Submitted:
16 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Morphological Analysis
2.3. Molecular Analysis
2.4. Phylogenetic Analysis
3. Results
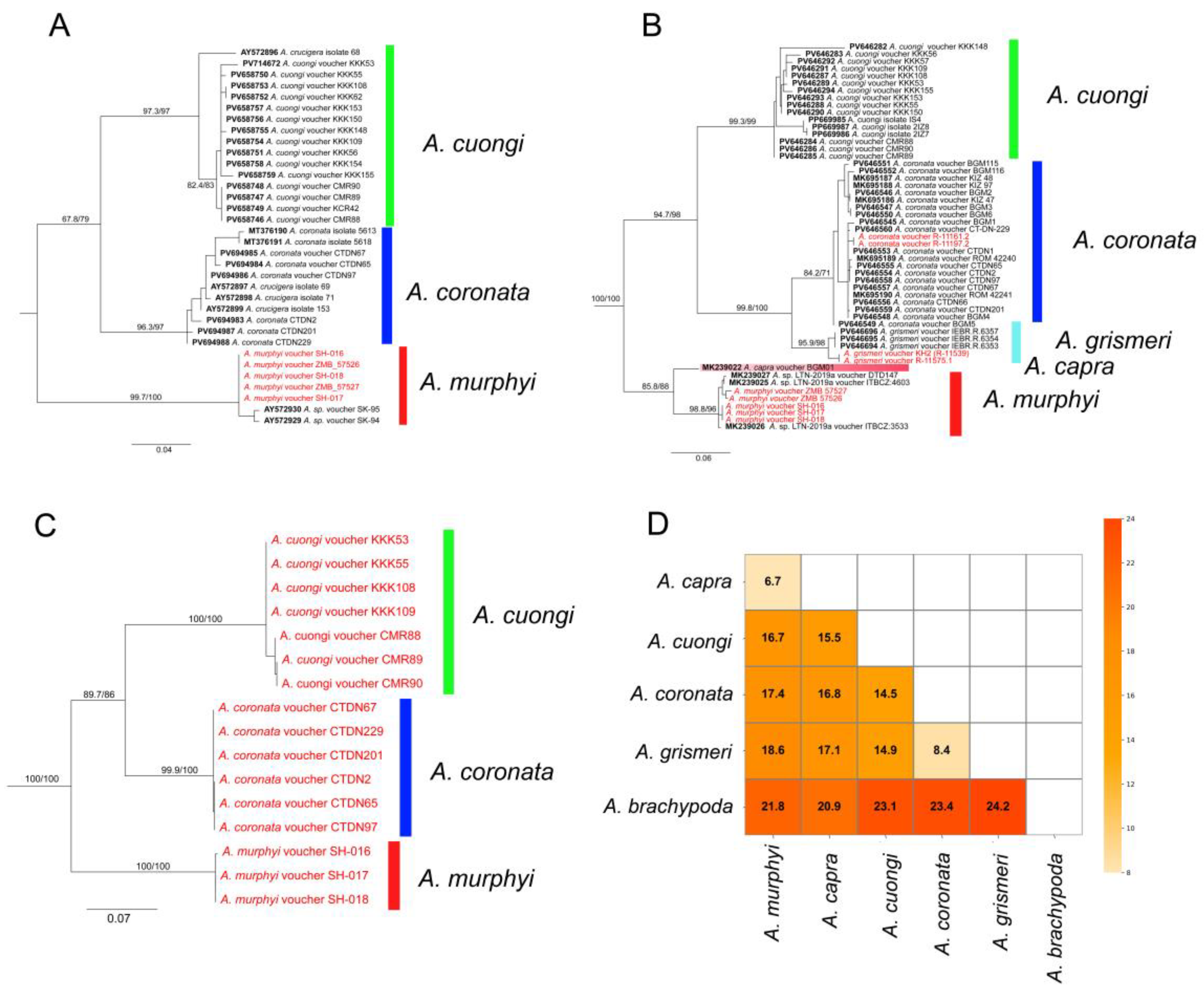
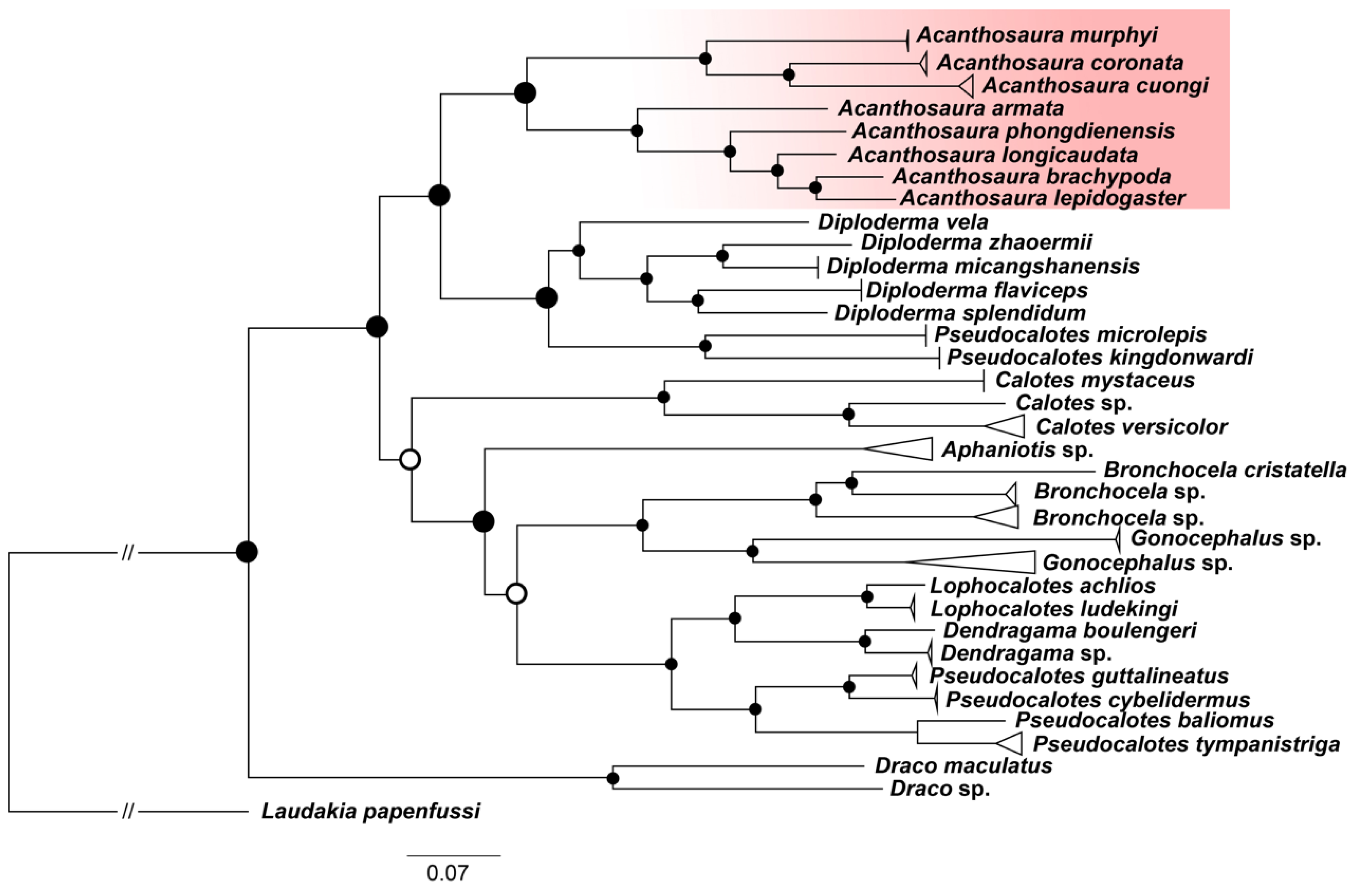
3.1. Phylogenetic Relationships
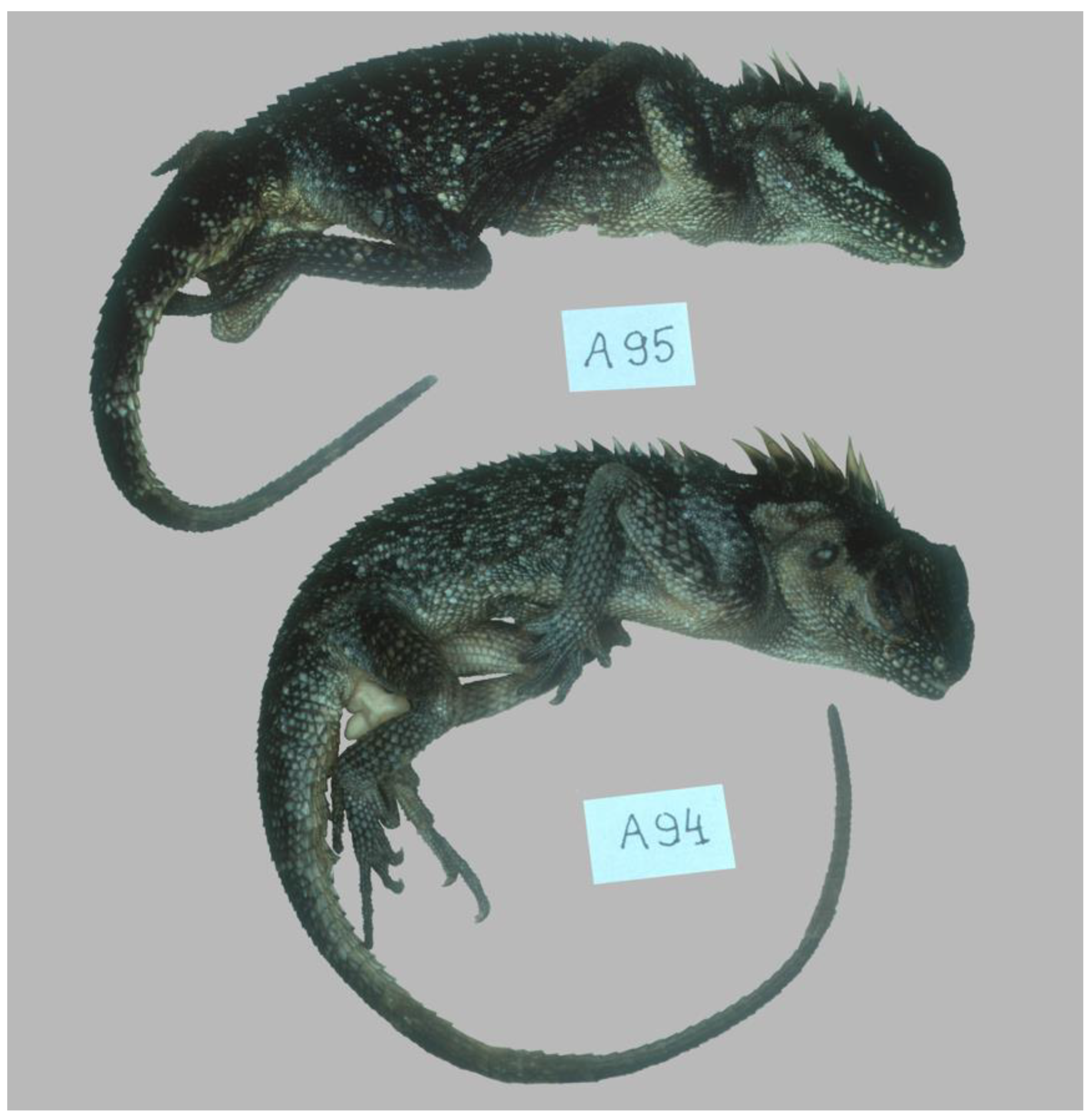
3.2. Morphology of Species from Acanthosaura coronata Complex
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HLMD | Hessisches Landesmuseum Darmstadt, Germany |
| ILS H | Herpetological collection of the Institute of Life Sciences (ISL), Vietnam Academy of Science and Technology, Ho Chi Minh City, Vietnam |
| ZMB | Universität Humboldt, Zoologisches Museum, Invalidenstrasse 43, 10115 Berlin, Germany |
| ZMMU | Zoological Museum of Moscow State University, 125009 B. Nikitskaya 2, Moscow, Russia |
References
- Uetz, P.; Freed, P.; Aquilar, R.; Reyes, F.; Kudera, J.; Hosek, J. (Eds.) The Reptile Database. Available online: http://www.reptile-database.org (accessed on 10 February 2026).
- Macey, J.R.; Schulte, J.A., II; Larson, A.; Ananjeva, N.B.; Wang, Y.; Pethiyagoda, R.; Rastegar-Pouyani, N.; Papenfuss, T.J. Evaluating trans-Tethys migration: an example using acrodont lizard phylogenetics. Syst. Biol. 2000, 49, 233–256. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Che, J.; Lin, S.; Deepak, V.; Aniruddha, D.-R.; Jiang, K.; Jin, J.; Chen, H.; Siler, C.D. Multilocus phylogeny and revised classification for mountain dragons of the genus Japalura s.l. (Reptilia: Agamidae: Draconinae) from Asia. Zool. J. Linn. Soc. 2018, 185, 246–267. [Google Scholar] [CrossRef]
- Shaney, K.J.; Maldonado, J.; Smart, U.; Thammachoti, P.; Fujita, M.; Hamidy, A.; Kurniawan, N.; Harvey, M.B.; Smith, E.N. Phylogeography of montane dragons could shed light on the history of forests and diversification processes on Sumatra. Mol. Phylogenet. Evol. 2020, 149, 106840. [Google Scholar] [CrossRef]
- Gowande, G.G.; Bhosale, H.S.; Phansalkar, P.U.; Sawant, M.; Mirza, Z.A. On the systematics and the phylogenetic position of the poorly known, montane dragon-lizard species Pseudocalotes austeniana (Annandale, 1908) (Squamata, Agamidae, Draconinae). Evol. Syst. 2021, 5, 141–150. [Google Scholar] [CrossRef]
- Harvey, M.B.; Larson, T.R.; Jacobs, J.L.; Shaney, K.; Streicher, J.W.; Hamidy, A.; Kurniawan, N.; Smith, E.N. Phoxophrys After 60 Years: Review of Morphology, Phylogeny, Status of Pelturagonia, and a New Species from Southeastern Kalimantan. Herpetol. Monogr. 2020, 33, 71–107, 37. [Google Scholar] [CrossRef]
- Ananjeva, N.B.; Ermakov, O.A.; Nguyen, S.N.; Nguyen, T.T.; Murphy, R.W.; Lukonina, S.A.; Orlov, N.L. A New Species of Acanthosaura Gray, 1831 (Squamata: Agamidae) from Central Highlands, Vietnam. Russ. J. Herpetol. 2020, 27, 217–230. [Google Scholar] [CrossRef]
- Ananjeva, N.B.; Matsiushova, M.I.; Svinin, A.O.; Bezman-Moseyko, O.S.; Nguyen, T.T.; Orlov, N.L. Unraveling Cryptic Diversity in Acanthosaura (Sauria: Squamata: Agamidae) Species Complexes in Vietnam: Research History and Current Status. Russ. J. Herpetol. 2025, 32, 211–232. [Google Scholar] [CrossRef]
- Honda, M.; Ota, H.; Kobayashi, M.; Nabhitabhata, J.; Yong, H.-S.; Hikida, T. Phylogenetic Relationships of the Flying Lizards, Genus Draco (Reptilia, Agamidae). Zool. Sci. 1999, 16, 535–549. [Google Scholar] [CrossRef]
- Huang, Y.; Li, H.; Wang, Y.; Li, M.; Hou, M.; Cai, B. Taxonomic review of the Calotes versicolor complex (Agamidae, Sauria, Squamata) in China, with description of a new species and subspecies. ZooKeys 2023, 1187, 63–89. [Google Scholar] [CrossRef]
- Ngo, H.N.; Le, L.T.H.; Nguyen, T.T.; Nguyen, T.M.; Nguyen, N.T.; Phan, T.Q.; Nguyen, T.Q.; Ziegler, T.; Do, D.T. A new species of Acanthosaura Gray, 1831 (Reptilia: Agamidae) from the Truong Son Mountain Range, Vietnam. Eur. J. Taxon. 2025, 976, 108–132. [Google Scholar] [CrossRef]
- Wood; Jr, P.L.; Grismer, L.L.; Grismer, J.L.; Neang, T.; Chav, T.; Holden, J. A new cryptic species of Acanthosaura Gray, 1831 (Squamata: Agamidae) from Thailand and Cambodia. Zootaxa 2010, 2488. [Google Scholar] [CrossRef]
- Xu, Y.; Gong, Y.; Hou, M.; Weng, S.; Liu, S.; Deng, J.; Hu, J.; Peng, L. A New Species of the Genus Pseudocalotes (Squamata: Agamidae) from Southwest Yunnan, China. Animals 2024, 14, 826. [Google Scholar] [CrossRef]
- Putra, C.A.; Thasun Amarasinghe, A.A.; Hikmatullah, D.; Scali, S.; Brinkman, J.J.; Manthey, U.; Ineich, I. Rediscovery of Modigliani’s nose-horned lizard, Harpesaurus modiglianii Vinciguerra, 1933 (Reptilia: Agamidae) after 129 years without any observation. Taprobanica 2020, 9, 3–11. [Google Scholar] [CrossRef]
- Liu, S.; Zhang, D.; Hou, M.; Orlov, N.L.; Rao, D.; Ananjeva, N.B.; Li, S. Taxonomic Assessment of Acanthosaura lepidogaster sensu lato (Reptilia: Agamidae) in China Through Extensive Sampling. Russ. J. Herp. 2023, 30, 127–143. [Google Scholar] [CrossRef]
- Kalyabina-Hauf, S.; Ananjeva, N.B.; Joger, U.; Lenk, P.; Murphy, R.W.; Stuart, B.L.; Orlov, N.L.; Ho, C.T.; Wink, M. Molecular Phylogeny of the Genus Acanthosaura (Agamidae). Curr. Herpetol. 2004, 23, 7–16. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Do, D.T.; Hoang, H.V.; Nguyen, T.T.; McCormack, T.E.M.; Nguyen, T.Q.; Orlov, N.L.; Nguyen, V.D.H.; Nguyen, S.N. A New Species of the Genus Acanthosaura Gray, 1831 (Reptilia: Agamidae) from Central Vietnam. Rus. J. Herp. 2018, 25, 259. [Google Scholar] [CrossRef]
- Ananjeva, N.B.; Orlov, N.L.; Kalyabina-Hauf, S.A. Species of Acanthosaura Gray, 1831 (Agamidae: Sauria, Reptilia) of Vietnam: results of molecular and morphological study. Biol. Bull. 2008, 35, 178–186. [Google Scholar] [CrossRef]
- Orlov, N.L.; Nguyen, T.Q.; Nguyen, S.V. A new Acanthosaura allied to A. capra Günther, 1861 (Agamidae, Sauria) from central Vietnam and southern Laos. Rus. J. Herp. 2006, 13, 61–76. [Google Scholar] [CrossRef]
- Liu, S.; Rao, D. A new species of the genus Acanthosaura from Yunnan, China (Squamata, Agamidae). Zookeys 2019, 888, 105–132. [Google Scholar] [CrossRef]
- Abramson, N.I.; Petrova, T.V.; Dokuchaev, N.E.; Obolenskaya, E.V.; Lissovsky, A.A. Phylogeography of the gray red-backed vole Craseomys rufocanus (Rodentia: Cricetidae) across the distribution range inferred from nonrecombining molecular markers. Russ. J. Theriol. 2012, 11, 137–156. [Google Scholar] [CrossRef]
- Irwin, D.M.; Kocher, T.D.; Wilson, A.C. Evolution of the cytochrome b gene of mammals. J. Mol. Evol. 1991, 32, 128–144. [Google Scholar] [CrossRef]
- Macey, J.R.; Larson, A.; Ananjeva, N.B.; Papenfuss, T.J. Replication slippage may cause parallel evolution in the secondary structures of mitochondrial transfer RNAs. Mol. Biol. Evol. 1997, 14, 30–39. [Google Scholar] [CrossRef]
- McGuire, J.A.; Heang, K.B. Phylogenetic systematics of Southeast Asian flying lizards (Iguania: Agamidae: Draco) as inferred from mitochondrial DNA sequence data. Biol. J. Linn. Soc. 2001, 72, 203–229. [Google Scholar] [CrossRef]
- Katoh, K. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.-T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; Von Haeseler, A.; Jermiin, L.S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Manthey, U. Agamid Lizards of Southern Asia Draconinae 1. In Terralog; Frankfurt am Main/Rodgau 2008: Edition Chimaira/Verlag ACS GmbH (AQUALOG), 2008; Vol 7a, pp. 1–161. [Google Scholar]



| Locus | Primer name | Sequence (5’ – 3’) | Reference |
| COI | VUTF | TGTAAAACGACGGCCAGTTCTCAACCAAYCAYAARGAYATYGG | [21] |
| VUTR | CAGGAAACAGCTATGACTARACTTCTGGRTGKCCRAARAAYCA | ||
| cyt b | L14841 | CCATCCAACATCTCAGCATGATGAAA | [22] |
| H151495 | GCCCCTCAGAATGATATTTGTCCTCA | ||
| ND2 | METF6 | AAGCTTTCGGGCCCATACC | [23,24] |
| ALAr.2m | AAAGTGTCTGAGTTGCATTCRG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




