Submitted:
30 March 2026
Posted:
31 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
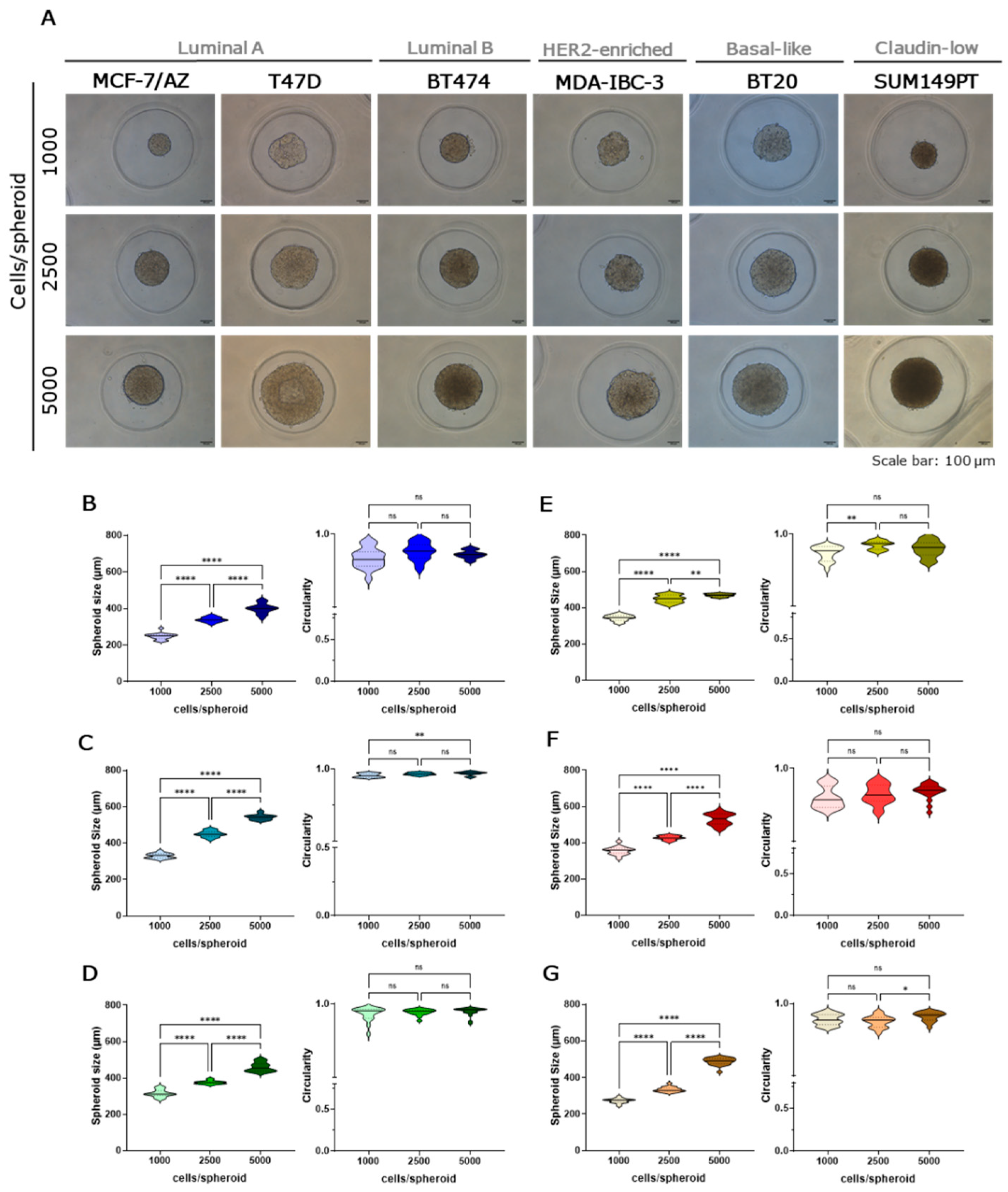
2.1. Establishment and Characterization of Breast Cancer Tumour Spheroids
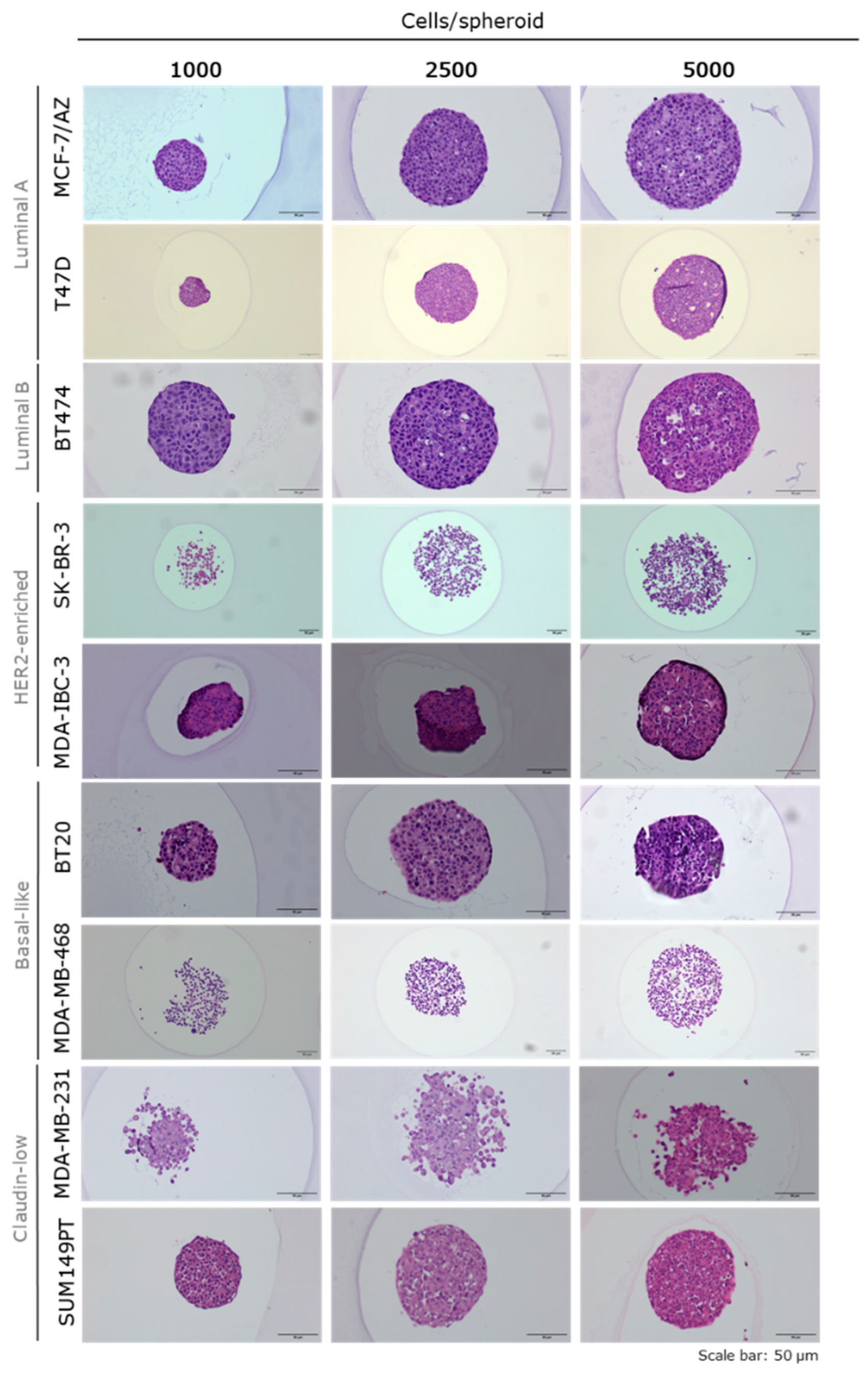
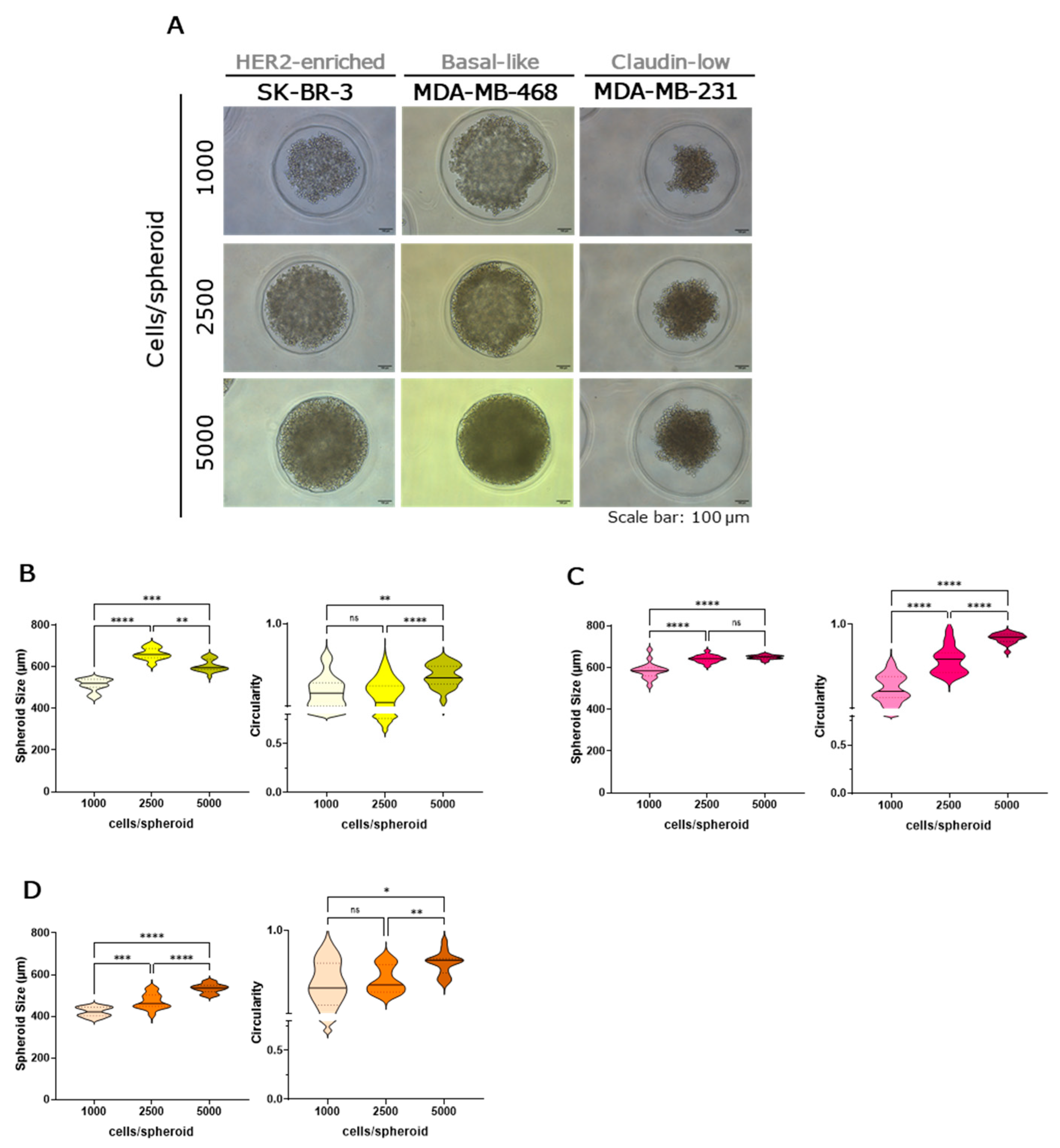
2.2. Morphological Characterization of Breast Cancer Tumour Spheroids
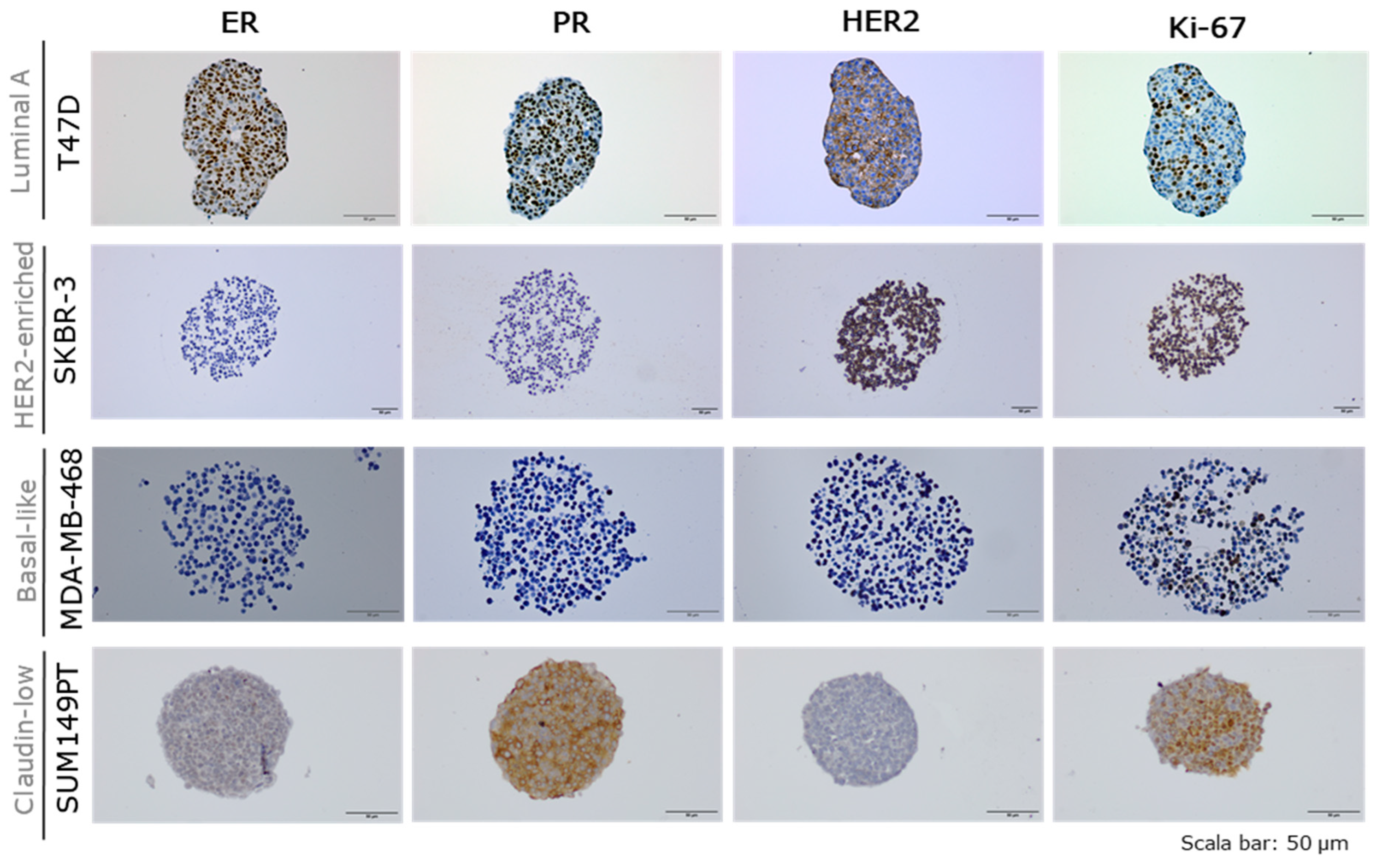
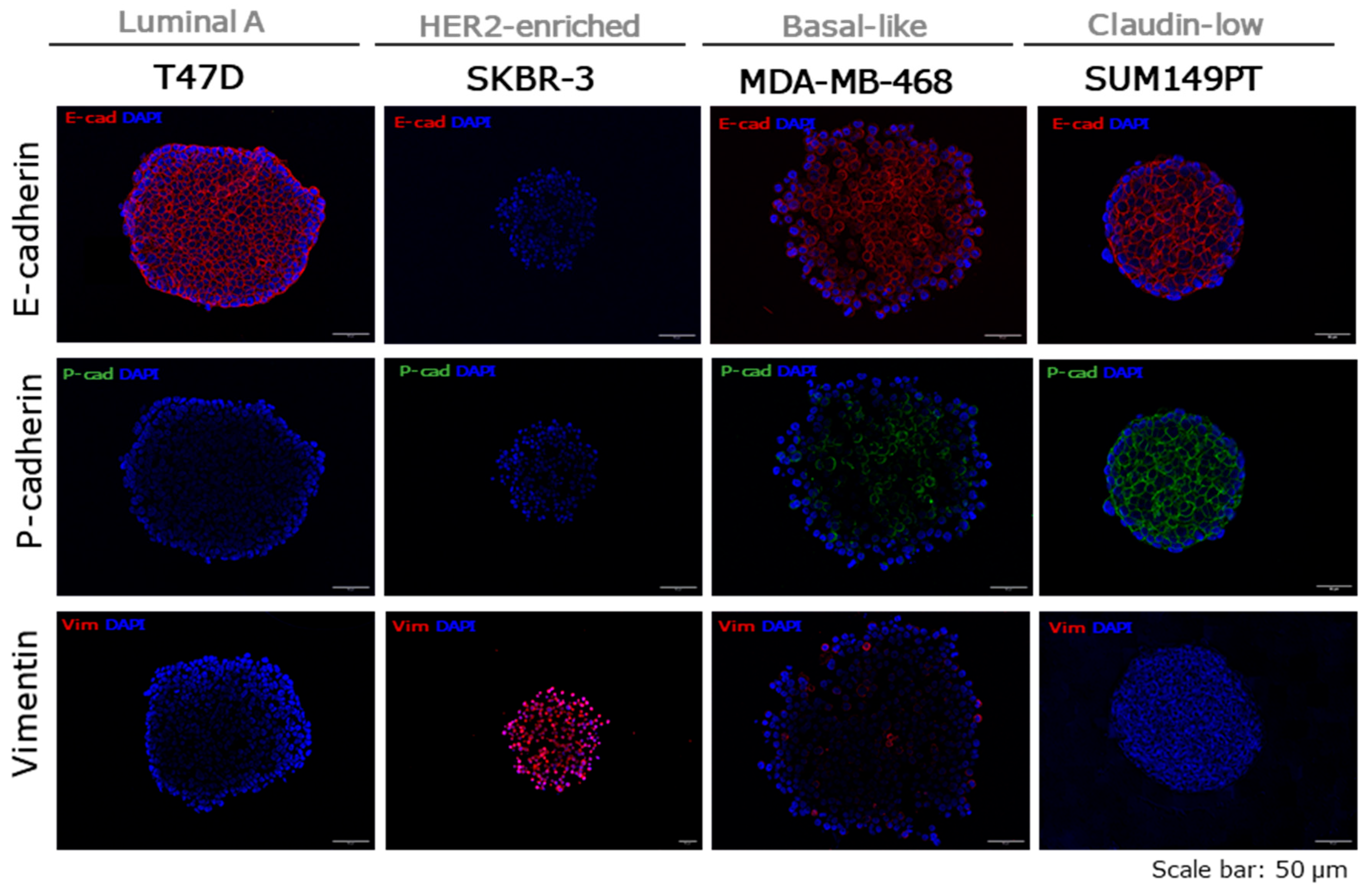
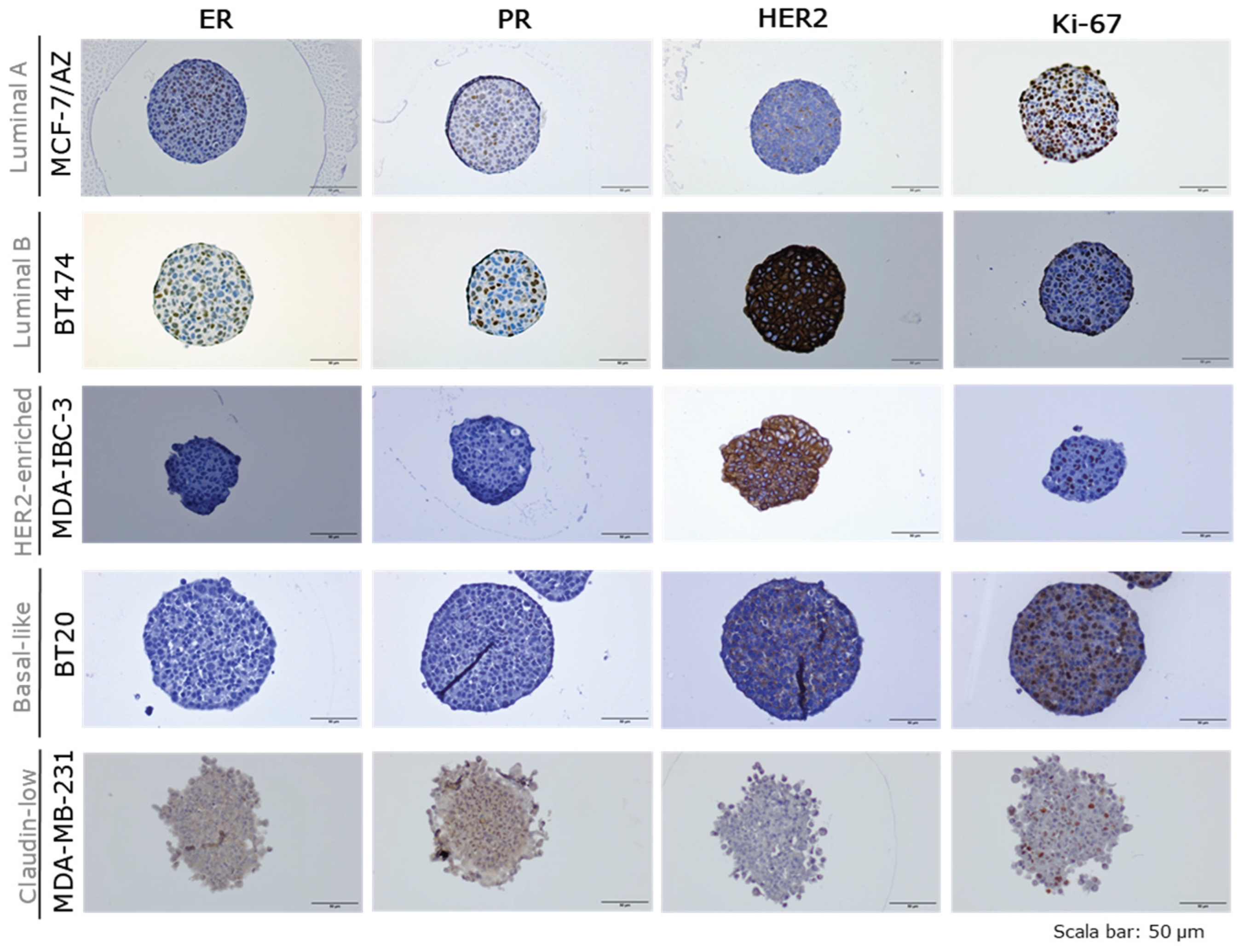
2.3. Molecular Characterization of Breast Cancer Tumour Spheroids
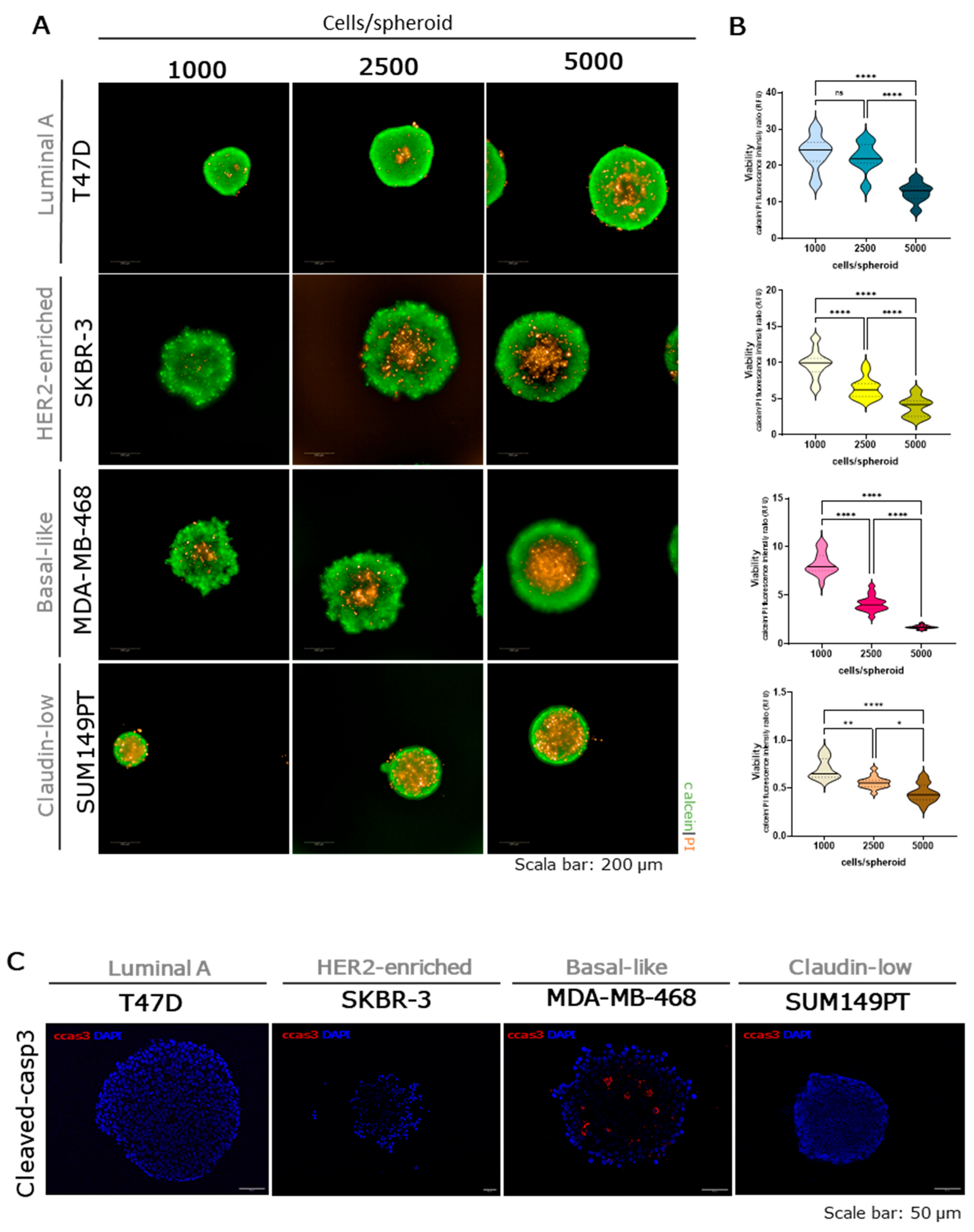
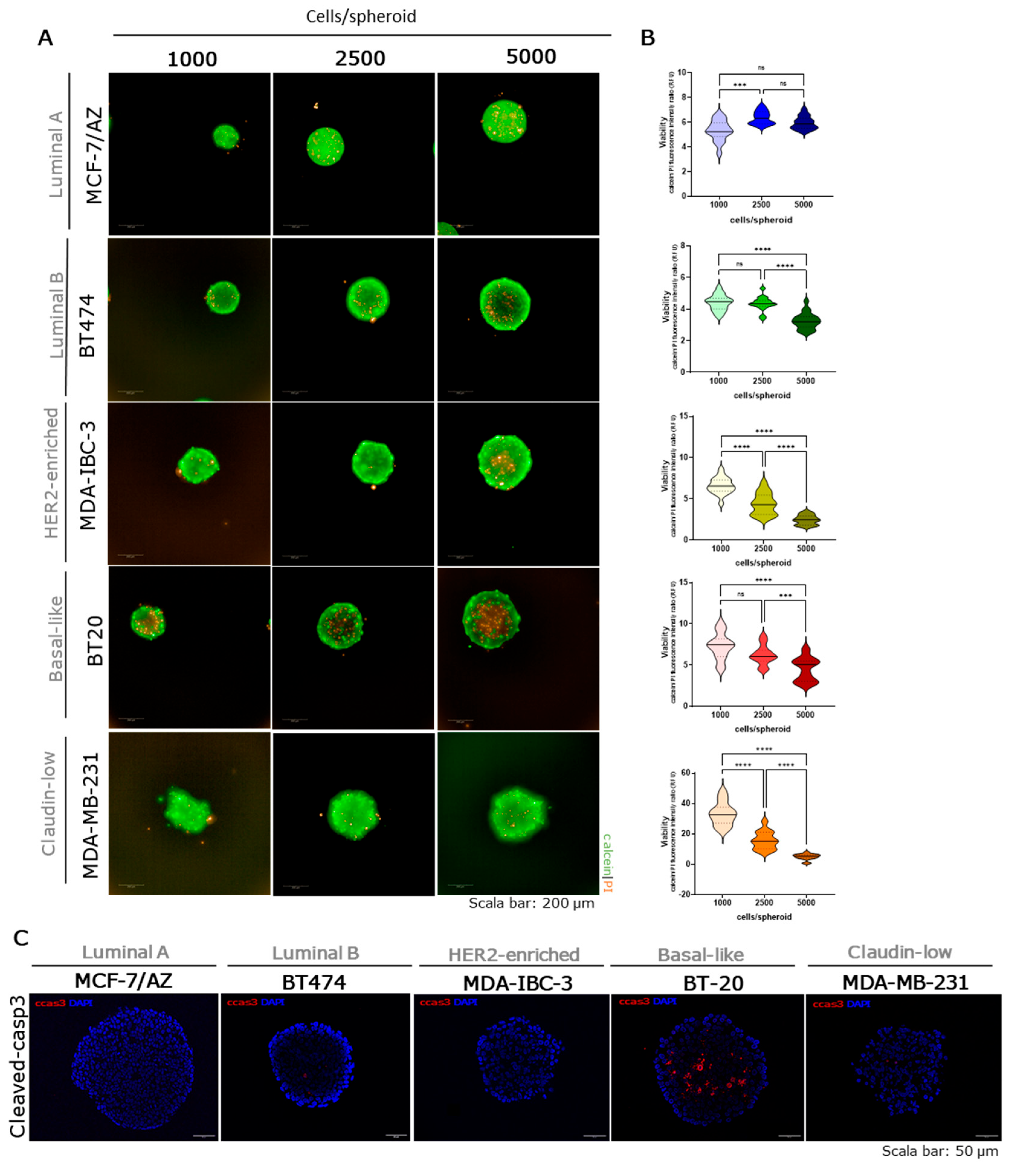
2.4. Viability and Cell Death in Breast Cancer Tumour Spheroids
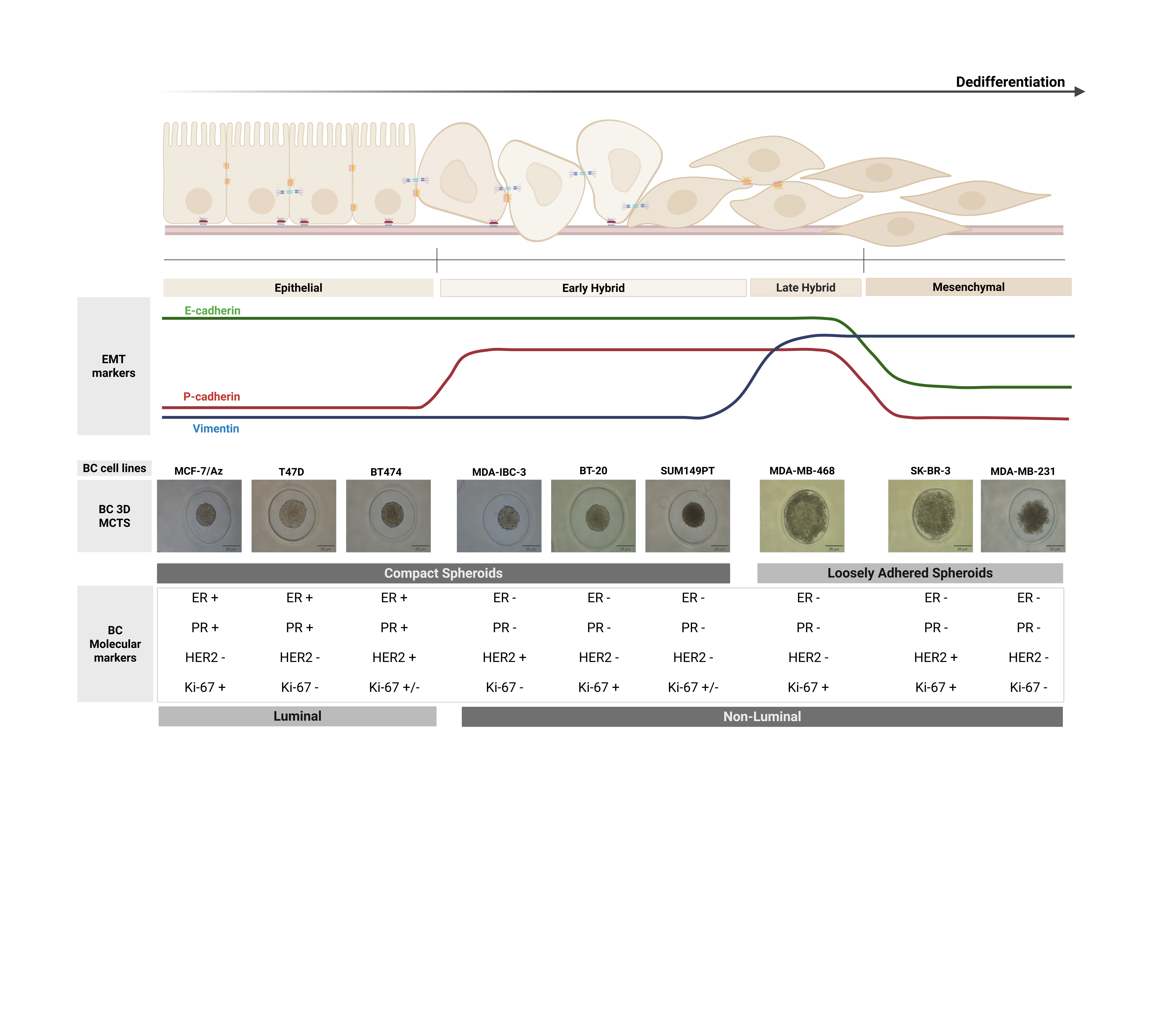
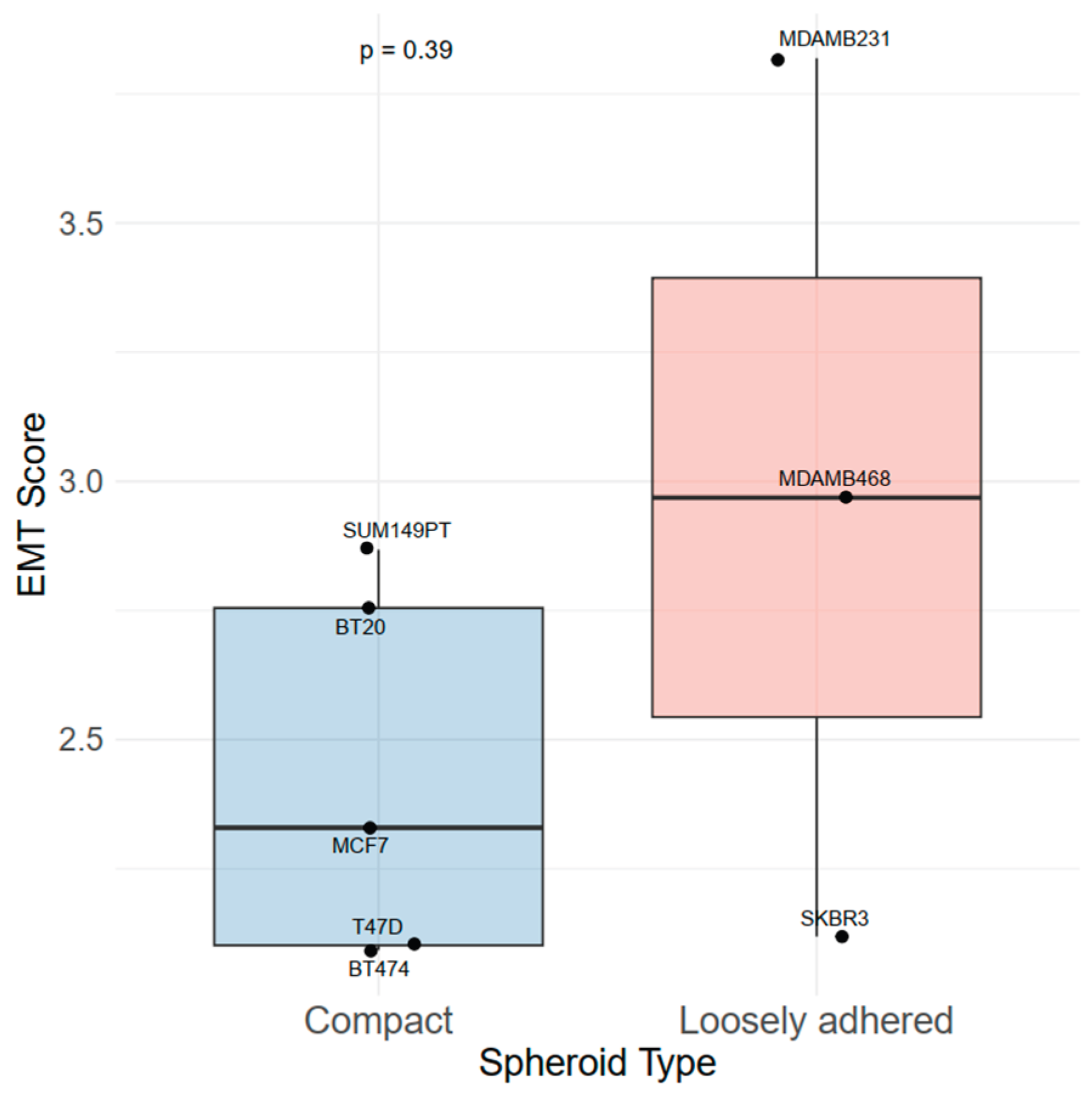
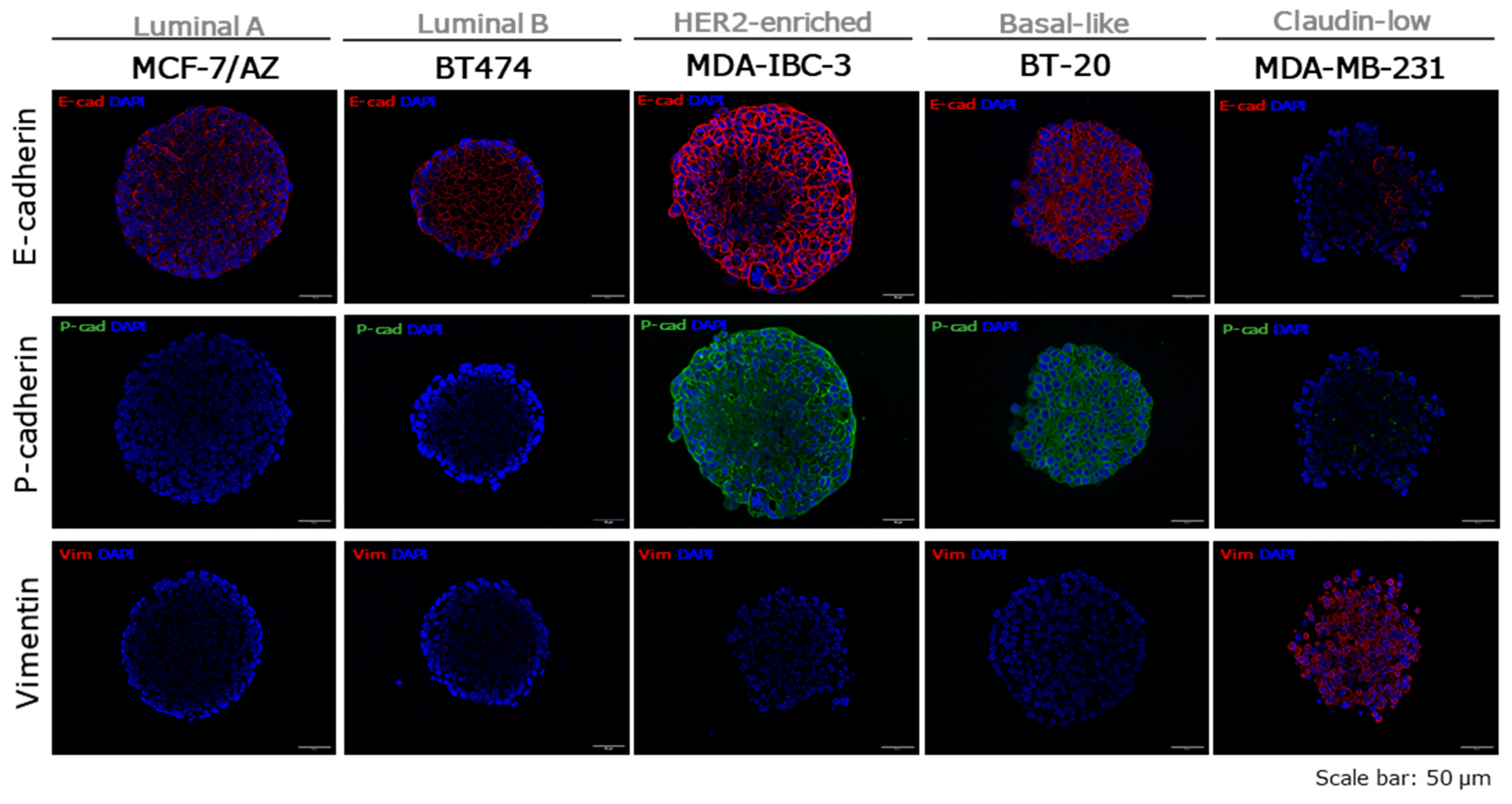
2.5. Epithelial-Mesenchymal Transition in Breast Cancer Tumour Spheroids
3. Discussion
4. Materials and Methods
4.1. Breast Cancer Cell Lines
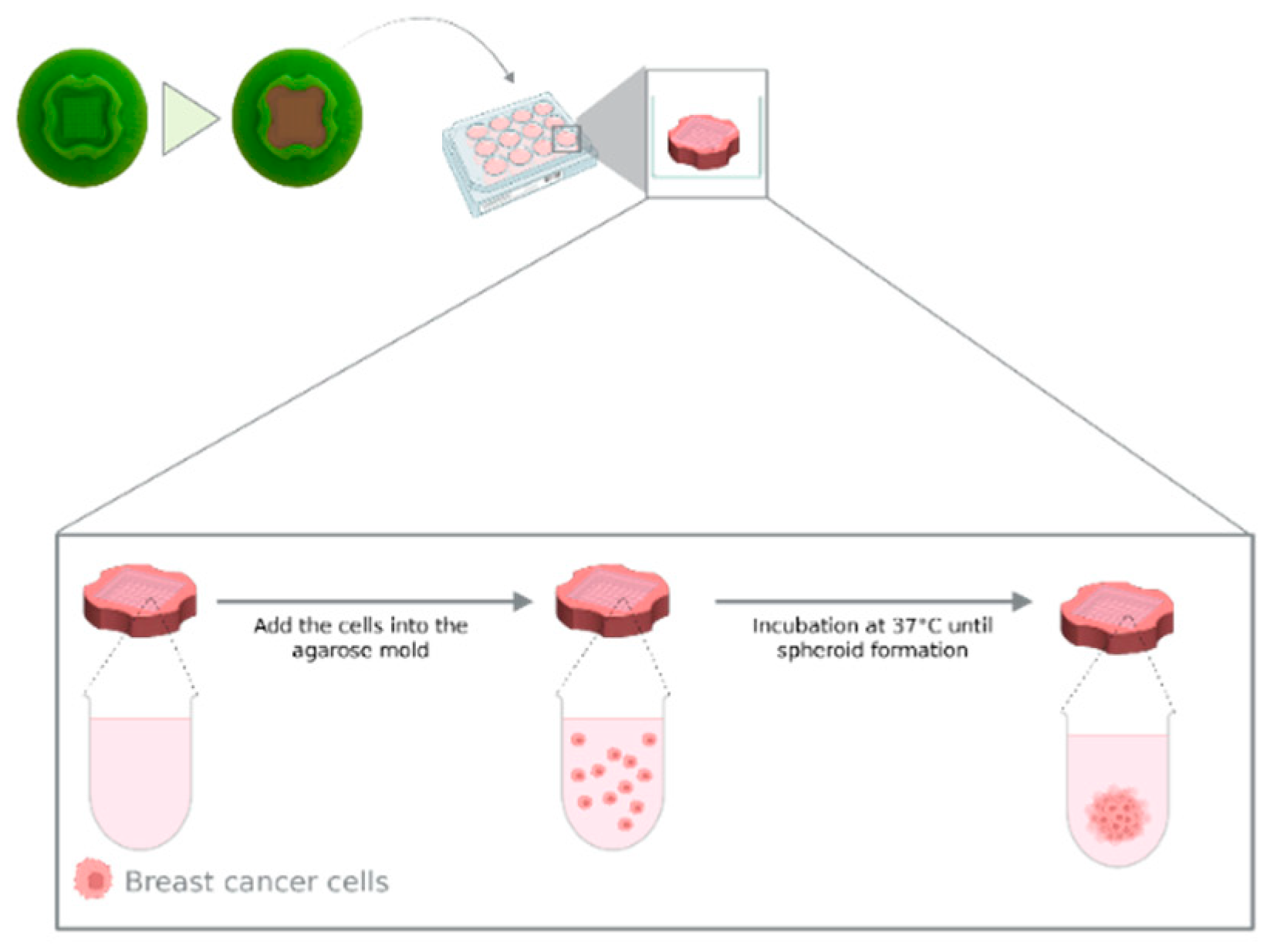
4.2. Breast Cancer Tumour Spheroids
4.3. Breast Cancer Spheroids’ Size Measurement
4.4. Breast Cancer Spheroids’ Histological Analysis
4.5. Breast Cancer Spheroids’ Immunohistochemistry
4.6. Cell Viability
4.7. Immunofluorescence
4.8. Bioinformatics Analysis
4.8.1. CCLE Dataset
4.8.2. EMT Gene Selection
4.8.3. EMT Score Calculation
4.8.4. Data Visualization
4.8.5. Data Processing
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Immunohistochemistry | ||||
| Protein | ER | PR | HER2 | Ki67 |
| Reference/Company | NCL-L-ER-6F11, Leica Biosystems | NCL-L-PGR, Leica Biosystems | Clone ab134182, Abcam | Clone MIB-1, Dako |
| Dilution | 1:50 | 1:100 | 1:500 | 1:250 |
| Immunofluorescence | ||||
| Protein | cCasp3 (asp175) (D3E9) | E-cadherin | P-cadherin | Vimentin |
| Reference/Company | 9579S, Cell Signaling, USA | 24E10, Cell Signaling, USA | Clone 56, BD Transduction, USA | D21H3, Cell Signaling, USA |
| Dilution | 1:200 | 1:100 | 1:100 | 1:400 |
References
- Bray, F; Laversanne, M; Sung, H; Ferlay, J; Siegel, RL; Soerjomataram, I; et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. In CA Cancer J Clin.; REQUESTEDJOURNAL:JOURNAL:15424863; STRING:PUBLICATION: WGROUP, 2024; Volume 74, pp. 229–263. [Google Scholar] [CrossRef]
- Xiong, X; Zheng, LW; Ding, Y; Chen, YF; Cai, YW; Wang, LP; et al. Breast cancer: pathogenesis and treatments. In Signal Transduct Target Ther.; SUBJMETA, 2025; Volume 10, pp. 1–33. [Google Scholar] [CrossRef]
- Perou, CM; Sørile, T; Eisen, MB; Van De Rijn, M; Jeffrey, SS; Ress, CA; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Sørlie, T; Perou, CM; Tibshirani, R; Aas, T; Geisler, S; Johnsen, H; et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A 2001, 98, 10869–10874. [Google Scholar] [CrossRef]
- Prat, A; Perou, CM. Deconstructing the molecular portraits of breast cancer. Mol Oncol. 2010, 5, 5. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, E; Canberk, S; Schmitt, F; Vale, N. Molecular Subtypes and Mechanisms of Breast Cancer: Precision Medicine Approaches for Targeted Therapies. Cancers 2025, Vol 17 17, 1102. [Google Scholar] [CrossRef]
- Lopez-Tarruella, S; Del Monte-Millán, M; Roche-Molina, M; Jerez, Y; Echavarria Diaz-Guardamino, I; Herrero López, B; et al. Correlation between breast cancer subtypes determined by immunohistochemistry and n-COUNTER PAM50 assay: a real-world study. Breast Cancer Res Treat. 2024, 203, 163–172. [Google Scholar] [CrossRef]
- Gutierrez, C; Schiff, R. HER2: Biology, Detection, and Clinical Implications. Arch Pathol Lab Med. 2011, 135, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Schettini, F; Prat, A. Dissecting the biological heterogeneity of HER2-positive breast cancer. The Breast 2021, 59, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Romond, EH; Perez, EA; Bryant, J; Suman, VJ; Geyer, CE; Davidson, NE; et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med. 2005, 353, 1673–1684. [Google Scholar] [CrossRef]
- Piccart-Gebhart, MJ; Procter, M; Leyland-Jones, B; Goldhirsch, A; Untch, M; Smith, I; et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005, 353, 1659–1672. [Google Scholar] [CrossRef]
- Podo, F; Buydens, LMC; Degani, H; Hilhorst, R; Klipp, E; Gribbestad, IS; et al. Triple-negative breast cancer: present challenges and new perspectives. Mol Oncol. 2010, 4, 209–229. [Google Scholar] [CrossRef]
- Nielsen, TO; Hsu, FD; Jensen, K; Cheang, M; Karaca, G; Hu, Z; et al. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res. 2004, 10, 5367–5374. [Google Scholar] [CrossRef]
- Prat, A; Parker, JS; Karginova, O; Fan, C; Livasy, C; Herschkowitz, JI; et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res 2010 125 2010, 12, R68. [Google Scholar] [CrossRef]
- Zanoni, M; Cortesi, M; Zamagni, A; Arienti, C; Pignatta, S; Tesei, A. Modeling neoplastic disease with spheroids and organoids. J Hematol Oncol. 2020, 13, 1–15. [Google Scholar] [CrossRef]
- Boix-Montesinos, P; Soriano-Teruel, PM; Armiñán, A; Orzáez, M; Vicent, MJ. The past, present, and future of breast cancer models for nanomedicine development. Adv Drug Deliv Rev. 2021, 173, 306–330. [Google Scholar] [CrossRef]
- Katt, ME; Placone, AL; Wong, AD; Xu, ZS; Searson, PC. In vitro tumor models: Advantages, disadvantages, variables, and selecting the right platform. Front Bioeng Biotechnol. 2016, 4. [Google Scholar] [CrossRef]
- Han, SJ; Kwon, S; Kim, KS. Challenges of applying multicellular tumor spheroids in preclinical phase. Cancer Cell Int. 2021, 21, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Fennema, E; Rivron, N; Rouwkema, J; van Blitterswijk, C; De Boer, J. Spheroid culture as a tool for creating 3D complex tissues. Trends Biotechnol. 2013, 31, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Pinto, B; Henriques, AC; Silva, PMA; Bousbaa, H. Three-Dimensional Spheroids as In Vitro Preclinical Models for Cancer Research. Pharm 2020, Vol 12 12, 1186. [Google Scholar] [CrossRef] [PubMed]
- Hirschhaeuser, F; Menne, H; Dittfeld, C; West, J; Mueller-Klieser, W; Kunz-Schughart, LA. Multicellular tumor spheroids: An underestimated tool is catching up again. J Biotechnol. 2010, 148, 3–15. [Google Scholar] [CrossRef]
- Neve, RM; Chin, K; Fridlyand, J; Yeh, J; Baehner, FL; Fevr, T; et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell. 2006, 10, 515–527. [Google Scholar] [CrossRef]
- Bauleth-Ramos, T; Feijão, T; Gonçalves, A; Shahbazi, MA; Liu, Z; Barrias, C; et al. Colorectal cancer triple co-culture spheroid model to assess the biocompatibility and anticancer properties of polymeric nanoparticles. J Control Release 2020, 323, 398–411. [Google Scholar] [CrossRef]
- Domingues, M; Leite Pereira, C; Sarmento, B; Castro, F. Mimicking 3D breast tumor-stromal interactions to screen novel cancer therapeutics. Eur J Pharm Sci. 2023, 190, 106560. [Google Scholar] [CrossRef] [PubMed]
- Pastushenko, I; Blanpain, C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol. 2019, 29, 212–226. [Google Scholar] [CrossRef]
- Lambert, AW; Weinberg, RA. Linking EMT programmes to normal and neoplastic epithelial stem cells. In Nat Rev Cancer; SUBJMETA, 2021; Volume 21, pp. 325–338. [Google Scholar] [CrossRef]
- Ribeiro, AS; Paredes, J. P-Cadherin Linking Breast Cancer Stem Cells and Invasion: A Promising Marker to Identify an “Intermediate/Metastable” EMT State. Front Oncol. 2015, 4, 371. [Google Scholar] [CrossRef]
- Jolly, MK; Boareto, M; Debeb, BG; Aceto, N; Farach-Carson, MC; Woodward, WA; et al. Inflammatory breast cancer: A model for investigating cluster-based dissemination. In npj Breast Cancer; SUBJMETA, 2017; Volume 3, pp. 1–8. [Google Scholar] [CrossRef]
- Canário, R; Ribeiro, AS; Morgado, I; Peixoto, A; Barbosa, A; Santos, C; et al. P-cadherin overexpression is associated with early transformation of the Fallopian tube epithelium and aggressiveness of tubo-ovarian high-grade serous carcinoma. Virchows Arch. cited. 2025. [CrossRef]
- Jolly, MK; Tripathi, SC; Jia, D; Mooney, SM; Celiktas, M; Hanash, SM; et al. Stability of the hybrid epithelial/mesenchymal phenotype. Oncotarget 2016, 7, 27067–27084. [Google Scholar] [CrossRef] [PubMed]
- Falkenberg, N; Höfig, I; Rosemann, M; Szumielewski, J; Richter, S; Schorpp, K; et al. Three-dimensional microtissues essentially contribute to preclinical validations of therapeutic targets in breast cancer. Cancer Med. 2016, 5, 703–710. [Google Scholar] [CrossRef] [PubMed]
- T47D Cell Line Spheroid Generation and Characterization for HT Assays | Thermo Fisher Scientific - PT. 6 Oct 2025. Available online: https://www.thermofisher.com/pt/en/home/references/protocols/cell-culture/3-d-cell-culture-protocol/t47d-cell-line-spheroid-generation.html?utm_source=chatgpt.com.
- Kenny, PA; Lee, GY; Myers, CA; Neve, RM; Semeiks, JR; Spellman, PT; et al. The morphologies of breast cancer cell lines in three-dimensional assays correlate with their profiles of gene expression. Mol Oncol. 2007, 1, 84–96. [Google Scholar] [CrossRef]
- Froehlich, K; Haeger, JD; Heger, J; Pastuschek, J; Photini, SM; Yan, Y; et al. Generation of Multicellular Breast Cancer Tumor Spheroids: Comparison of Different Protocols. J Mammary Gland Biol Neoplasia 2016, 21, 89–98. [Google Scholar] [CrossRef]
- Gong, X; Lin, C; Cheng, J; Su, J; Zhao, H; Liu, T; et al. Generation of Multicellular Tumor Spheroids with Microwell-Based Agarose Scaffolds for Drug Testing. PLoS One 2015, 10, e0130348. [Google Scholar] [CrossRef]
- Cavaco, M; Fraga, P; Valle, J; Andreu, D; Castanho, MARB; Neves, V. Development of breast cancer spheroids to evaluate cytotoxic response to an anticancer peptide. Pharmaceutics 2021, 13, 1863. [Google Scholar] [CrossRef]
- Lee, GY; Kenny, PA; Lee, EH; Bissell, MJ. Three-dimensional culture models of normal and malignant breast epithelial cells Three-dimensional. Nat Methods 2010, 4, 359–365. [Google Scholar] [CrossRef]
- Hofmann, S; Cohen-Harazi, R; Maizels, Y; Koman, I. Patient-derived tumor spheroid cultures as a promising tool to assist personalized therapeutic decisions in breast cancer. Transl Cancer Res. 2022, 11, 134–147. [Google Scholar] [CrossRef]
- Koedoot, E; Wolters, L; Smid, M; Stoilov, P; Burger, GA; Herpers, B; et al. Differential reprogramming of breast cancer subtypes in 3D cultures and implications for sensitivity to targeted therapy. In Sci Rep.; SUBJMETA, 2021; Volume 11, pp. 1–13. [Google Scholar] [CrossRef]
- Pickl, M; Ries, CH. Comparison of 3D and 2D tumor models reveals enhanced HER2 activation in 3D associated with an increased response to trastuzumab. Oncogene KWRD. 2009, 28, 461–468. [Google Scholar] [CrossRef]
- Krause, S; Maffini, M V.; Soto, AM; Sonnenschein, C. The microenvironment determines the breast cancer cells’ phenotype: Organization of MCF7 cells in 3D cultures. BMC Cancer 2010, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Imamura, Y; Mukohara, T; Shimono, Y; Funakoshi, Y; Chayahara, N; Toyoda, M; et al. Comparison of 2D- and 3D-culture models as drug-testing platforms in breast cancer. Oncol Rep. 2015, 33, 1837–1843. [Google Scholar] [CrossRef] [PubMed]
- Peirsman, A; Blondeel, E; Ahmed, T; Anckaert, J; Audenaert, D; Boterberg, T; et al. MISpheroID: a knowledgebase and transparency tool for minimum information in spheroid identity. Nat Methods 2021, 18, 1294. [Google Scholar] [CrossRef] [PubMed]
- Kirkegaard, T; Hansen, SK; Larsen, SL; Reiter, BE; Sørensen, BS; Lykkesfeldt, AE. T47D breast cancer cells switch from ER/HER to HER/c-Src signaling upon acquiring resistance to the antiestrogen fulvestrant. Cancer Lett. 2014, 344, 90–100. [Google Scholar] [CrossRef]
- Singh, M; Mukundan, S; Jaramillo, M; Oesterreich, S; Sant, S. Three-dimensional breast cancer models mimic hallmarks of size-induced tumor progression. Cancer Res. 2016, 76, 3732. [Google Scholar] [CrossRef]
- Park, HJ; Helfman, DM. Up-regulated fibronectin in 3D culture facilitates spreading of triple negative breast cancer cells on 2D through integrin β-5 and Src. Sci Rep. TECHMETA. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Yersal, O; Barutca, S. Biological subtypes of breast cancer: Prognostic and therapeutic implications. World J Clin Oncol. 2014, 5, 412–424. [Google Scholar] [CrossRef]
- Leung, BM; Lesher-Perez, SC; Matsuoka, T; Moraes, C; Takayama, S. Media additives to promote spheroid circularity and compactness in hanging drop platform. Biomater Sci. 2015, 3, 336–344. [Google Scholar] [CrossRef]
- Zanoni, M; Piccinini, F; Arienti, C; Zamagni, A; Santi, S; Polico, R; et al. 3D tumor spheroid models for in vitro therapeutic screening: a systematic approach to enhance the biological relevance of data obtained. Sci Reports 2016 61 2016, 6, 19103. [Google Scholar] [CrossRef]
- Mazloomi, MA; Hamishehkar, H; Jahanban Esfahlan, R. SpheroidSync as edge cutting transfer strategy for uniform and robust MCF7 spheroids in 3D culture. Sci Reports 2025 151 2025, 15, 41237. [Google Scholar] [CrossRef]
- Zhu, S; Yin, J; Lu, X; Jiang, D; Chen, R; Cui, K; et al. Influence of experimental variables on spheroid attributes. Sci Reports 2025 151 2025, 15, 9751. [Google Scholar] [CrossRef]
- Lerma, E; Peiro, G; Ramón, T; Fernandez, S; Martinez, D; Pons, C; et al. Immunohistochemical heterogeneity of breast carcinomas negative for estrogen receptors, progesterone receptors and Her2/neu (basal-like breast carcinomas). Mod Pathol. 2007, 20, 1200–1207. [Google Scholar] [CrossRef]
- Hollestelle, A; Elstrodt, F; Timmermans, M; Sieuwerts, AM; Klijn, JGM; Foekens, JA; et al. Four human breast cancer cell lines with biallelic inactivating α-catenin gene mutations. Breast Cancer Res Treat. 2010, 122, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Mangani, S; Koutsakis, C; Koletsis, NE; Piperigkou, Z; Franchi, M; Götte, M; et al. Spheroid-Based 3D Models to Decode Cell Function and Matrix Effectors in Breast Cancer. Cancers 2025, Vol 17, 17. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z; Yu, P; Tang, J. Characterization of Triple-Negative Breast Cancer MDA-MB-231 Cell Spheroid Model. Onco Targets Ther. 2020, 13, 5395–5405. [Google Scholar] [CrossRef]
- Arnaldi, P; Zotti, VD; Bellese, G; Gagliani, MC; Orecchia, P; Castagnola, P; et al. 3D Breast Cancer Spheroids Reveal Architecture-Dependent HER2 Expression and Signaling. Biol 2025, Vol 14 14, 1654. [Google Scholar] [CrossRef]
- Piwocka, O; Sterzyńska, K; Malińska, A; Suchorska, WM; Kulcenty, K. Development of tetraculture spheroids as a versatile 3D model for personalized breast cancer research. In Sci Rep.; SUBJMETA, 2025; Volume 15, pp. 1–14. [Google Scholar] [CrossRef]
- Ahvaraki, A; Gheytanchi, E; Behroodi, E; Latifi, H; Vakhshiteh, F; Bagheri, Z; et al. Advanced co-culture 3D breast cancer model to study cell death and nanodrug sensitivity of tumor spheroids. Biochem Eng J 2024, 209, 109400. [Google Scholar] [CrossRef]
- Diosdi, A; Piccinini, F; Boroczky, T; Dobra, G; Castellani, G; Buzas, K; et al. Single-cell light-sheet fluorescence 3D images of tumour-stroma spheroid multicultures. Sci Data 2025 121 2025, 12, 492. [Google Scholar] [CrossRef]
- Saraiva, DP; Matias, AT; Braga, S; Jacinto, A; Cabral, MG. Establishment of a 3D Co-culture With MDA-MB-231 Breast Cancer Cell Line and Patient-Derived Immune Cells for Application in the Development of Immunotherapies. Front Oncol. 2020, 10, 561826. [Google Scholar] [CrossRef] [PubMed]
- Ortiz, E; Thway, KH; Ortiz-Soto, G; Yao, P; Kelber, JA. Hetero-multicellular stromal cells incorporate into scaffold-free 3D cultures of epithelial cancer cells to drive invasion. bioRxiv 2025, 2025.01.21.634082. [Google Scholar] [CrossRef]
- Chen, G; Liu, W; Yan, B; Chen, G; Liu, W; Yan, B. Breast Cancer MCF-7 Cell Spheroid Culture for Drug Discovery and Development. J Cancer Ther. 2022, 13, 117–130. [Google Scholar] [CrossRef] [PubMed]
- Coelho, LL; Vianna, MM; da Silva, DM; Gonzaga, BM de S; Ferreira, RR; Monteiro, AC; et al. Spheroid Model of Mammary Tumor Cells: Epithelial–Mesenchymal Transition and Doxorubicin Response. Biol 2024, Vol 13 13. [Google Scholar] [CrossRef]
- Lin, YN; Nasir, A; Camacho, S; Berry, DL; Schmidt, MO; Pearson, GW; et al. Monitoring cancer cell invasion and t-cell cytotoxicity in 3d culture. J Vis Exp. 2020, 2020, 1–19. [Google Scholar] [CrossRef]
- Silveira, MJ; Martins, C; Cardoso, AP; Ankone, MJK; Blyth, RRR; Oliveira, MJ; et al. Immunostimulatory effects of IL-12 targeted pH-responsive nanoparticles in macrophage-enriched 3D immuno-spheroids in vitro model. In Drug Deliv Transl Res; 2025; pp. 1–20. [Google Scholar] [CrossRef]
- Kessel, S; Cribbes, S; Déry, O; Kuksin, D; Sincoff, E; Qiu, J; et al. High-Throughput 3D Tumor Spheroid Screening Method for Cancer Drug Discovery Using Celigo Image Cytometry. SLAS Technol. 2017, 22, 454–465. [Google Scholar] [CrossRef]
| Molecular subtype | |||||
| Cell line | Disease | Source | Molecular markers | Neve et al& | Prat A et al#* |
| MCF-7/AZ | Invasive breast carcinoma |
Pleural Effusion |
ER+/PR+/HER2- |
Luminal |
Luminal |
| T47D | Invasive breast carcinoma |
Pleural Effusion |
ER+/PR+/HER2- | ||
| BT474 | Invasive breast carcinoma |
Primary tumour |
ER+/PR+/HER2+ | HER2-enriched | |
| SK-BR-3 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2+ | HER2-enriched | |
| MDA-IBC-3 | Breast inflammatory carcinoma | Pleural effusion |
ER-/PR-/HER2+ | - | HER2-enriched |
| BT20 | Invasive breast carcinoma |
Primary tumour |
ER-/PR-/HER2- | Basal A | Basal |
| MDA-MB-468 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2- | Basal A | Basal |
| SUM149PT | Breast inflammatory carcinoma | Primary tumour |
ER-/PR-/HER2- | Basal B | Basal/Claudin-low |
| MDA-MB-231 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2- | Basal B | Claudin-low |
| Molecular subtype | |||||
| Cell line | Disease | Source | Molecular markers | Neve et al& | Prat A et al#* |
| MCF-7/AZ | Invasive breast carcinoma |
Pleural Effusion |
ER+/PR+/HER2- |
Luminal |
Luminal |
| T47D | Invasive breast carcinoma |
Pleural Effusion |
ER+/PR+/HER2- | ||
| BT474 | Invasive breast carcinoma |
Primary tumour |
ER+/PR+/HER2+ | HER2-enriched | |
| SK-BR-3 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2+ | HER2-enriched | |
| MDA-IBC-3 | Breast inflammatory carcinoma | Pleural effusion |
ER-/PR-/HER2+ | - | HER2-enriched |
| BT20 | Invasive breast carcinoma |
Primary tumour |
ER-/PR-/HER2- | Basal A | Basal |
| MDA-MB-468 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2- | Basal A | Basal |
| SUM149PT | Breast inflammatory carcinoma | Primary tumour |
ER-/PR-/HER2- | Basal B | Basal/Claudin-low |
| MDA-MB-231 | Breast adenocarcinoma |
Pleural effusion |
ER-/PR-/HER2- | Basal B | Claudin-low |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




