Submitted:
16 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
Keywords:
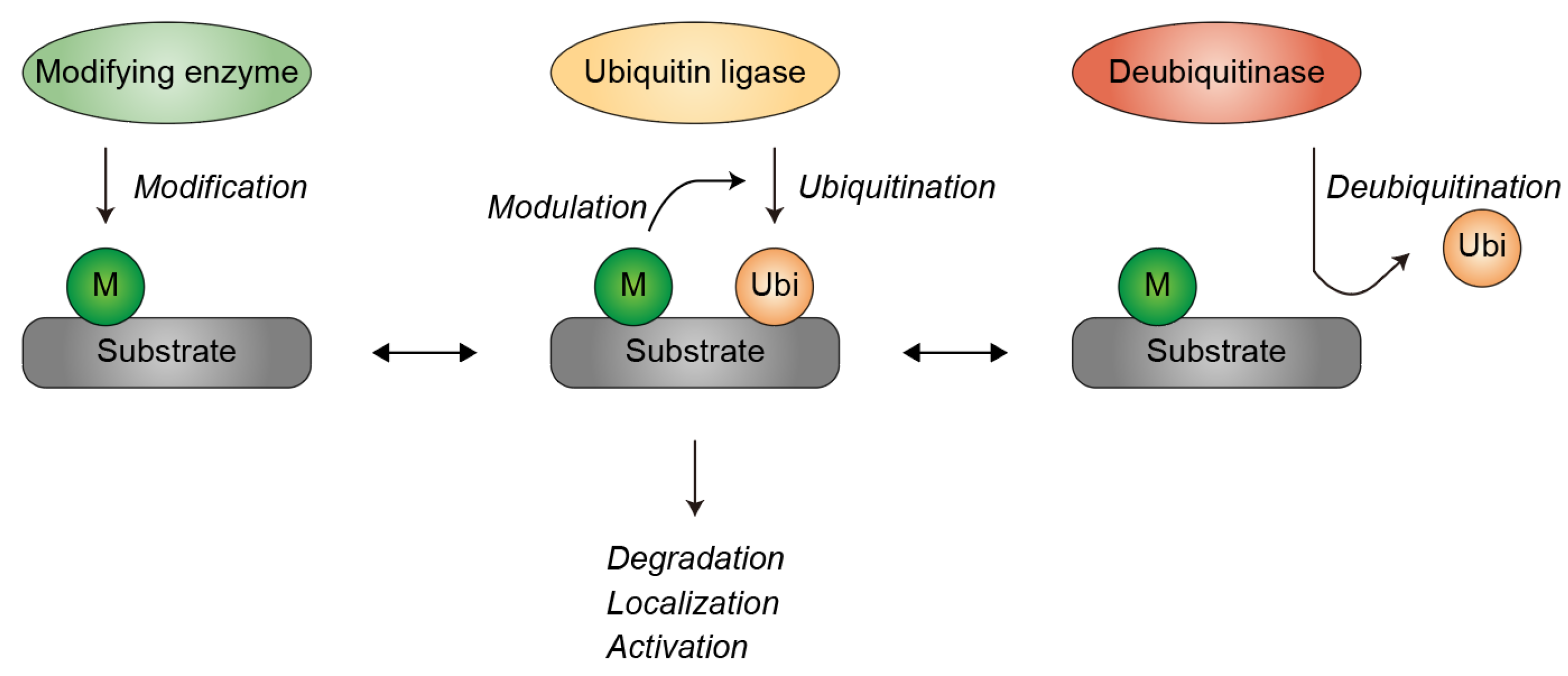
1. Introduction
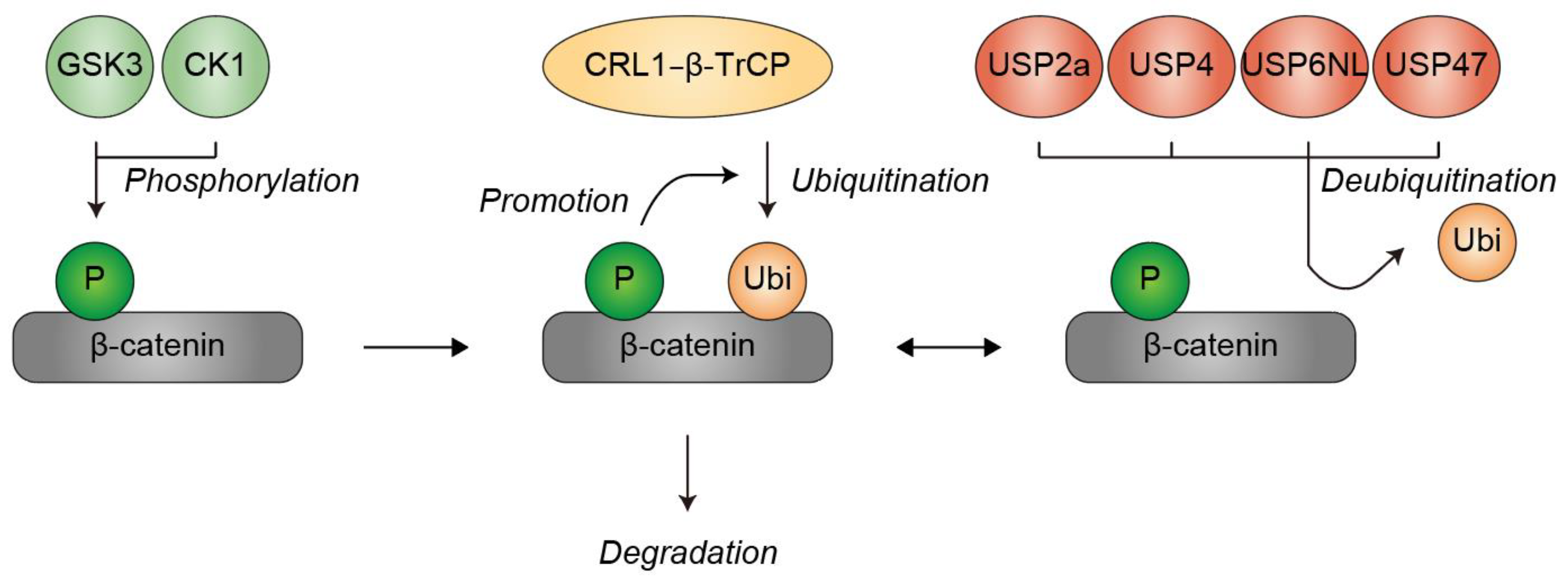
2. Ubiquitination of β-Catenin
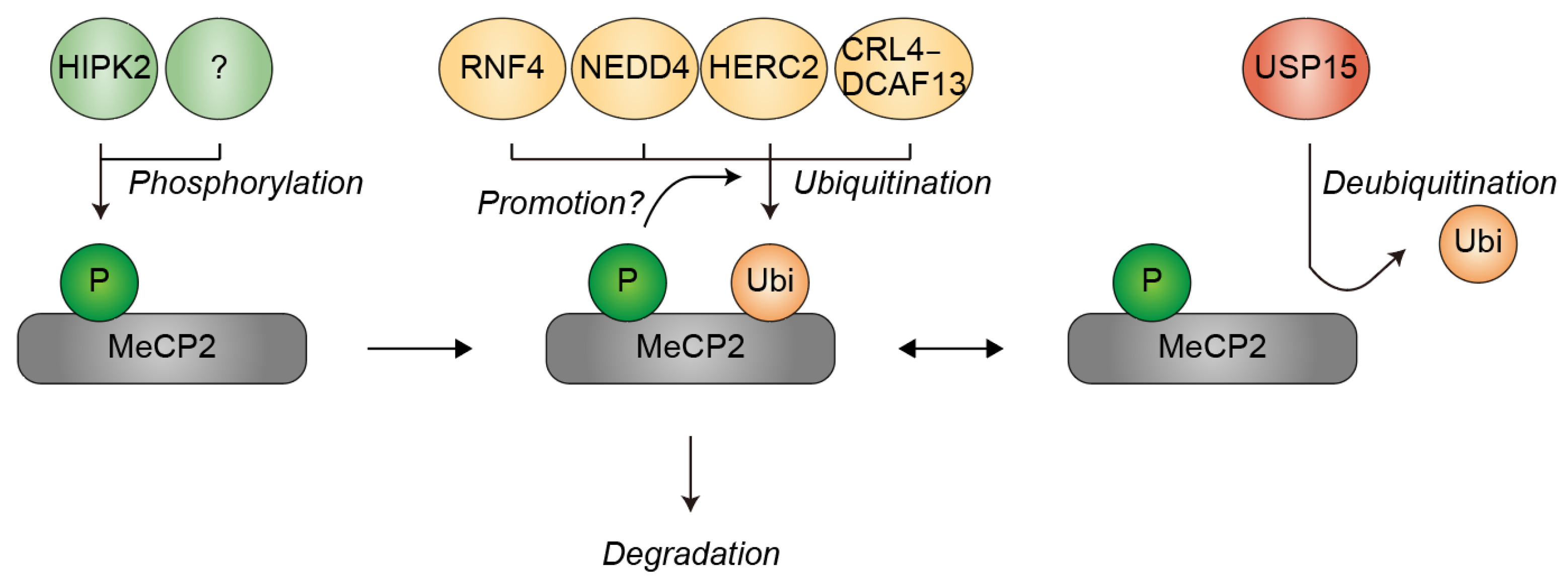
3. Ubiquitination of MeCP2
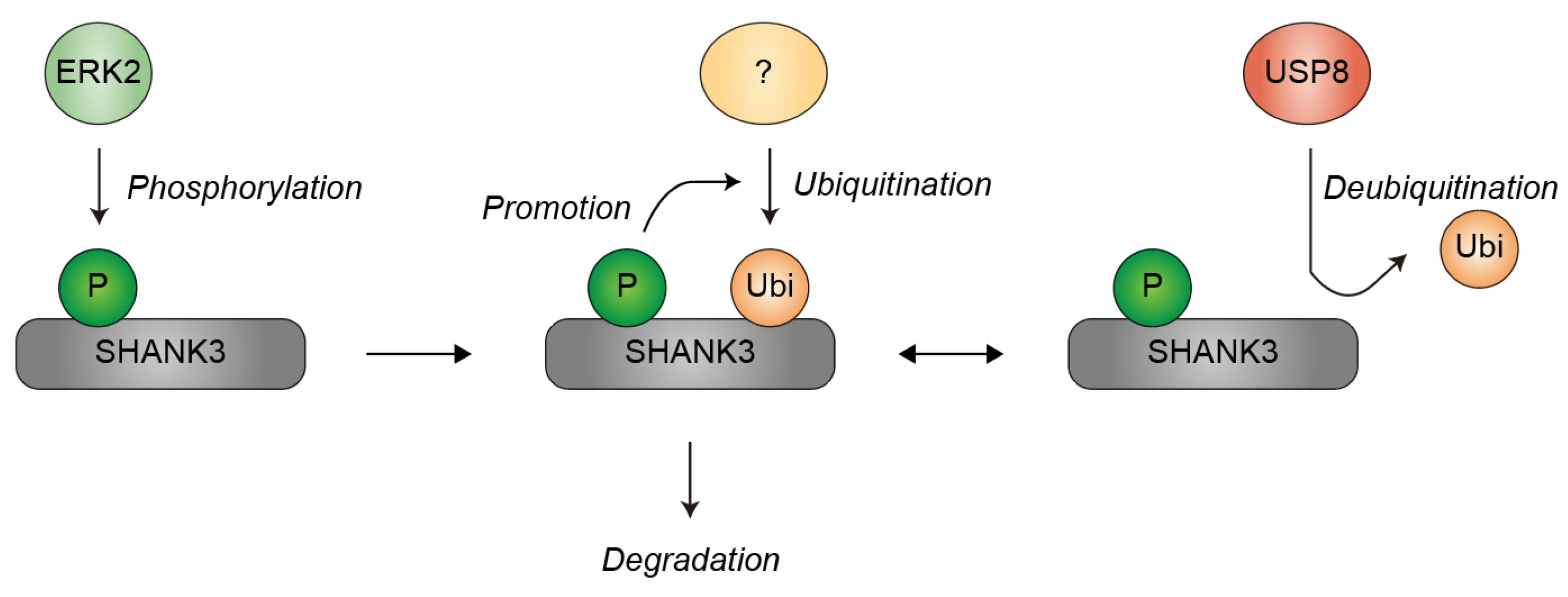
4. Ubiquitination of SHANK3
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Thapar, A., M. Cooper, and M. Rutter, Neurodevelopmental disorders. Lancet Psychiatry, 2017. 4(4): p. 339-346.
- Morris-Rosendahl, D.J. and M.A. Crocq, Neurodevelopmental disorders-the history and future of a diagnostic concept. Dialogues Clin Neurosci, 2020. 22(1): p. 65-72.
- Gao, S., et al., Genetic advances in neurodevelopmental disorders. Med Rev (2021), 2025. 5(2): p. 139-151. [CrossRef]
- Zheng, X., J. Li, and X. Jin, Functional Neurogenomics to Dissect Disease Mechanisms Across Models. Annu Rev Genomics Hum Genet, 2025. 26(1): p. 189-216. [CrossRef]
- Jeste, S.S. and D.H. Geschwind, Disentangling the heterogeneity of autism spectrum disorder through genetic findings. Nat Rev Neurol, 2014. 10(2): p. 74-81. [CrossRef]
- Chaste, P., et al., A genome-wide association study of autism using the Simons Simplex Collection: Does reducing phenotypic heterogeneity in autism increase genetic homogeneity? Biol Psychiatry, 2015. 77(9): p. 775-84. [CrossRef]
- Geschwind, D.H. and M.W. State, Gene hunting in autism spectrum disorder: on the path to precision medicine. Lancet Neurol, 2015. 14(11): p. 1109-20. [CrossRef]
- Liao, Y., et al., The role of ubiquitination in health and disease. MedComm (2020), 2024. 5(10): p. e736.
- Ohno, M., S. Wakatsuki, and T. Araki, The essential role of E3 ubiquitin ligases in the pathogenesis of neurodevelopmental and psychiatric disorders: Cul3, Cul4, Ube3a, and ZNRF1. Biochem Biophys Res Commun, 2025. 763: p. 151798. [CrossRef]
- Yang, X., et al., Targeting ubiquitination in disease and therapy. Signal Transduct Target Ther, 2025. 10(1): p. 424.
- Nakagawa, M. and T. Nakagawa, CUL4-Based Ubiquitin Ligases in Chromatin Regulation: An Evolutionary Perspective. Cells, 2025. 14(2). [CrossRef]
- Pinto, M.J., D. Tomé, and R.D. Almeida, The Ubiquitinated Axon: Local Control of Axon Development and Function by Ubiquitin. J Neurosci, 2021. 41(13): p. 2796-2813. [CrossRef]
- Ambrozkiewicz, M.C. and S. Lorenz, Understanding ubiquitination in neurodevelopment by integrating insights across space and time. Nat Struct Mol Biol, 2025. 32(1): p. 14-22. [CrossRef]
- Ashitomi, H., et al., Cullin-RING Ubiquitin Ligases in Neurodevelopment and Neurodevelopmental Disorders. Biomedicines, 2025. 13(4). [CrossRef]
- Shigenaka, A., et al., Defective Neural Stem and Progenitor Cell Proliferation in Neurodevelopmental Disorders. J Dev Biol, 2025. 13(4). [CrossRef]
- Haglund, K. and I. Dikic, Ubiquitylation and cell signaling. Embo j, 2005. 24(19): p. 3353-9.
- Komander, D. and M. Rape, The ubiquitin code. Annu Rev Biochem, 2012. 81: p. 203-29. [CrossRef]
- Mattiroli, F. and T.K. Sixma, Lysine-targeting specificity in ubiquitin and ubiquitin-like modification pathways. Nat Struct Mol Biol, 2014. 21(4): p. 308-16. [CrossRef]
- Nakagawa, T., et al., Phosphodegrons in Health and Disease: From Cellular Homeostasis to Therapeutic Potential. Kinases and Phosphatases, 2025. 3(1): 3. [CrossRef]
- Hayat, R., M. Manzoor, and A. Hussain, Wnt signaling pathway: A comprehensive review. Cell Biol Int, 2022. 46(6): p. 863-877. [CrossRef]
- Rim, E.Y., H. Clevers, and R. Nusse, The Wnt Pathway: From Signaling Mechanisms to Synthetic Modulators. Annu Rev Biochem, 2022. 91: p. 571-598. [CrossRef]
- Xue, C., et al., Wnt signaling pathways in biology and disease: mechanisms and therapeutic advances. Signal Transduct Target Ther, 2025. 10(1): p. 106. [CrossRef]
- de Ligt, J., et al., Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med, 2012. 367(20): p. 1921-9.
- Zhuang, W., et al., CTNNB1 in neurodevelopmental disorders. Front Psychiatry, 2023. 14: p. 1143328.
- Kayumi, S., et al., Genomic and phenotypic characterization of 404 individuals with neurodevelopmental disorders caused by CTNNB1 variants. Genet Med, 2022. 24(11): p. 2351-2366. [CrossRef]
- Sudnawa, K.K., et al., Clinical phenotypic spectrum of CTNNB1 neurodevelopmental disorder. Clin Genet, 2024. 105(5): p. 523-532. [CrossRef]
- MacDonald, B.T., K. Tamai, and X. He, Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell, 2009. 17(1): p. 9-26. [CrossRef]
- Shi, J., et al., Deubiquitinase USP47/UBP64E Regulates β-Catenin Ubiquitination and Degradation and Plays a Positive Role in Wnt Signaling. Mol Cell Biol, 2015. 35(19): p. 3301-11. [CrossRef]
- Yun, S.I., et al., Ubiquitin specific protease 4 positively regulates the WNT/β-catenin signaling in colorectal cancer. Mol Oncol, 2015. 9(9): p. 1834-51. [CrossRef]
- Kim, J., et al., Ubiquitin-specific peptidase 2a (USP2a) deubiquitinates and stabilizes β-catenin. Am J Cancer Res, 2018. 8(9): p. 1823-1836.
- Sun, K., et al., Tre2 (USP6NL) promotes colorectal cancer cell proliferation via Wnt/β-catenin pathway. Cancer Cell Int, 2019. 19: p. 102. [CrossRef]
- Yang, S.W., et al., USP47 and C terminus of Hsp70-interacting protein (CHIP) antagonistically regulate katanin-p60-mediated axonal growth. J Neurosci, 2013. 33(31): p. 12728-38. [CrossRef]
- Anckar, J. and A. Bonni, Regulation of neuronal morphogenesis and positioning by ubiquitin-specific proteases in the cerebellum. PLoS One, 2015. 10(1): p. e0117076. [CrossRef]
- Srikanta, S.B., K. Stojkovic, and N. Cermakian, Behavioral phenotyping of mice lacking the deubiquitinase USP2. PLoS One, 2021. 16(2): p. e0241403. [CrossRef]
- Lopes, K.P., et al., Genetic analysis of the human microglial transcriptome across brain regions, aging and disease pathologies. Nat Genet, 2022. 54(1): p. 4-17. [CrossRef]
- Alexander, J.M., A. Pirone, and M.H. Jacob, Excessive β-Catenin in Excitatory Neurons Results in Reduced Social and Increased Repetitive Behaviors and Altered Expression of Multiple Genes Linked to Human Autism. Front Synaptic Neurosci, 2020. 12: p. 14. [CrossRef]
- Gulmez Karaca, K., D.V.C. Brito, and A.M.M. Oliveira, MeCP2: A Critical Regulator of Chromatin in Neurodevelopment and Adult Brain Function. Int J Mol Sci, 2019. 20(18). [CrossRef]
- Tillotson, R. and A. Bird, The Molecular Basis of MeCP2 Function in the Brain. J Mol Biol, 2020. 432(6): p. 1602-1623. [CrossRef]
- Liu, Y., et al., Exploring the complexity of MECP2 function in Rett syndrome. Nat Rev Neurosci, 2025. 26(7): p. 379-398. [CrossRef]
- Gold, W.A., et al., Rett syndrome. Nat Rev Dis Primers, 2024. 10(1): p. 84.
- Bricker, K. and B.V. Vaughn, Rett syndrome: a review of clinical manifestations and therapeutic approaches. Front Sleep, 2024. 3: p. 1373489. [CrossRef]
- Sandweiss, A.J., V.L. Brandt, and H.Y. Zoghbi, Advances in understanding of Rett syndrome and MECP2 duplication syndrome: prospects for future therapies. Lancet Neurol, 2020. 19(8): p. 689-698. [CrossRef]
- D’Mello, S.R., 3rd, MECP2 and the biology of MECP2 duplication syndrome. J Neurochem, 2021. 159(1): p. 29-60. [CrossRef]
- Kalani, L., et al., MeCP2 ubiquitination and sumoylation, in search of a function†. Hum Mol Genet, 2023. 33(1): p. 1-11. [CrossRef]
- Partscht, P., et al., The HIPK2/CDC14B-MeCP2 axis enhances the spindle assembly checkpoint block by promoting cyclin B translation. Sci Adv, 2023. 9(3): p. eadd6982. [CrossRef]
- Wang, Y., RING finger protein 4 (RNF4) derepresses gene expression from DNA methylation. J Biol Chem, 2014. 289(49): p. 33808-13. [CrossRef]
- Ren, P., et al., CRL4(DCAF13) E3 ubiquitin ligase targets MeCP2 for degradation to prevent DNA hypermethylation and ensure normal transcription in growing oocytes. Cell Mol Life Sci, 2024. 81(1): p. 165. [CrossRef]
- Yang, J., et al., Mecp2 fine-tunes quiescence exit by targeting nuclear receptors. Elife, 2024. 12.
- Zhou, B., et al., The Ubiquitin Ligase HERC2 Promotes Ang II-Induced Cardiac Hypertrophy Through Destabilization of MeCP2 to Enhance Lin28a Expression. J Cardiovasc Pharmacol, 2025. 85(2): p. 145-155. [CrossRef]
- Zhang, Z.T., et al., USP15 inhibits hypoxia-induced IL-6 signaling by deubiquitinating and stabilizing MeCP2. Febs j, 2025. 292(1): p. 153-167. [CrossRef]
- Bellini, E., et al., MeCP2 post-translational modifications: a mechanism to control its involvement in synaptic plasticity and homeostasis? Front Cell Neurosci, 2014. 8: p. 236. [CrossRef]
- Sánchez-Lafuente, C.L., et al., The Role of MeCP2 in Regulating Synaptic Plasticity in the Context of Stress and Depression. Cells, 2022. 11(4). [CrossRef]
- Mabb, A.M. and M.D. Ehlers, Ubiquitination in postsynaptic function and plasticity. Annu Rev Cell Dev Biol, 2010. 26: p. 179-210. [CrossRef]
- Patrick, M.B., et al., The ubiquitin-proteasome system and learning-dependent synaptic plasticity - A 10 year update. Neurosci Biobehav Rev, 2023. 152: p. 105280. [CrossRef]
- Monteiro, P. and G. Feng, SHANK proteins: roles at the synapse and in autism spectrum disorder. Nat Rev Neurosci, 2017. 18(3): p. 147-157. [CrossRef]
- Kaizuka, T. and T. Takumi, Postsynaptic density proteins and their involvement in neurodevelopmental disorders. J Biochem, 2018. 163(6): p. 447-455. [CrossRef]
- Durand, C.M., et al., Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet, 2007. 39(1): p. 25-7. [CrossRef]
- Kolevzon, A., et al., Phelan-McDermid syndrome: a review of the literature and practice parameters for medical assessment and monitoring. J Neurodev Disord, 2014. 6(1): p. 39. [CrossRef]
- Leblond, C.S., et al., Meta-analysis of SHANK Mutations in Autism Spectrum Disorders: a gradient of severity in cognitive impairments. PLoS Genet, 2014. 10(9): p. e1004580. [CrossRef]
- De Rubeis, S., et al., Delineation of the genetic and clinical spectrum of Phelan-McDermid syndrome caused by SHANK3 point mutations. Mol Autism, 2018. 9: p. 31.
- Wang, L., et al., A kinome-wide RNAi screen identifies ERK2 as a druggable regulator of Shank3 stability. Mol Psychiatry, 2020. 25(10): p. 2504-2516. [CrossRef]
- Kerrisk Campbell, M. and M. Sheng, USP8 Deubiquitinates SHANK3 to Control Synapse Density and SHANK3 Activity-Dependent Protein Levels. J Neurosci, 2018. 38(23): p. 5289-5301. [CrossRef]
- Dhawka, L., et al., Post-ischemic ubiquitination at the postsynaptic density reversibly influences the activity of ischemia-relevant kinases. Commun Biol, 2024. 7(1): p. 321. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



