Submitted:
13 March 2026
Posted:
16 March 2026
You are already at the latest version
Abstract
Keywords:
1. INTRODUCTION
2. METHODS
2.1. Datasets and Preprocessing
2.2. AENetMoX Architecture
- Centrality (8): Degree, weighted degree, betweenness, closeness, eigenvector, PageRank, clustering coefficient, and average neighbor degree.
- Neighborhood aggregates (6): The six expression statistics of neighbor TFs.
- Contrasts (3): TF variance vs. neighborhood variance (activity weighted degree, expression contrast, and stability contrast)
2.3. Experimental Setup
2.3.1. Timepoint Partitioning
2.3.2. Baseline Methods
- CLR (Faith et al., 2007): A statistical method based on Mutual Information with context likelihood normalization.
- GRNBoost2 (Moerman et al., 2019): A regression-based method using gradient boosting.
- SCENIC (Aibar et al., 2017): An ensemble framework that combines GRNBoost2 with motif-based pruning,
- - motifs-v10nr_clust-nr.hgnc-m0.001-o0.0.tbl
- hg38__refseq-r80__10kb_up_and_down_tss.mc9nr.genes_vs_motifs.rankings.feather
- hg38__refseq-r80__500bp_up_and_100bp_down_tss.mc9nr.genes_vs_motifs.rankings.feather
- motifs-v10nr_clust-nr.hgnc-m0.001-o0.0.tbl
2.3.3. Ablation Studies
4. RESULTS AND DISCUSSION
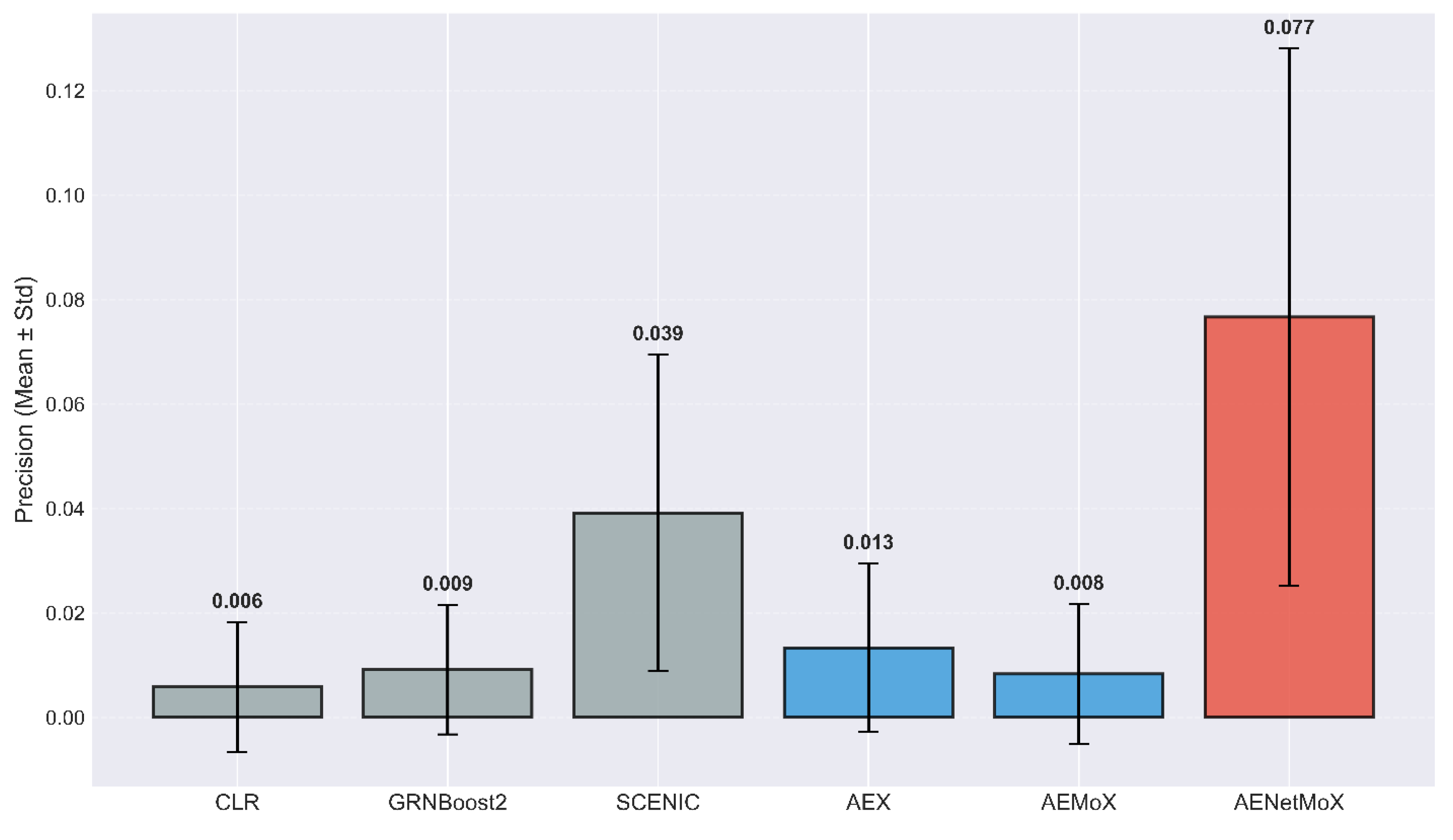
4.1. Performance Summary
Supplementary Materials
Funding
Acknowledgments
Conflicts of Interest
Availability and Implementation
References
- Agrawal, A.; et al. WikiPathways 2024: next generation pathway database. Nucleic Acids Res 2024, 52, D679–D689. [Google Scholar] [CrossRef]
- Aibar, S.; et al. SCENIC: single-cell regulatory network inference and clustering. Nat Methods 2017, 14, 1083–1086. [Google Scholar] [CrossRef]
- Aleksander, S.A.; et al. The Gene Ontology knowledgebase in 2026. Nucleic Acids Res. 2025. [Google Scholar]
- Bateman, A.; et al. UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res 2025, 53, D609–D617. [Google Scholar]
- Birtele, M.; et al. Modelling human brain development and disease with organoids. Nat Rev Mol Cell Biol 2025, 26, 389–412. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Liu, Z.-P. Graph attention network for link prediction of gene regulations from single-cell RNA-sequencing data. Bioinformatics 2022, 38, 4522–4529. [Google Scholar] [CrossRef]
- Dennis, D.J.; et al. bHLH transcription factors in neural development, disease, and reprogramming. Brain Res 2019, 1705, 48–65. [Google Scholar] [CrossRef] [PubMed]
- Faith, J.J.; et al. Large-Scale Mapping and Validation of Escherichia coli Transcriptional Regulation from a Compendium of Expression Profiles. PLoS Biol 2007, 5, e8–e8. [Google Scholar] [CrossRef]
- Feng, C.; et al. KnockTF 2.0: a comprehensive gene expression profile database with knockdown/knockout of transcription (co-)factors in multiple species. Nucleic Acids Res 2024, 52, D183–D193. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Alonso, L.; et al. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res 2019, 29, 1363–1375. [Google Scholar] [CrossRef]
- Hammal, F.; et al. ReMap 2022: a database of Human, Mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res 2022, 50, D316–D325. [Google Scholar] [CrossRef]
- Han, H.; et al. TRRUST v2: an expanded reference database of human and mouse transcriptional regulatory interactions. Nucleic Acids Res 2018, 46, D380–D386. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res 2019, 47, D33–D38. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; et al. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res 2021, 49, D545–D551. [Google Scholar] [CrossRef]
- Kartha, V.K.; et al. Functional inference of gene regulation using single-cell multi-omics. Cell Genomics 2022, 2, 100166. [Google Scholar] [CrossRef]
- Kuleshov, M. V.; et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 2016, 44, W90–W97. [Google Scholar] [CrossRef] [PubMed]
- Lambert, S.A.; et al. The Human Transcription Factors. Cell 2018, 172, 650–665. [Google Scholar] [CrossRef]
- Meng, X.; et al. CRISPR screens in human neural organoids and assembloids. Nat Protoc. 2025. [Google Scholar] [CrossRef]
- Meylan, P.; et al. EPD in 2020: enhanced data visualization and extension to ncRNA promoters. Nucleic Acids Res. 2019. [Google Scholar] [CrossRef]
- Milacic, M.; et al. The Reactome Pathway Knowledgebase 2024. Nucleic Acids Res 2024, 52, D672–D678. [Google Scholar] [CrossRef]
- Moerman, T.; et al. GRNBoost2 and Arboreto: efficient and scalable inference of gene regulatory networks. Bioinformatics 2019, 35, 2159–2161. [Google Scholar] [CrossRef]
- Mudge, J.M.; et al. GENCODE 2025: reference gene annotation for human and mouse. Nucleic Acids Res 2025, 53, D966–D975. [Google Scholar] [CrossRef] [PubMed]
- Omidi, A.; et al. AlphaFold-Multimer accurately captures interactions and dynamics of intrinsically disordered protein regions. In Proceedings of the National Academy of Sciences, 2024; p. 121. [Google Scholar]
- Pratapa, A.; et al. Benchmarking algorithms for gene regulatory network inference from single-cell transcriptomic data. Nat Methods 2020, 17, 147–154. [Google Scholar] [CrossRef]
- Quadrato, G.; et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 2017, 545, 48–53. [Google Scholar] [CrossRef]
- Rauluseviciute, I.; et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res 2024, 52, D174–D182. [Google Scholar] [CrossRef]
- Sokpor, G.; et al. Chromatin Remodeling BAF (SWI/SNF) Complexes in Neural Development and Disorders. Front Mol Neurosci 2017, 10. [Google Scholar] [CrossRef]
- Stuart, T.; et al. Comprehensive Integration of Single-Cell Data. Cell 2019, 177, 1888–1902.e21. [Google Scholar] [CrossRef]
- Szklarczyk, D.; et al. The STRING database in 2025: protein networks with directionality of regulation. Nucleic Acids Res 2025, 53, D730–D737. [Google Scholar] [CrossRef]
- Wolf, F.A.; et al. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 2018, 19, 15. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Bar-Joseph, Z. Deep learning for inferring gene relationships from single-cell expression data. Proceedings of the National Academy of Sciences 2019, 116, 27151–27158. [Google Scholar] [CrossRef] [PubMed]
| Metric | AENetMoX | Best Baseline | vs Best Baseline | p-value | |
|---|---|---|---|---|---|
| Precision | SCENIC: | ||||
| AUPRC | SCENIC: | ||||
| ChIP-seq Precision | SCENIC: | ||||
| ChIP-seq F1 | CLR: | ||||
| ChIP-seq F1 vs SCENIC |
| Method | Features | vs AEX) | AUPRC vs AEX) | ChIP-seq Precision vs AEX) | vs AEX) |
|---|---|---|---|---|---|
| AENetMoX | Expression + Motif + PPI |
|
|
|
|
| AEMoX | Expression + Motif |
|
|
|
|
| AEX | Expression only |
| Method | Coherence (% TFs) | Enrichment (%) | ChIP-seq Precision (%) | Hub TFs (%) | Largest Component (%) |
|---|---|---|---|---|---|
| CLR | |||||
| GRNBoost2 | |||||
| SCENIC | |||||
| AEX | |||||
| AEMoX | |||||
| AENetMoX |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




