Submitted:
31 March 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
The alkaline pectate lyase A from Paenibacillus barcinonensis, encoded by pelA (GenBank accession no. CAB40884), is an enzyme with high activity on pectin and potential application in sustainable industrial biotechnology. In this study, pelA was expressed in Saccharomyces cerevisiae by using different domains of the cell wall protein Pir4 as translational fusion partners. Given the presence of five potential N-glycosylation sites in the amino acid sequence coded by pelA, two of them in conserved regions of class III pectate lyases, the effect of glycosylation on the enzymatic activity of the recombinant enzyme was investigated by expressing the recombinant fusion proteins in both, standard and glycosylation deficient strains of S. cerevisiae. Correct targeting of the recombinant fusion proteins was confirmed by Western blot analysis using Pir-specific antibodies, whilst enzymatic activity on polygalacturonic acid was demonstrated on both plate assays and colorimetric assays. Hyper- glycosylation of the enzyme when expressed in the standard strain of S. cerevisiae did not occur, however maximum activities were over two and a half times higher when the enzyme was expressed in the glycosylation deficient strain, suggesting a better adaptation of this strain to the secretion of the functional enzyme. Notably, pectate lyase activity was approximately fourfold higher when the pelA gene was expressed in this yeast strain compared to its expression in a prokaryotic host such as Bacillus subtilis or Escherichia coli.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Strains and Media
2.2. Reagents
2.3. Transformation of Strains and DNA Isolation
2.4. Construction of the Gene Fusion Between PIR4 and pelA
2.5. Isolation of Cell Wall Mannoproteins
2.6. SDS-Polyacrylamide Gels and Western Blot Analysis
2.7. Determination of Pectate Lyase Activity
3. Results
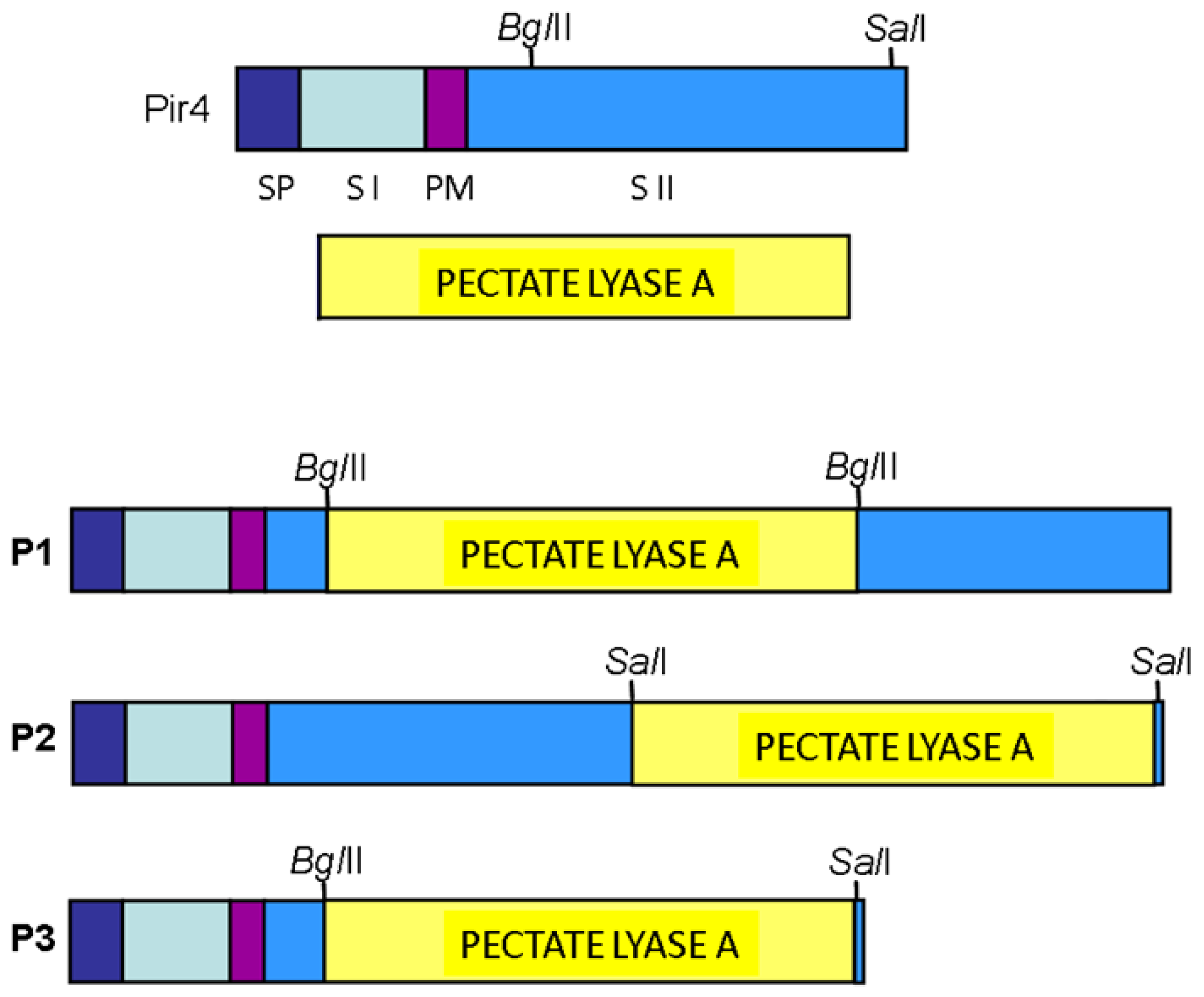
3.1. PIR4/pelA Gene Fusion Strategies and Expression in Saccharomyces cerevisiae
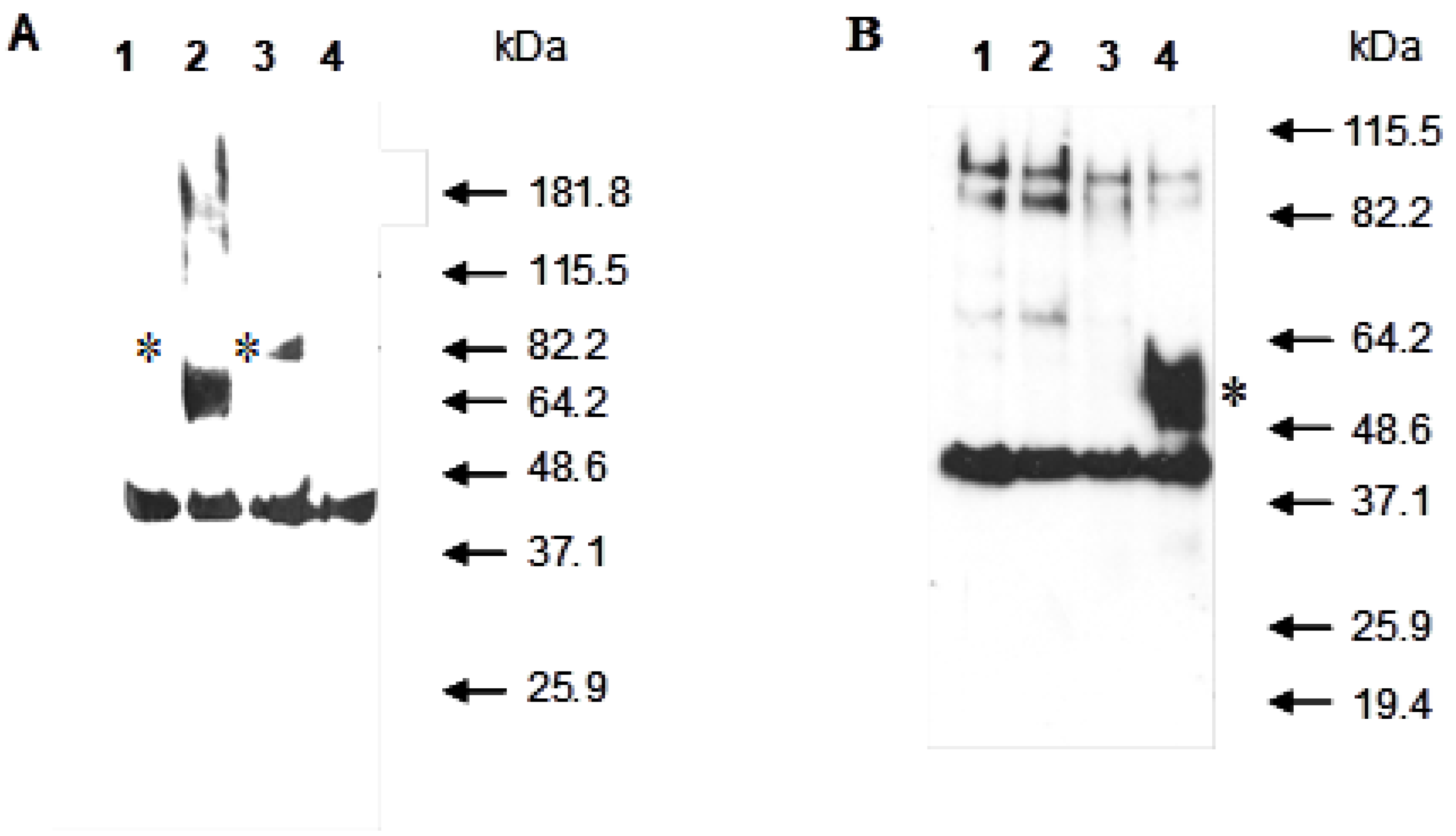
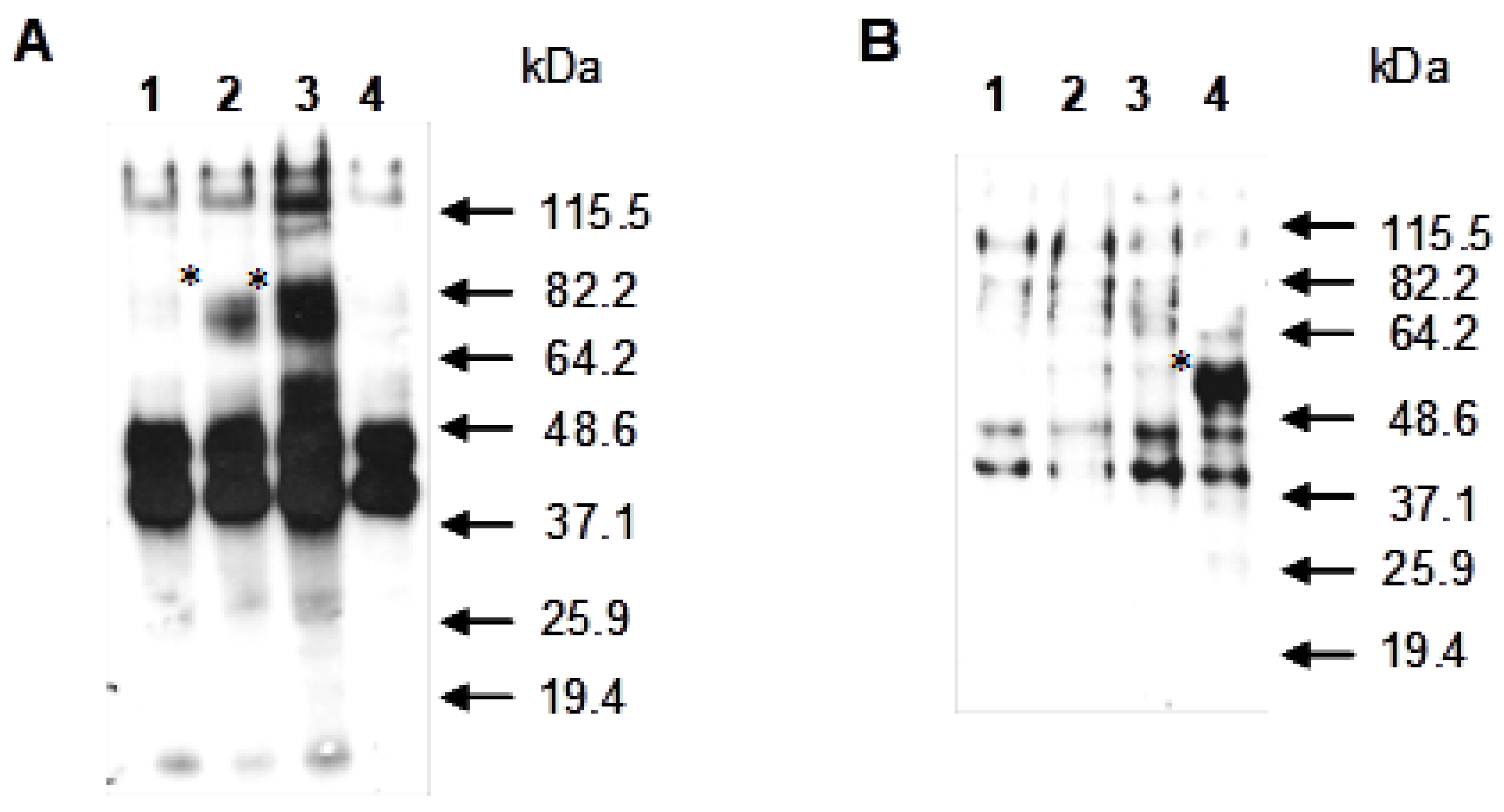
3.2. Study of the Localization of the Recombinant Pir4-PelA Fusion Proteins by Western Blot Analysis
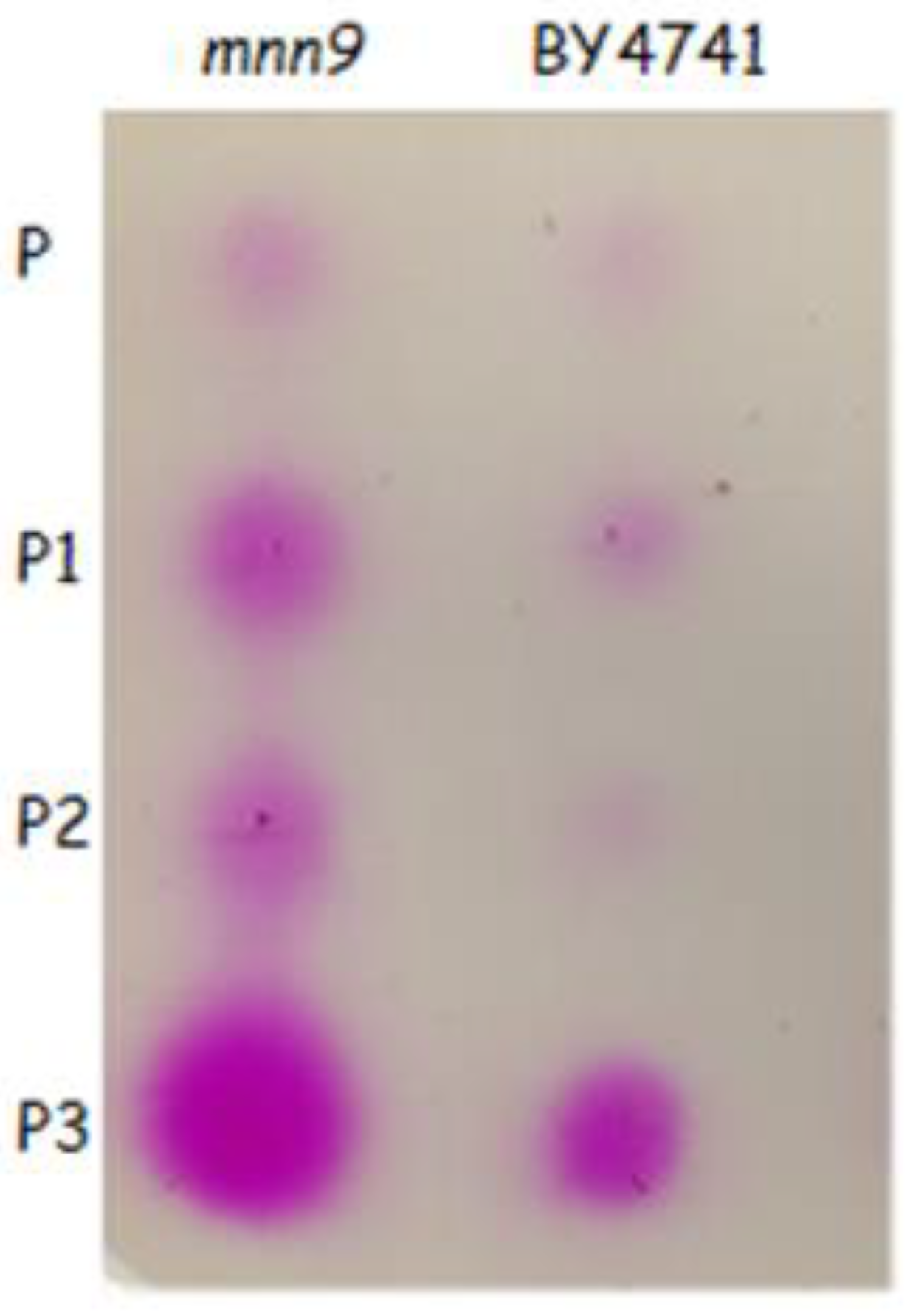
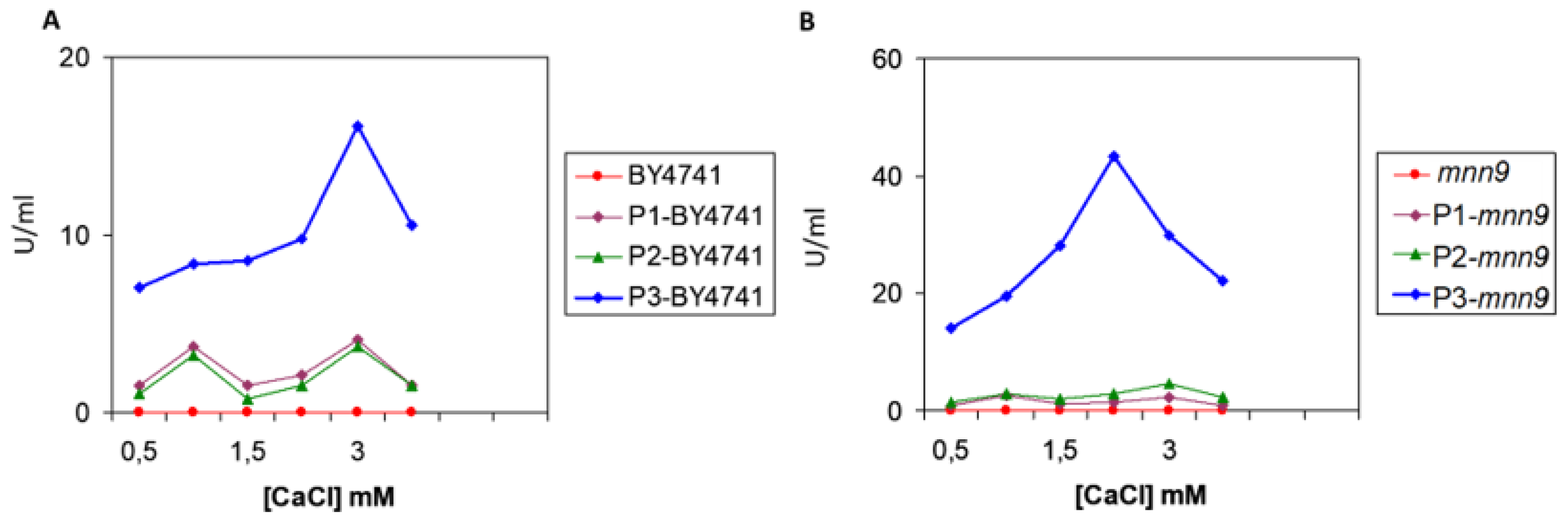
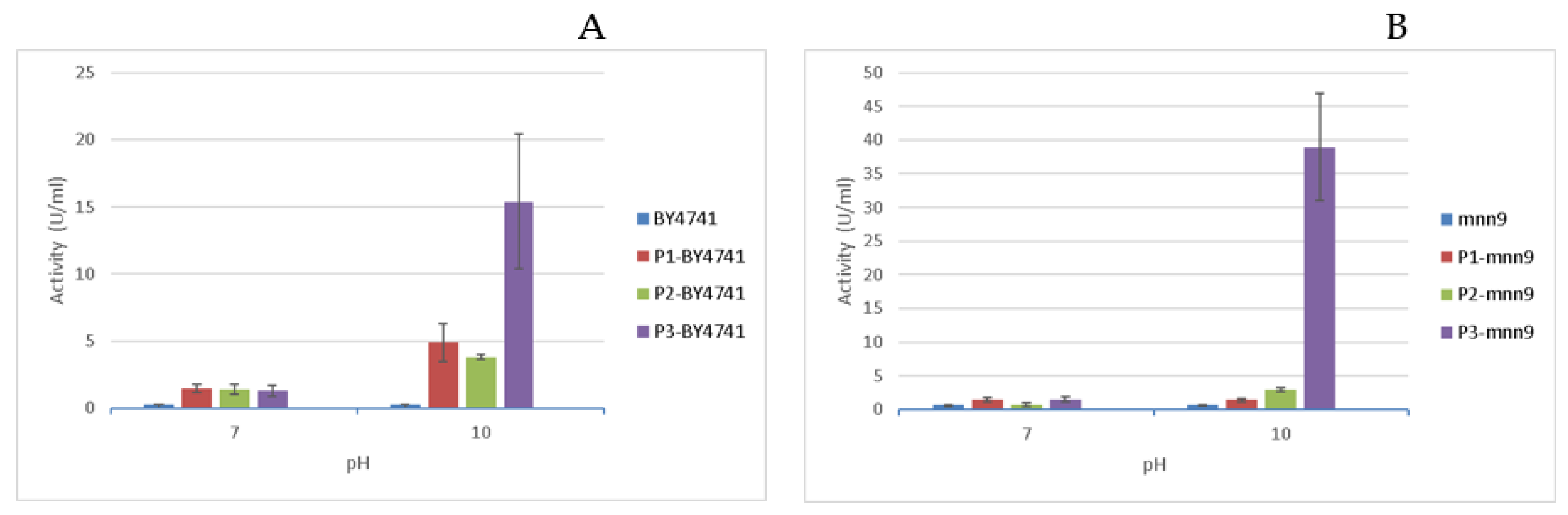
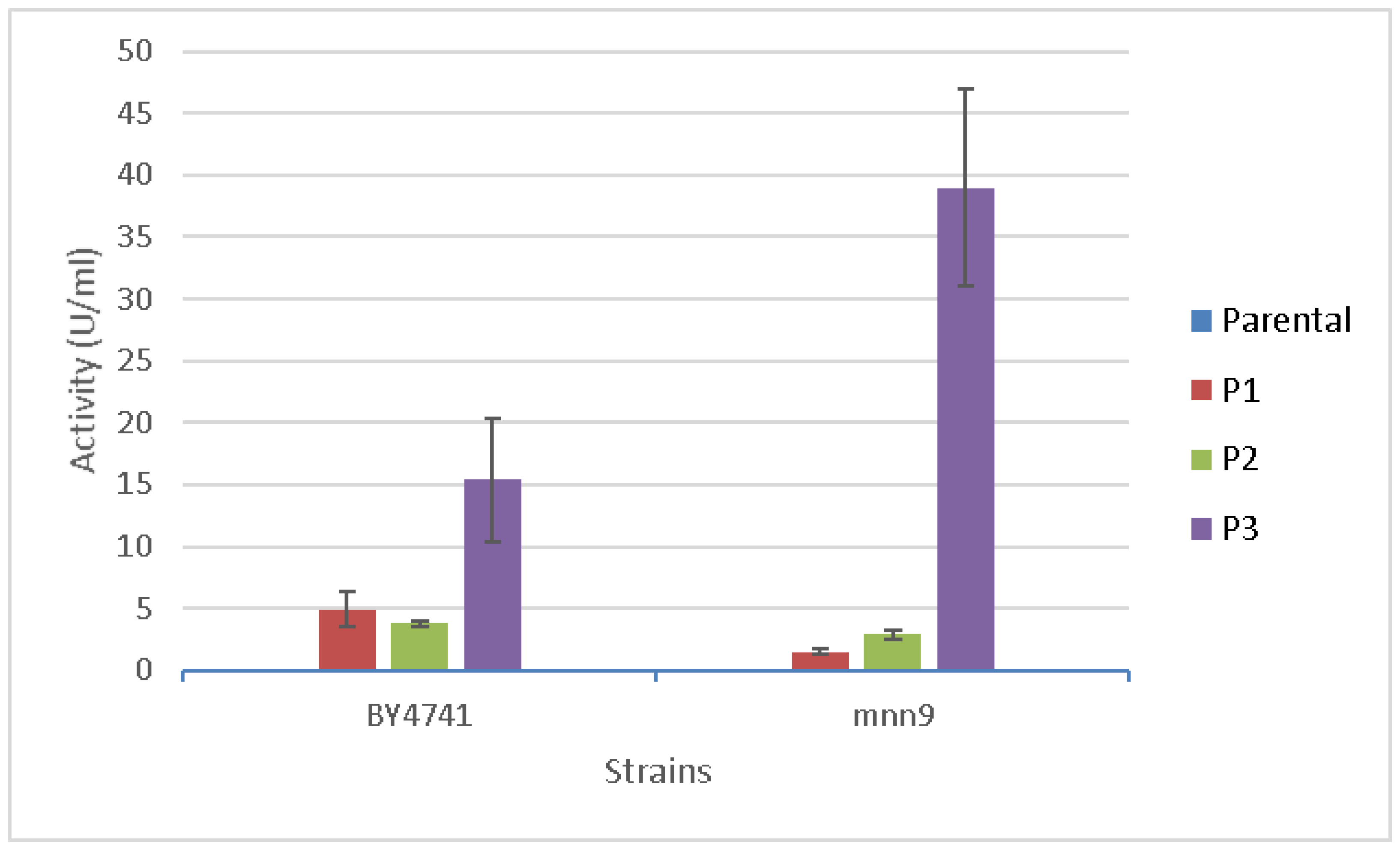
3.3. Quantification of Pectate Lyase Activity Associated to the Different Strains
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gummadi, SN; Panda, T. Purification and biochemical properties of microbial pectinases: a review. Process Biochem. 2003, 38(7), 987–996. [Google Scholar] [CrossRef]
- Soriano, M; Blanco, A; Díaz, P; Pastor, FIJ. An unusual pectate lyase from a Bacillus sp. with high activity on pectin: cloning and characterization. Microbiology (Reading) 2000, 146 Pt 1, 89–95. [Google Scholar] [CrossRef]
- Gummadi, SN; Kumar, DS. Microbial pectic transeliminases. Biotechnol Lett 2005, 27(7), 451–8. [Google Scholar] [CrossRef] [PubMed]
- Jayani, RS; Saxena, S; Gupta, R. Microbial pectinolytic enzymes: a review. Process Biochem. 2005, 40(9), 2931–2944. [Google Scholar] [CrossRef]
- Alkorta, I; Garbisu, C; Llama, MJ; Serra, JL. Industrial applications of pectic enzymes: a review. Process Biochem. 1998, 33(1), 21–28. [Google Scholar] [CrossRef]
- Sakai, T; Sakamoto, T; Hallaert, J; Vandamme, EJ. Pectin, pectinase and protopectinase: production, properties, and applications. Adv Appl Microbiol. 1993, 39, 213–94. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y; Huang, CH; Liu, W; Ko, TP; Xue, Y; Zhou, C; Guo, RT; Ma, Y. Crystal structure and substrate-binding mode of a novel pectate lyase from alkaliphilic Bacillus sp. N16-5. Biochem Biophys Res Commun 2012, 420(2), 269–74. [Google Scholar] [CrossRef]
- Thakur, P; Singh, A. K; Singh, M; Mukherjee, G. Extracellular alkaline pectinases production: a review. Journal of Microbiology, Biotechnology and Food Sciences 2022, 11(5), e3745. [Google Scholar] [CrossRef]
- Bera, S; Rajan, E; Shakya, S; Perera, I; Wijewarna, S; Semini, M. Revisiting microbial pectinases: an Understanding between struc-ture-functional relationship in the arena of genetic engineering. J Appl Biology Biotechnol. 2023. [Google Scholar] [CrossRef]
- Satpathy, A; Mukherjee, K; Nigam, VK. Improved production of thermostable pectinase from novel Bacillus siamensis (BIOS-MNF45) using computational-conventional approach. Sci Rep 2025, 15(1), 11489. [Google Scholar] [CrossRef]
- Kaissar, FZ; Benine, ML; Saadi, SA; Khaldi, A; Missouri, A; Laraki, M; Barberini, S; Emiliani, G; Lebouachera, SEI. Alkaline adapted pectinase from polyextremotolerant Bacillus amyloliquefaciens via agro-wastes valorization: dual biocatalyst and multifunctional bioinoculant. Int Microbiol 2025, 28(8), 3243–3257. [Google Scholar] [CrossRef] [PubMed]
- Strauss, ML; Jolly, NP; Lambrechts, MG; van Rensburg, P. Screening for the production of extracellular hydrolytic enzymes by non-Saccharomyces wine yeasts. J Appl Microbiol. 2001, 91(1), 182–90. [Google Scholar] [CrossRef] [PubMed]
- Busto, MD; García-Tramontín, KE; Ortega, N; Perez-Mateos, M. Preparation and properties of an immobilized pectinlyase for the treatment of fruit juices. Bioresour Technol. 2006, 97(13), 1477–83. [Google Scholar] [CrossRef]
- Van Rensburg, P; Strauss, ML; Lambrechts, MG; Cordero Otero, RR; Pretorius, IS. The heterologous expression of polysaccharidase-encoding genes with oenological relevance in Saccharomyces cerevisiae. J Appl Microbiol. 2007, 103(6), 2248–57. [Google Scholar] [CrossRef]
- Kaul, K; Rajauria, G; Singh, R. Valorization of agro-industrial waste for pectinase production and its influence on circular econ¬omy. Food Bioprod Process 2024, 148, 141–153. [Google Scholar] [CrossRef]
- Haile, S; Ayele, A. Pectinase from microorganisms and its industrial applications. ScientificWorldJournal 2022, 1881305. [Google Scholar] [CrossRef]
- Gazaloğlu, M; Camarasa, C; Nevoigt, E. Exploring pectinolytic yeast diversity: toward effective polygalacturonase producers for applications in wine-making. FEMS Yeast Res. 2025, 25, foae033. [Google Scholar] [CrossRef]
- Zhao, M; Chen, J; Pan, X; Zabed, HM; Arsalan, A; Qi, X. Advances in Pectinase Engineering for Food Bioprocessing: Novel Sources, Mechanisms, and Optimization Strategies. J Agric Food Chem. 2025, 73(37), 23078–23097. [Google Scholar] [CrossRef]
- Kashyap, DR; Vohra, PK; Chopra, S; Tewari, R. Applications of pectinases in the commercial sector: a review. Bioresour Technol 2001, 77(3), 215–27. [Google Scholar] [CrossRef]
- Angayarkanni, J; Palaniswamy, M; Murugesan, S; Swaminathan, K. Improvement of tea leaves fermentation with Aspergillus spp. pectinase. J Biosci Bioeng. 2002, 94(4), 299–303. [Google Scholar] [CrossRef] [PubMed]
- Masoud, W; Jespersen, L. Pectin degrading enzymes in yeasts involved in fermentation of Coffea arabica in East Africa. Int J Food Microbiol 2006, 110(3), 291–6. [Google Scholar] [CrossRef] [PubMed]
- Silva, M. E. d. S.; Oliveira, R. L. d.; Moraes, M. M. d.; Camara, C. A. G. d.; Silva, S. P. d.; Porto, T. S. Application of Commercial Pectinase as a Biocatalyst During Self-Induced Anaerobic Fermentation of Coffee (Coffea arabica L. var. Typica). Fermentation 2025, 11(7), 361. [Google Scholar] [CrossRef]
- Zhang, C; Yao, J; Zhou, C; Mao, L; Zhang, G; Ma, Y. The alkaline pectate lyase PEL168 of Bacillus subtilis heterologously expressed in Pichia pastoris is more stable and efficient for degumming ramie fiber. BMC Biotechnol. 2013, 13, 26. [Google Scholar] [CrossRef]
- Li, X.; Zhou, Y.; Zheng, F. Alkaline pectate lyases for textile degumming: Advances and applications. Applied Microbiology and Biotechnology 2014, 98(20), 8725–8737. [Google Scholar] [CrossRef]
- Zheng, X; Zhang, Y; Liu, X; Li, C; Lin, Y; Liang, S. High-Level Expression and Biochemical Properties of A Thermo-Alkaline Pectate Lyase From Bacillus sp. RN1 in Pichia pastoris With Potential in Ramie Degumming. Front Bioeng Biotechnol 2020, 8, 850. [Google Scholar] [CrossRef] [PubMed]
- Mushtaq, B; Nawab, Y; Ahmad, F; Ahmad, S. An enzymatic modification of sisal fiber macromolecules (cellulose/lignin) for spinnable fibers used in value-added textile applications. Int J Biol Macromol 2025, 323 Pt 1, 147046. [Google Scholar] [CrossRef]
- Tzanov, T; Paar, A; Gudelj, M; Gübitz, GM; Cavaco-Paulo, A. Immobilization of catalases from Bacillus SF on alumina for the treatment of textile bleaching effluents. Enzyme Microb Technol 2001, 28(9-10), 815–819. [Google Scholar] [CrossRef]
- Hoondal, GS; Tiwari, RP; Tewari, R; Dahiya, N; Beg, QK. Microbial alkaline pectinases and their industrial applications: a review. Appl Microbiol Biotechnol 2002, 59(4-5), 409–18. [Google Scholar] [CrossRef]
- Lenting, HBM; Warmoeskerken, MMCG. A fast, continuous enzyme-based pretreatment process concept for cotton containing textiles. Biocatal Biotransformation 2004, 22(5-6), 361–368. [Google Scholar] [CrossRef]
- Zou, M; Li, X; Zhao, J; Qu, Y. Characteristics of polygalacturonate lyase C from Bacillus subtilis 7-3-3 and its synergistic action with PelA in enzymatic degumming. PLoS One 2013, 8(11), e79357. [Google Scholar] [CrossRef]
- Zhou, C; Ye, J; Xue, Y; Ma, Y. Directed Evolution and Structural Analysis of Alkaline Pectate Lyase from the Alkaliphilic Bacterium Bacillus sp. Strain N16-5 To Improve Its Thermostability for Efficient Ramie Degumming. Appl Environ Microbiol 2015, 81(17), 5714–23. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D; Bhardwaj, R; Jassal, S; Goyal, T; Khullar, A; Gupta, N. Application of enzymes for an eco-friendly approach to textile processing. Environ Sci Pollut Res Int. 2023, 30(28), 71838–71848. [Google Scholar] [CrossRef]
- Zhou, C; Xue, Y; Ma, Y. Cloning, evaluation, and high-level expression of a thermo-alkaline pectate lyase from alkaliphilic Bacillus clausii with potential in ramie degumming. Appl Microbiol Biotechnol 2017, 101(9), 3663–3676. [Google Scholar] [CrossRef]
- Shi, A; Hu, H; Zheng, F; Long, L; Ding, S. Biochemical characteristics of an alkaline pectate lyase PelA from Volvariella volvacea: roles of the highly conserved N-glycosylation site in its secretion and activity. Appl Microbiol Biotechnol 2015, 99(8), 3447–58. [Google Scholar] [CrossRef]
- Wu, P; Yang, S; Zhan, Z; Zhang, G. Origins and features of pectate lyases and their applications in industry. Appl Microbiol Biotechnol 2020, 104(17), 7247–7260. [Google Scholar] [CrossRef]
- Han, Y; Peng, XB; Wei, SY; Chen, QG; Yang, JK. Molecular Characterization of a Novel Alkaline Endo-Pectate Lyase from Paenibacillus borealis and Over-Production in Bioreactor Realized by Constructing the Tandem Expression Cassettes in Host Genome. Molecules 2025, 30(17), 3612. [Google Scholar] [CrossRef]
- Hernández, LM; Ballou, L; Alvarado, E; Gillece-Castro, BL; Burlingame, AL; Ballou, CE. A new Saccharomyces cerevisiae mnn mutant N-linked oligosaccharide structure. J Biol Chem. 1989, 264(20), 11849–56. [Google Scholar] [CrossRef] [PubMed]
- Sambrook J, Fritsch EF, Maniatis T. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory. Cold Spring Harbor N. Y.
- Ito, H; Fukuda, Y; Murata, K; Kimura, A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983, 153(1), 163–8. [Google Scholar] [CrossRef] [PubMed]
- Gietz, RD; Sugino, A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene 1988, 74(2), 527–34. [Google Scholar] [CrossRef]
- Andrés, I; Zueco, J; Parascandola, P. Immobilization of Saccharomyces cerevisiae cells to protein G-Sepharose by cell wall engineering. J Mol Microbiol Biotechnol. 2003, 5(3), 161–6. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227(5259), 680–5. [Google Scholar] [CrossRef]
- Burnette, WN. Western blotting": electrophoretic transfer of proteins from sodium dodecyl sulfate--polyacrylamide gels to unmodified nitrocellulose and radiographic detection with antibody and radioiodinated protein A. Anal Biochem 1981, 112(2), 195–203. [Google Scholar] [CrossRef]
- Moukadiri, I; Jaafar, L; Zueco, J. Identification of two mannoproteins released from cell walls of a Saccharomyces cerevisiae mnn1 mnn9 double mutant by reducing agents. J Bacteriol. 1999, 181(16), 4741–5. [Google Scholar] [CrossRef]
- McKay, AM. A plate assay method for the detection of fungal polygalacturonase secretion. FEMS Microbiol. Lett. 1988, 56(3), 355–358. [Google Scholar] [CrossRef]
- Gognies, S; Gainvors, A; Aigle, M; Belarbi, A. Cloning, sequence analysis and overexpression of a Saccharomyces cerevisiae endopolygalacturonase-encoding gene (PGL1). Yeast 1999, 15(1), 11–22. [Google Scholar] [CrossRef]
- Collmer, A; Ried, JL; Mount, MS. Assay methods for pectic enzymes. Methods. Enzymol. 1988, 161, 329–335. [Google Scholar] [CrossRef]
- Moukadiri, I; Zueco, J. Evidence for the attachment of Hsp150/Pir2 to the cell wall of Saccharomyces cerevisiae through disulfide bridges. FEMS Yeast Res. 2001, 1(3), 241–5. [Google Scholar] [CrossRef]
- Andrés, I; Gallardo, O; Parascandola, P; Javier Pastor, FI; Zueco, J. Use of the cell wall protein Pir4 as a fusion partner for the expression of Bacillus sp. BP-7 xylanase A in Saccharomyces cerevisiae. Biotechnol Bioeng. 2005, 89(6), 690–7. [Google Scholar] [CrossRef] [PubMed]
- Andrés, I; Rodríguez-Díaz, J; Buesa, J; Zueco, J. Yeast expression of the VP8* fragment of the rotavirus spike protein and its use as immunogen in mice. Biotechnol Bioeng. 2006, 93(1), 89–98. [Google Scholar] [CrossRef]
- Mormeneo, M; Andrés, I; Bofill, C; Díaz, P; Zueco, J. Efficient secretion of Bacillus subtilis lipase A in Saccharomyces cerevisiae by translational fusion to the Pir4 cell wall protein. Appl Microbiol Biotechnol 2008, 80(3), 437–45. [Google Scholar] [CrossRef]
- Paciello, L; de Alteriis, E; Mazzoni, C; Palermo, V; Zueco, J; Parascandola, P. Performance of the auxotrophic Saccharomyces cerevisiae BY4741 as host for the production of IL-1beta in aerated fed-batch reactor: role of ACA supplementation, strain viability, and maintenance energy. Microb Cell Fact 2009, 8, 70. [Google Scholar] [CrossRef]
- Mormeneo, M; Pastor, FJ; Zueco, J. Efficient expression of a Paenibacillus barcinonensis endoglucanase in Saccharomyces cerevisiae. J Ind Microbiol Biotechnol. 2012, 39(1), 115–23. [Google Scholar] [CrossRef] [PubMed]
- Yang, N; Yu, Z; Jia, D; Xie, Z; Zhang, K; Xia, Z; Lei, L; Qiao, M. The contribution of Pir protein family to yeast cell surface display. Appl Microbiol Biotechnol 2014, 98(7), 2897–905. [Google Scholar] [CrossRef]
- Martinić Cezar, T.; Lozančić, M.; Novačić, A.; Matičević, A.; Matijević, D.; Vallée, B.; Mrša, V.; Teparić, R.; Žunar, B. Streamlining N-terminally anchored yeast surface display via structural insights into S. cerevisiae Pir proteins. Microbial Cell Factories 2023, 22, Article 174. [Google Scholar] [CrossRef] [PubMed]
- Soriano, M; Diaz, P; Pastor, FI. Pectinolytic systems of two aerobic sporogenous bacterial strains with high activity on pectin. Curr Microbiol 2005, 50(2), 114–8. [Google Scholar] [CrossRef] [PubMed]
- Soriano, M; Diaz, P; Pastor, FIJ. Pectate lyase C from Bacillus subtilis: a novel endo-cleaving enzyme with activity on highly methylated pectin. Microbiology (Reading) 2006, 152 Pt 3, 617–625. [Google Scholar] [CrossRef]
- de Nobel, JG; Klis, FM; Priem, J; Munnik, T; van den Ende, H. The glucanase-soluble mannoproteins limit cell wall porosity in Saccharomyces cerevisiae. Yeast 1990, 6(6), 491–9. [Google Scholar] [CrossRef] [PubMed]
- Sieiro, C; Poza, M; Vilanova, M; Villa, TG. Heterologous expression of the Saccharomyces cerevisiae PGU1 gene in Schizosaccharomyces pombe yields an enzyme with more desirable properties for the food industry. Appl Environ Microbiol 2003, 69(3), 1861–5. [Google Scholar] [CrossRef]
- Louw, C; La Grange, D; Pretorius, IS; van Rensburg, P. The effect of polysaccharide-degrading wine yeast transformants on the efficiency of wine processing and wine flavour. J Biotechnol. 2006, 125(4), 447–61. [Google Scholar] [CrossRef]
| S. cerevisiae strains | Genotype | Origin |
|---|---|---|
| BY4741 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his3Δ1 | EUROSCARF |
| mnn9 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his 3Δ1, ypl050c::kanMX4 | EUROSCARF |
| P1-BY4741 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his3Δ1, (YEplac195-PIR4/pelA-BglII) | This study |
| P2-BY4741 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his3Δ1, (YEplac195-PIR4/pelA-SalI) | This study |
| P3-BY4741 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his3Δ1, (YEplac195-PIR4/pelA-BglII-SalI) | This study |
| P1-mnn9 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his 3Δ1, ypl050c::kanMX4, (YEplac195-PIR4/pelA-BglII) | This study |
| P2-mnn9 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his3Δ1, ypl050c::kanMX4, (YEplac195-PIR4/pelA-SalI) | This study |
| P3-mnn9 | MATα, ura3Δ0, leu2Δ0, met15Δ0, his 3Δ1, ypl050c::kanMX4, (YEplac195-PIR4/pelA-BglII-SalI) | This study |
| PRIMERS | SEQUENCES |
|---|---|
| PIR5 | TGCATTCCATACGATTTCCACGGG |
| PIR3 | GTGTATATTAAAGGCTGCATGTGG |
| PB5 | TATATAAGATCTTAGCGCCAACCGTCGTCAATTC |
| PB3 | TATAGGAGATCTTATACTGTGTATTTCCGGACTG |
| PS5 | AAGCCTCTCGAGGCGCCAACCGTCGTCAATTC |
| PS3 | GATCCCCTCGAGATACTGTGTATTTCCGGACTGG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




