Submitted:
01 April 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
Keywords:
Introduction
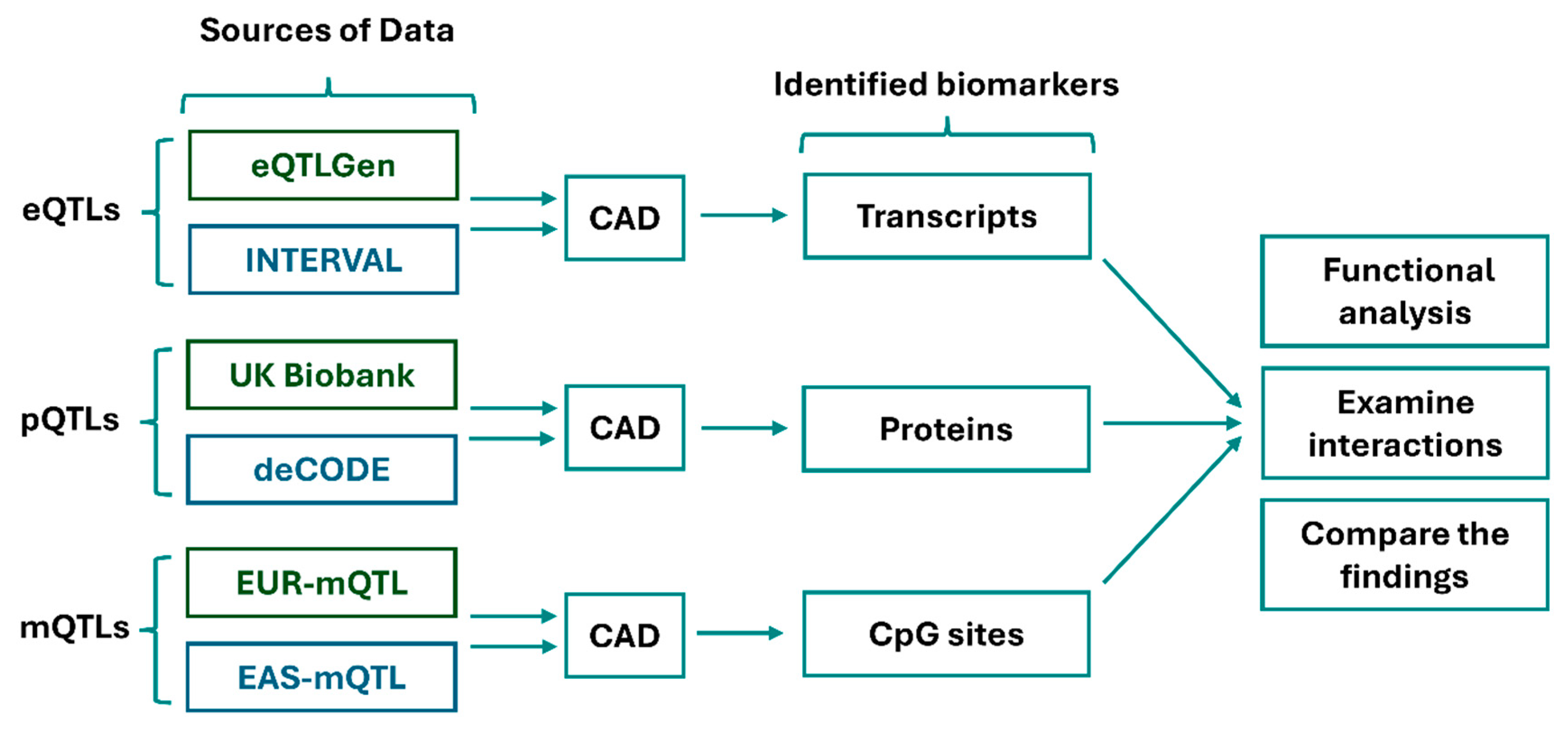
Methods
GWAS Data:
eQTL Data:
pQTL Data:
mQTL Data:
Analyses:
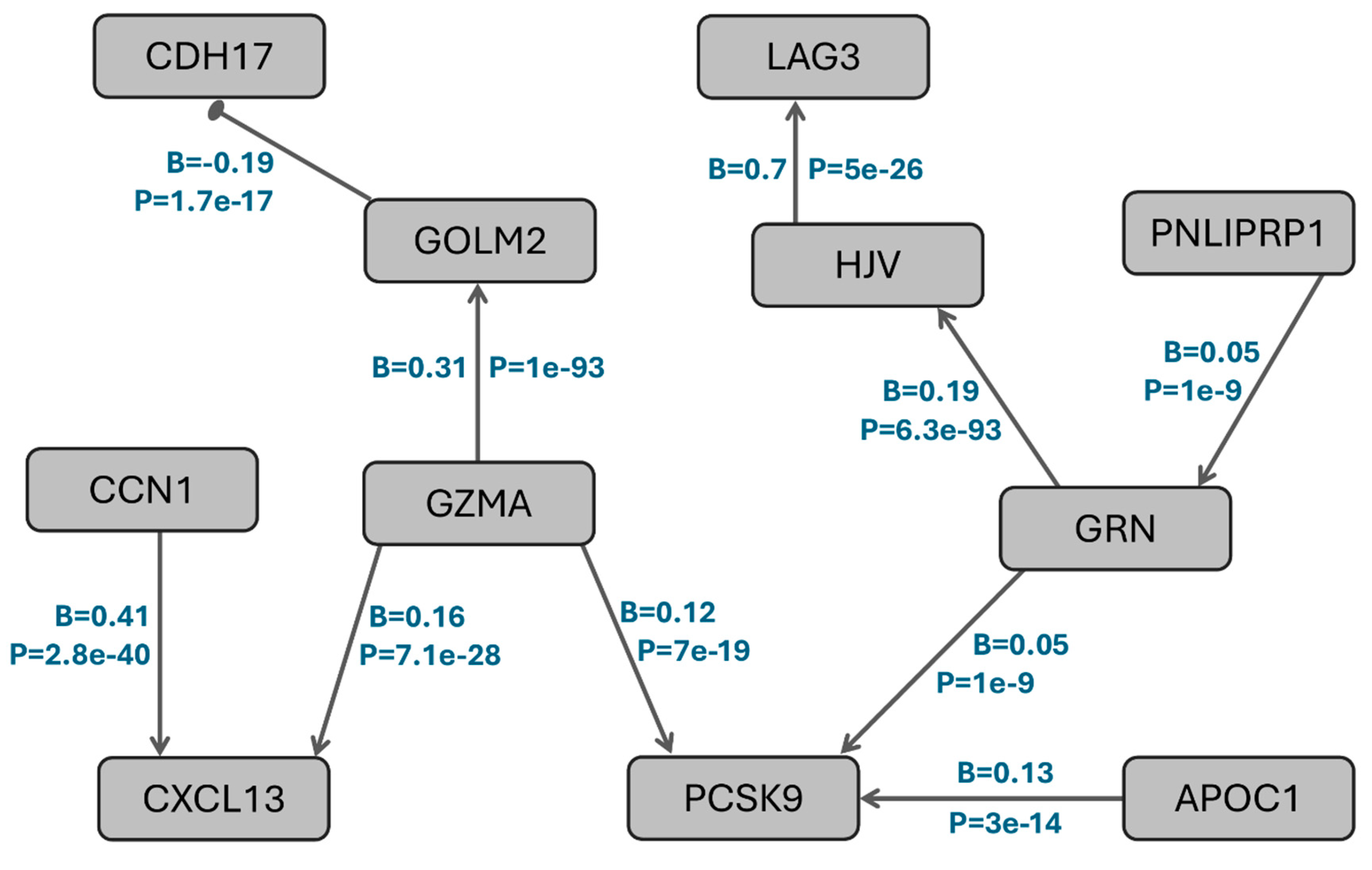
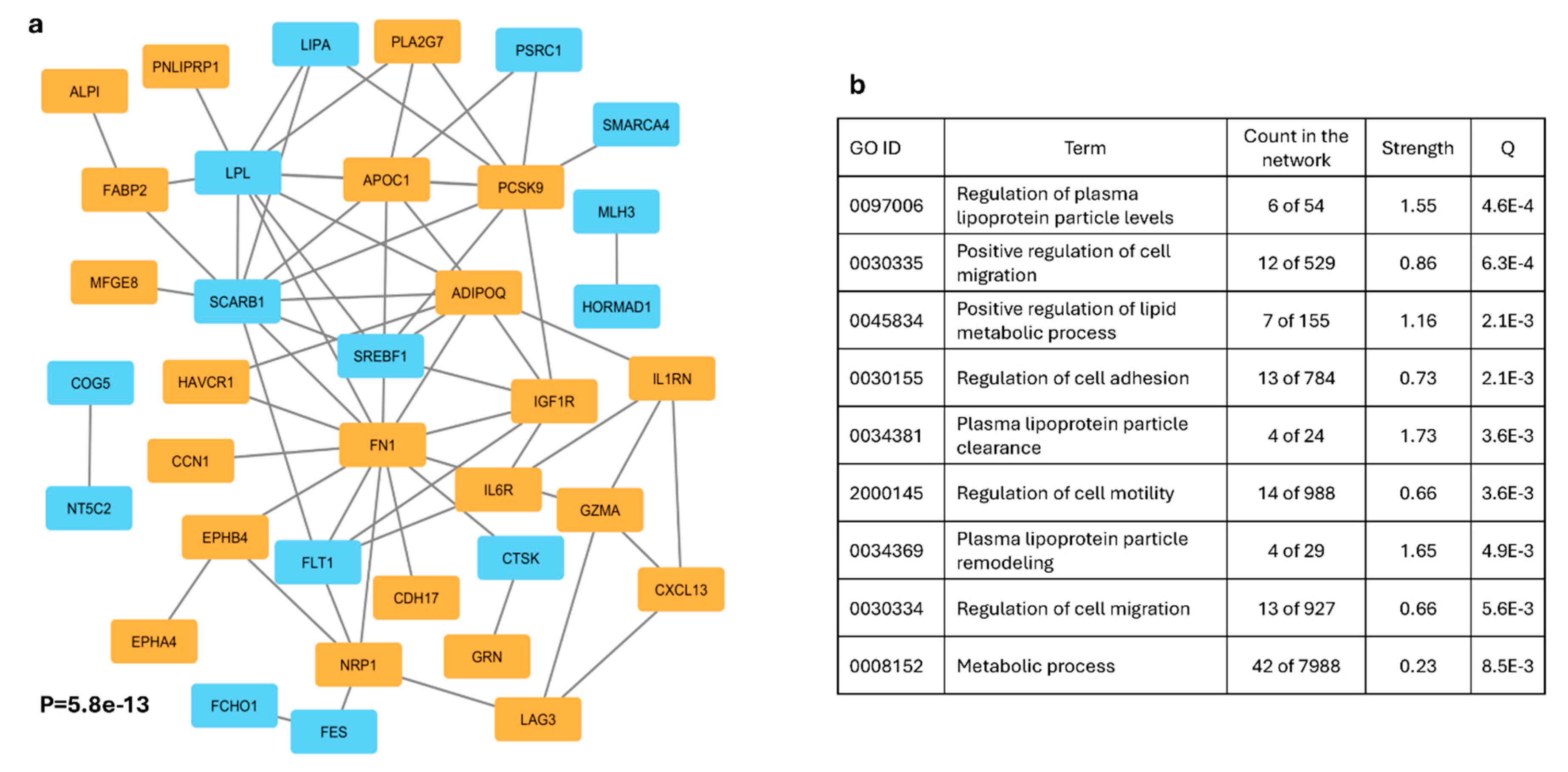
Functional Interaction:
Conditional Analysis:
Results
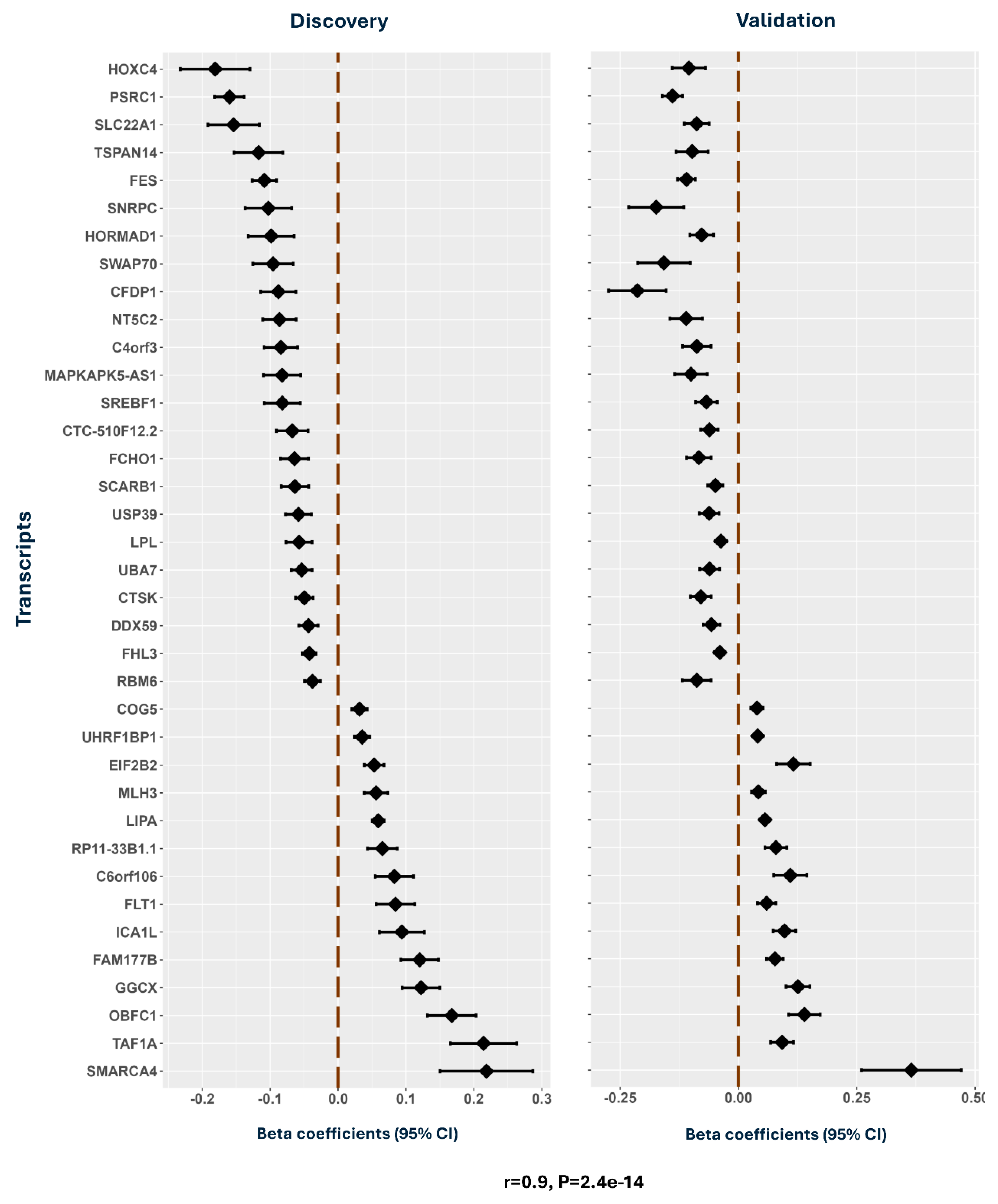
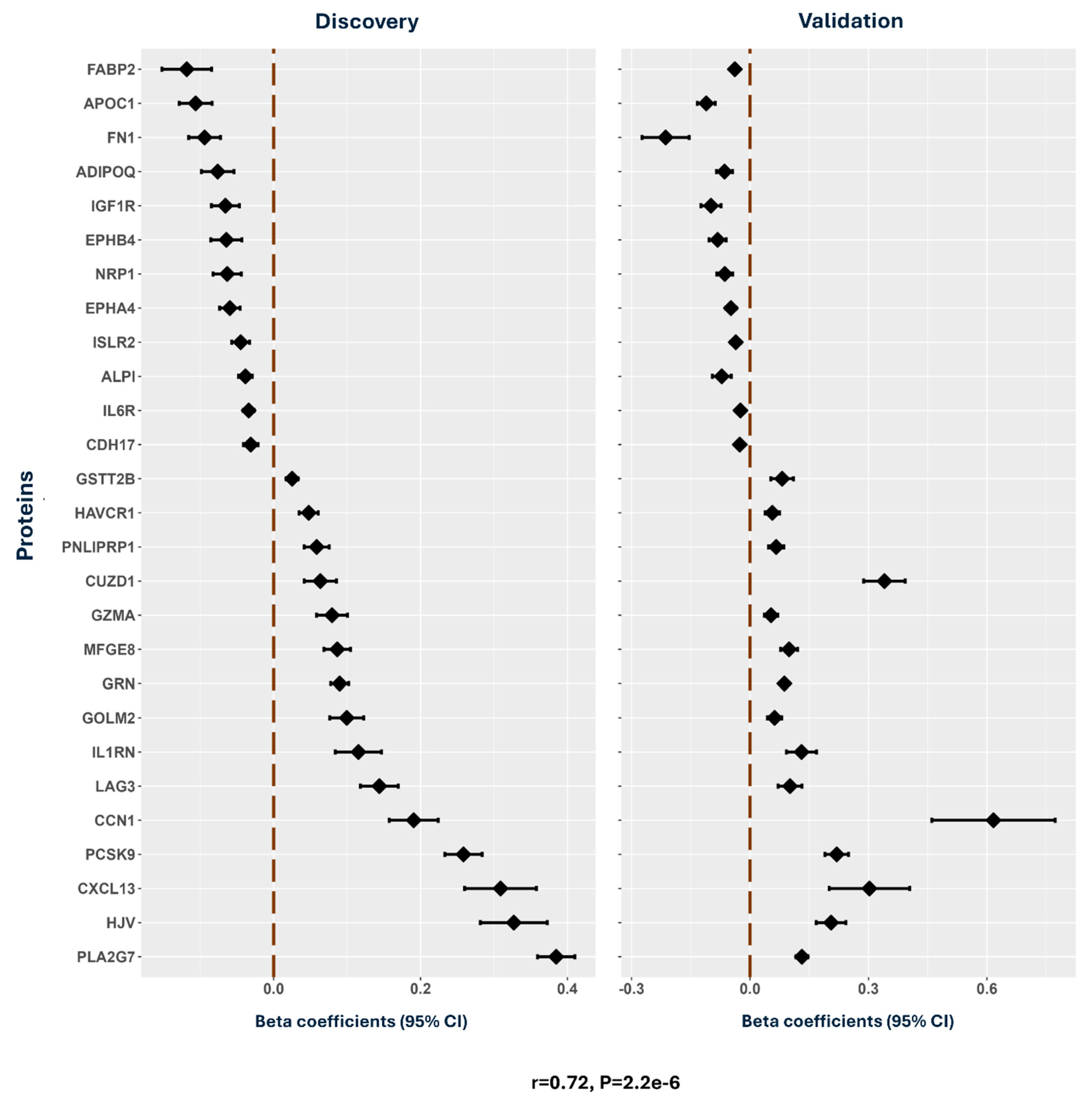
Transcriptome-Wide Association Study:
Discussion
Supplementary Materials
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Aragam, K. G.; et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Nat. Genet. 2022, 54, 1803–1815. [Google Scholar] [CrossRef] [PubMed]
- Võsa, U.; et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 2021, 53, 1300–1310. [Google Scholar] [CrossRef]
- Xu, Y.; et al. An atlas of genetic scores to predict multi-omic traits. Nature 2023, 616, 123–131. [Google Scholar] [CrossRef]
- Sun, B. B.; et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 2023, 622, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Ferkingstad, E.; et al. Large-scale integration of the plasma proteome with genetics and disease. Nat Genet 2021, 53, 1712–1721. [Google Scholar] [CrossRef]
- Hatton, A. A.; et al. Genetic control of DNA methylation is largely shared across European and East Asian populations. Nat. Commun. 2024, 15, 2713. [Google Scholar] [CrossRef]
- Zhu, Z.; et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 2018, 9, 224. [Google Scholar] [CrossRef]
- Szklarczyk, D.; et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef]
- Johanns, M.; et al. Genetic deletion of soluble 5′-nucleotidase II reduces body weight gain and insulin resistance induced by a high-fat diet. Mol. Genet. Metab. 2019, 126, 377–387. [Google Scholar] [CrossRef] [PubMed]
- Messina, G.; et al. The human Cranio Facial Development Protein 1 (Cfdp1) gene encodes a protein required for the maintenance of higher-order chromatin organization. Sci. Rep. 2017, 7, 45022. [Google Scholar] [CrossRef] [PubMed]
- Giardoglou, P.; Deloukas, P.; Dedoussis, G.; Beis, D. Cfdp1 Is Essential for Cardiac Development and Function. Cells 2023, 12, 1994. [Google Scholar] [CrossRef]
- Wensink, A. C.; Hack, C. E.; Bovenschen, N. Granzymes Regulate Proinflammatory Cytokine Responses. J. Immunol. 2015, 194, 491–497. [Google Scholar] [CrossRef]
- Saeedi-Boroujeni, A.; et al. Progranulin (PGRN) as a regulator of inflammation and a critical factor in the immunopathogenesis of cardiovascular diseases. J. Inflamm. 2023, 20, 1. [Google Scholar] [CrossRef]
- Berbée, J. F. P.; et al. Apolipoprotein CI enhances the biological response to LPS via the CD14/TLR4 pathway by LPS-binding elements in both its N- and C-terminal helix. J. Lipid Res. 2010, 51, 1943–1952. [Google Scholar] [CrossRef]
- Glerup, S.; Schulz, R.; Laufs, U.; Schlüter, K.-D. Physiological and therapeutic regulation of PCSK9 activity in cardiovascular disease. Basic Res. Cardiol. 2017, 112, 32. [Google Scholar] [CrossRef] [PubMed]
- Lisignoli, G.; et al. IL1beta and TNFalpha differently modulate CXCL13 chemokine in stromal cells and osteoblasts isolated from osteoarthritis patients: evidence of changes associated to cell maturation. Exp. Gerontol. 2004, 39, 659–665. [Google Scholar] [CrossRef]
- Barutcu, A. R.; et al. SMARCA4 regulates gene expression and higher-order chromatin structure in proliferating mammary epithelial cells. Genome Res. 2016, 26, 1188–1201. [Google Scholar] [CrossRef] [PubMed]
- Salma, N.; Xiao, H.; Mueller, E.; Imbalzano, A. N. Temporal recruitment of transcription factors and SWI/SNF chromatin-remodeling enzymes during adipogenic induction of the peroxisome proliferator-activated receptor gamma nuclear hormone receptor. Mol. Cell. Biol. 2004, 24, 4651–4663. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; et al. (Apo)Lipoprotein Profiling with Multi-Omics Analysis Identified Medium-HDL-Targeting PSRC1 with Therapeutic Potential for Coronary Artery Disease. Adv. Sci. 2025, 12, 2413491. [Google Scholar] [CrossRef]
- Karamanavi, E.; et al. The FES Gene at the 15q26 Coronary-Artery-Disease Locus Inhibits Atherosclerosis. Circ. Res. 2022, 131, 1004–1017. [Google Scholar] [CrossRef]
- Wegler, C.; et al. Global variability analysis of mRNA and protein concentrations across and within human tissues. NAR Genomics Bioinforma 2020, 2, lqz010. [Google Scholar] [CrossRef] [PubMed]
- Vogel, C.; Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 2012, 13, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Edfors, F.; et al. Gene-specific correlation of RNA and protein levels in human cells and tissues. Mol. Syst. Biol. 2016, 12, 883. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; et al. Comparative studies of 2168 plasma proteins measured by two affinity-based platforms in 4000 Chinese adults. Nat. Commun. 2025, 16, 1869. [Google Scholar] [CrossRef]
- Raffield, L. M.; et al. Comparison of Proteomic Assessment Methods in Multiple Cohort Studies. PROTEOMICS 2020, 20, 1900278. [Google Scholar] [CrossRef]
- Pietzner, M.; et al. Synergistic insights into human health from aptamer- and antibody-based proteomic profiling. Nat. Commun. 2021, 12, 6822. [Google Scholar] [CrossRef]
- Rooney, M. R.; et al. Comparison of Proteomic Measurements Across Platforms in the Atherosclerosis Risk in Communities (ARIC) Study. Clin. Chem. 2023, 69, 68–79. [Google Scholar] [CrossRef]
- Villicaña, S.; et al. Genetic impacts on DNA methylation help elucidate regulatory genomic processes. Genome Biol. 2023, 24, 176. [Google Scholar] [CrossRef]
- Kanchibhotla, S. C.; et al. Heritability of Gene Expression Measured from Peripheral Blood in Older Adults. Genes 2024, 15, 495. [Google Scholar] [CrossRef]
- Drouard, G.; et al. Twin Study Provides Heritability Estimates for 2321 Plasma Proteins and Assesses Missing SNP Heritability. J. Proteome Res. 24, 2689–2697. [CrossRef]
- Christiansen, S. N.; et al. Reproducibility of the Infinium methylationEPIC BeadChip assay using low DNA amounts. Epigenetics 17, 1636–1645. [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



