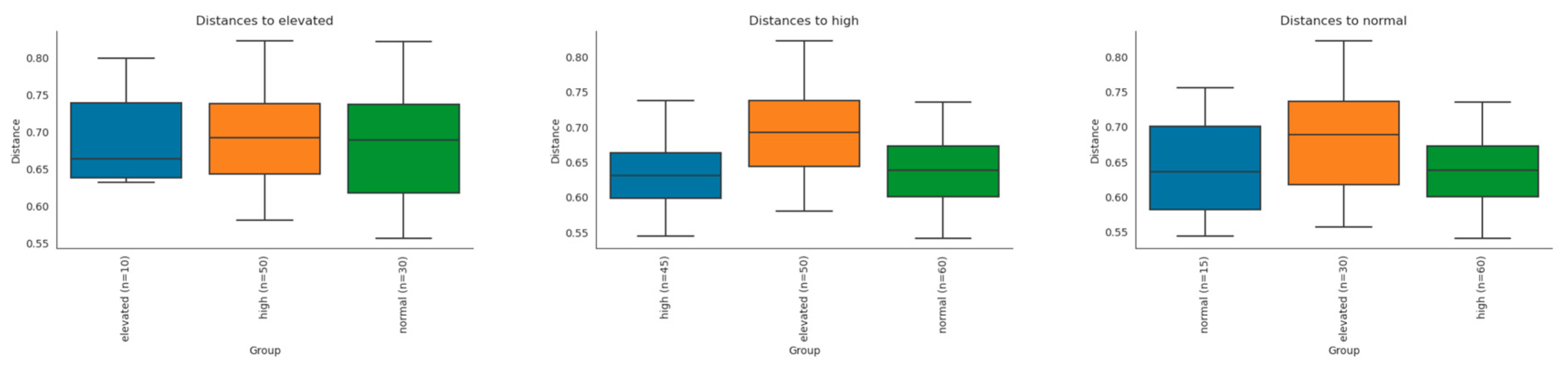
- Visceral Adipose Tissue (VAT): classified into three categories: normal, elevated, and high.
- Phase Angle: grouped into three ranges: decreased (< 5°), normal (5–7°), and increased (> 7°).
- Fat-Free Mass Index (FFMI): segmented by gender with specific cutoff points.
- ○
- For females: below average (< 14 kg/m²), good muscle mass (14–18 kg/m²), and high muscle mass (> 18 kg/m²).
- ○
- For males: below average (< 18 kg/m²), average (18–25 kg/m²), and high (> 25 kg/m²).
3. Results
4. Discussion
5. Conclusions
6. Strengths
7. Limitations
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Feret W, Safranow K, Ciechanowski K, Kwiatkowska Ewa. How Is Body Composition and Nutrition Status Associated with Erythropoietin Response in Hemodialyzed Patients? A Single-Center Prospective Cohort Study. J. Clin. Med.2022, 11(9), 2426;
- Kalantar-Zadeh K, Ikizler TA, Block G, Avram MM, Kopple JD. Malnutrition–inflammation complex syndrome in dialysis patients: causes and consequences. Am J Kidney Dis. 2003;42(5):864–81.
- Meijers B, Evenepoel P. The gut–kidney axis: indoxyl sulfate, p-cresyl sulfate and CKD progression. Nephrol Dial Transplant. 2011;26(3):759–61.
- Vaziri ND, Wong J, Pahl M, et al. Chronic kidney disease alters intestinal microbial flora and impairs intestinal barrier. Kidney Int. 2013;83(2):308–15.
- Ramezani A, Raj DS. The gut microbiome, kidney disease, and targeted interventions. Nat Rev Nephrol. 2014;10(7):383–97.
- Davenport A. Improving the outcomes of vascular access infections in haemodialysis patients. Nephrol Dial Transplant. 2014;29(11):1941–6.
- Evenepoel P, Poesen R, Meijers B. The gut–kidney axis. Pediatr Nephrol. 2017;32(11):2005–14.
- Vaziri ND. Emerging role of gut microbiome in chronic kidney disease: current evidence and potential therapies. Nephron. 2020;144(6):527–33.
- Rysz J, Gluba-Brzózka A, et al. Gut microbiota, a regulator of systemic inflammation in chronic kidney disease. Int J Mol Sci. 2021;22(18):9995.
- Kaysen GA. The microinflammatory state in uremia: causes and potential consequences. J Am Soc Nephrol. 2001;12(7):1549–57.
- Stenvinkel P. Inflammation in end-stage renal disease: the hidden enemy. Nephrol Dial Transplant. 2006;21(11):3163–5.
- Kalantar-Zadeh K, et al. Management of patients at risk of protein–energy wasting: ESPEN guideline on clinical nutrition in chronic kidney disease. Clin Nutr. 2019;38(1):355–72.
- Tsukamoto T., Matsubara T., Akashi Y., Kondo M., Yanagita M. Annual Iron Loss Associated with Hemodialysis. Am. J. Nephrol. 2016;43:32–38. doi: 10.1159/000444335. [DOI].
- Harvinder G.S., Swee W.C.S., Karupaiah T., Sahathevan S., Chinna K., Ahmad G., Bavanandan S., Goh B.L. Dialysis Malnutrition and Malnutrition Inflammation Scores: Screening Tools for Prediction of Dialysis—Related Protein-Energy Wasting in Malaysia. Asia Pac. J. Clin. Nutr.
- Wehedy E, Shatat IF, Khodor Sal. The Human Microbiome in Chronic Kidney Disease: A Double-Edged Sword. Front Med (Lausanne) 2022 Jan 17:8:790783.
- Huang H, Chen M. Exploring the Preventive and Therapeutic Mechanisms of Probiotics in Chronic Kidney Disease through the Gut-Kidney Axis. J Agric Food Chem 2024 Apr 17;72(15):8347-8364.
- Pantazi AC, Kassim Kassim MA, Nori W et al. Clinical Perspectives of Gut Microbiota in Patients with Chronic Kidney Disease and End-Stage Kidney Disease: Where Do We Stand? Biomedicines 2023 Sep 7;11(9):2480.
- Laiola M, Koppe L, Larabi A. Toxic microbiome and progression of chronic kidney disease: insights from a longitudinal CKD-Microbiome Study. Gut . 2025 Sep 8;74(10):1624-1637.
- Seca mBCA User Manual. [(accessed on 7 March 2020)]. Available online: https://www.seca.com/fileadmin/documents/manual/seca_man_525_535_en.pdf.
- Coin A, Sergi G, Minicuci N et al. Fat-free mass and fat mass reference values by dual-energy X-ray absorptiometry (DEXA) in a 20-80 year-old Italian population Clin Nutr . 2008 Feb;27(1):87-94.
- Kalantar-Zadeh K, Kopple JD, Humphreys MH, Block G. Comparing outcome predictability of markers of malnutrition–inflammation complex syndrome in haemodialysis patients Nephrol Dial Transplant (2004) 19: 1507–1519.
- Li, H. (2018). Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics, 34(18), 3094.
- De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M., & Van Broeckhoven, C. (2018). NanoPack: visualizing and processing long-read sequencing data. Bioinformatics, 34(15), 2666–2669.
- Ewels, P., Magnusson, M., Lundin, S., & Käller, M. (2016). MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics, 32(19), 3047–3048.
- Rognes, T., Flouri, T., Nichols, B., Quince, C., & Mahé, F. (2016). VSEARCH: A versatile open source tool for metagenomics. PeerJ, 2016(10), e2584.
- Robeson, M. S., O’Rourke, D. R., Kaehler, B. D., Ziemski, M., Dillon, M. R., Foster, J. T., & Bokulich, N. A. (2021). RESCRIPt: Reproducible sequence taxonomy reference database management. PLOS Computational Biology, 17(11), e1009581.
- Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution, 30(4), 772–780.
- Price, M. N., Dehal, P. S., & Arkin, A. P. (2009). FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Molecular Biology and Evolution, 26(7), 1641–1650.
- Faith, D. P. (1992). Conservation evaluation and phylogenetic diversity. Biological Conservation, 61(1), 1–10.
- Lozupone, C., & Knight, R. (2005). UniFrac: A new phylogenetic method for comparing microbial communities. Applied and Environmental Microbiology, 71(12), 8228–8235.
- Lozupone, C. A., Hamady, M., Kelley, S. T., & Knight, R. (2007). Quantitative and qualitative β diversity measures lead to dif-ferent insights into factors that structure microbial communities. Applied and Environmental Microbiology, 73(5), 1576–1585.
- Bray, J. R., & Curtis, J. T. (1957). An Ordination of the Upland Forest Communities of Southern Wisconsin. Ecological Monographs, 27(4), 325–349.
- Shannon, C. E. (1948). A Mathematical Theory of Communication. Bell System Technical Journal, 27(3), 379–423.
- Simpson, E. H. (1949). Measurement of Diversity. Nature 1949 163:4148, 163(4148), 688–688.
- Vázquez-Baeza, Y., Pirrung, M., Gonzalez, A., & Knight, R. (2013). EMPeror: A tool for visualizing high-throughput microbial community data. GigaScience, 2(1).
- Vázquez-Baeza, Y., Gonzalez, A., Smarr, L., McDonald, D., Morton, J. T., Navas-Molina, J. A., & Knight, R. (2017). Bringing the Dynamic Microbiome to Life with Animations. Cell Host & Microbe, 21(1), 7–10.
- Ondov, B. D., Bergman, N. H., & Phillippy, A. M. (2011). Interactive metagenomic visualization in a Web browser. BMC Bioin-formatics, 12(1), 385.
- Krukowski, H., Valkenburg, S., Madella, A. M., Garssen, J., van Bergenhenegouwen, J., Overbeek, S. A., Huys, G. R. B., Raes, J., & Glorieux, G. (2023a). Gut microbiome studies in CKD: opportunities, pitfalls and therapeutic potential. Nature Reviews. Nephrology, 19(2), 87–101.
- Lim, X., Ooi, L., Ding, U., Wu, H. H. L., & Chinnadurai, R. (2024). Gut Microbiota in Patients Receiving Dialysis: A Review. Pathogens (Basel, Switzerland), 13(9).
- Vaziri, N. D., Wong, J., Pahl, M., Piceno, Y. M., Yuan, J., Desantis, T. Z., Ni, Z., Nguyen, T. H., & Andersen, G. L. (2013). Chronic kidney disease alters intestinal microbial flora. Kidney International, 83(2), 308–315.
- Yang, T., Richards, E. M., Pepine, C. J., & Raizada, M. K. (2018). The gut microbiota and the brain-gut-kidney axis in hyperten-sion and chronic kidney disease. Nature Reviews. Nephrology, 14(7), 442–456.
- Stadlbauer, V., Horvath, A., Ribitsch, W., Schmerböck, B., Schilcher, G., Lemesch, S., Stiegler, P., Rosenkranz, A. R., Fickert, P., & Leber, B. (2017). Structural and functional differences in gut microbiome composition in patients undergoing haemo-dialysis or peritoneal dialysis. Scientific Reports 2017 7:1, 7(1), 15601.
- Vriend, E. M. C., Galenkamp, H., Herrema, H., Nieuwdorp, M., van den Born, B. J. H., & Verhaar, B. J. H. (2024). Machine learning analysis of sex and menopausal differences in the gut microbiome in the HELIUS study. Npj Biofilms and Microbiomes 2024 10:1, 10(1), 152.
- Ghosh, T. S., Shanahan, F., & O’Toole, P. W. (2022). The gut microbiome as a modulator of healthy ageing. Nature Reviews Gastroenterology & Hepatology 2022 19:9, 19(9), 565–584.
- Procházková, N., Laursen, M. F., La Barbera, G., Tsekitsidi, E., Jørgensen, M. S., Rasmussen, M. A., Raes, J., Licht, T. R., Dragsted, L. O., & Roager, H. M. (2024a). Gut physiology and environment explain variations in human gut microbiome composition and metabolism. Nature Microbiology 2024 9:12, 9(12), 3210–3225.
- Keshet, A., & Segal, E. (2024a). Identification of gut microbiome features associated with host metabolic health in a large pop-ulation-based cohort. Nature Communications 2024 15:1, 15(1), 9358.
- Shantaram, D., Hoyd, R., Blaszczak, A. M., Antwi, L., Jalilvand, A., Wright, V. P., Liu, J., Smith, A. J., Bradley, D., Lafuse, W., Liu, Y. Z., Williams, N. F., Snyder, O., Wheeler, C., Needleman, B., Brethauer, S., Noria, S., Renton, D., Perry, K. A., … Hsueh, W. A. (2024). Obesity-associated microbiomes instigate visceral adipose tissue inflammation by recruitment of distinct neu-trophils. Nature Communications 2024 15:1, 15(1), 5434.
- Keshet, A., & Segal, E. (2024b). Identification of gut microbiome features associated with host metabolic health in a large pop-ulation-based cohort. Nature Communications 2024 15:1, 15(1), 9358.
- Keshet, A., & Segal, E. (2024c). Identification of gut microbiome features associated with host metabolic health in a large pop-ulation-based cohort. Nature Communications 2024 15:1, 15(1), 9358.
- Procházková, N., Laursen, M. F., La Barbera, G., Tsekitsidi, E., Jørgensen, M. S., Rasmussen, M. A., Raes, J., Licht, T. R., Dragsted, L. O., & Roager, H. M. (2024b). Gut physiology and environment explain variations in human gut microbiome composition and metabolism. Nature Microbiology 2024 9:12, 9(12), 3210–3225.
- Evenepoel, P., Stenvinkel, P., Shanahan, C., & Pacifici, R. (2023). Inflammation and gut dysbiosis as drivers of CKD-MBD. Nature Reviews. Nephrology, 19(10), 646–657.
- Krukowski, H., Valkenburg, S., Madella, A. M., Garssen, J., van Bergenhenegouwen, J., Overbeek, S. A., Huys, G. R. B., Raes, J., & Glorieux, G. (2023b). Gut microbiome studies in CKD: opportunities, pitfalls and therapeutic potential. Nature Reviews. Nephrology, 19(2), 87–101.
| Variable | Overall participants |
| Overall participants | n = 96 |
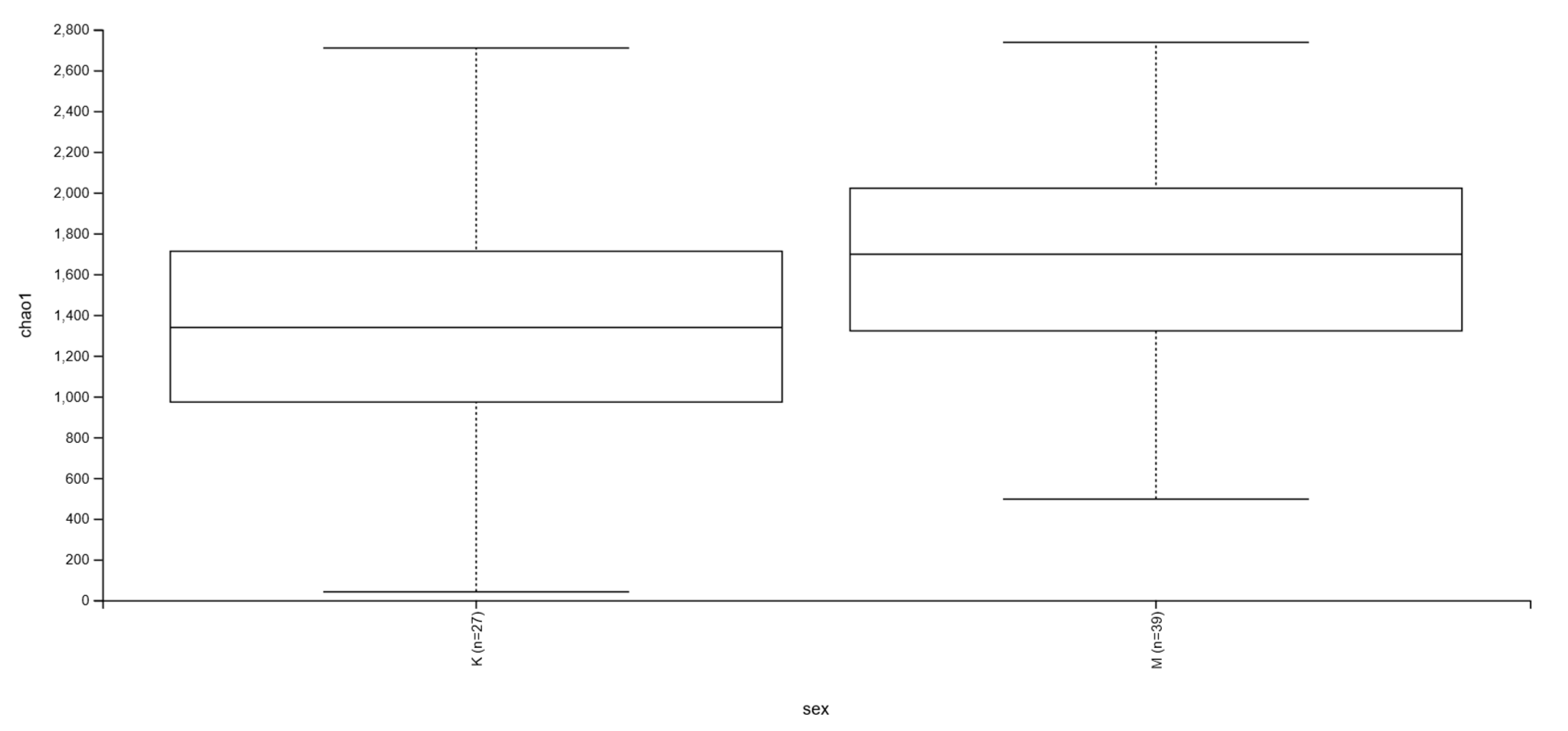
| Women | n = 34 (35.4%) |
| Men | n = 62 (64.6%) |
| Age [years] | Median: 66; IQR = 51.0-73.3 |
| Dialysis vintage [months] | Median: 25.5; IQR = 14.0-46.3 |
| Patients’ nutrition by BMI [%] | Underweight: n = 0 (0.0%); Normal weight: n = 43 (44.8%); Overweight: n = 31 (32.3%); Obese: n = 22 (22.9%) |
| FMI [kg/m²] | Median: 8.2; IQR = 4.0-11.5 |
| FFMI [kg/m²] | Median: 19.3; IQR = 17.2-21.4 |
| SMM [kg] | Median: 25.1; IQR = 19.9-29.8 |
| VAT [L] | Median: 2.3; IQR = 0.9-4.4 |
| PhA | Median: 4.5; IQR = 3.9-5.5 |
| TBW [%] | Median: 52.7; IQR = 46.0-61.1 |
| ERI [IU/kg/g/dL/week] | Median: 9.27; IQR = 4.14-18.68 |
| IL-6 [pg/mL] | Median: 6.92; IQR = 3.90-12.29 |
| hsCRP [mg/L] | Median: 4.5; IQR = 2.3-17.3 |
| IL-1β [pg/mL] | Median: 0.04; IQR = 0.00-0.16 |
| TNF-α [pg/mL] | Median: 2.77; IQR = 2.24-3.65 |
| Albumin [g/L] | Median: 40; IQR = 37.0-41.0 |
| Transferrin [g/L] | Median: 1.76; IQR = 1.55-2.02 |
| Kt/V | Mean: 1.29 ± 0.25 |
| Total MIS score | Median: 5; IQR = 4.0-9.0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




