Submitted:
31 March 2026
Posted:
02 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Dataset
- (1)
- Only samples involving single amino acid substitution mutations were retained;
- (2)
- Abnormal complex structures lacking valid protein contacts around the ligand were removed;
- (3)
- PDB records with ambiguous structural mapping caused by multichain assemblies or duplicated residue numbering were excluded;
- (4)
- Samples with incomplete or unresolvable structural information were discarded.
2.2. Protein Representation
2.3. Mutation-Site Indicator Encoding
2.4. Ligand Representation
2.4.1. Vectorized Ligand Representation
2.4.2. Graph-Based Ligand Representation
2.5. Protein Feature Construction Schemes
2.6. Graph Construction
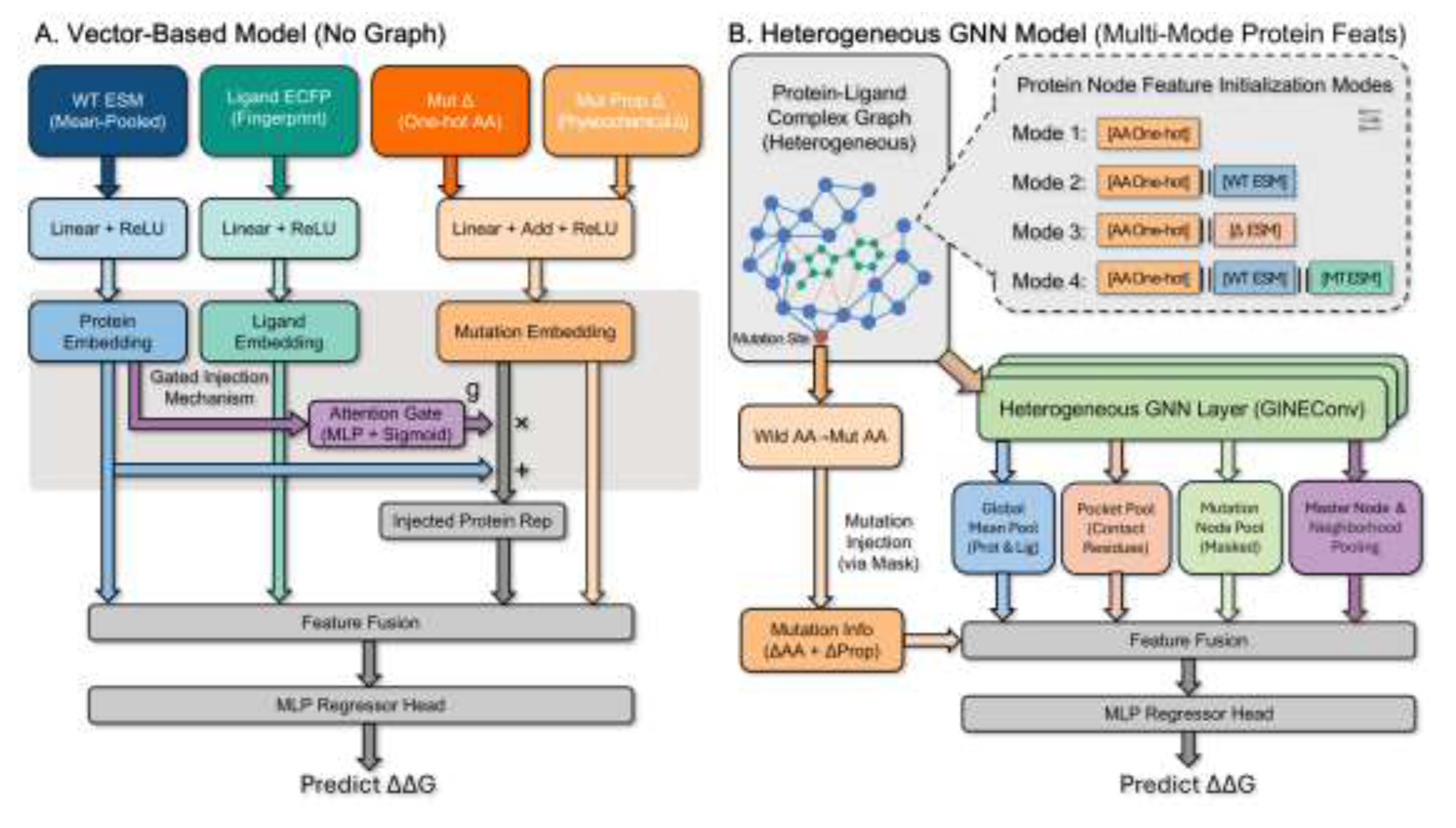
2.7. Model Architecture
2.8. Training Strategy
3. Results and Discussion
3.1. Overall Predictive Performance under Different Data Splitting Strategies
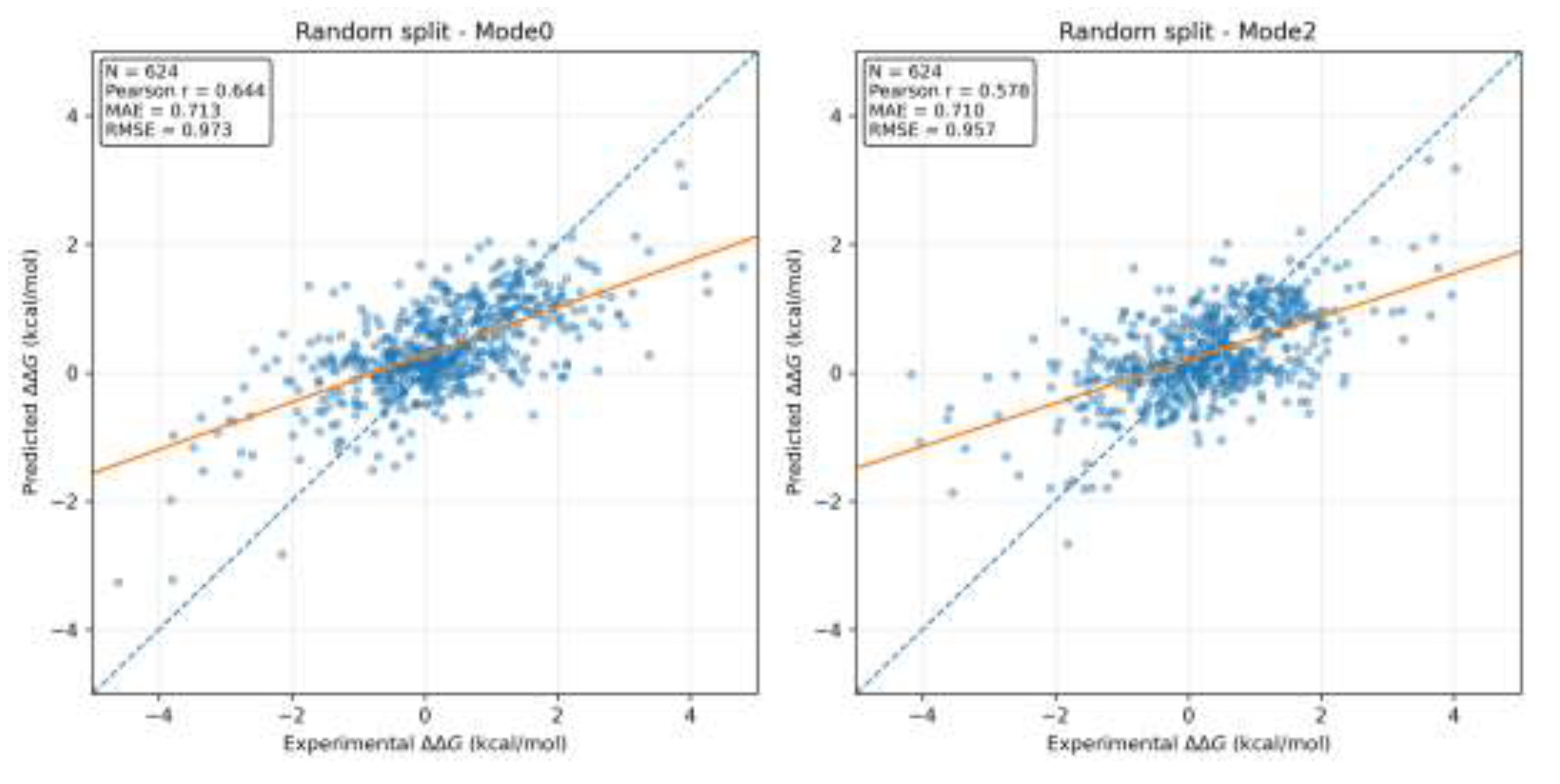
3.2. Scatter Plot and Regression Trend Analysis
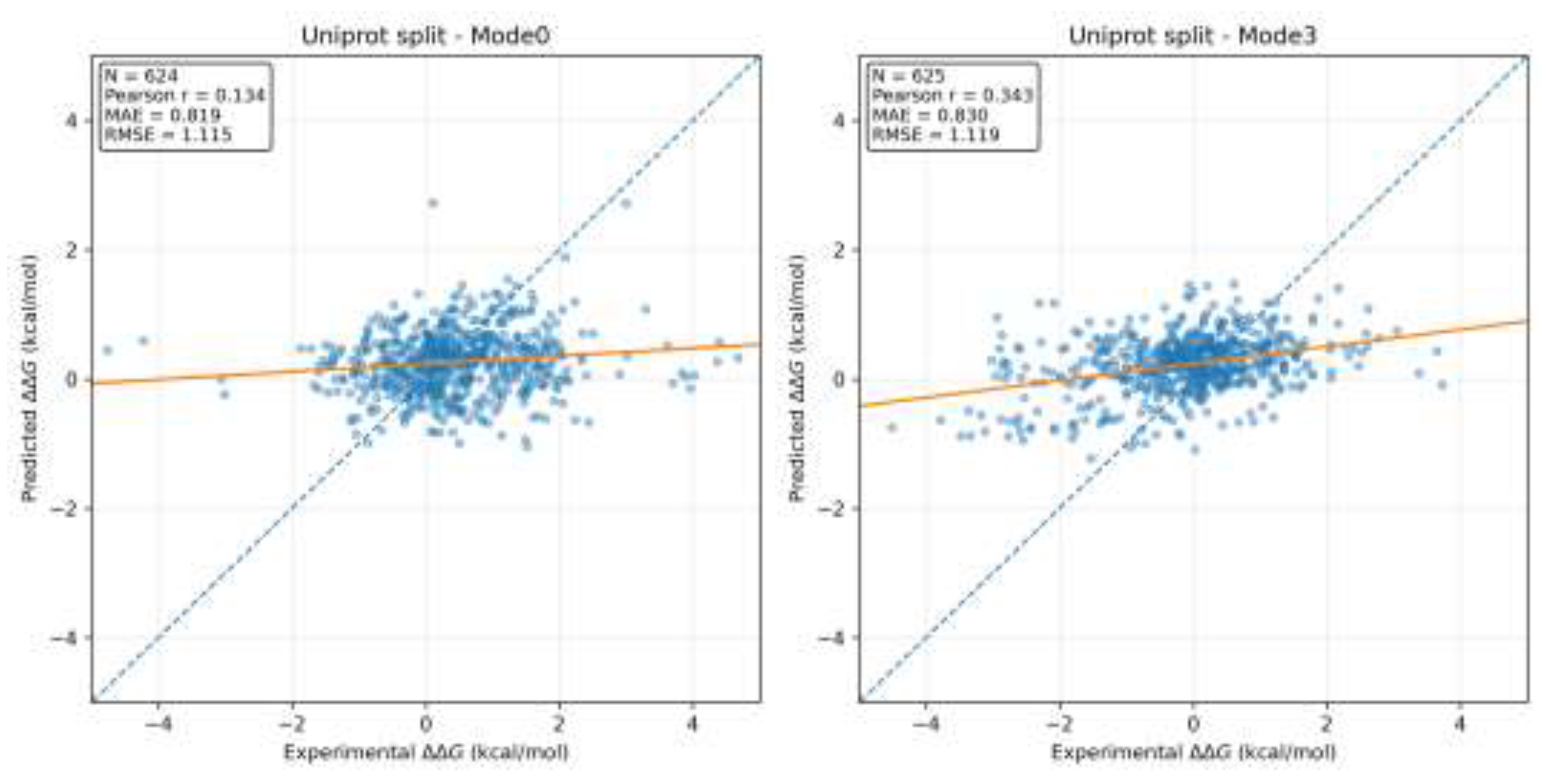
3.3. Discussion of Generalization and Data Splitting Strategies
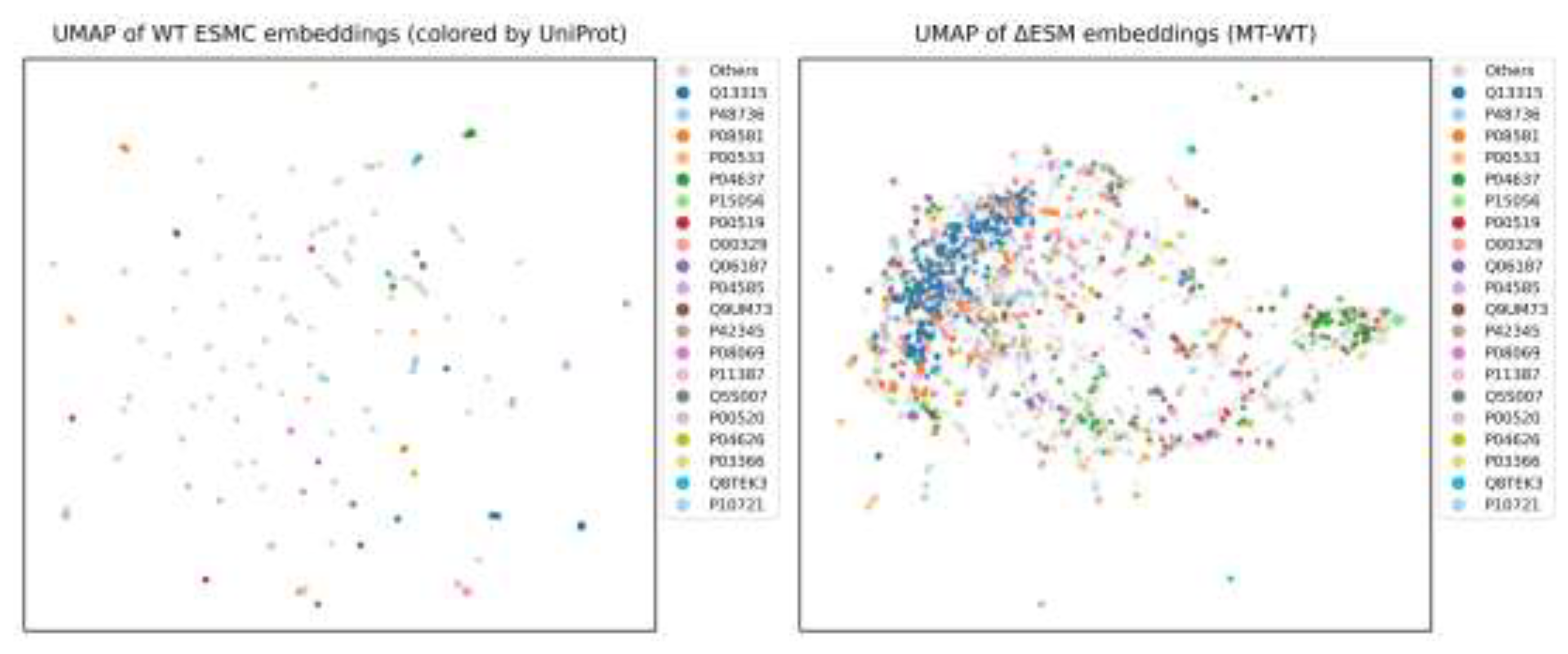
3.4. Representation Space Analysis of Wild-Type ESM and ΔESM Embeddings
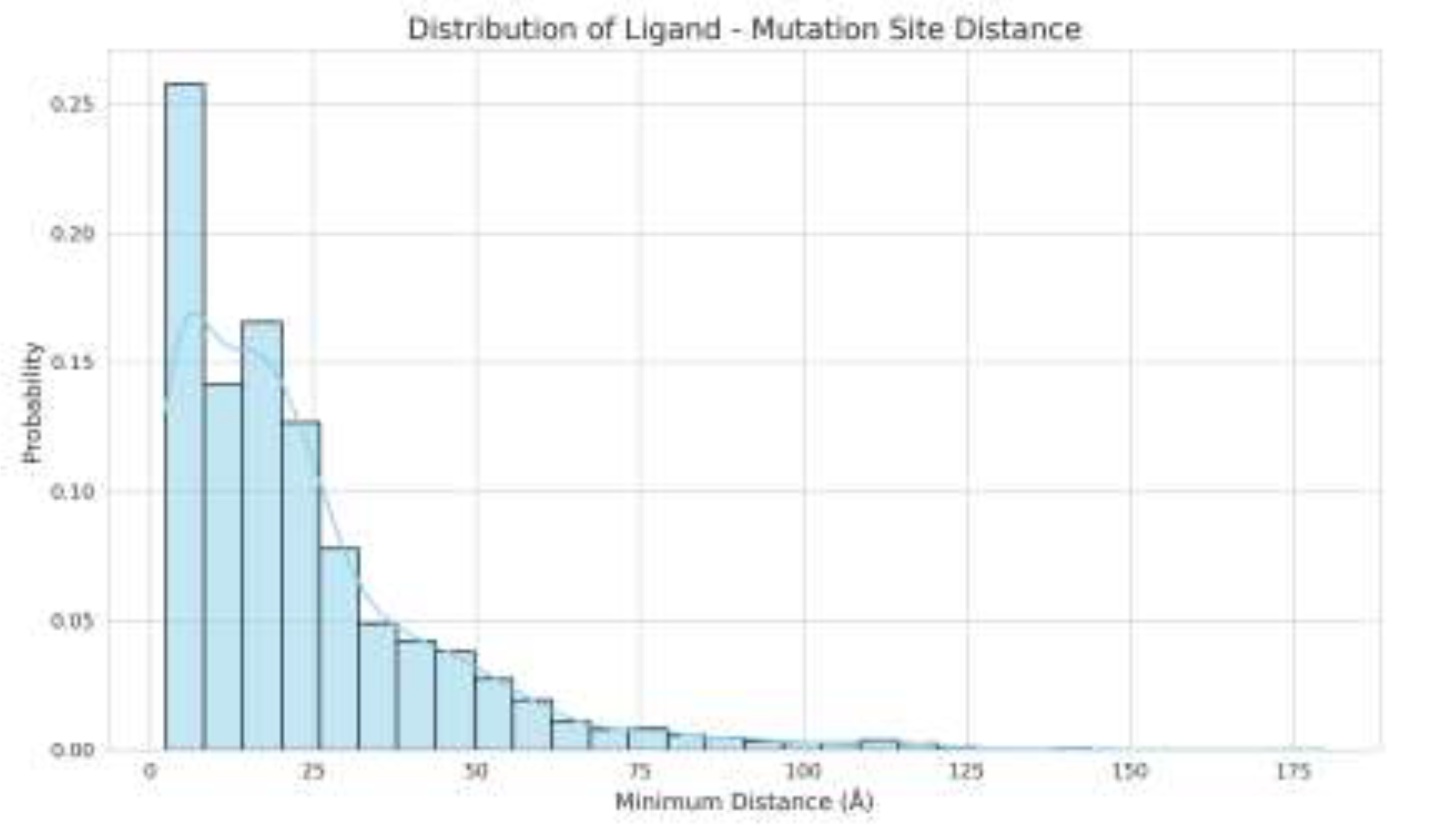
3.5. Analysis of Dataset Characteristics and Factors Limiting Model Performance
4. Conclusions
5. Data Availability
Supplementary Materials
References
- Vasan N, Baselga J, Hyman DM. A view on drug resistance in cancer. Nature. 2019 Nov 1;575(7782):299–309. [CrossRef]
- Guerois R, Nielsen JE, Serrano L. Predicting Changes in the Stability of Proteins and Protein Complexes: A Study of More Than 1000 Mutations. Journal of Molecular Biology. 2002 Jul 5;320(2):369–87. [CrossRef]
- Cournia Z, Allen B, Sherman W. Relative Binding Free Energy Calculations in Drug Discovery: Recent Advances and Practical Considerations. J Chem Inf Model. 2017 Dec 26;57(12):2911–37. [CrossRef]
- Wang L, Wu Y, Deng Y, Kim B, Pierce L, Krilov G, et al. Accurate and Reliable Prediction of Relative Ligand Binding Potency in Prospective Drug Discovery by Way of a Modern Free-Energy Calculation Protocol and Force Field. J Am Chem Soc. 2015 Feb 25;137(7):2695–703. [CrossRef]
- Atz K, Grisoni F, Schneider G. Geometric deep learning on molecular representations. Nat Mach Intell. 2021 Dec;3(12):1023–32. [CrossRef]
- Gaudelet T, Day B, Jamasb AR, Soman J, Regep C, Liu G, et al. Utilizing graph machine learning within drug discovery and development. Brief Bioinform. 2021 Nov 1;22(6):bbab159. [CrossRef]
- Ferreira FJN, Carneiro AS. AI-Driven Drug Discovery: A Comprehensive Review. ACS Omega. 10(23):23889–903. PubMed PMID: 40547666; PubMed Central PMCID: PMC12177741. [CrossRef]
- Wang H. Prediction of protein–ligand binding affinity via deep learning models. Brief Bioinform. 2024 Mar 1;25(2):bbae081. [CrossRef]
- Zhao L, Zhu Y, Wang J, Wen N, Wang C, Cheng L. A brief review of protein–ligand interaction prediction. Computational and Structural Biotechnology Journal. 2022 Jan 1;20:2831–8. [CrossRef]
- Verburgt J, Jain A, Kihara D. Recent Deep-Learning Applications to Structure-Based Drug Design. Methods Mol Biol. 2024;2714:215–34. PubMed PMID: 37676602; PubMed Central PMCID: PMC10578466. [CrossRef]
- Yu G, Bi X, Ma T, Li Y, Wang J. CATH-ddG: towards robust mutation effect prediction on protein–protein interactions out of CATH homologous superfamily. Bioinformatics. 2025 Jul 1;41(Supplement_1):i362–72. [CrossRef]
- Zhou Y, Pan Q, Pires DEV, Rodrigues CHM, Ascher DB. DDMut: predicting effects of mutations on protein stability using deep learning. Nucleic Acids Res. 2023 Jul 5;51(W1):W122–8. [CrossRef]
- Pires DEV, Blundell TL, Ascher DB. mCSM-lig: quantifying the effects of mutations on protein-small molecule affinity in genetic disease and emergence of drug resistance. Sci Rep. 2016 Jul 7;6(1):29575. [CrossRef]
- Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021 Aug;596(7873):583–9. [CrossRef]
- Pak MA, Markhieva KA, Novikova MS, Petrov DS, Vorobyev IS, Maksimova ES, et al. Using AlphaFold to predict the impact of single mutations on protein stability and function. PLoS One. 2023 Mar 16;18(3):e0282689. PubMed PMID: 36928239; PubMed Central PMCID: PMC10019719. [CrossRef]
- Buel GR, Walters KJ. Can AlphaFold2 predict the impact of missense mutations on structure? Nat Struct Mol Biol. 2022 Jan;29(1):1–2. PubMed PMID: 35046575; PubMed Central PMCID: PMC11218004. [CrossRef]
- Nguyen T, Le H, Quinn TP, Nguyen T, Le TD, Venkatesh S. GraphDTA: predicting drug–target binding affinity with graph neural networks. Bioinformatics. 2021 May 23;37(8):1140–7. [CrossRef]
- Li S, Zhou J, Xu T, Huang L, Wang F, Xiong H, et al. Structure-aware Interactive Graph Neural Networks for the Prediction of Protein-Ligand Binding Affinity. In: Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining [Internet]. New York, NY, USA: Association for Computing Machinery; 2021 [cited 2026 Mar 4]. p. 975–85. (KDD ’21). Available from: https://dl.acm.org/doi/10.1145/3447548.3467311. [CrossRef]
- Satorras VG, Hoogeboom E, Welling M. E(n) Equivariant Graph Neural Networks [Internet]. arXiv; 2022 [cited 2026 Mar 5]. Available from: http://arxiv.org/abs/2102.09844. [CrossRef]
- Graber D, Stockinger P, Meyer F, Mishra S, Horn C, Buller R. Resolving data bias improves generalization in binding affinity prediction. Nat Mach Intell. 2025 Oct;7(10):1713–25. [CrossRef]
- Cao H, Wang J, He L, Qi Y, Zhang JZ. DeepDDG: Predicting the Stability Change of Protein Point Mutations Using Neural Networks. J Chem Inf Model. 2019 Apr 22;59(4):1508–14. [CrossRef]
- Xie L, Bao G, Zhang D, Xu L, Xu X, Chang S. Evaluating data partitioning strategies for accurate prediction of protein-ligand binding free energy changes in mutated proteins. Computational and Structural Biotechnology Journal. 2025 Jan 1;27:4418–30. PubMed PMID: 41169760. [CrossRef]
- Yang Z, Ye Z, Qiu J, Feng R, Li D, Hsieh C, et al. A mutation-induced drug resistance database (MdrDB). Commun Chem. 2023 Jun 14;6(1):123. [CrossRef]
- Rives A, Meier J, Sercu T, Goyal S, Lin Z, Liu J, et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences. 2021 Apr 13;118(15):e2016239118. [CrossRef]
- Lin Z, Akin H, Rao R, Hie B, Zhu Z, Lu W, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023 Mar 17;379(6637):1123–30. [CrossRef]
- Hayes T, Rao R, Akin H, Sofroniew NJ, Oktay D, Lin Z, et al. Simulating 500 million years of evolution with a language model. Science. 2025 Feb 21;387(6736):850–8. [CrossRef]
- Rogers D, Hahn M. Extended-Connectivity Fingerprints. J Chem Inf Model. 2010 May 24;50(5):742–54. [CrossRef]
- Wang K, Zhou R, Tang J, Li M. GraphscoreDTA: optimized graph neural network for protein–ligand binding affinity prediction. Bioinformatics. 2023 Jun 1;39(6):btad340. [CrossRef]
- Yang D, Kuang L, Hu A. Edge-enhanced interaction graph network for protein-ligand binding affinity prediction. PLOS ONE. 2025 Apr 8;20(4):e0320465. [CrossRef]
- Samudrala MV, Dandibhotla S, Kaneriya A, Dakshanamurthy S. PLAIG: Protein–Ligand Binding Affinity Prediction Using a Novel Interaction-Based Graph Neural Network Framework. ACS Bio Med Chem Au. 2025 Jun 18;5(3):447–63. [CrossRef]
- Landrum G. RDKit: Open-source cheminformatics [Internet]. 2016. Available from: https://www.rdkit.org.
- Xu* K, Hu* W, Leskovec J, Jegelka S. How Powerful are Graph Neural Networks? In. 2018 [cited 2026 Mar 4]. Available from: https://openreview.net/forum?id=ryGs6iA5Km.
- Schütt K, Kindermans PJ, Sauceda Felix HE, Chmiela S, Tkatchenko A, Müller KR. SchNet: A continuous-filter convolutional neural network for modeling quantum interactions. In: Advances in Neural Information Processing Systems [Internet]. Curran Associates, Inc.; 2017 [cited 2026 Mar 5]. Available from: https://proceedings.neurips.cc/paper/2017/hash/303ed4c69846ab36c2904d3ba8573050-Abstract.html.
- Huber PJ. Robust Estimation of a Location Parameter. The Annals of Mathematical Statistics. 1964 Mar;35(1):73–101. [CrossRef]
- Loshchilov I, Hutter F. Decoupled Weight Decay Regularization. In. 2018 [cited 2026 Mar 5]. Available from: https://openreview.net/forum?id=Bkg6RiCqY7.
- Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, et al. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In: Advances in Neural Information Processing Systems [Internet]. Curran Associates, Inc.; 2019 [cited 2026 Mar 4]. Available from: https://papers.neurips.cc/paper_files/paper/2019/hash/bdbca288fee7f92f2bfa9f7012727740-Abstract.html.
- Fey M, Lenssen JE. Fast Graph Representation Learning with PyTorch Geometric [Internet]. arXiv; 2019 [cited 2026 Mar 5]. Available from: http://arxiv.org/abs/1903.02428. [CrossRef]
- Bernett J, Blumenthal DB, List M. Cracking the black box of deep sequence-based protein–protein interaction prediction. Brief Bioinform. 2024 Mar 5;25(2):bbae076. PubMed PMID: 38446741; PubMed Central PMCID: PMC10939362. [CrossRef]
- Wallach I, Heifets A. Most Ligand-Based Classification Benchmarks Reward Memorization Rather than Generalization. J Chem Inf Model. 2018 May 29;58(5):916–32. [CrossRef]
- Bushuiev A, Bushuiev R, Sedlar J, Pluskal T, Damborsky J, Mazurenko S, et al. Revealing data leakage in protein interaction benchmarks [Internet]. arXiv; 2024 [cited 2026 Mar 5]. Available from: http://arxiv.org/abs/2404.10457. [CrossRef]
- McInnes L, Healy J, Saul N, Großberger L. UMAP: Uniform Manifold Approximation and Projection. Journal of Open Source Software. 2018 Sep 2;3(29):861. [CrossRef]
- Kramer C, Kalliokoski T, Gedeck P, Vulpetti A. The Experimental Uncertainty of Heterogeneous Public Ki Data. J Med Chem. 2012 Jun 14;55(11):5165–73. [CrossRef]
- Gilson MK, Zhou HX. Calculation of Protein-Ligand Binding Affinities*. Annual Review of Biophysics. Annual Reviews; 2007. p. 21–42. [CrossRef]
- Cournia Z, Chipot C, Roux B, York DM, Sherman W. Free Energy Methods in Drug Discovery—Introduction. In: Free Energy Methods in Drug Discovery: Current State and Future Directions [Internet]. American Chemical Society; 2021 [cited 2026 Mar 4]. p. 1–38. (ACS Symposium Series; no. 1397). Available from: . [CrossRef]
- Mobley DL, Dill KA. Binding of Small-Molecule Ligands to Proteins: “What You See” Is Not Always “What You Get.” Structure. 2009 Apr 15;17(4):489–98. [CrossRef]
- Boehr DD, Nussinov R, Wright PE. The role of dynamic conformational ensembles in biomolecular recognition. Nat Chem Biol. 2009 Nov;5(11):789–96. [CrossRef]
| Mode | Pearson | Spearman | MAE | RMSE |
| Mode0 | 0.5545±0.0581 | 0.5411±0.0399 | 0.7121±0.0158 | 0.9941±0.0303 |
| Mode1 | 0.4873±0.0156 | 0.4554±0.0290 | 0.7567±0.0359 | 1.0475±0.0392 |
| Mode2 | 0.5549±0.0271 | 0.5255±0.0411 | 0.7223±0.0283 | 0.9971±0.0525 |
| Mode3 | 0.5194±0.0385 | 0.4896±0.0275 | 0.7471±0.0340 | 1.0281±0.0525 |
| Mode4 | 0.5388±0.0438 | 0.5076±0.0500 | 0.7356±0.0387 | 1.0130±0.0574 |
| Mode | Pearson | Spearman | MAE | RMSE |
| Mode0 | 0.0439±0.0774 | 0.0436±0.0830 | 0.9308±0.1600 | 1.3101±0.3731 |
| Mode1 | 0.0897±0.1472 | 0.1418±0.0973 | 0.8679±0.0588 | 1.2558±0.2423 |
| Mode2 | 0.1001±0.1108 | 0.0906±0.1160 | 0.8645±0.0383 | 1.2536±0.2421 |
| Mode3 | 0.1526±0.1481 | 0.0863±0.1724 | 0.8838±0.0907 | 1.2539±0.2749 |
| Mode4 | 0.1033±0.1163 | 0.0960±0.1336 | 0.8535±0.0405 | 1.2458±0.2357 |
| UniProt ID | Mutation | Drug | ΔΔG (Source 1) | ΔΔG (Source 2) | Abs. Diff |
| P00533 (EGFR) | L858R | Gefitinib | -4.04 | -1.6 | 2.44 |
| P00533 (EGFR) | G719S | Gefitinib | -1.53 | 0.48 | 2.01 |
| P15056 (BRAF) | L597V | PLX-4720 | -1.65 | 0.35 | 2.00 |
| P15056 (BRAF) | A728V | PLX-4720 | -2.51 | -0.64 | 1.87 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




