Submitted:
17 March 2026
Posted:
18 March 2026
You are already at the latest version
Abstract
Polyhydroxyalkanoates (PHAs) are biodegradable microbial polyesters that represent a promising sustainable alternative to petroleum-based plastics. Salterns, hypersaline environments, are recognized as significant sources of halotolerant microorganisms that can produce PHAs in high-salinity conditions; however, Vietnamese saltern ecosystems have not been extensively investigated. This research aimed to isolate and initially characterize salt-tolerant bacteria capable of synthesizing PHAs from the Hon Khoi saltern in Khanh Hoa province, Vietnam. A total of 37 halotolerant bacterial isolates were obtained, and potential PHA-producing strains were initially screened using Sudan Black B and Nile Blue A. TEM microscopy was then employed to confirm the existence of PHA granules. Furthermore, FTIR spectroscopy and GC–MS/MS spectrometry were utilized to analyze the chemical structure and monomer composition of the extracted polymers. Six isolates were identified as PHA-producing bacteria, including Salinivibrio sp. HK101 and HK116, Halomonas sp. HK105, Priestia sp. HK125 and HK142, and Bacillus sp. HK130. These strains exhibited growth across 3–10% NaCl and temperatures from 25 to 45 °C. Priestia sp. HK142 and Salinivibrio sp. HK101 exhibited the most substantial PHA accumulation, achieving 50.72 ± 1.83% and 42.07 ± 1.8% of DCW, respectively. These results indicate that the Hon Khoi saltern represents a promising source of halotolerant PHA-producing bacteria with potential relevance for future biopolymer production studies.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Isolation of Bacteria
2.2. Primary Screening for PHA Accumulation
2.3. Transmission Electron Microscopy
2.4. Identification and Characterization of PHA-Producing Isolates
2.5. Fermentation of PHA
2.6. Determination of Dry Cell Weight
2.7. Extraction and Purification of PHA
2.8. Determination of PHA Content and Monomer Composition
2.9. Structural Characterization of PHA
2.10. Statistical Analysis
3. Results
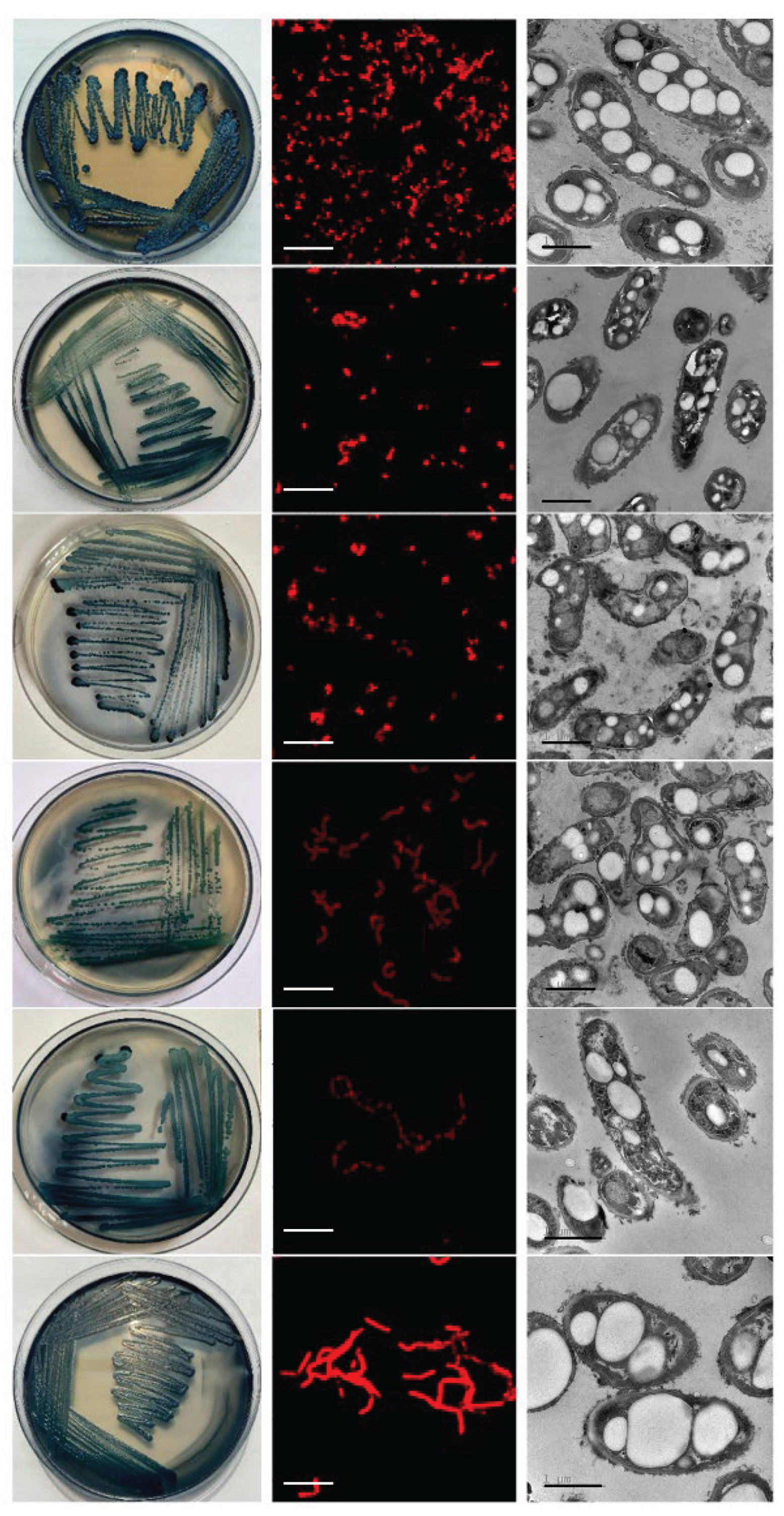
3.1. Isolation and Primary Screening of Halotolerant PHA-Producing Bacteria from the Hon Khoi Salt Fields
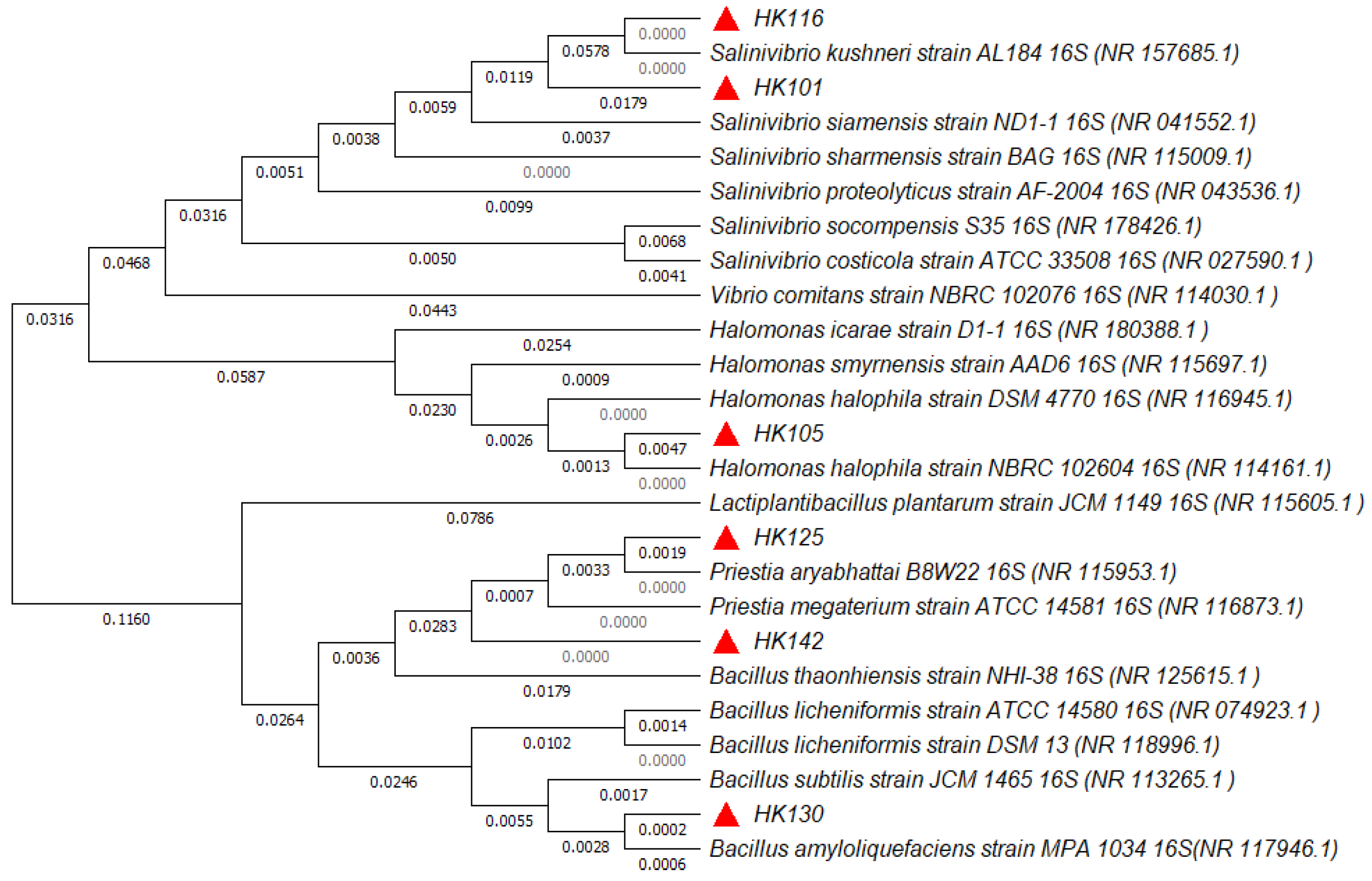
3.2. Characterization and Identification of Selected Strains
3.3. PHA Production of Selected Strains
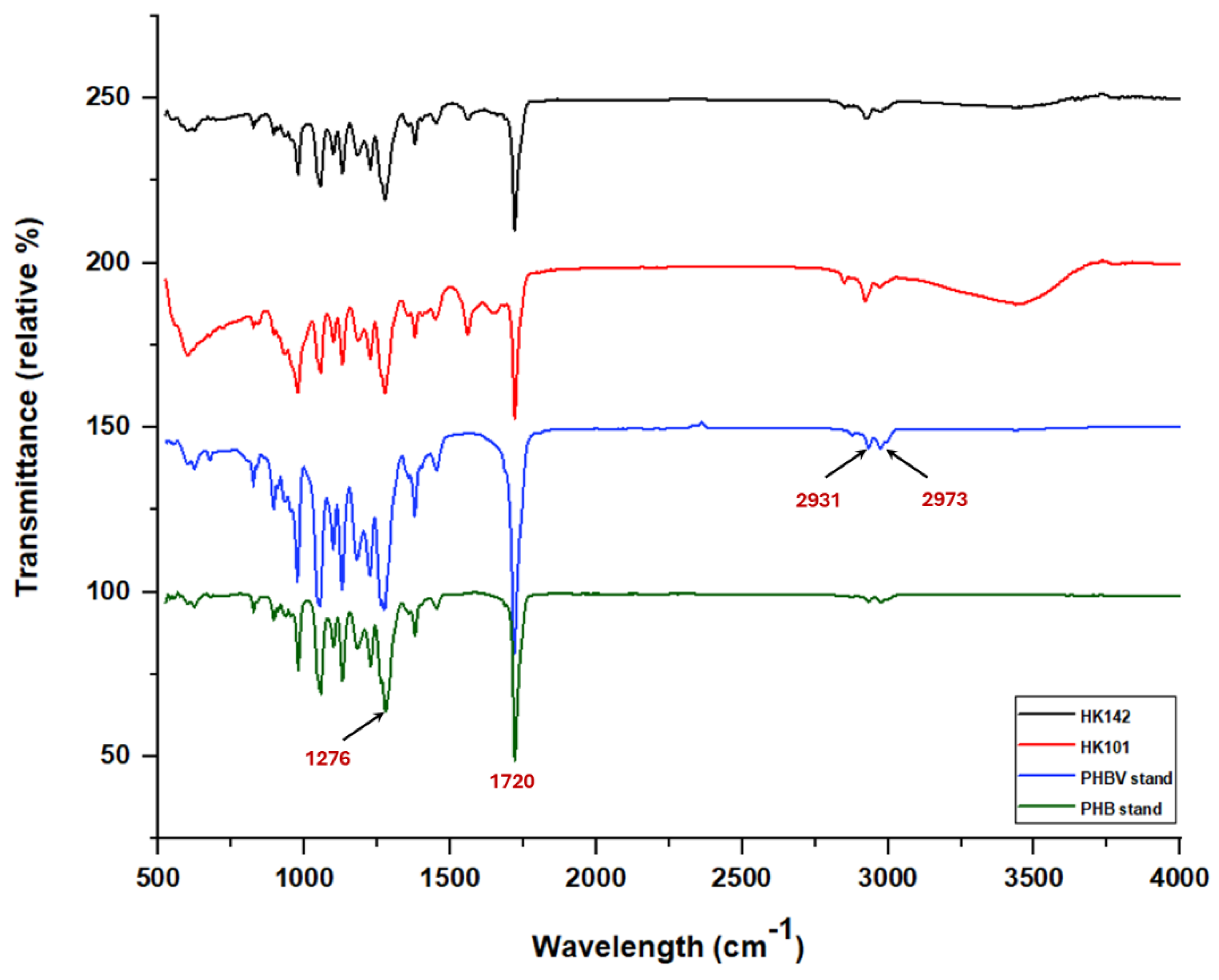
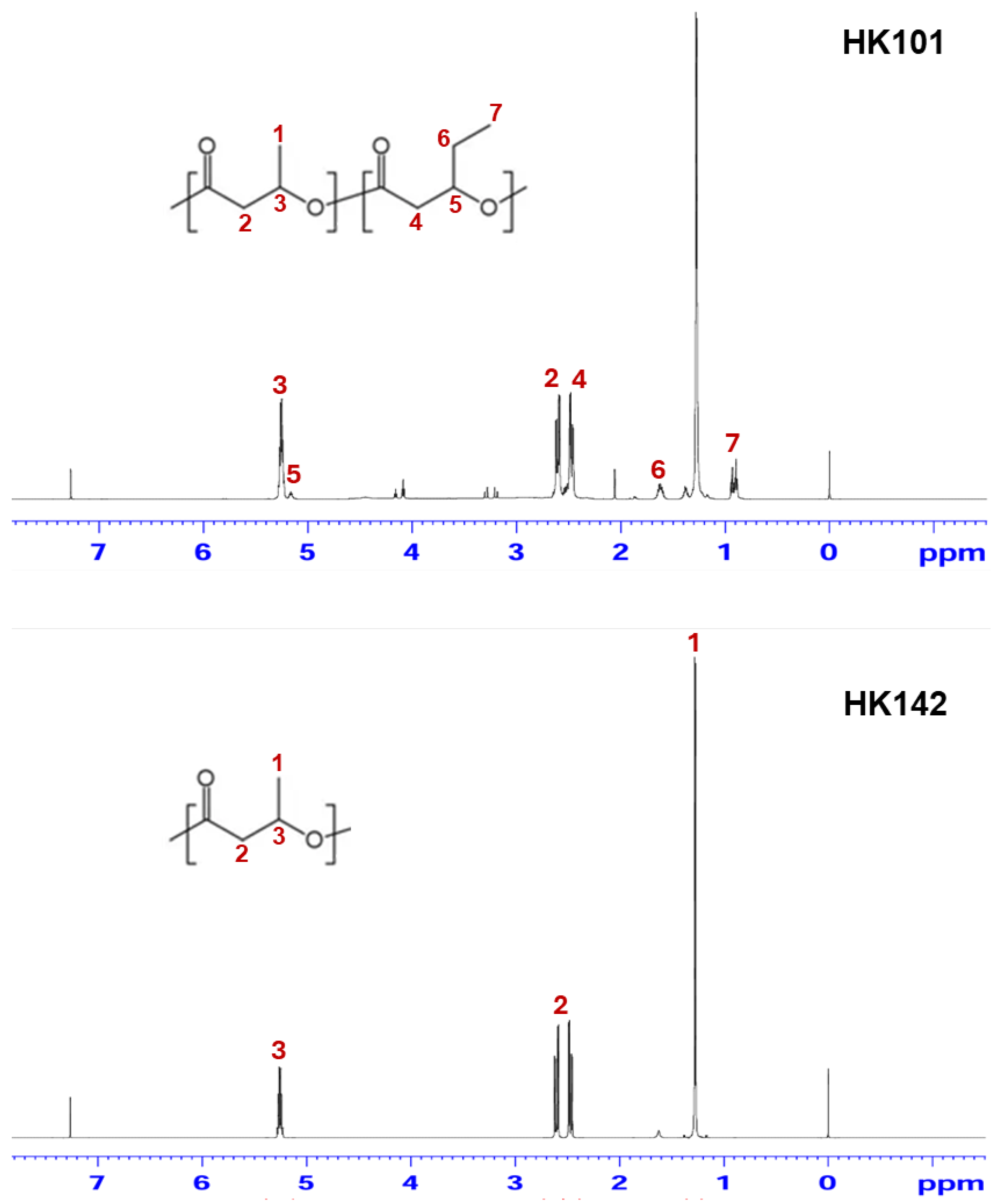
3.4. PHA Characterization of HK101 and HK142 Strains
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| PHA | PolyHydroxyAlkanoates |
| 3HB | 3-HydroxyButyrate |
| 3HV | 3-HydroxyValerate |
| PHB | Poly(3-HydroxyButyrate) |
| PHBV | Poly(3-HydroxyButyrate-co-3-hydroxyValerate) |
| TEM | Transmission Electron Microscopy |
| GC–MS/MS | Gas Chromatography-Tandem Mass Spectrometry |
| FTIR | Fourier-Transform Infrared Spectroscopy |
| DCW | Dry Cell Weight |
References
- Muigano, M.N.; Anami, S.E.; Onguso, J.M.; Omare, G.M. The isolation, screening, and characterization of polyhydroxyalkanoate-producing bacteria from hypersaline lakes in Kenya. Bacteria 2023, 2, 81-97.
- Martínez-Gutiérrez, C.A.; Latisnere-Barragán, H.; García-Maldonado, J.Q.; López-Cortés, A. Screening of polyhydroxyalkanoate-producing bacteria and PhaC-encoding genes in two hypersaline microbial mats from Guerrero Negro, Baja California Sur, Mexico. PeerJ 2018, 6, e4780. [CrossRef]
- Novackova, I.; Hrabalova, V.; Slaninova, E.; Sedlacek, P.; Samek, O.; Koller, M.; Krzyzanek, V.; Hrubanova, K.; Mrazova, K.; Nebesarova, J.; et al. The role of polyhydroxyalkanoates in adaptation of Cupriavidus necator to osmotic pressure and high concentration of copper ions. Int J Biol Macromol 2022, 206, 977-989. [CrossRef]
- Vicente, D.; Proença, D.N.; Morais, P.V. The role of bacterial Polyhydroalkanoate (PHA) in a sustainable future: A review on the biological diversity. Int J Environ Res Public Health 2023, 20, 2959. [CrossRef]
- Ibrahim Osman, A.; Noble, B.; Percy, L.; Basnett, P. Development of a highly degenerate primer-based molecular tool for detecting and classifying the four major classes of polyhydroxyalkanoate synthase (phaC) genes in bacteria. Microbial cell factories 2025, 24, 205. [CrossRef]
- Mahato, R.P.; Kumar, S.; Singh, P. Optimization of growth conditions to produce sustainable polyhydroxyalkanoate bioplastic by Pseudomonas aeruginosa EO1. Front Microbiol 2021, 12, 711588. [CrossRef]
- Thu, N.T.T.; Hoang, L.H.; Cuong, P.K.; Viet-Linh, N.; Nga, T.T.H.; Kim, D.D.; Leong, Y.K.; Nhi-Cong, L.T. Evaluation of polyhydroxyalkanoate (PHA) synthesis by Pichia sp. TSLS24 yeast isolated in Vietnam. Sci Rep 2023, 13, 3137. [CrossRef]
- Baria, D.; Dodiya, K.; Shaikh, J.; Rana, J.; Patel, J.; Raval, V. Isolation, screening, and quantification of polyhydroxyalkanoates (PHA) producing bacteria aided by FTIR analysis. VIDYA 2025, 4, 430-442. [CrossRef]
- Song, T.; Liang, Q.; Du, Z.; Wang, X.; Chen, G.; Du, Z.; Mu, D. Salinity gradient controls microbial community structure and assembly in coastal solar salterns. Genes 2022, 13, 385.
- Koller, M. Polyhydroxyalkanoate biosynthesis at the edge of water activitiy-haloarchaea as biopolyester factories. Bioengineering (Basel, Switzerland) 2019, 6, 34. [CrossRef]
- Van-Thuoc, D.; Huu-Phong, T.; Thi-Binh, N.; Thi-Tho, N.; Minh-Lam, D.; Quillaguamán, J. Polyester production by halophilic and halotolerant bacterial strains obtained from mangrove soil samples located in Northern Vietnam. MicrobiologyOpen 2012, 1, 395-406. [CrossRef]
- Tan, D.; Wang, Y.; Tong, Y.; Chen, G.-Q. Grand challenges for industrializing Polyhydroxyalkanoates (PHAs). Trends Biotechnol. 2021, 39, 953-963. [CrossRef]
- Mitra, R.; Xu, T.; Xiang, H.; Han, J. Current developments on polyhydroxyalkanoates synthesis by using halophiles as a promising cell factory. Microbial cell factories 2020, 19, 86. [CrossRef]
- Thomas, T.; Elain, A.; Bazire, A.; Bruzaud, S. Complete genome sequence of the halophilic PHA-producing bacterium Halomonas sp. SF2003: insights into its biotechnological potential. World journal of microbiology & biotechnology 2019, 35, 50. [CrossRef]
- Gholami, A.; Younes, G.; Kazemi, R.; Abootalebi, S.; Irajie, C.; Ireji, A.; Omidifar, N. Bacterial strain isolated from high-salt environments can produce large amounts of new polyhydroxyalkanoate (PHA). J Environ Eng 2020, 8, 1268-1273. [CrossRef]
- Neagu, S.; Stancu, M.M. Novel halotolerant bacteria from saline environments: isolation and biomolecule production. BioTech 2025, 14, 49.
- Ljutov, V. Technique of Voges-Proskauer test. Acta Pathol Microbiol Scand 1963, 58, 325-335. [CrossRef]
- Heuer, H.; Krsek, M.; Baker, P.; Smalla, K.; Wellington, E.M. Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 1997, 63, 3233-3241. [CrossRef]
- Hahn, S.K.; Chang, Y.K.; Kim, B.S.; Chang, H.N. Optimization of microbial poly(3-hydroxybutyrate) recover using dispersions of sodium hypochlorite solution and chloroform. Biotechnol Bioeng 1994, 44, 256-261. [CrossRef]
- Samrot, A.V.; Samanvitha, S.K.; Shobana, N.; Renitta, E.R.; Senthilkumar, P.; Kumar, S.S.; Abirami, S.; Dhiva, S.; Bavanilatha, M.; Prakash, P.; et al. The synthesis, characterization and applications of polyhydroxyalkanoates (PHAs) and PHA-based nanoparticles. Polymers 2021, 13, 3302. [CrossRef]
- Schlegel, H.G.; Lafferty, R.; Krauss, I. The isolation of mutants not accumulating poly-beta-hydroxybutyric acid. Archiv fur Mikrobiologie 1970, 71, 283-294. [CrossRef]
- Ostle, A.G.; Holt, J.G. Nile blue A as a fluorescent stain for poly-beta-hydroxybutyrate. Appl Environ Microbiol 1982, 44, 238-241. [CrossRef]
- Ibrahim, R.; Aranjani, J.M.; Prasanna, N.; Biswas, A.; Gayam, P.K.R. Production, isolation, optimization, and characterization of microbial PHA from Bacillus australimaris. Sci Rep 2025, 15, 8395. [CrossRef]
- Mohammed, S.; Panda, A.N.; Ray, L. An investigation for recovery of polyhydroxyalkanoates (PHA) from Bacillus sp. BPPI-14 and Bacillus sp. BPPI-19 isolated from plastic waste landfill. Int J Biol Macromol 2019, 134, 1085-1096. [CrossRef]
- Kanavaki, I.; Drakonaki, A.; Geladas, E.D.; Spyros, A.; Xie, H.; Tsiotis, G. Polyhydroxyalkanoate (PHA) production in Pseudomonas sp. PHDV1 strain grown on phenol as carbon sources. Microorganisms 2021, 9, 1636.
- Hobmeier, K.; Cantone, M.; Nguyen, Q.A.; Pflüger-Grau, K.; Kremling, A.; Kunte, H.J.; Pfeiffer, F.; Marin-Sanguino, A. Adaptation to varying salinity in Halomonas elongata: much more than ectoine accumulation. Front Microbiol 2022, 13, 846677. [CrossRef]
- Yoo, Y.; Young Kwon, D.; Jeon, M.; Lee, J.; Kwon, H.; Lee, D.; Seong Khim, J.; Choi, Y.-E.; Kim, J.-J. Enhancing poly(3-hydroxybutyrate) production in halophilic bacteria through improved salt tolerance. Biores Technol 2024, 394, 130175. [CrossRef]
- Li, J.; Xiao, X.; Zhou, M.; Zhang, Y. Strategy for the adaptation to stressful conditions of the novel isolated conditional piezophilic strain Halomonas titanicae ANRCS81. Appl Environ Microbiol 2023, 89, e01304-01322. [CrossRef]
- Dickinson, A.W.; Gault, S.; Cane, R.; Cockell, C.S. The multiple extremes of temperature, salt and pH define narrower limits to microbial growth in Halomonas hydrothermalis than individual extremes. Int J Astrobiol 2025, 24, e19. [CrossRef]
- Thomas, T.; Sudesh, K.; Bazire, A.; Elain, A.; Tan, H.T.; Lim, H.; Bruzaud, S. PHA production and PHA synthases of the halophilic bacterium Halomonas sp. SF2003. Bioengineering (Basel, Switzerland) 2020, 7, 29. [CrossRef]
- Aytar Celik, P.; Barut, D.; Enuh, B.M.; Erdogan Gover, K.; Nural Yaman, B.; Burcin Mutlu, M.; Cabuk, A. A novel higher polyhydroxybutyrate producer Halomonas halmophila 18H with unique cell factory attributes. Biores Technol 2023, 372, 128669. [CrossRef]
- Christensen, M.; Chiciudean, I.; Lascu, I.; Jablonski, P.; Shapaval, V.; Zimmermann, B.; Tanase, A.M.; Hansen, H. Halomonas sp. MC140, a polyhydroxyalkanoate (PHA) producer isolated from the Arctic environment. Sci Rep 2025, 15, 23744. [CrossRef]
- Chamroensaksri, N.; Tanasupawat, S.; Akaracharanya, A.; Visessanguan, W.; Kudo, T.; Itoh, T. Salinivibrio siamensis sp. nov., from fermented fish (pla-ra) in Thailand. Int J Syst Evol Microb 2009, 59, 880-885. [CrossRef]
- Galisteo, C.; Sánchez-Porro, C.; de la Haba, R.R.; López-Hermoso, C.; Fernández, A.B.; Farias, M.E.; Ventosa, A. Characterization of Salinivibrio socompensis sp. nov., A new halophilic bacterium isolated from the high-altitude hypersaline lake Socompa, Argentina. Microorganisms 2019, 7, 241. [CrossRef]
- López-Hermoso, C.; de la Haba, R.R.; Sánchez-Porro, C.; Ventosa, A. Salinivibrio kushneri sp. nov., a moderately halophilic bacterium isolated from salterns. Syst Appl Microbiol 2018, 41, 159-166. [CrossRef]
- Pu, N.; Wang, M.-R.; Li, Y.; Li, Z.-J. Metabolic engineering of Salinivibrio sp. TGB10 for PHBV biosynthesis with a high 3-hydroxyvalerate fraction from starch and propionate. Int J Biol Macromol 2025, 308, 142359. [CrossRef]
- Tao, G.-B.; Tian, L.; Pu, N.; Li, Z.-J. Efficient production of poly-3-hydroxybutyrate from acetate and butyrate by halophilic bacteria Salinivibrio spp. TGB4 and TGB19. Int J Biol Macromol 2022, 221, 1365-1372. [CrossRef]
- Masmoudi, F.; Naimi, L.; Trigui, M.; Safran, M.; Tounsi, S.; Saadaoui, I. Novel thermo-halotolerant bacteria Bacillus cabrialesii native to Qatar desert: enhancing seedlings’ growth, halotolerance, and antifungal defense in tomato. Journal of plant growth regulation 2024, 44, 587-604. [CrossRef]
- Wael, D.; El-Amier, Y.; Saber, W.I.A.; Elsayed, A. Plant-associated halotolerant bacteria improving growth of Vicia faba L. Mariout-2 under salinity conditions. Sci Rep 2024, 14, 16737. [CrossRef]
- Cueva-Almendras, L.C.; Alvarado, J.C.M.; Fuentes-Olivera, A.J.; Llontop-Bernabé, K.S.; Cerna, C.E.Q.; Rodríguez-Soto, J.C.; Cruz-Monzón, J.A.; Alvarez, M.A.Q. Production of polyhydroxyalkanoate by Bacillus thuringiensis Isolated from agricultural soils of Cascas-Peru. Braz Arch Biol Technol 2022, 65, e22220107.
- Cal, A.J.; Chan, V.J.; Luo, W.K.; Lee, C.C. Polyhydroxyalkanoate production in Priestia megaterium strains from glycerol feedstock. PloS one 2025, 20, e0322838. [CrossRef]
- Shi, L.; Zhu, X.; Qian, T.; Du, J.; Du, Y.; Ye, J. Mechanism of salt tolerance and plant growth promotion in Priestia megaterium zs-3 revealed by cellular metabolism and whole-genome studies. International journal of molecular sciences 2023, 24, 15751.
- Hwang, H.-H.; Chien, P.-R.; Huang, F.-C.; Yeh, P.-H.; Hung, S.-H.W.; Deng, W.-L.; Huang, C.-C. A plant endophytic bacterium Priestia megaterium strain BP-R2 isolated from the halophyte Bolboschoenus planiculmis enhances plant growth under salt and drought stresses. Microorganisms 2022, 10, 2047.
- Obruča, S.; Dvořák, P.; Sedláček, P.; Koller, M.; Sedlář, K.; Pernicová, I.; Šafránek, D. Polyhydroxyalkanoates synthesis by halophiles and thermophiles: towards sustainable production of microbial bioplastics. Biotechnology advances 2022, 58, 107906. [CrossRef]
- Sooksawat, T.; Attapong, M.; Saengsakun, W.; Siripornadulsil, S.; Siripornadulsil, W. Optimization of polyhydroxybutyrate (PHB) production by Priestia megaterium ASL11 and glycerol and thermoplastic properties of PHB-based films. Biocatal Agric Biotechnol 2023, 54, 102951. [CrossRef]
- Lee, J.Y.; Kim, M.H.; Kim, J.S.; Yun, B.R.; Kim, D.Y.; Chung, C.W. Biotransformation of d-xylose-rich rice husk hydrolysate by a rice paddy soil bacterium, Priestia sp. strain JY310, to low molecular weight poly(3-hydroxybutyrate). Biomolecules 2023, 13, 131. [CrossRef]
- Bai, X.; Xu, L.; Li, K.; Zhang, G.; Zhang, M.; Huang, Y. Unlocking efficient polyhydroxyalkanoate production by Gram-positive Priestia megaterium using waste-derived feedstocks. Microbial cell factories 2025, 24, 210. [CrossRef]
- Tao, G.-B.; Tan, B.-W.; Li, Z.-J. Production of polyhydroxyalkanoates by a moderately halophilic bacterium of Salinivibrio sp. TGB10. Int J Biol Macromol 2021, 186, 574-579. [CrossRef]
- Zhou, W.; Colpa, D.I.; Geurkink, B.; Euverink, G.-J.W.; Krooneman, J. The impact of carbon to nitrogen ratios and pH on the microbial prevalence and polyhydroxybutyrate production levels using a mixed microbial starter culture. Sci Total Environ 2022, 811, 152341. [CrossRef]
- Sánchez Valencia, A.I.; Rojas Zamora, U.; Meraz Rodríguez, M.; Álvarez Ramírez, J.; Salazar Peláez, M.L.; Fajardo Ortiz, C. Effect of C/N ratio on the PHA accumulation capability of microbial mixed culture fed with leachates from the organic fraction of municipal solid waste (OFMSW). J Water Proc Eng 2021, 40, 101975. [CrossRef]
| Characteristic | HK101 | HK105 | HK116 | HK125 | HK130 | HK142 |
|---|---|---|---|---|---|---|
| Colony morphology | Round, smooth, convex, opaque, cream colonies (2.0–2.5 mm). | Round, smooth, convex, opaque, yellow colonies (0.5–1.0 mm) | Round, smooth, convex, opaque, light cream colonies (0.8-1.2 mm) | Round, irregular surfaced, opaque, creamy white colonies (2.0–3.0 mm). | Rounds, irregular surfaced, opaque, grayish-white colonies (2.0–2.6 mm) | Round, undulate margin, opaque, cream colonies (2.0–2.5 mm) |
| Cell morphology | Curved rods | Rods | Curved rod | Rods | Rods | Rods |
| Motility | + | + | + | + | + | + |
| Spore formation | - | - | - | + | + | + |
| Gram staining | - | - | - | + | + | + |
| Oxidase | + | + | + | - | + | + |
| Catalase | + | + | + | + | + | ± |
| Voges-Proskauer | + | - | + | + | + | - |
| Growth at 0% NaCl | - | - | - | - | + | - |
| 3% NaCl | + | + | + | + | + | + |
| 5% NaCl | + | + | + | + | + | + |
| 10% NaCl | + | + | - | + | - | - |
| 15% NaCl | - | + | - | + | - | - |
| Growth at 25oC | + | + | + | + | + | + |
| 30oC | + | + | + | + | + | + |
| 37oC | + | + | + | + | + | + |
| 40oC | + | + | + | + | + | + |
| 45oC | + | - | + | ± | + | + |
| Sugar fermentation: | ||||||
| D-glucose | + | + | + | + | + | + |
| Sucrose | + | - | + | + | + | + |
| Maltose | + | + | + | + | + | + |
| D-galactose | - | + | - | + | + | + |
| L-arabinose | - | + | - | ± | - | + |
| D-xylose | + | + | - | - | - | + |
| D-mannose | + | + | + | + | + | - |
| D-mannitol | + | - | + | ± | + | + |
| Hydrolysis: | ||||||
| Soluble starch | + | - | + | + | + | + |
| Casein | + | - | - | + | + | + |
| CMC | ± | - | + | + | - | + |
| Strain | Carbon source | DCW (g/L) |
PHA content (% DCW) ** |
PHA component (%mol) ** | |
| 3HB | 3HV | ||||
| HK101 | Glucose | 1.69 ± 0.07 | 44.14 ± 3.28 | 100 | 0 |
| Sucrose | 1.71 ± 0.08 | 47.37 ± 2.57 | 100 | 0 | |
| Glycerol | 1.59 ± 0.05 | 30.82 ± 1.59 | 100 | 0 | |
| Mixture* | 1.45 ± 0.04 | 42.07 ± 1.80 | 93.6 | 6.4 | |
| HK105 | Glucose | 1.57 ± 0.07 | 27.77 ± 3.90 | 100 | 0 |
| Sucrose | 1.61 ± 0.08 | 28.32 ± 2.01 | 100 | 0 | |
| Glycerol | 1.53 ± 0.06 | 28.10 ± 1.71 | 100 | 0 | |
| Mixture* | 1.30 ± 0.06 | 32.31 ± 2.06 | 100 | 0 | |
| HK116 | Glucose | 1.59 ± 0.09 | 26.33 ± 2.42 | 100 | 0 |
| Sucrose | 1.66 ± 0.07 | 28.52 ± 1.91 | 100 | 0 | |
| Glycerol | 1.45 ± 0.04 | 15.17 ± 0.81 | 100 | 0 | |
| Mixture* | 1.50 ± 0.05 | 26.67 ± 2.12 | 100 | 0 | |
| HK125 | Glucose | 1.64 ± 0.04 | 31.65 ± 2.08 | 100 | 0 |
| Sucrose | 1.59 ± 0.07 | 30.78 ± 1.47 | 100 | 0 | |
| Glycerol | 1.56 ± 0.08 | 26.92 ± 1.88 | 100 | 0 | |
| Mixture* | 1.47 ± 0.08 | 31.97 ± 2.42 | 100 | 0 | |
| HK130 | Glucose | 1.61 ± 0.06 | 30.07 ± 2.41 | 100 | 0 |
| Sucrose | 1.50 ± 0.07 | 24.70 ± 1.71 | 100 | 0 | |
| Glycerol | 1.42 ± 0.08 | 25.35 ± 2.01 | 100 | 0 | |
| Mixture* | 1.48 ± 0.07 | 22.97 ± 1.28 | 96.8 | 3.2 | |
| HK142 | Glucose | 1.84 ± 0.09 | 48.45 ± 4.16 | 100 | 0 |
| Sucrose | 1.72 ± 0.07 | 30.78 ± 1.84 | 100 | 0 | |
| Glycerol | 1.62 ± 0.09 | 24.07 ± 1.82 | 100 | 0 | |
| Mixture* | 1.59 ± 0.08 | 26.42 ± 1.74 | 100 | 0 | |
| Strain | C/N ratio (mol/mol) | DCW (g/L) |
PHA content (% DCW) * |
PHA component (%mol) * | |
| 3HB | 3HV | ||||
| HK101 | 20:1 | 1.78 ± 0.05 | 23.60 ± 1.30 | 97.3 | 2.7 |
| 25:1 | 1.85 ± 0.06 | 35.68 ± 1.99 | 94.2 | 5.8 | |
| 30:1 | 1.45 ± 0.04 | 42.07 ± 1.80 | 93.6 | 6.4 | |
| 35:1 | 1.52 ± 0.07 | 30.26 ± 1.75 | 95.8 | 4.2 | |
| HK105 | 20:1 | 1.84 ± 0.06 | 14.13 ± 0.71 | 100 | 0 |
| 25:1 | 1.75 ± 0.07 | 26.06 ± 2.01 | 100 | 0 | |
| 30:1 | 1.61 ± 0.08 | 28.32 ± 2.01 | 100 | 0 | |
| 35:1 | 1.62 ± 0.07 | 20.99 ± 1.33 | 100 | 0 | |
| HK116 | 20:1 | 1.88 ± 0.06 | 17.55 ± 0.77 | 100 | 0 |
| 25:1 | 1.82 ± 0.06 | 26.65 ± 1.87 | 100 | 0 | |
| 30:1 | 1.66 ± 0.07 | 28.52 ± 1.91 | 100 | 0 | |
| 35:1 | 1.66 ± 0.08 | 28.43 ± 1.66 | 100 | 0 | |
| HK125 | 20:1 | 1.92 ± 0.04 | 24.48 ± 1.16 | 100 | 0 |
| 25:1 | 1.89 ± 0.06 | 28.10 ± 1.82 | 100 | 0 | |
| 30:1 | 1.64 ± 0.04 | 31.65 ± 2.08 | 100 | 0 | |
| 35:1 | 1.73 ± 0.09 | 29.94 ± 1.92 | 100 | 0 | |
| HK130 | 20:1 | 1.68 ± 0.05 | 16.67 ± 0.78 | 98.1 | 1.9 |
| 25:1 | 1.68 ± 0.08 | 29.17 ± 1.83 | 95.6 | 4.4 | |
| 30:1 | 1.48 ± 0.07 | 22.97 ± 1.28 | 96.8 | 3.2 | |
| 35:1 | 1.71 ± 0.07 | 19.88 ± 0.97 | 97.3 | 2.7 | |
| HK142 | 20:1 | 1.96 ± 0.07 | 29.08 ± 1.46 | 100 | 0 |
| 25:1 | 2.09 ± 0.09 | 50.72 ± 1.83 | 100 | 0 | |
| 30:1 | 1.84 ± 0.09 | 48.45 ± 4.16 | 100 | 0 | |
| 35:1 | 1.77 ± 0.06 | 50.28 ± 1.97 | 100 | 0 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




