Submitted:
26 March 2026
Posted:
27 March 2026
You are already at the latest version
Abstract
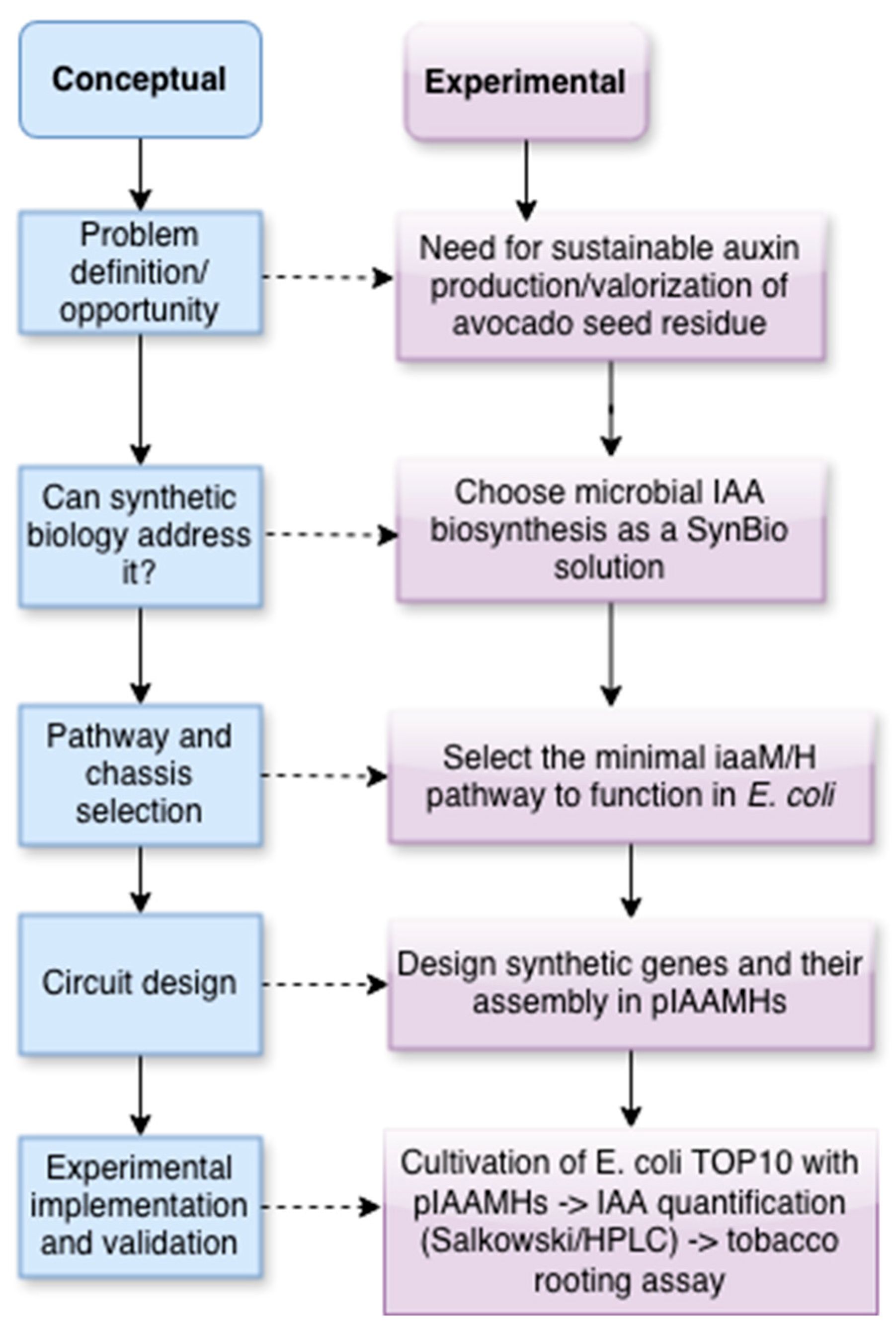
Indole-3-acetic acid (IAA) is the main natural auxin and a key regulator of plant growth. However, most commercial auxins are synthetically produced from non-renewable resources. Here, we present a minimal synthetic biology platform for microbial IAA production that also serves as a teaching model for genetic circuit design and bioprocess development. We developed codon-optimized versions of the iaaM and iaaH genes, which encode tryptophan 2-monooxygenase and indole-3-acetamide hydrolase, and assembled them into a compact expression cassette in Escherichia coli TOP10. Correct expression of both enzymes was confirmed by SDS-PAGE. The engineered strain was cultivated in a low-cost medium made from avocado seed hydrolysate, an agro-industrial waste, supplemented with tryptophan as a precursor. IAA levels of about 300 µg/mL were measured after 48 hours using the Salkowski assay and HPLC, with the medium costing five times less locally than traditional LB. The supernatants containing biosynthetic IAA induced strong root formation in tobacco leaf explants, confirming biological activity. Since this workflow follows the Design–Build–Test–Learn (DBTL) cycle: Design (pathway selection and codon optimization), Build (plasmid assembly), Test (protein expression, metabolite quantification, plant bioassays), and Learn (medium and process optimization), it provides a sustainable production method and an accessible educational platform for synthetic biology.
Keywords:
1. Introduction
2. Results
2.1. Assembly of the Genetic Circuit for IAA Synthesis in E. coli
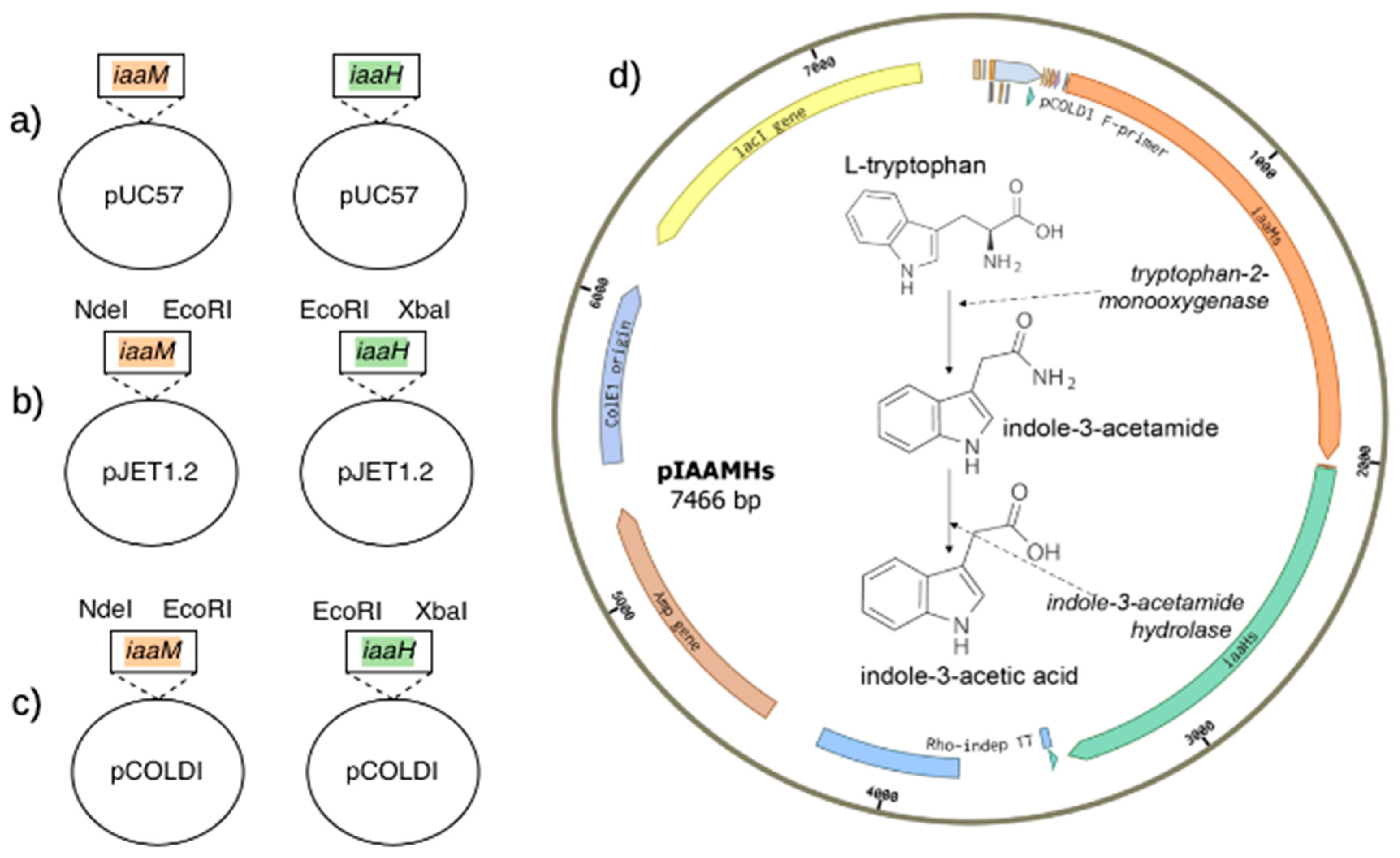
2.1.1. Design to Complete the Metabolic Pathway for IAA Synthesis from Tryptophan Precursor in E. coli
2.1.2. Construction of the Plasmid pIAAMHs
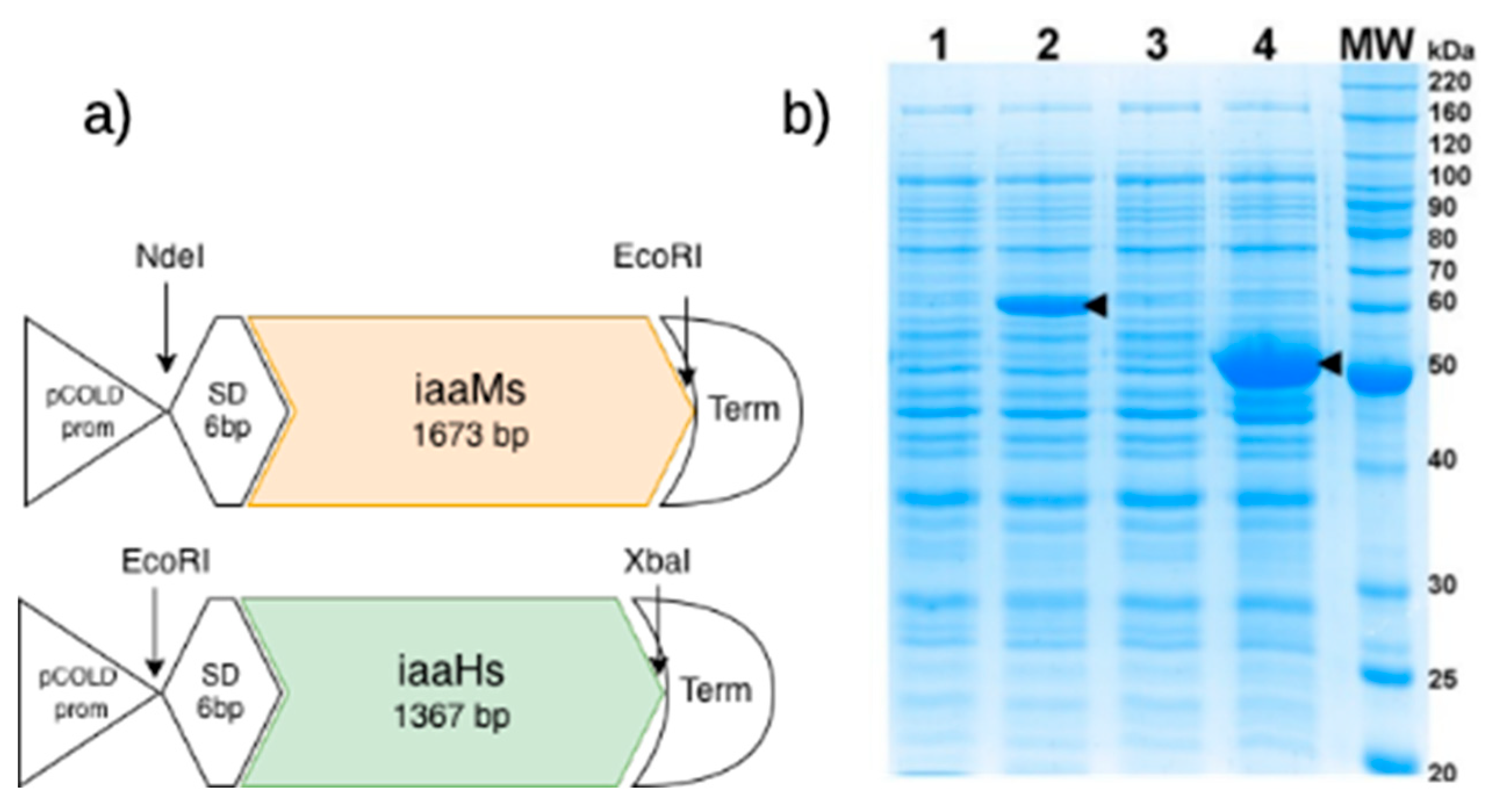
2.2. Verification of Correct Protein Expression for IAA Synthesis in E. coli
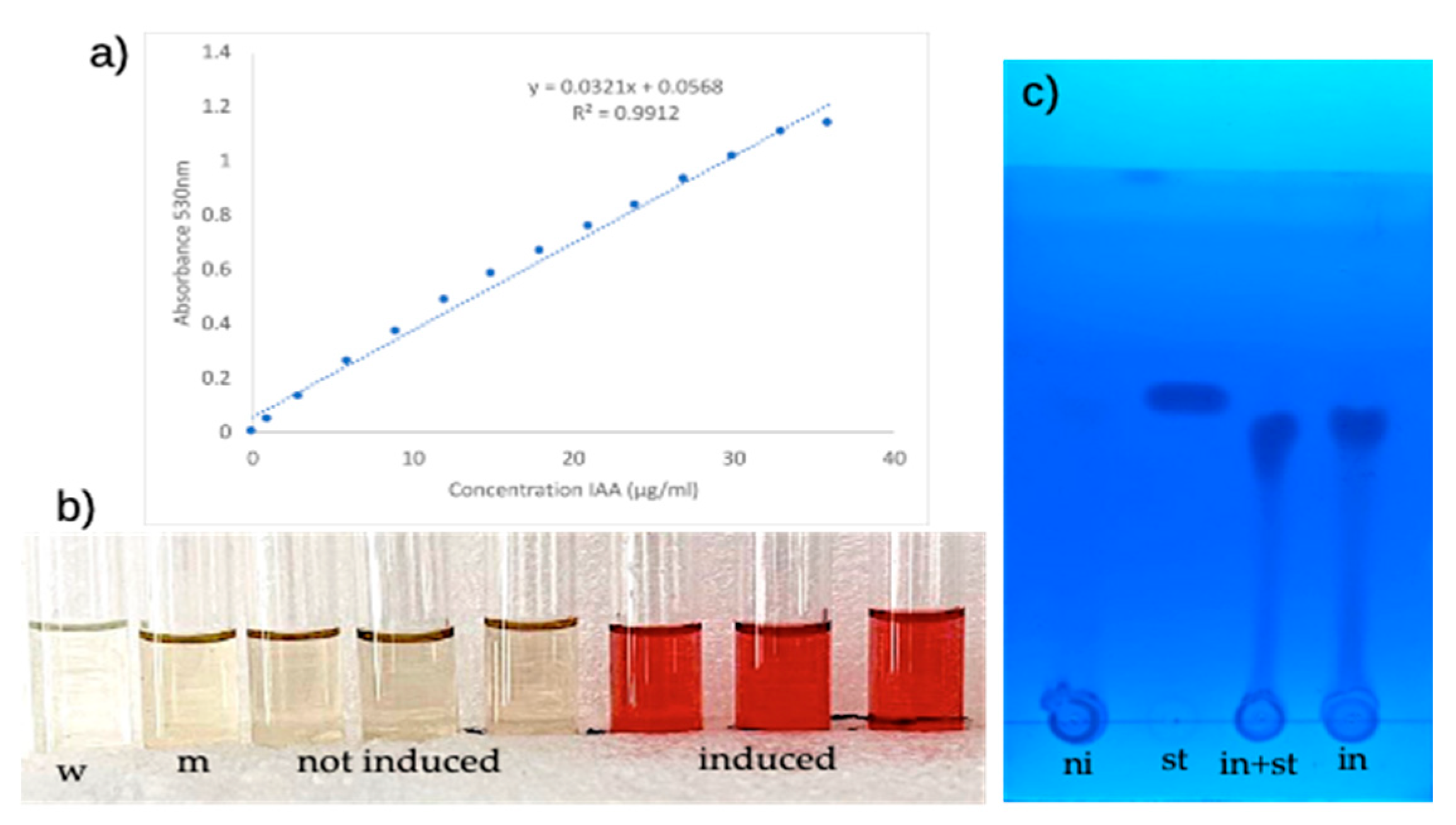
2.3. IAA Production in Escherichia coli
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains and Media
4.2. Gene Synthesis
4.3. Bacterial Culture for Protein Expression
4.4. Protein Electrophoresis
4.5. Bacterial Culture for IAA Production
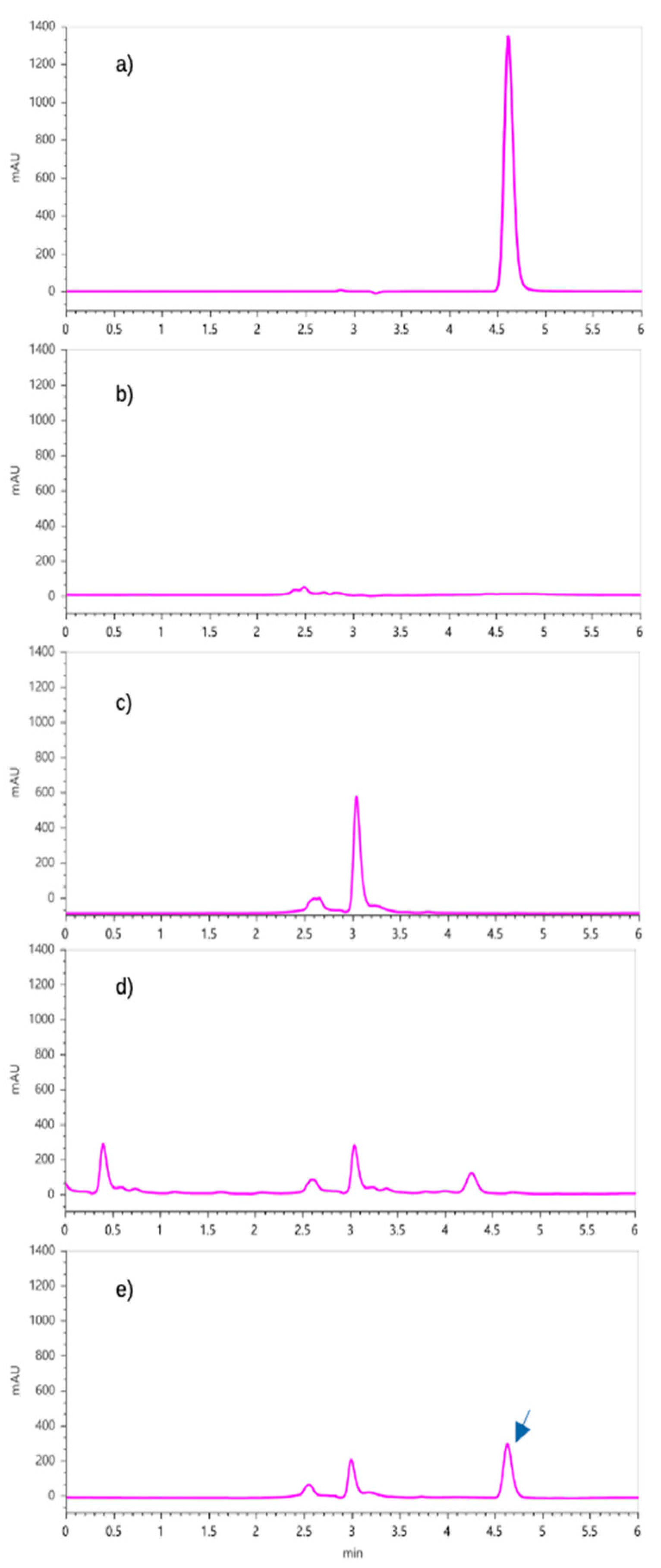
4.6. Qualitative Identification of Produced IAA
4.7. HPLC Quantification of IAA
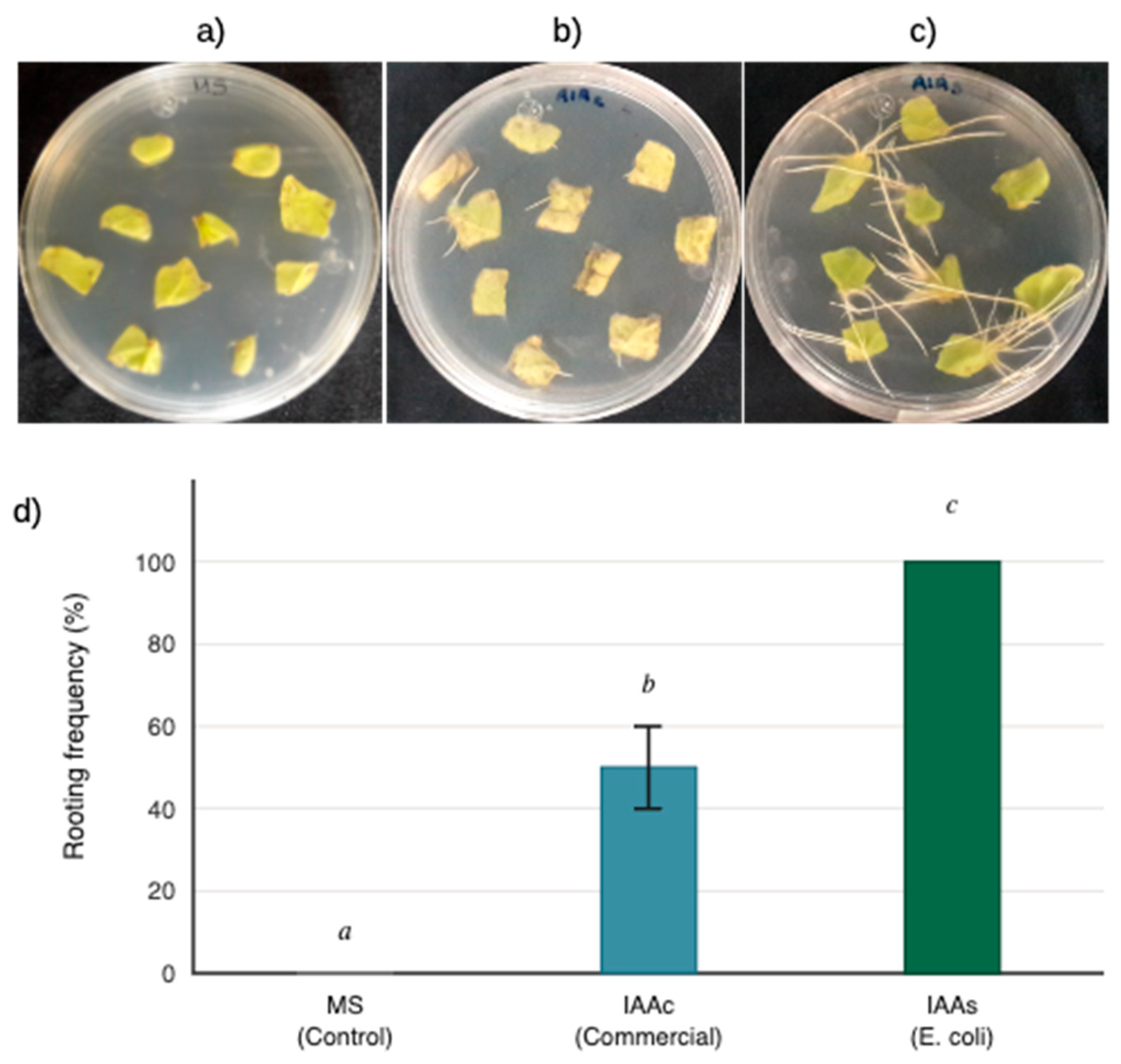
4.8. In Vitro Rooting Activity of IAA
5. Conclusions
Data Availability
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Islam, S. Agriculture, Food Security, and Sustainability: A Review. Explor Foods Foodomics. 2025, 3, 101082. [Google Scholar] [CrossRef]
- Wang, L.; Garland, G.M.; Ge, T.; Guo, S.; Kebede, E.A.; He, C.; Hijri, M.; Plaza-Bonilla, D.; Stringer, L.C.; Davis, K.F.; et al. Integrated Strategies for Enhancing Agrifood Productivity, Lowering Greenhouse Gas Emissions, and Improving Soil Health. The Innovation 2025, 6, 101006. [Google Scholar] [CrossRef]
- Clauer, P.; Nou, A. X.; Toth, T.; Yu, Q.; Chemla, Y.; Boo, A.; Voigt, C. Synthetic Biology of Plants and Microbes for Agriculture, Environment, and Future Applications. Chemical Reviews 2026, 126(2), 895–1109. [Google Scholar] [CrossRef]
- Han, S.-W.; Yoshikuni, Y. Microbiome Engineering for Sustainable Agriculture: Using Synthetic Biology to Enhance Nitrogen Metabolism in Plant-Associated Microbes. Current Opinion in Microbiology 2022, 68, 102172. [Google Scholar] [CrossRef] [PubMed]
- Spaepen, S.; Vanderleyden, J.; Remans, R. Indole-3-Acetic Acid in Microbial and Microorganism-Plant Signaling. FEMS Microbiol Rev 2007, 31, 425–448. [Google Scholar] [CrossRef]
- Straits Research. Plant Growth Regulators Market Size, Share & Growth Report 2033. Available online: https://straitsresearch.com/report/plant-growth-regulators-market (accessed on 18 March 2026).
- Lyubushkina, I.V.; Kirichenko, K.A.; Polyakova, M.S.; Polyanskaya, I.V.; Zabanova, N.S.; Korsukova, A.V.; Pobezhimova, T.P.; Dudareva, L.V.; Rikhvanov, E.G.; Grabelnych, O.I. Synthetic Auxins Toxicity: Effects on Growth and Fatty Acid Composition in Etiolated and Green Spring Wheat Seedlings. Molecules 2025, 30. [Google Scholar] [CrossRef] [PubMed]
- European Commission. Proposal for a Regulation on the Sustainable Use of Plant Protection Products (SUR). Available online: https://www.pan-europe.info/eu-legislation/eu-pesticide-reduction-sustainable-use-regulation-sur (accessed on 18 March 2026).
- Wu, H.; Yang, J.; Shen, P.; Li, Q.; Wu, W.; Jiang, X.; Qin, L.; Huang, J.; Cao, X.; Qi, F. High-Level Production of Indole-3-Acetic Acid in the Metabolically Engineered Escherichia Coli. J. Agric. Food Chem. 2021, 69, 1916–1924. [Google Scholar] [CrossRef] [PubMed]
- Romasi, E.F.; Lee, J. Development of Indole-3-Acetic Acid-Producing Escherichia Coli by Functional Expression of IpdC, AspC, and Iad1. J Microbiol Biotechnol 2013, 23, 1726–1736. [Google Scholar] [CrossRef]
- Guo, D.; Kong, S.; Chu, X.; Li, X.; Pan, H. De Novo Biosynthesis of Indole-3-Acetic Acid in Engineered Escherichia Coli. J. Agric. Food Chem. 2019, 67, 8186–8190. [Google Scholar] [CrossRef]
- Cuadrado-Osorio, P.D.; Ramírez-Mejía, J.M.; Mejía-Avellaneda, L.F.; Mesa, L.; Bautista, E.J. Agro-Industrial Residues for Microbial Bioproducts: A Key Booster for Bioeconomy. Bioresource Technology Reports 2022, 20, 101232. [Google Scholar] [CrossRef]
- Liu, S.; Xu, J.-Z.; Zhang, W.-G. Advances and Prospects in Metabolic Engineering of Escherichia Coli for L-Tryptophan Production. World J Microbiol Biotechnol 2022, 38, 22. [Google Scholar] [CrossRef]
- Menon, N.; Richmond, D.; Rahman, M.R.; Menon, B.R.K. Versatile and Facile One-Pot Biosynthesis for Amides and Carboxylic Acids in E. Coli by Engineering Auxin Pathways of Plant Microbiomes. ACS Catal. 2022, 12, 2309–2319. [Google Scholar] [CrossRef]
- iGEM Registry of Standard Biological Parts. BBa_K515100: Auxin Biosynthesis Cassette (iaaM/iaaH). Available online: https://parts.igem.org/Part:BBa_K515100:Design (accessed on 21 March 2026).
- Woo, S.-G.; Kim, S.K.; Lee, S.-G.; Lee, D.-H. Engineering Probiotic Escherichia Coli for Inflammation-Responsive Indoleacetic Acid Production Using RiboJ-Enhanced Genetic Circuits. J Biol Eng 2025, 19, 10. [Google Scholar] [CrossRef]
- Mapinta, S.; Kongjaroon, S.; Trisrivirat, D.; Kesornpun, C.; Wu, J.; Chaiyen, P.; Weeranoppanant, N. Continuous-Flow Chemo-Enzymatic Gram-Scale Synthesis of Indole-3-Acetic Acid. Green Chem. 2025, 27, 793–803. [Google Scholar] [CrossRef]
- Tzintzun-Camacho, O.; Sánchez-Segura, L.; Minchaca-Acosta, A.Z.; Rosales-Colunga, L.M.; Hernández-Orihuela, A.; Martínez-Antonio, A. Development of Bacterial Culture Medium from Avocado Seed Waste. Revista Mexicana de Ingeniería Química 2016, 15, 831–842. Available online: https://www.redalyc.org/pdf/620/62048168014.pdf. [CrossRef]
- Carbonell, P.; Jervis, A.J.; Robinson, C.J.; Yan, C.; Dunstan, M.; Swainston, N.; Vinaixa, M.; Hollywood, K.A.; Currin, A.; Rattray, N.J.W.; et al. An Automated Design-Build-Test-Learn Pipeline for Enhanced Microbial Production of Fine Chemicals. Commun Biol 2018, 1, 66. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Palenzuela, P.; Matas, I.M.; Murillo, J.; López-Solanilla, E.; Bardaji, L.; Pérez-Martínez, I.; Rodríguez-Moskera, M.E.; Penyalver, R.; López, M.M.; Quesada, J.M.; et al. Annotation and Overview of the Pseudomonas Savastanoi Pv. Savastanoi NCPPB 3335 Draft Genome Reveals the Virulence Gene Complement of a Tumour-Inducing Pathogen of Woody Hosts. Environmental Microbiology 2010, 12, 1604–1620. [Google Scholar] [CrossRef]
- Davies, K. From the Bench to Benchling. GEN Edge 2020, 2, genedge.2.1.52. [Google Scholar] [CrossRef]
- Pacheco-Villalobos, D.; Díaz-Moreno, S.M.; van der Schuren, A.; Tamaki, T.; Kang, Y.H.; Gujas, B.; Novak, O.; Jaspert, N.; Li, Z.; Wolf, S.; et al. The Effects of High Steady State Auxin Levels on Root Cell Elongation in Brachypodium[OPEN]. Plant Cell 2016, 28, 1009–1024. [Google Scholar] [CrossRef]
- Indole-3-Acetic Acid (IAA) 98% TC Plant Growth Regulator. Available online: https://www.pomais.com/product/indole-3-acetic-acid-iaa-98-tc/ (accessed on 19 March 2026).
- Ramos-Valdovinos, M.A.; Salas-Navarrete, P.C.; Amores, G.R.; Hernández-Orihuela, A.L.; Martínez-Antonio, A. Qualitative Perturbation Analysis and Machine Learning: Elucidating Bacterial Optimization of Tryptophan Production. Algorithms 2024, 17. [Google Scholar] [CrossRef]
- Martínez-Moreno, F. Avocado Seed Hydrolysate as an Alternative Growth Medium for Fungi Hidrolizado de Semilla de Aguacate Como Medio de Cultivo Alternativo Para Hongos. 2021, 20, 569–580. Available online: https://rmiq.org/iqfvp/Numbers/V20/No2/Bio1951.pdf.
- Palmerín-Carreño, D.M.; Hernández-Orihuela, A.L.; Martínez-Antonio, A. Production of D-Lactate from Avocado Seed Hydrolysates by Metabolically Engineered Escherichia Coli JU15. Fermentation 2019, 5. [Google Scholar] [CrossRef]
- Sierra-Ibarra, E.; Leal-Reyes, L.J.; Huerta-Beristain, G.; Hernández-Orihuela, A.L.; Gosset, G.; Martínez-Antonio, A.; Martinez, A. Limited Oxygen Conditions as an Approach to Scale-up and Improve d and l-Lactic Acid Production in Mineral Media and Avocado Seed Hydrolysates with Metabolically Engineered Escherichia Coli. Bioprocess Biosyst Eng 2021, 44, 379–389. [Google Scholar] [CrossRef]
- Espinel-Ríos, S.; Palmerín-Carreño, D.M.; Hernández-Orihuela, A.L.; Martínez-Antonio, A. A Plackett-Burman Design for Substituting MRS Medium Components with Avocado Seed Hydrolysate for Growth and Lactic Acid Production by Lactobacillus Sp. Revista Mexicana de Ingeniería Química 2019, 18, 131–141. Available online: https://rmiq.org/iqfvp/Numbers/V18/No1/Bio108.pdf. [CrossRef]
- Johnson, K.C.; Sabel, J.L.; Cole, J.; Pruett, C.L.; Plymale, R.; Reyna, N.S. From Genetics to Biotechnology: Synthetic Biology as a Flexible Course-Embedded Research Experience. Biochem Mol Biol Educ 2022, 50, 580–591. [Google Scholar] [CrossRef]
- Beach, D.L.; Alvarez, C.J. Biotechnology by Design: An Introductory Level, Project-Based, Synthetic Biology Laboratory Program for Undergraduate Students. J Microbiol Biol Educ 2015, 16, 237–246. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.W.; Garcia-Morales, L.; Martins dos Santos, V.A.P.; Saccenti, E. Research-Driven Education: An Introductory Course to Systems and Synthetic Biology. Front. Syst. Biol. 2022, 2. Erratum in Front. Syst. Biol. 2022, 2,1114700. [Google Scholar] [CrossRef]
- Nsanzabera, F.; Liu, B. Multiplex Metabolic Engineering for Enhanced Indole-3-Acetic Acid Production via Optimized Biosynthetic Pathways in E. Coli. Process Biochemistry 2025, 157, 147–161. [Google Scholar] [CrossRef]
- Khani, M.-H.; Bagheri, M. Skimmed Milk as an Alternative for IPTG in Induction of Recombinant Protein Expression. Protein Expr Purif 2020, 170, 105593. [Google Scholar] [CrossRef]
- Briand, L.; Marcion, G.; Kriznik, A.; Heydel, J.M.; Artur, Y.; Garrido, C.; Seigneuric, R.; Neiers, F. A Self-Inducible Heterologous Protein Expression System in Escherichia Coli. Sci Rep 2016, 6, 33037. [Google Scholar] [CrossRef]
- Current Protocols in Molecular Biology - National Library of Medicine Institution. Available online: https://catalog.nlm.nih.gov/discovery/fulldisplay/alma996618043406676/01NLM_INST:01NLM_INST (accessed on 19 March 2026).
- Addgene: Vector Database - pCOLD-I. Available online: https://www.addgene.org/vector-database/2261/ (accessed on 19 March 2026).
| Reference | Strain | Pathway | Key genes | Medium | Trp added (g/L) | IAA titer Δ | Yield (μg/mg Trp) | Didactic accessibility |
|---|---|---|---|---|---|---|---|---|
| [10] | DH5α | IPyA | ipdC, aspC, iad1 | LB + 2 g/L Trp | 2.0 | ~1.1 mg/mL | ~550 | Low (3-gene, specialized promoters) |
| [9] | MG1655MIA-6 | IAM | iaaM, iaaH (heterologous) | Whole cell + 10 g/L Trp | 10.0 | 7.10 mg/mL | ~710 | Low (extensive rewiring, metabolic engineering) |
| [14] | Lemo21(DE3) | IAM | iaaM, iaaH (plant microbiome) | M9 + no Trp added | 0 | ~1.2 µg /mL | — | Moderate (2-gene, SUMO-fusion, biocatalytic focus) |
| [11] | BL21(DE3) | IPyA | ipdC, trpABCDE, aldH | Glucose (de novo) | 0 | ~0.7 mg/mL | — | Low (full Trp pathway + de novo) |
| [15] | DH5α | IAM | iaaM, iaaH (BioBrick) | LB + Trp | ND | ~55 μM (~9.6 µg /mL) | ND | Moderate (BioBrick format, LB-based) |
| [16] | E. coli Nissle 1917 | IPyA | aspC, ipdC, iad1 + RiboJ | LB + IPTG | ND | µg/mL range | ND | Moderate (3-gene, specialized chassis, biosensor integration) |
| [17] | Cell-free (TMO enzyme) | IAM (chemo-enzymatic) | TMO (iaaM homolog) + acid hydrolysis | Continuous flow, L-Trp feed | ND | 11.16 mg/mL/ day | 48.50 % | Low (requires purified enzyme, flow reactor) |
| This work | TOP10 | IAM | iaaMs, iaaHs (codon-optimized) | ASH + M9 salts + 0.5 g/L Trp | 0.5 | ~303 μg/mL | ~60.6 | High (2-gene, standard strain, low-cost medium, full DBTL) |
| Module | DBTL phase | Key activities | Learning objectives | Level | Estimated sessions |
|---|---|---|---|---|---|
| M1: Circuit design and codon optimization | Design | Pathway selection (IAM vs. IPyA); sequence retrieval from NCBI; codon optimization using online tools (e.g., Benchling, IDT); Shine-Dalgarno design; primer design with restriction sites | Apply rational design principles to a heterologous metabolic pathway; understand codon usage bias and its effect on translation; design oligonucleotides for directional cloning | UG/PG | 2–3 (dry lab) |
| M2: Modular cloning and plasmid assembly | Build | PCR amplification of synthetic genes; agarose gel verification; restriction digestion and ligation; transformation into TOP10; colony selection and Sanger sequencing verification | Execute a complete multi-step cloning workflow; interpret gel electrophoresis results; apply troubleshooting logic to cloning failures; understand vector-insert compatibility | UG/PG | 4–6 (wet lab) |
| M3: Protein expression verification | Test | Bacterial growth and IPTG/cold-shock induction; cell lysis in Laemmli buffer; SDS-PAGE and Coomassie staining; molecular weight estimation from gel | Verify recombinant protein production; interpret SDS-PAGE results; compare induced vs. uninduced expression profiles; understand induction conditions and their limitations | UG/PG | 2–3 (wet lab) |
| M4: IAA production and quantification | Test | Shake-flask cultivation in ASH medium; Salkowski colorimetric assay and standard curve; TLC with Rf determination; HPLC fluorescence quantification | Apply colorimetric and chromatographic methods for metabolite quantification; calculate production titers and conversion yields; compare analytical methods for accuracy and sensitivity | UG/PG | 3–4 (wet lab) |
| M5: Biological validation | Test | Tobacco leaf explant preparation and aseptic inoculation; MS medium preparation with IAA dilution series; incubation and rooting scoring; statistical analysis (Kruskal–Wallis) | Design and execute a plant bioassay with appropriate controls; apply non-parametric statistics to bioassay data; interpret auxin dose-response in tissue culture | UG/PG | 4–5 (wet lab + observation period) |
| M6: Circuit redesign — promoter/RBS variants | Learn → Design | Comparative promoter analysis (constitutive vs. cold-shock); design of alternative RBS sequences; cloning of variant constructs; repeat M3–M4 with new design | Formulate and test design hypotheses; understand how regulatory elements affect expression; compare performance of multiple circuit variants; practice iterative engineering logic | PG | 4–6 (dry + wet lab) |
| M7: Substrate optimization and metabolome analysis | Learn → Design | Comparison of ASH batches and alternative agro-industrial substrates; time-course IAA quantification; LC-MS/MS identification of co-produced indolic compounds | Understand the effect of medium composition on microbial metabolism; apply mass spectrometry for compound identification; discuss circular bioeconomy implications | PG | 4–5 (wet lab + instrument access) |
| Primer name | 5´-3´ DNA sequences * | descriptions |
|---|---|---|
| iaaMrbsF | TCTCATATGCGTCGAGATATAAGGAGATATACATG | Forward primer with the NdeI restriction site |
| iaaMR | GGAATTCGATGGTTAATAGCGATACGATACGAT | Reverse primer with EcoRI restriction site |
| iaaHrbsF | CCGGAATTCGATACCCAATAAGGAGATATACATG | Forward primer with the EcoRI restriction site |
| iaaHR | TGCGTCTAGATATTGGGTCTTAGGTGGTATGCGG | Reverse primer with XbaI for restriction site |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




