Submitted:
14 March 2026
Posted:
16 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
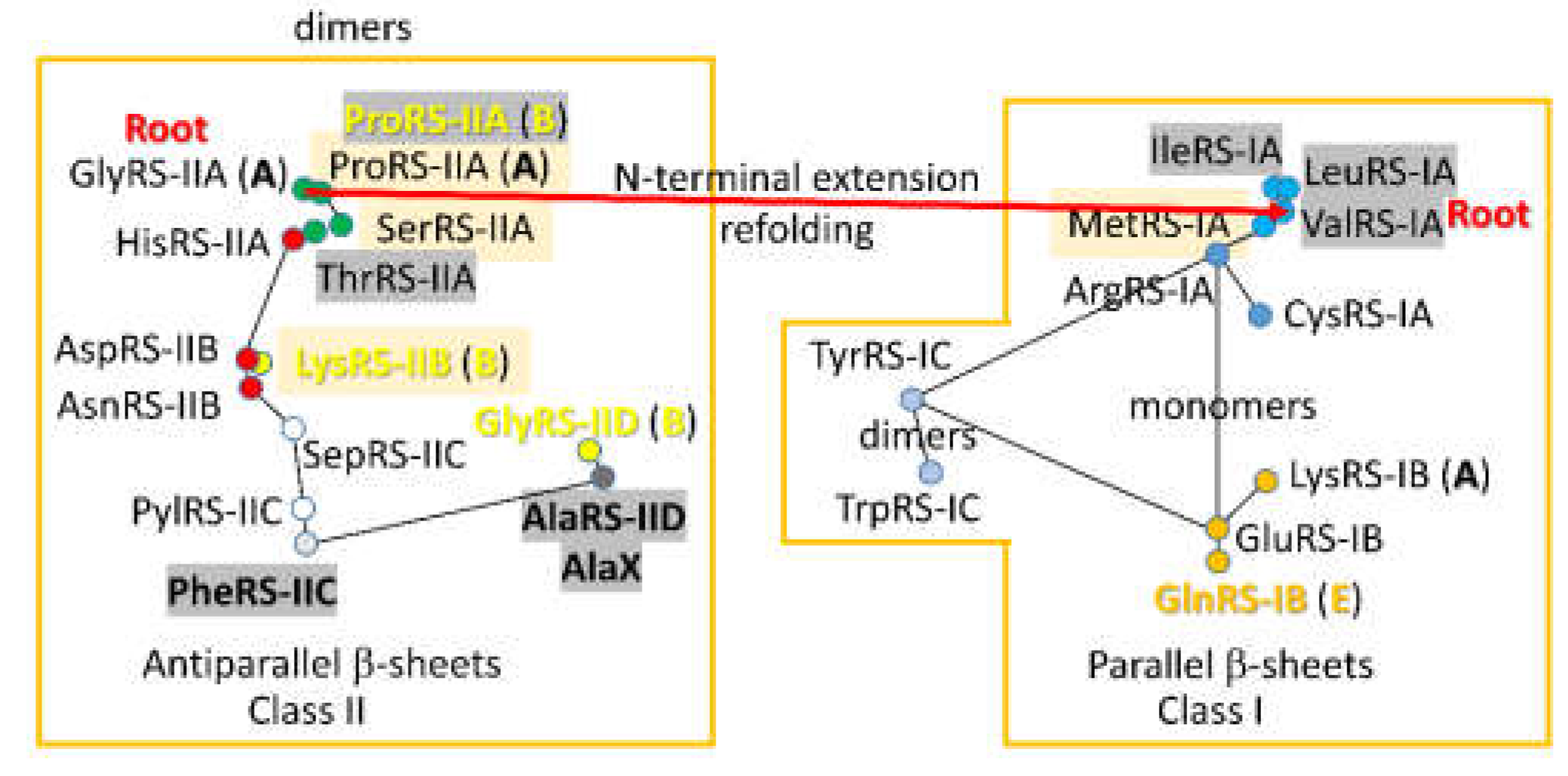
3. AARS Enzymes at the Base of Code Evolution
3.1. The AARS Mechanism
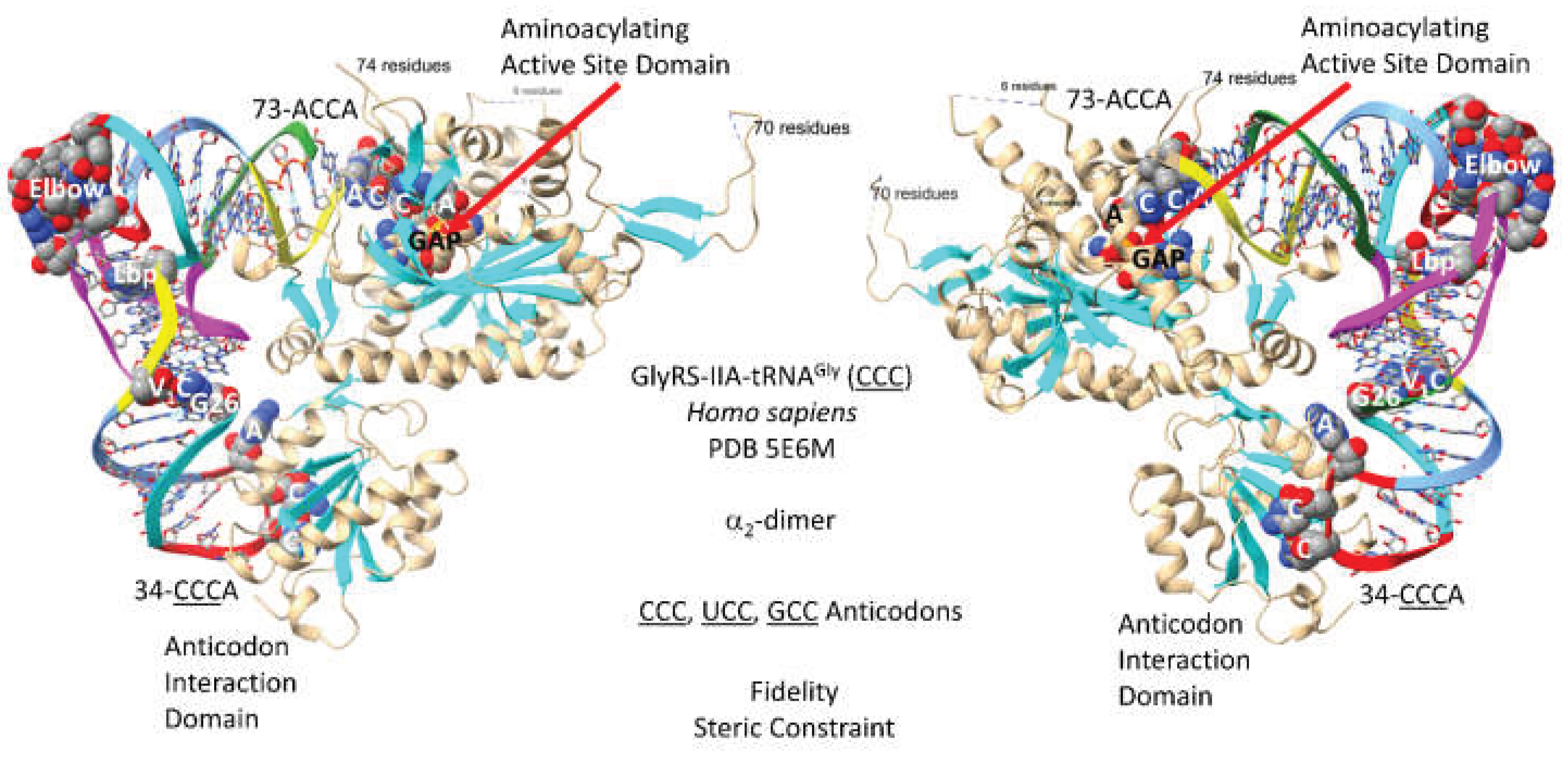
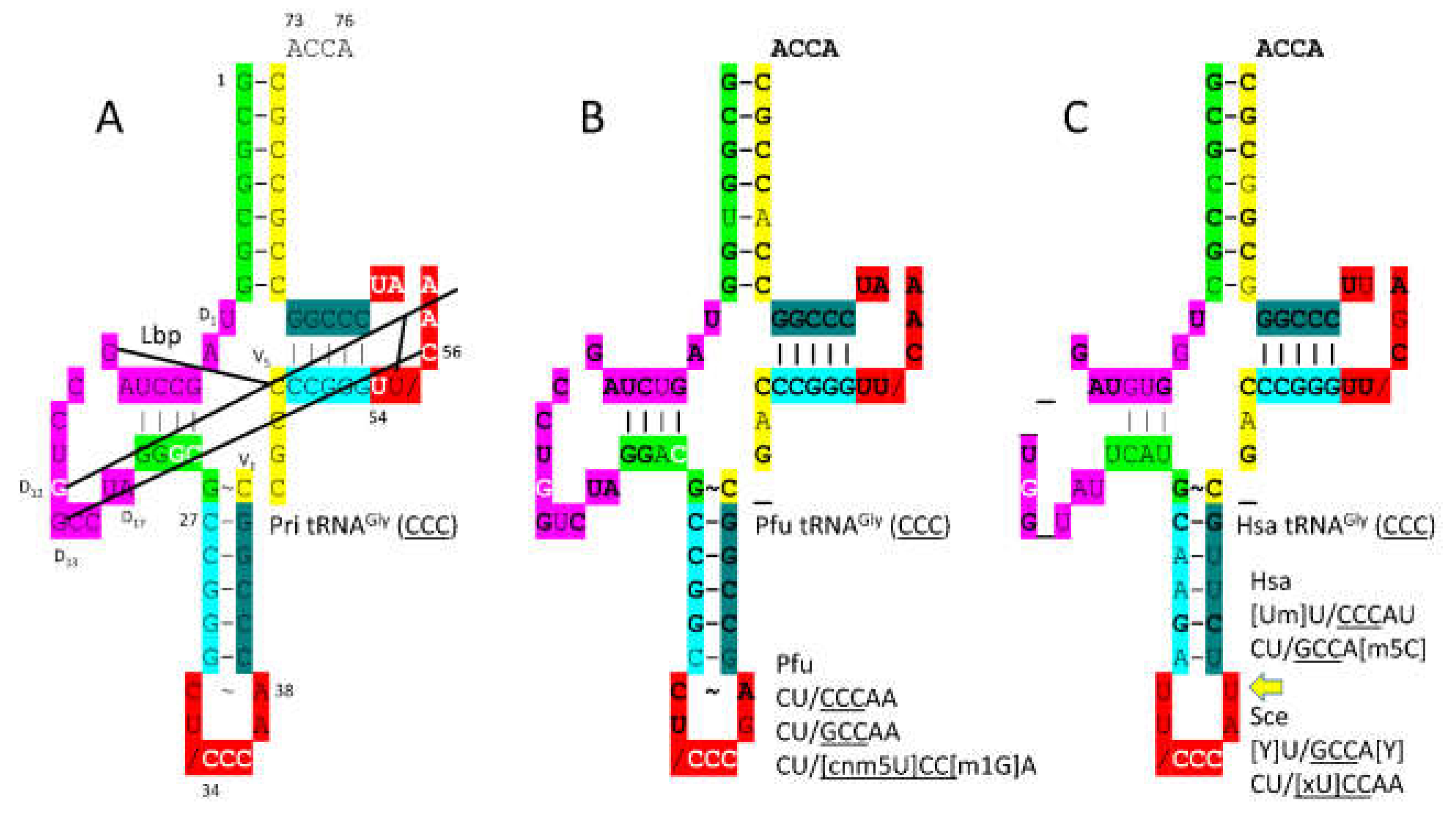
3.2. GlyRS-IIA
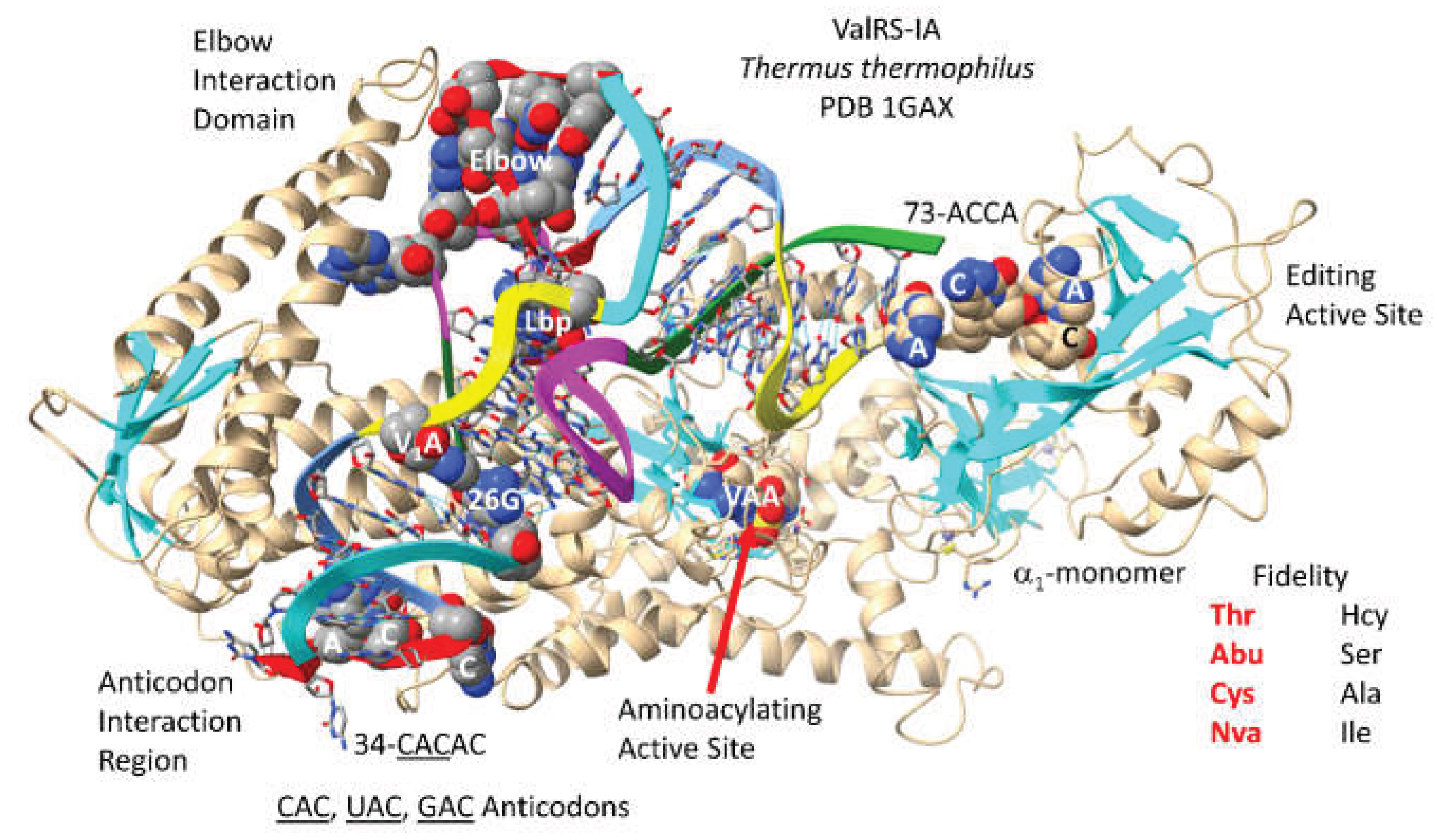
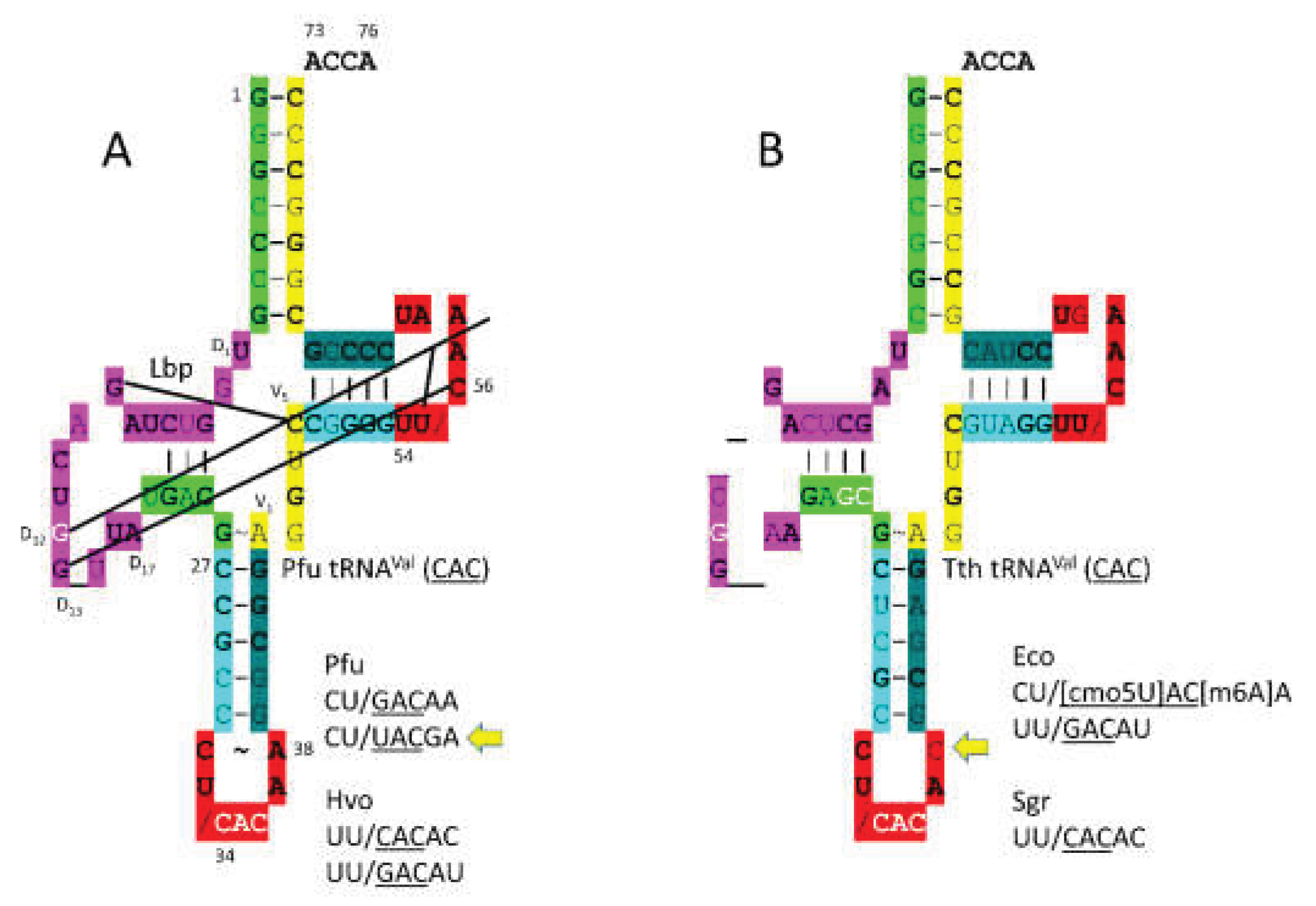
3.3. ValRS-IA
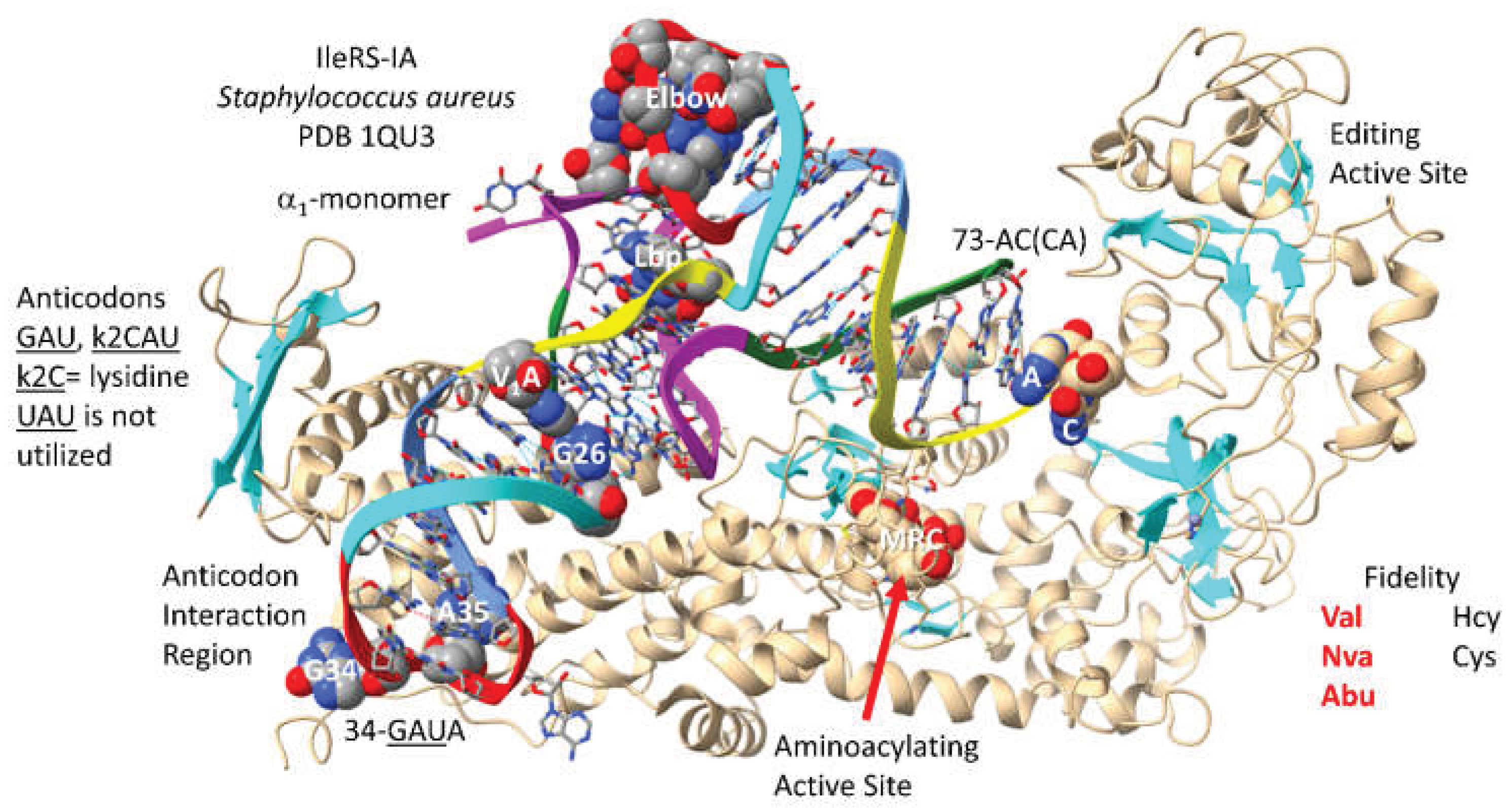
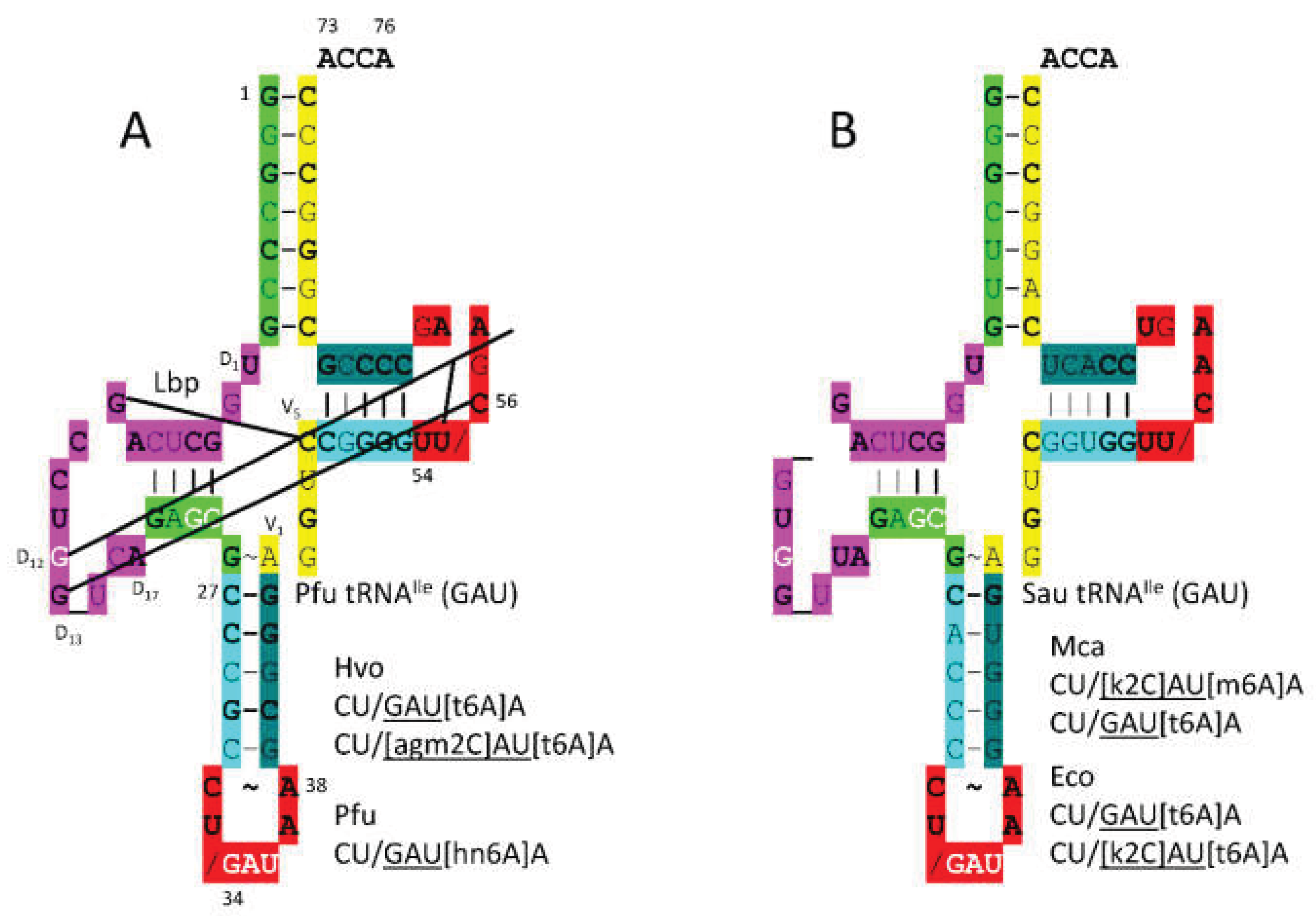
3.4. IleRS-IA
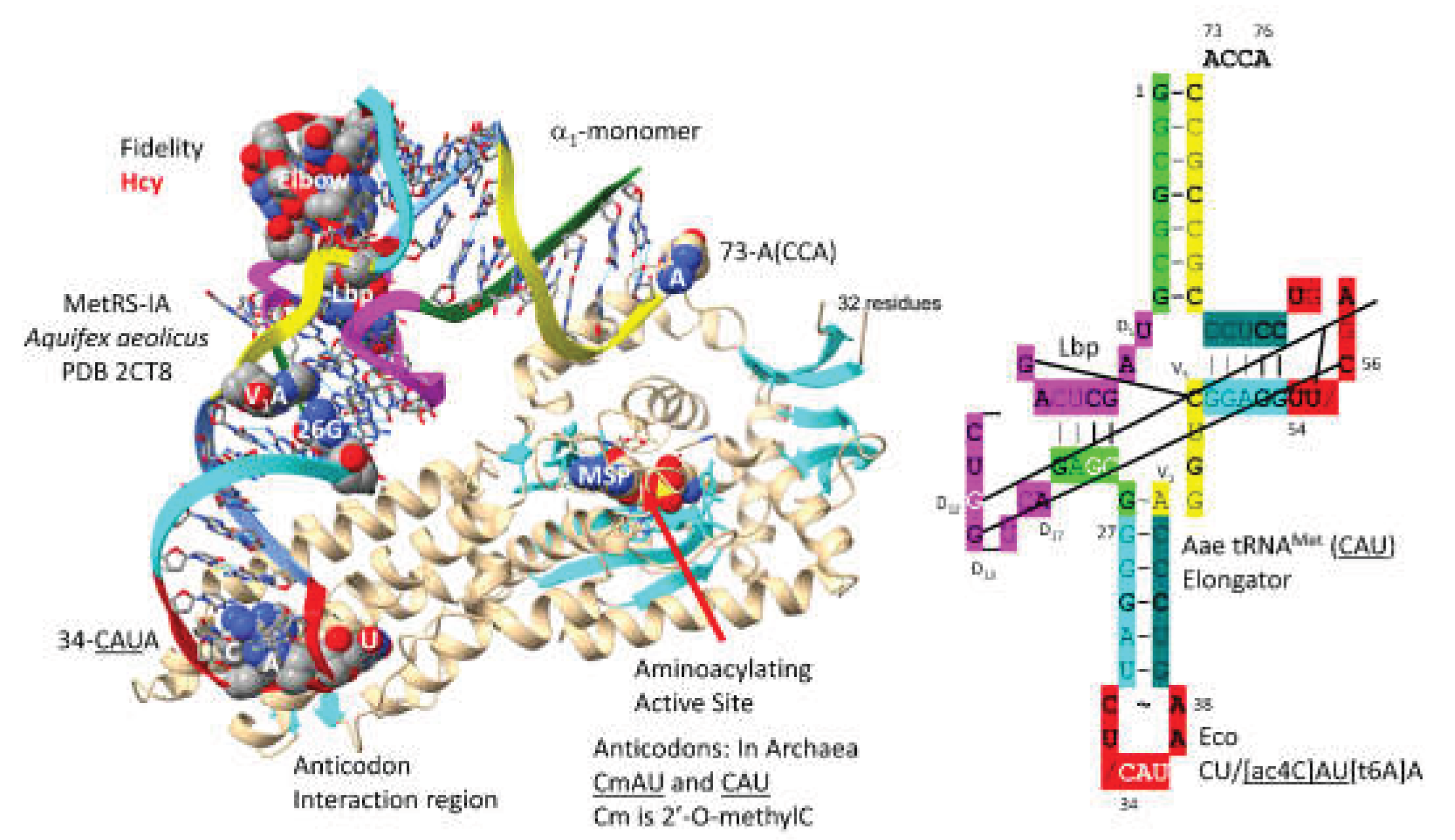
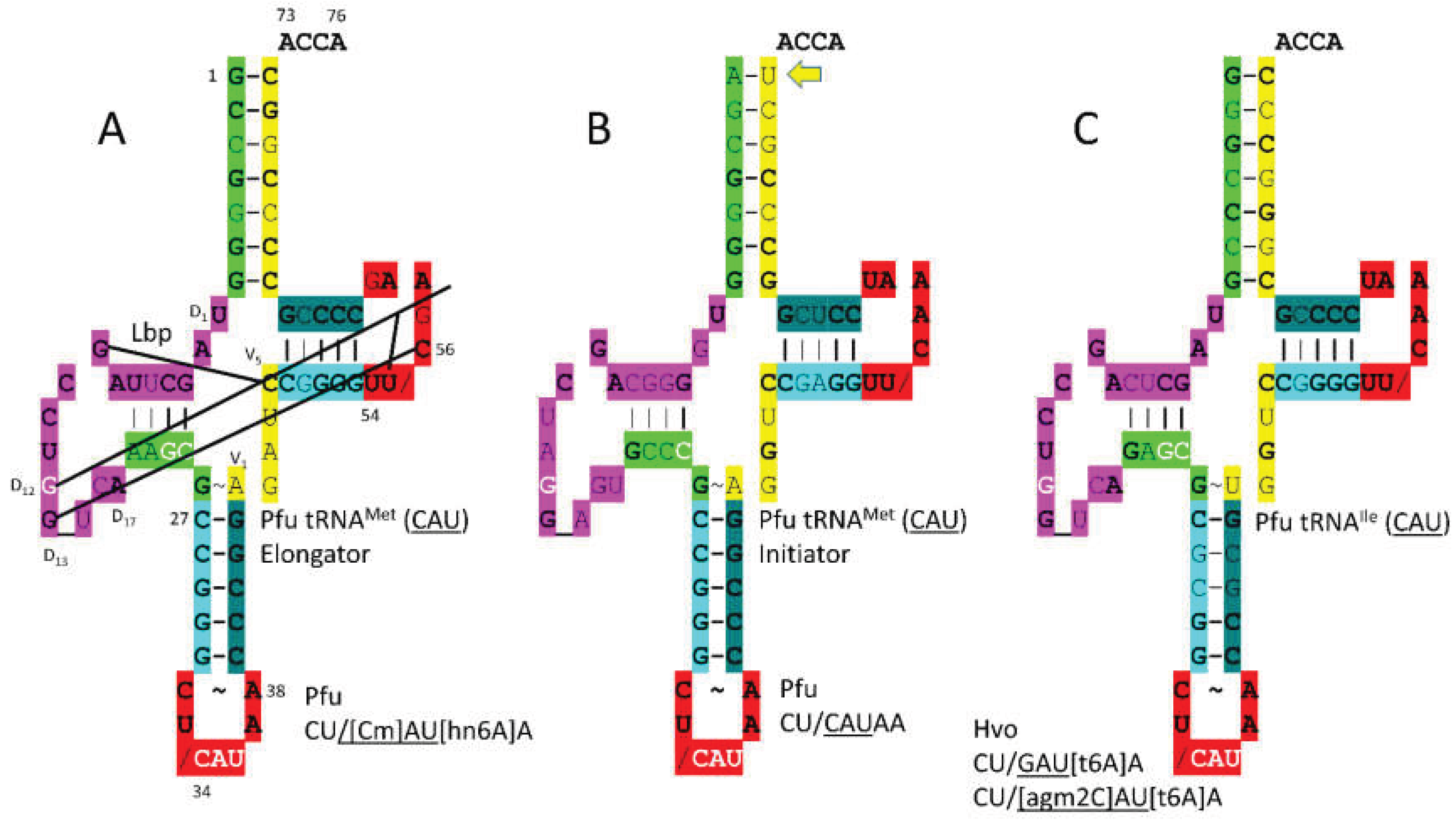
3.5. MetRS-IA
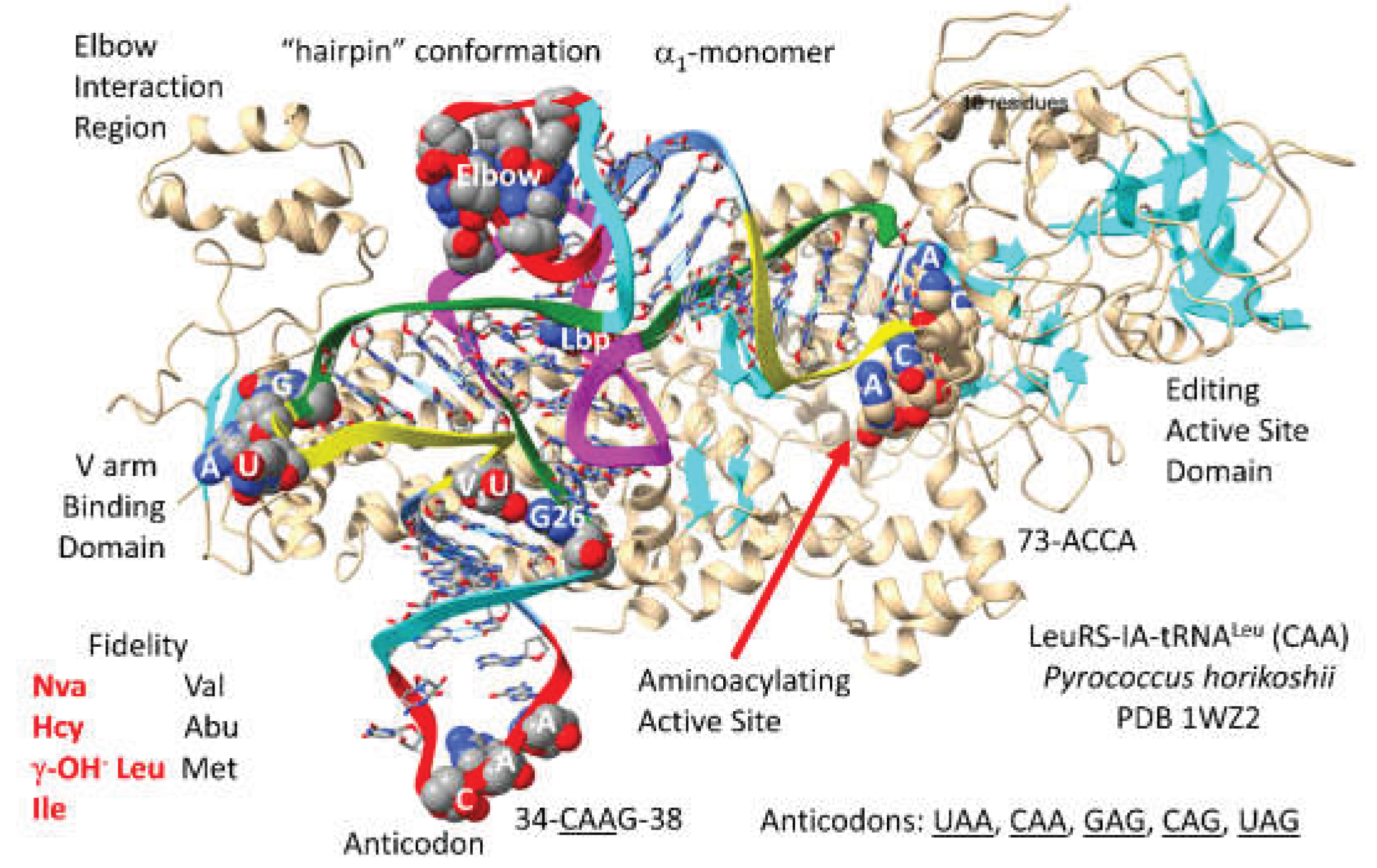
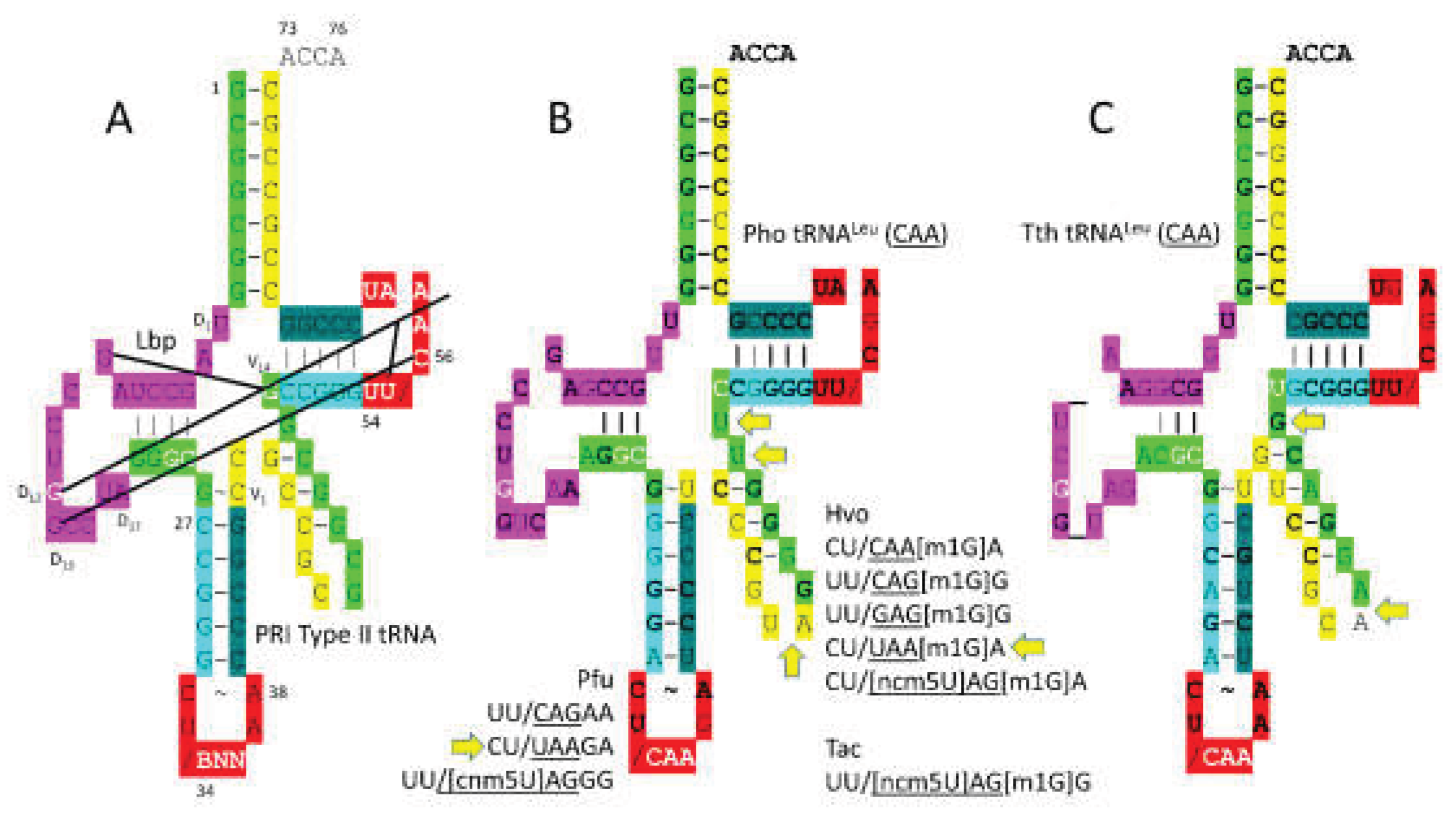
3.6. LeuRS-IA
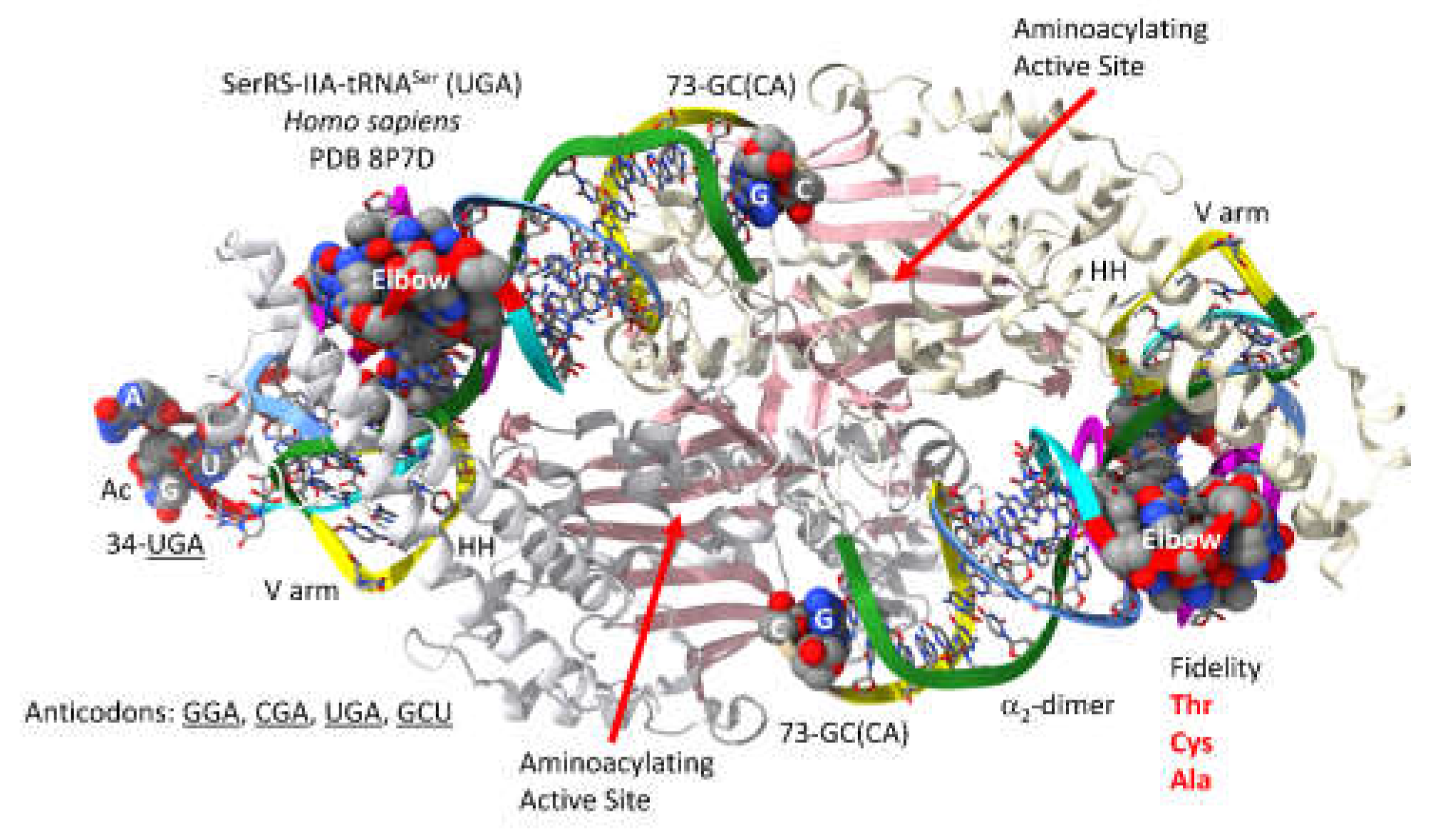
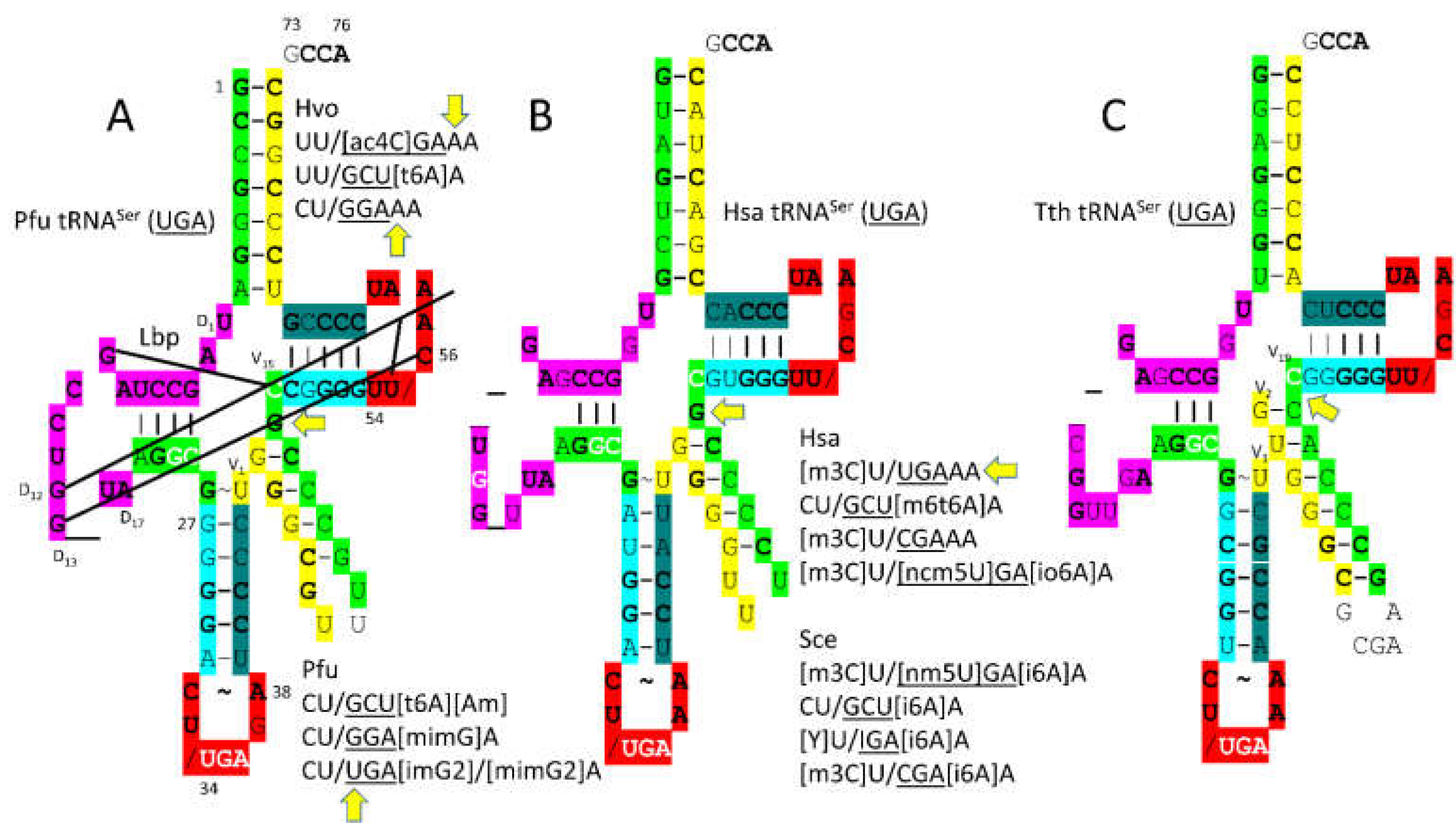
3.7. SerRS-IIA
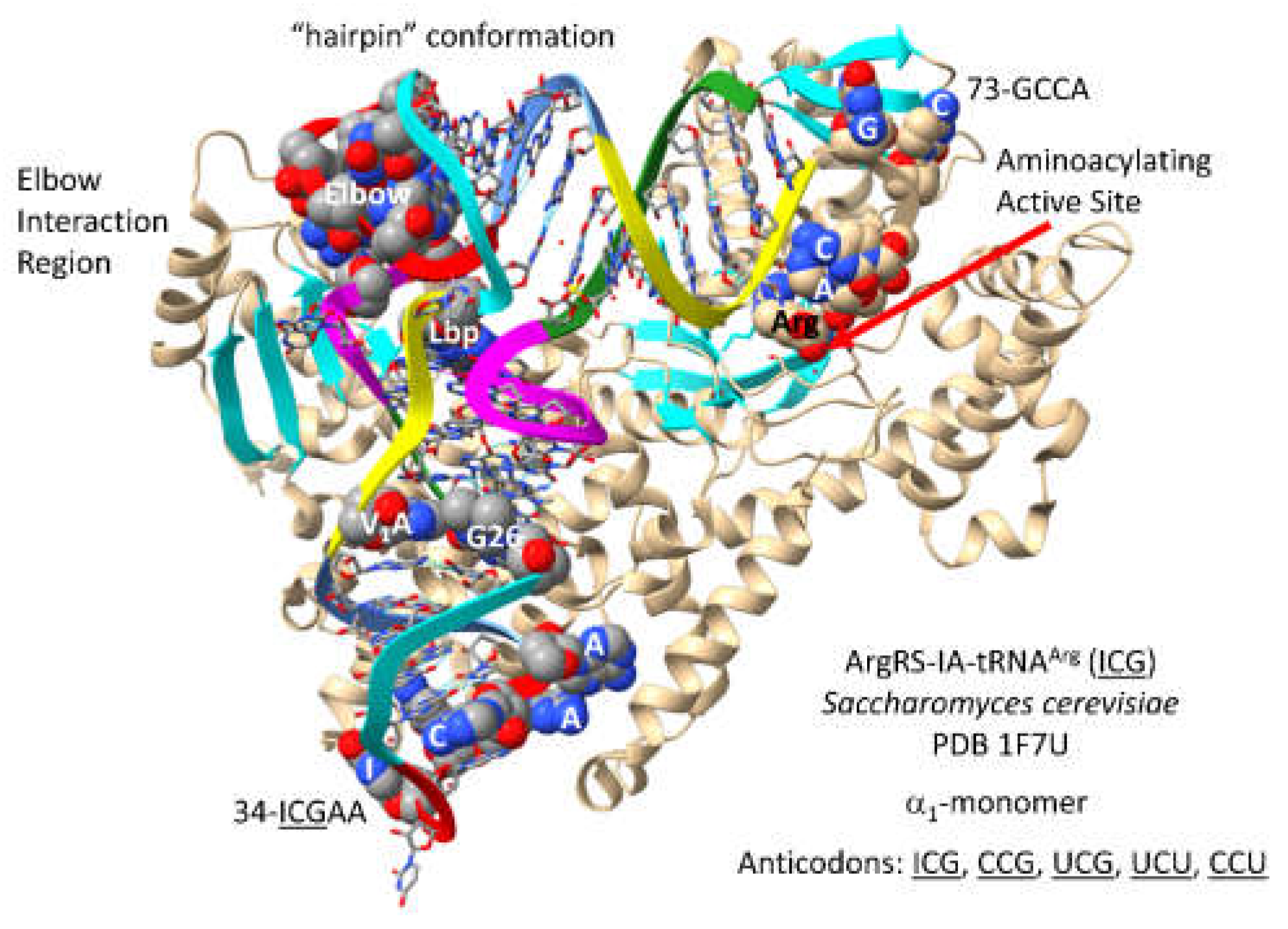
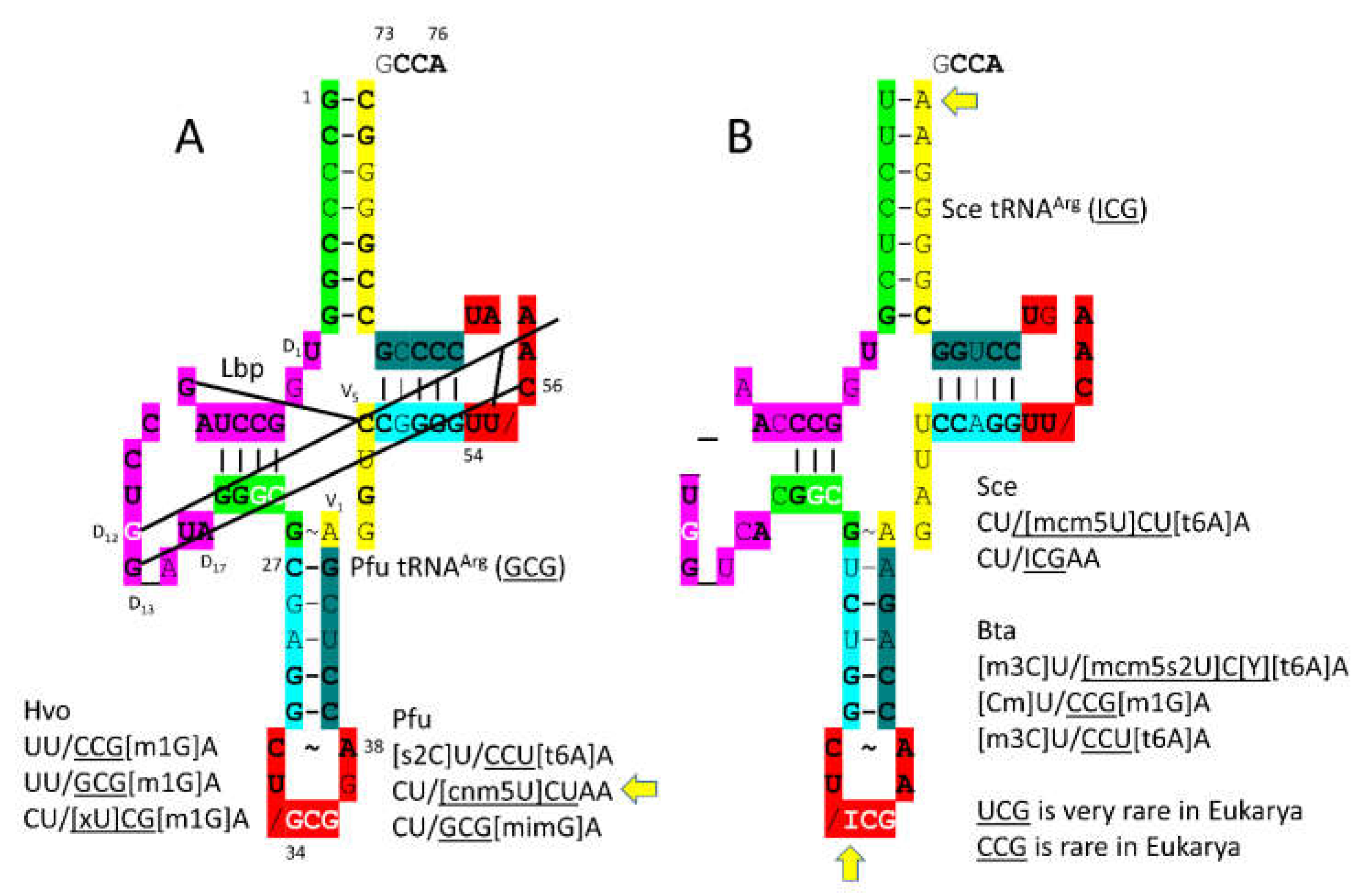
3.8. ArgRS-IA
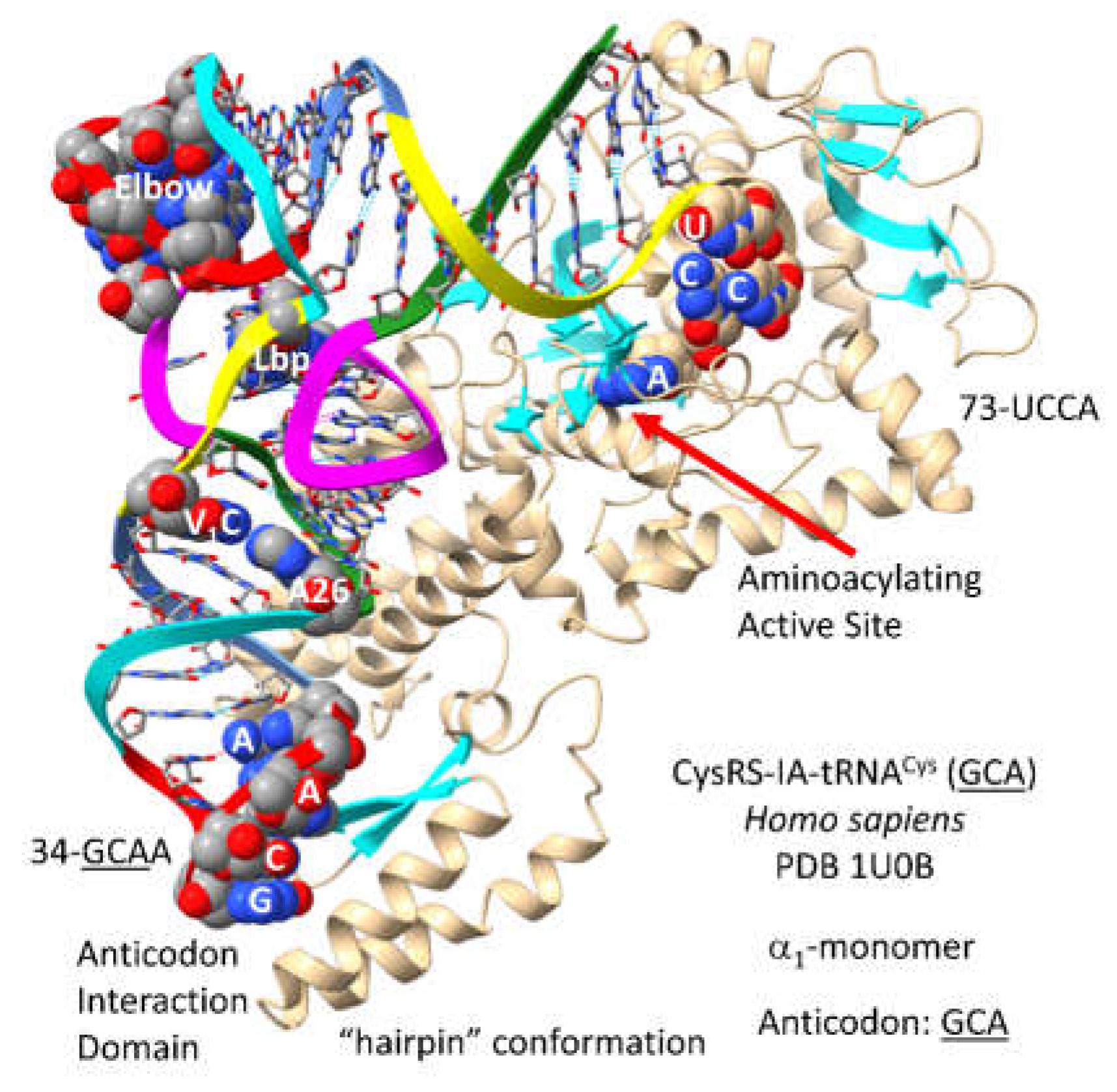
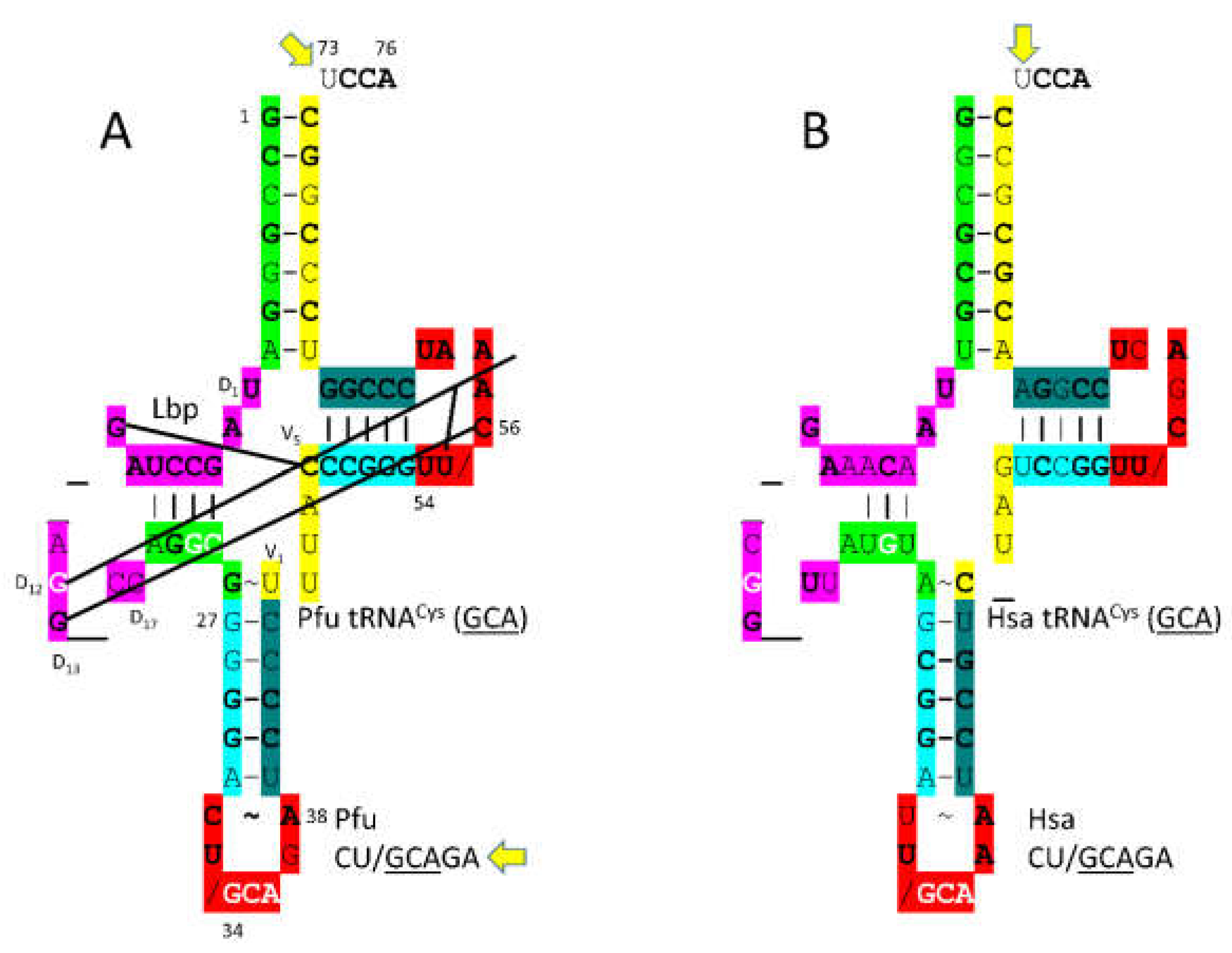
3.9. CysRS-IA
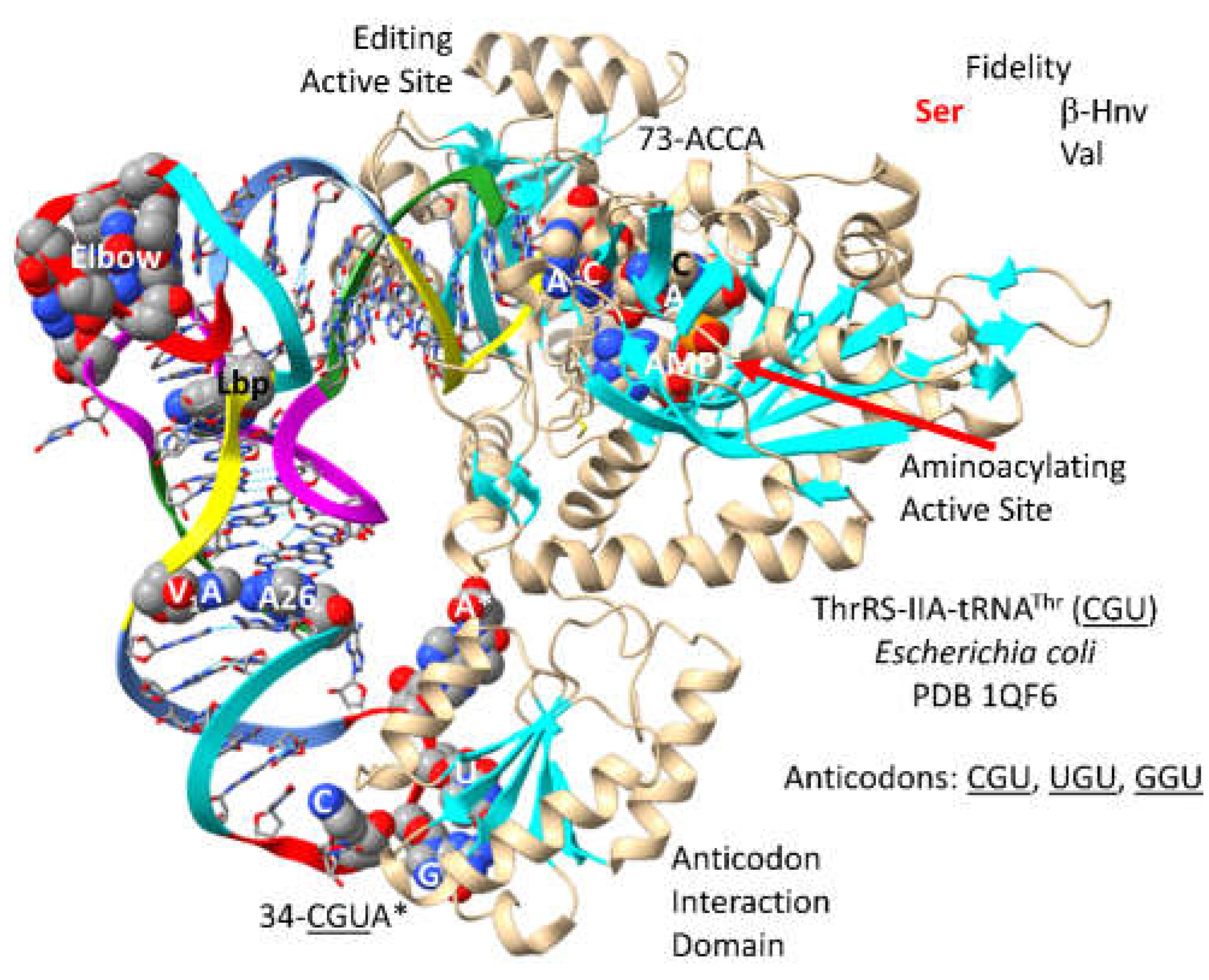
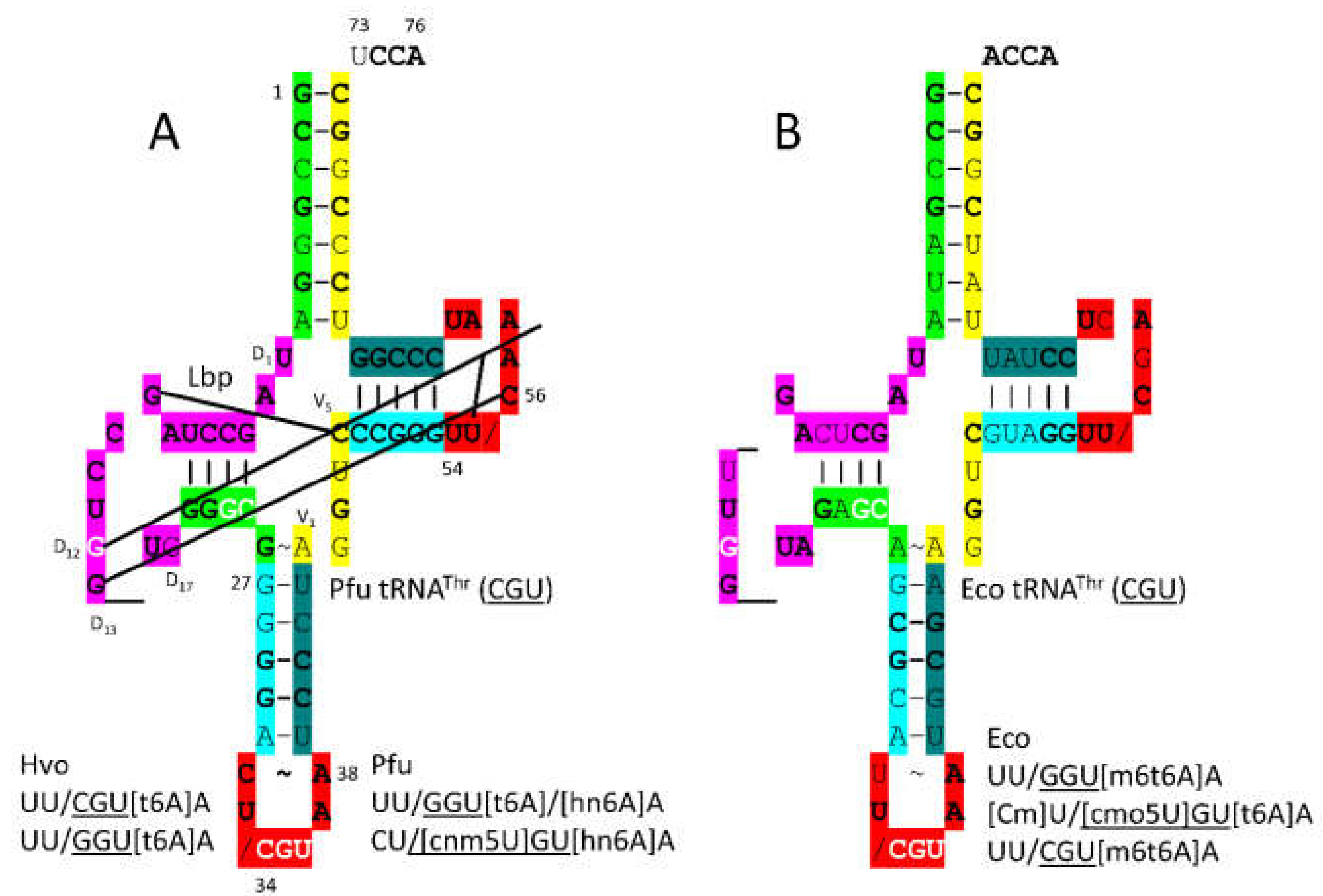
3.10. ThrRS-IIA
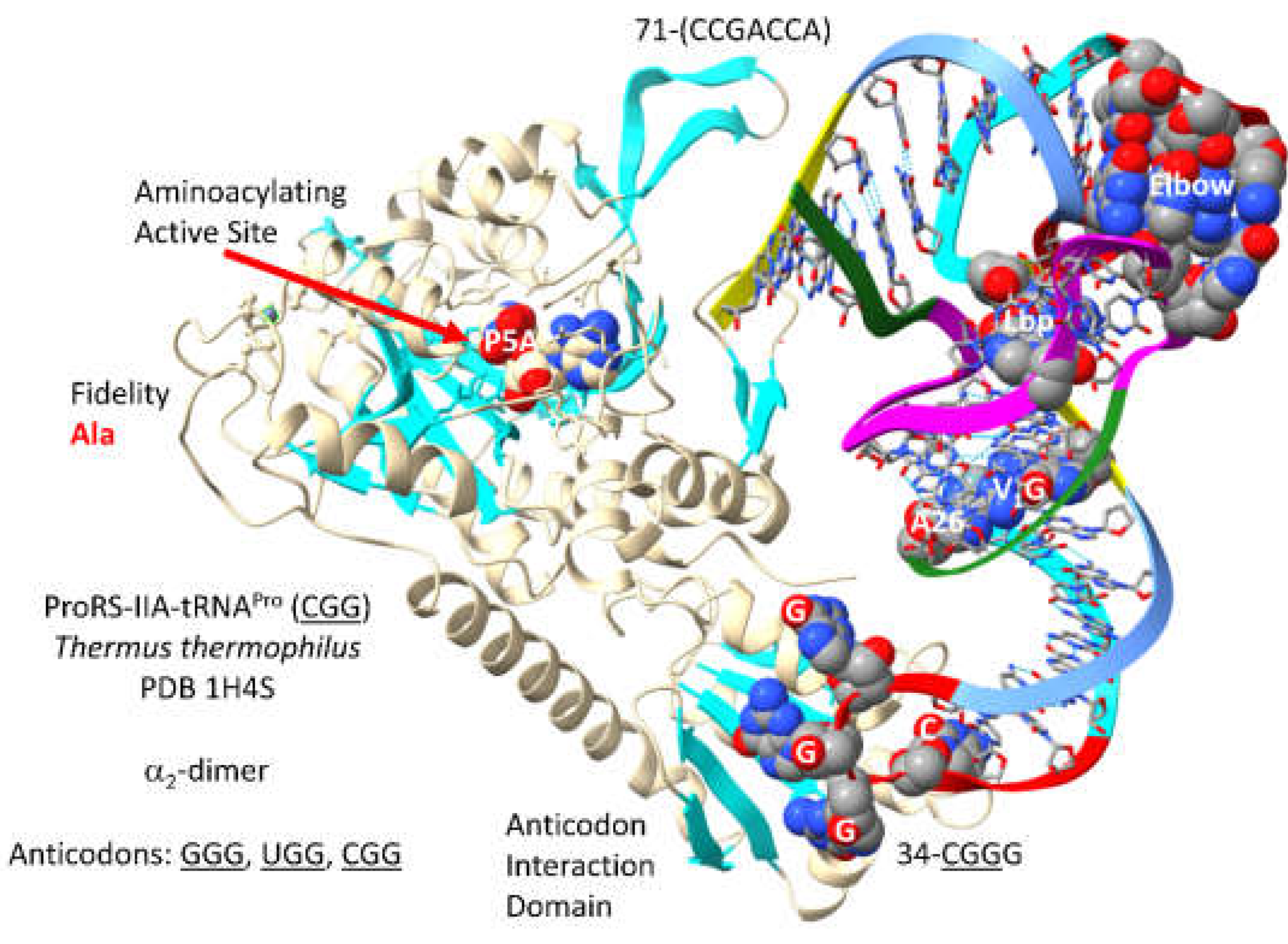
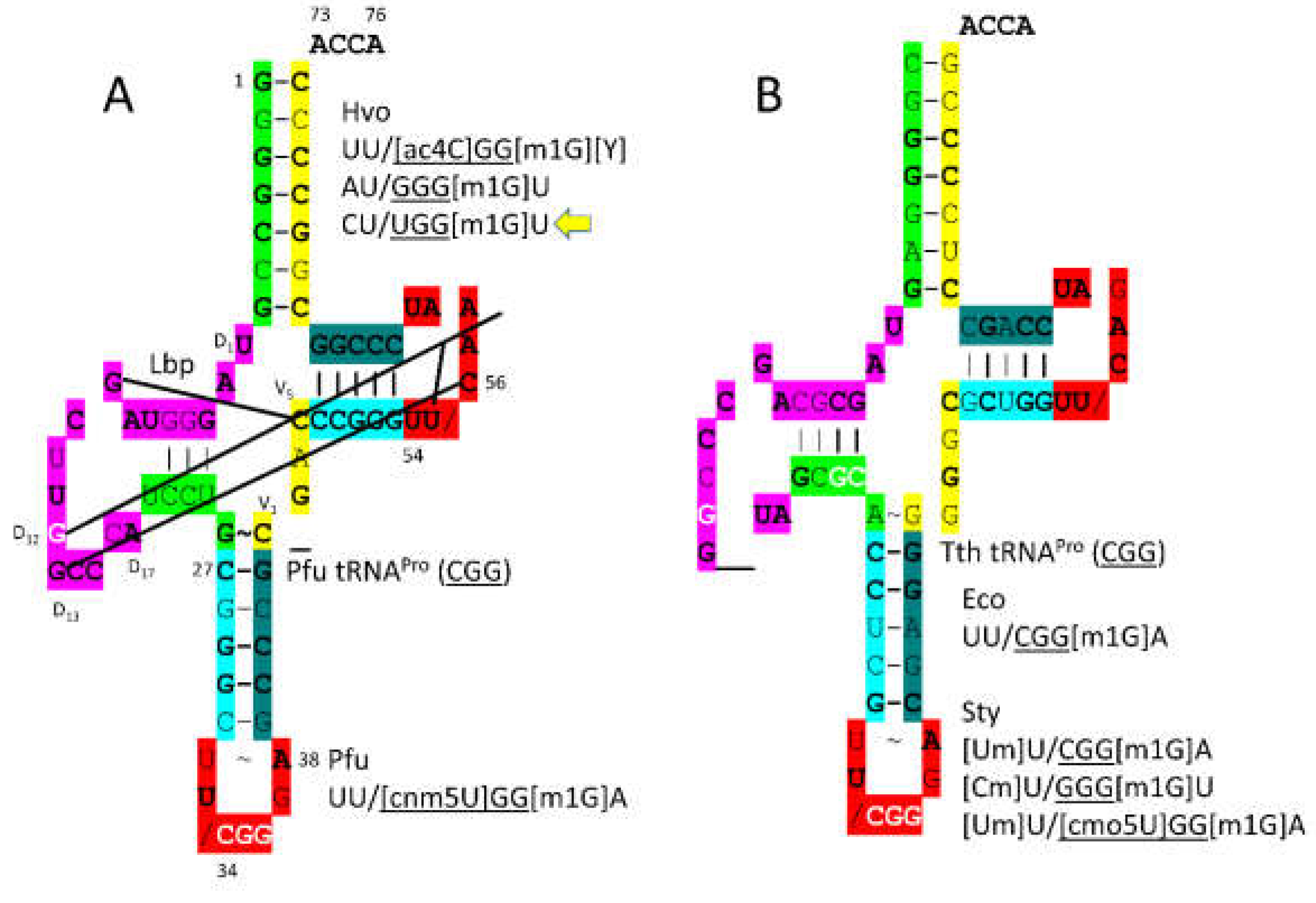
3.11. ProRS-IIA
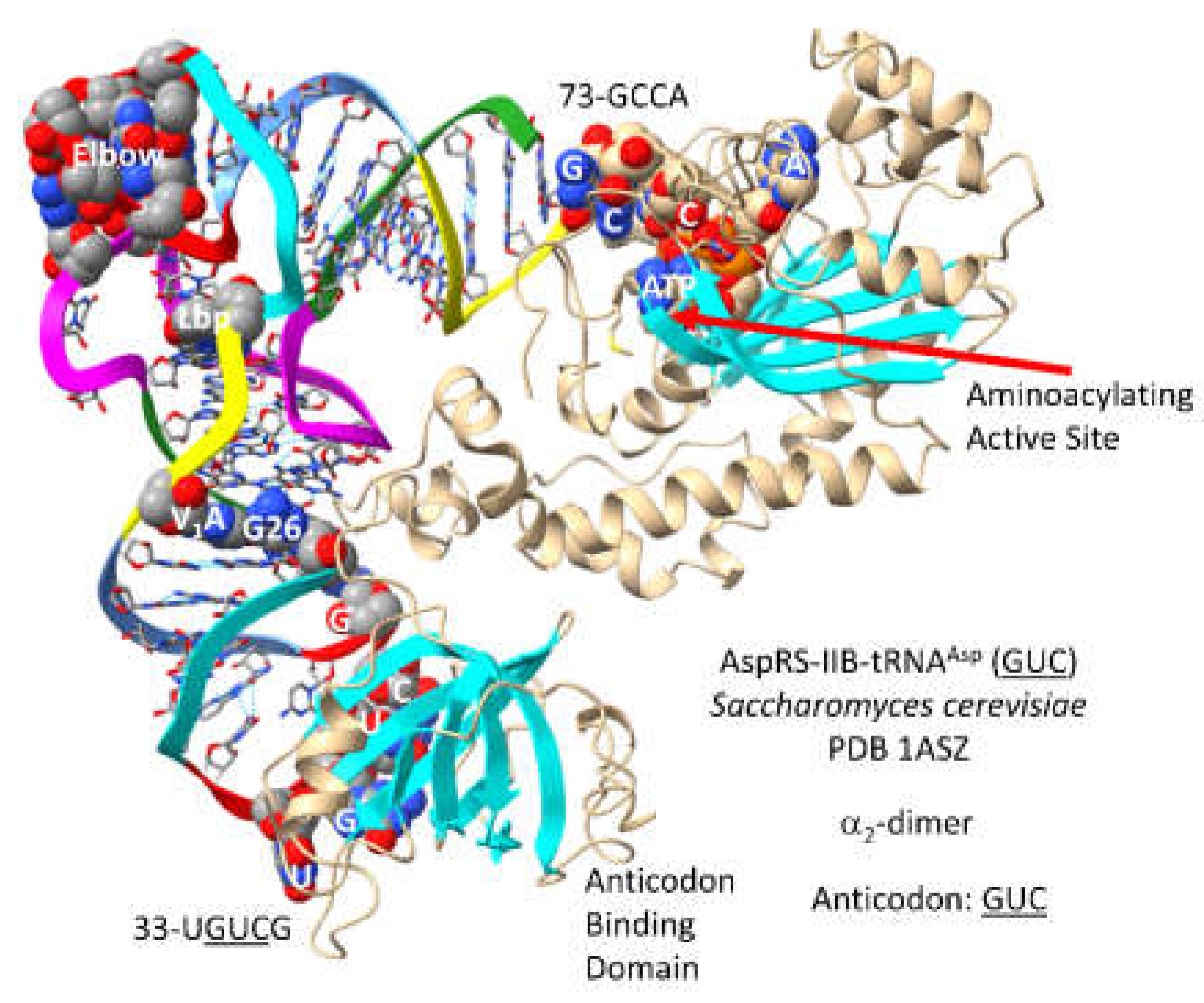
3.12. AspRS-IIB
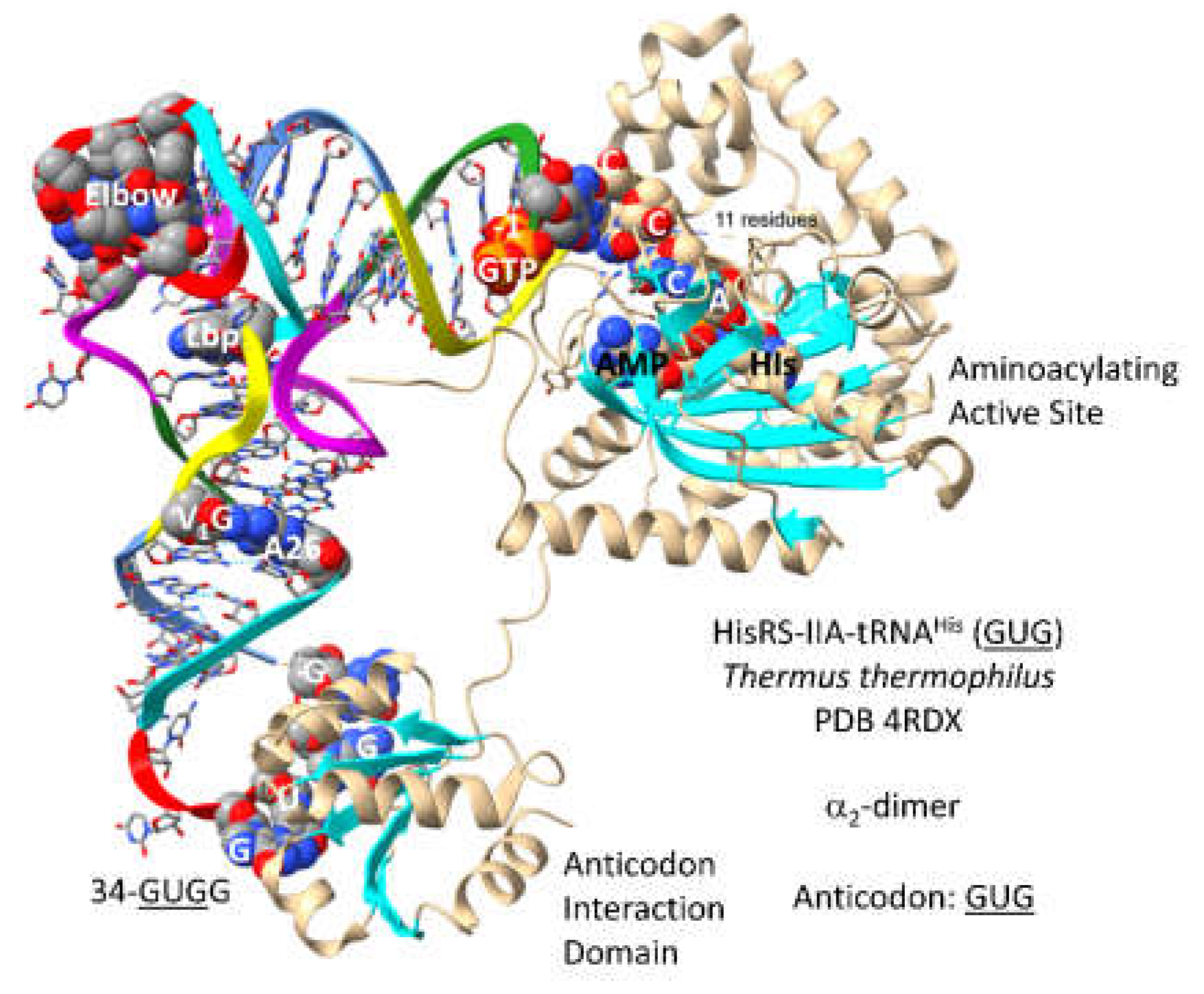
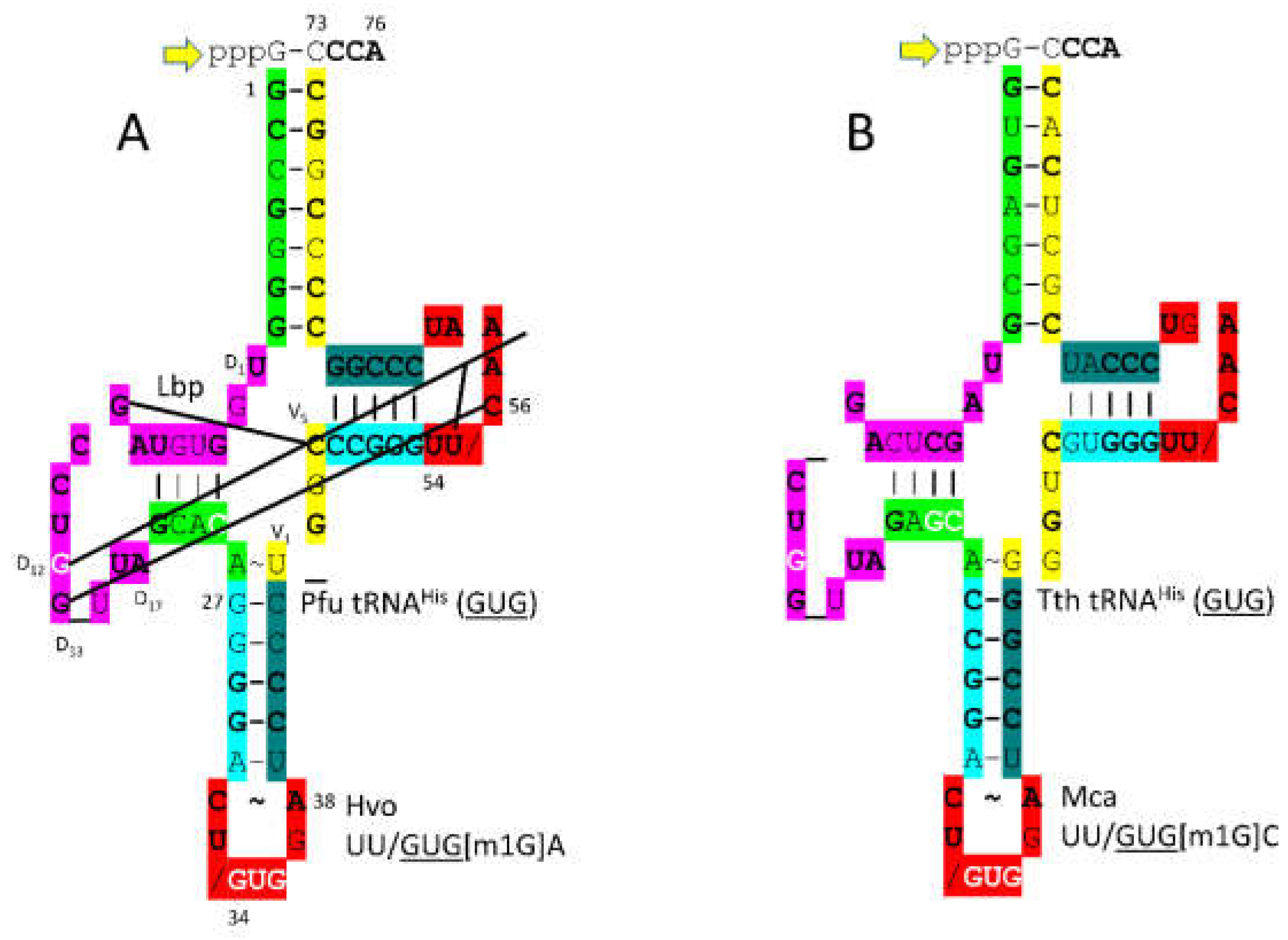
3.13. HisRS-IIA
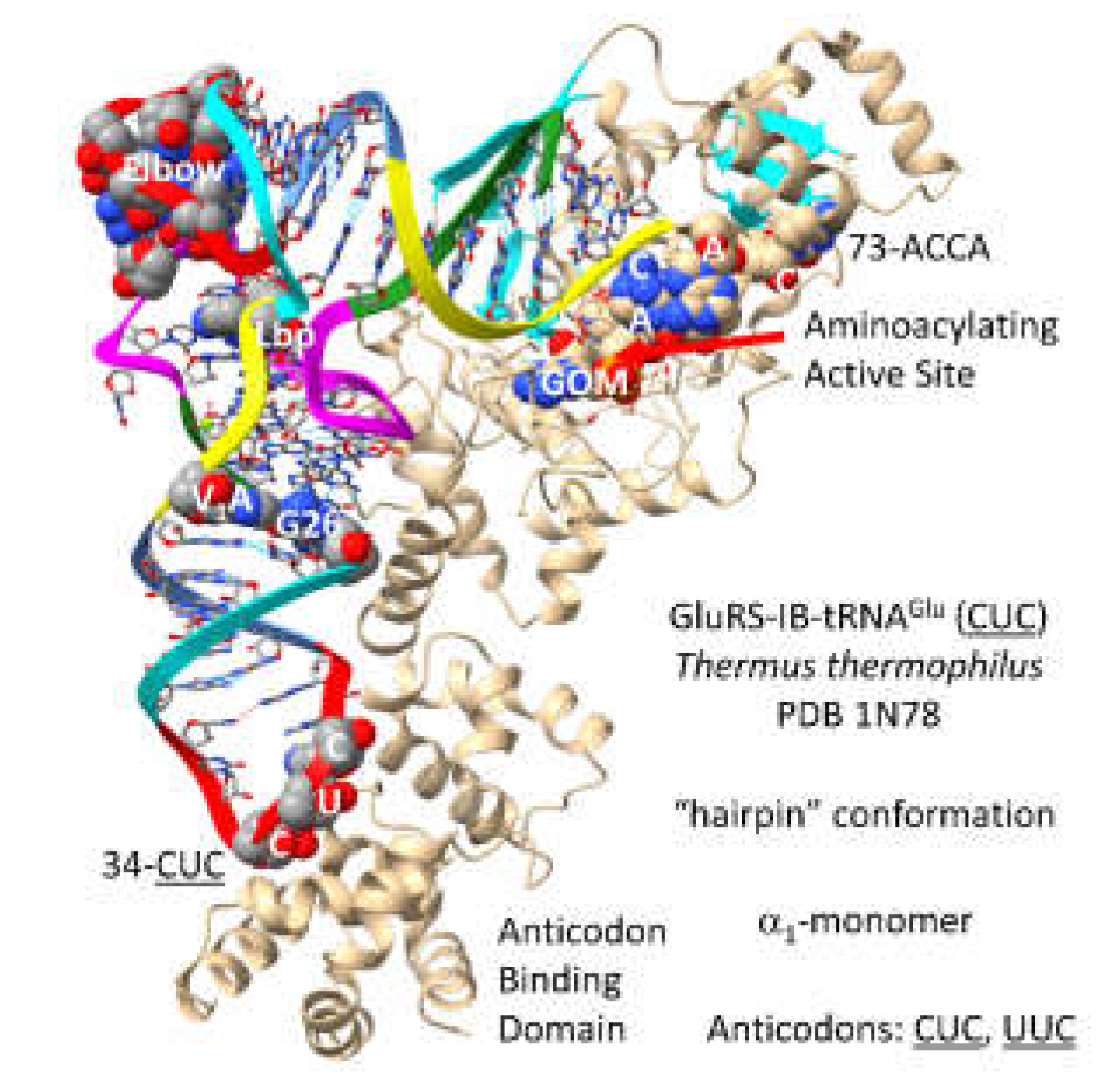
3.14. GluRS-IB
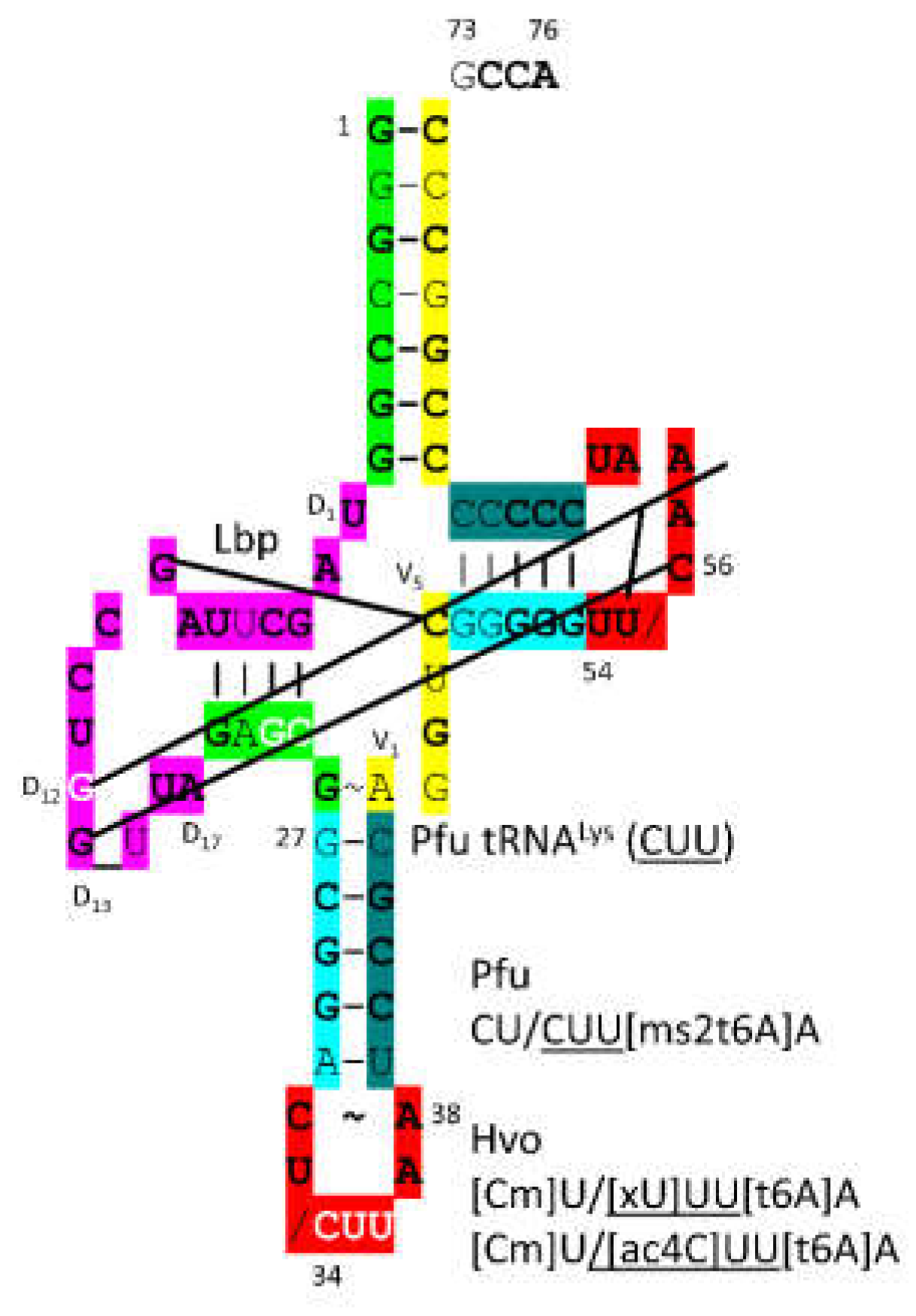
3.15. LysRS-IB
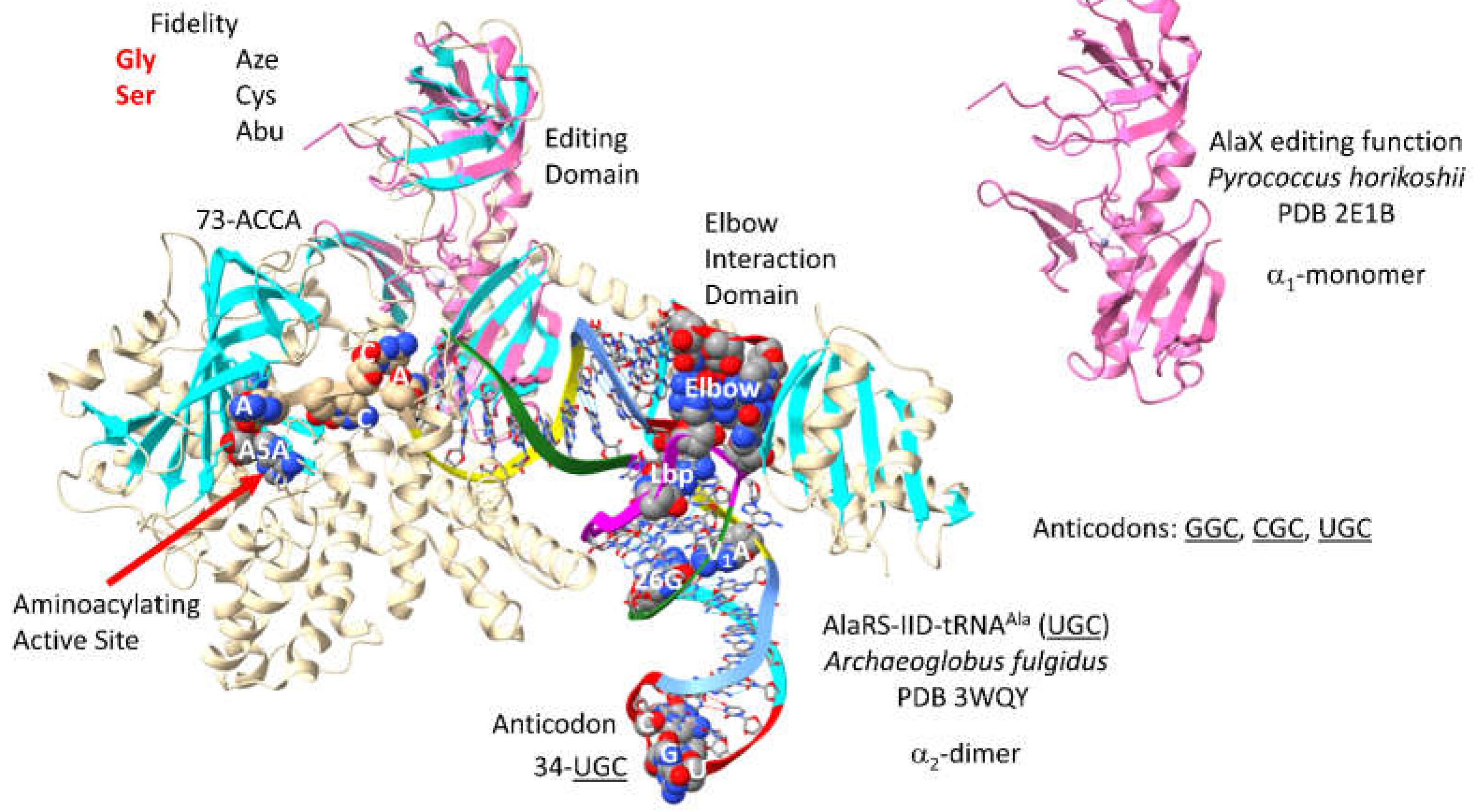
3.16. AlaRS-IID
3.17. PheRS-IIC
3.18. TyrRS-IC
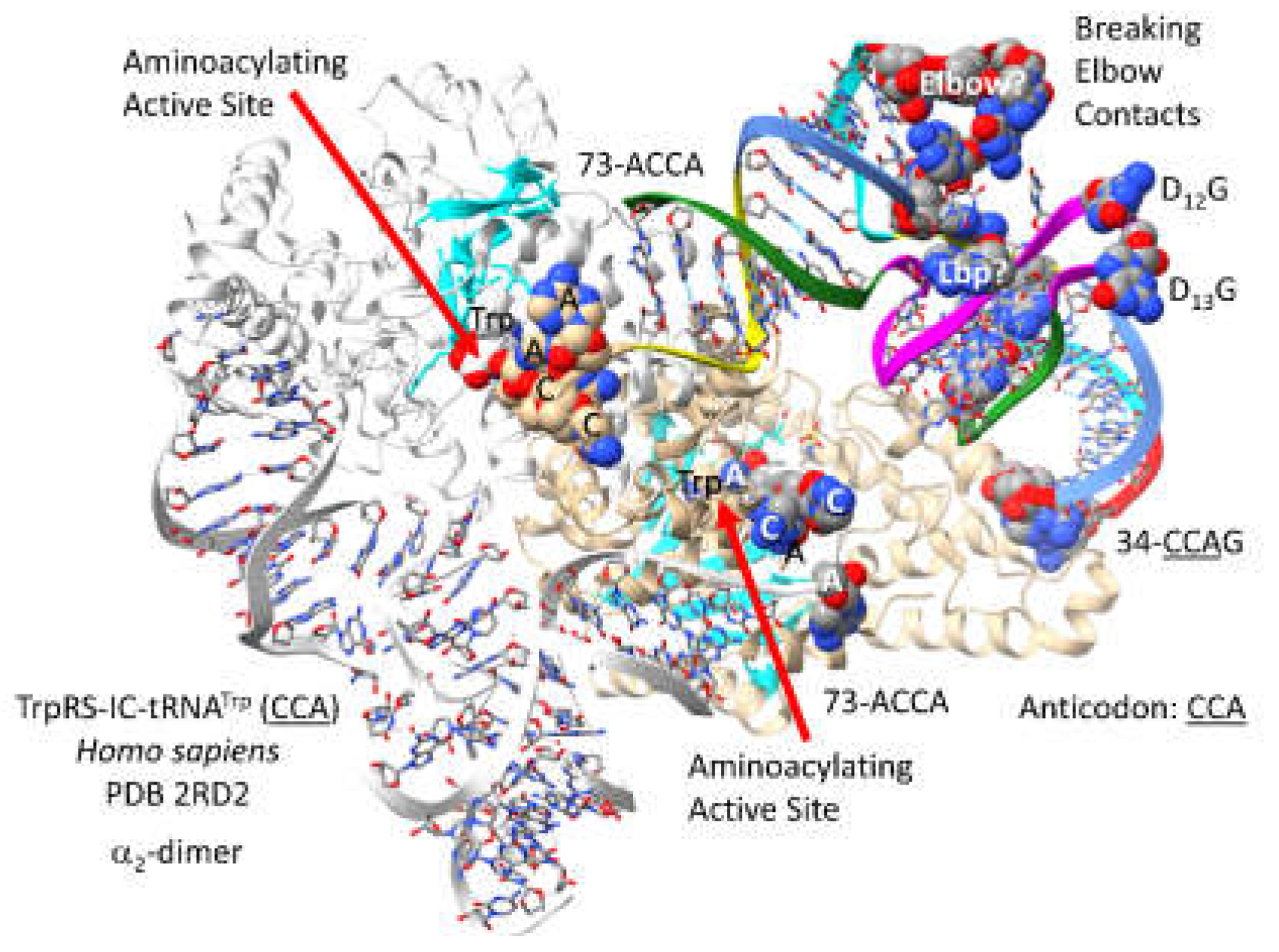
3.19. TrpRS-IC
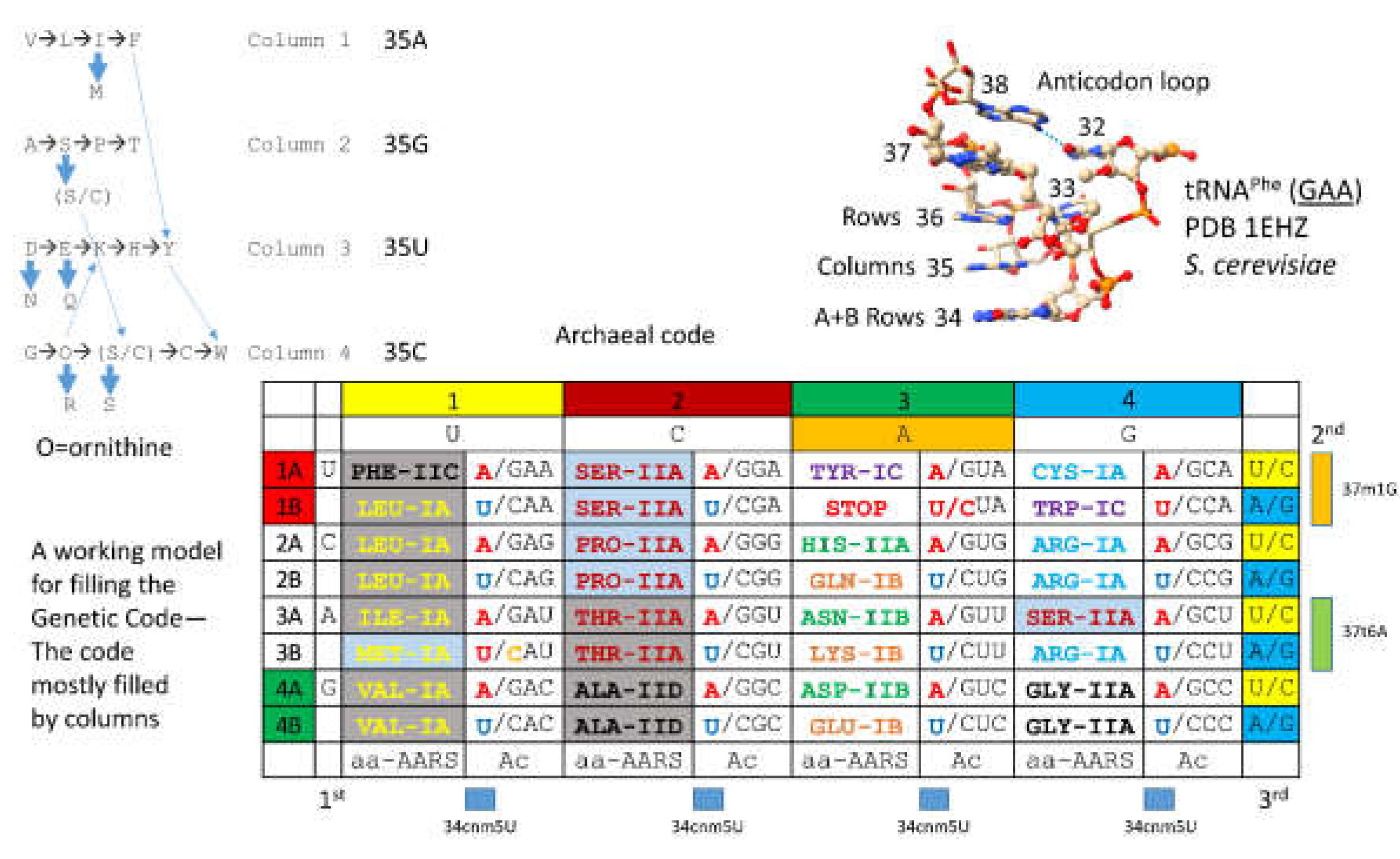
4. The Genetic Code
4.1. Column 1
4.2. Column 2
4.3. Column 3
4.4. Column 4
4.5. Disfavored Row 1
4.6. Stop Codons and Evolution of Translational Fidelity
5. Radiation of AARSomes
6. Evolution of Complex Life
7. Discussion
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AARS | Aminoacyl-tRNA synthetase |
| Aae | Aquifex aeolicus |
| Bta | Bos taurus |
| Eco | Escherichia coli |
| Hvo | Haloferax volcanii |
| Hsa | Homo sapiens |
| Lbp | Levitt base pair |
| Lla | Lactobacillus lactis |
| LUCA | Last Universal Common (cellular) Ancestor |
| Mca Pri |
Mycoplasma capricolum Primordial |
| Pfu | Pyrococcus furiosus |
| Sau | Staphylococcus aureus |
| Sgr | Streptomyces griseus |
| Tca | Thermoplasma acidophilum |
| Tth | Thermus thermophilus |
References
- Pavlinova, P.; Lambert, C.N.; Malaterre, C.; Nghe, P. Abiogenesis through gradual evolution of autocatalysis into template-based replication. FEBS Lett 2023, 597, 344–379. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Linderoth, J.; Baum, D.A. The hierarchical organization of autocatalytic reaction networks and its relevance to the origin of life. PLoS Comput Biol 2022, 18, e1010498. [Google Scholar] [CrossRef] [PubMed]
- Freeland, S. Undefining life's biochemistry: implications for abiogenesis. J R Soc Interface 2022, 19, 20210814. [Google Scholar] [CrossRef]
- Williamson, M.P. Autocatalytic Selection as a Driver for the Origin of Life. Life (Basel) 2024, 14. [Google Scholar] [CrossRef] [PubMed]
- Prosdocimi, F.; de Farias, S.T. Origin of life: Drawing the big picture. Prog Biophys Mol Biol 2023, 180-181, 28–36. [Google Scholar] [CrossRef]
- Farias, S.T.; Prosdocimi, F. RNP-world: The ultimate essence of life is a ribonucleoprotein process. Genet Mol Biol 2022, 45, e20220127. [Google Scholar] [CrossRef]
- de Farias, S.T.; Rego, T.G.; Jose, M.V. Origin of the 16S Ribosomal Molecule from Ancestor tRNAs. J Mol Evol 2021, 89, 249–256. [Google Scholar] [CrossRef]
- de Farias, S.T.; Jose, M.V. Transfer RNA: The molecular demiurge in the origin of biological systems. Prog Biophys Mol Biol 2020, 153, 28–34. [Google Scholar] [CrossRef]
- de Farias, S.T.; Rego, T.G.; Jose, M.V. tRNA Core Hypothesis for the Transition from the RNA World to the Ribonucleoprotein World. Life (Basel) 2016, 6. [Google Scholar] [CrossRef]
- de Farias, S.T.; do Rego, T.G.; Jose, M.V. Evolution of transfer RNA and the origin of the translation system. Front Genet 2014, 5, 303. [Google Scholar] [CrossRef]
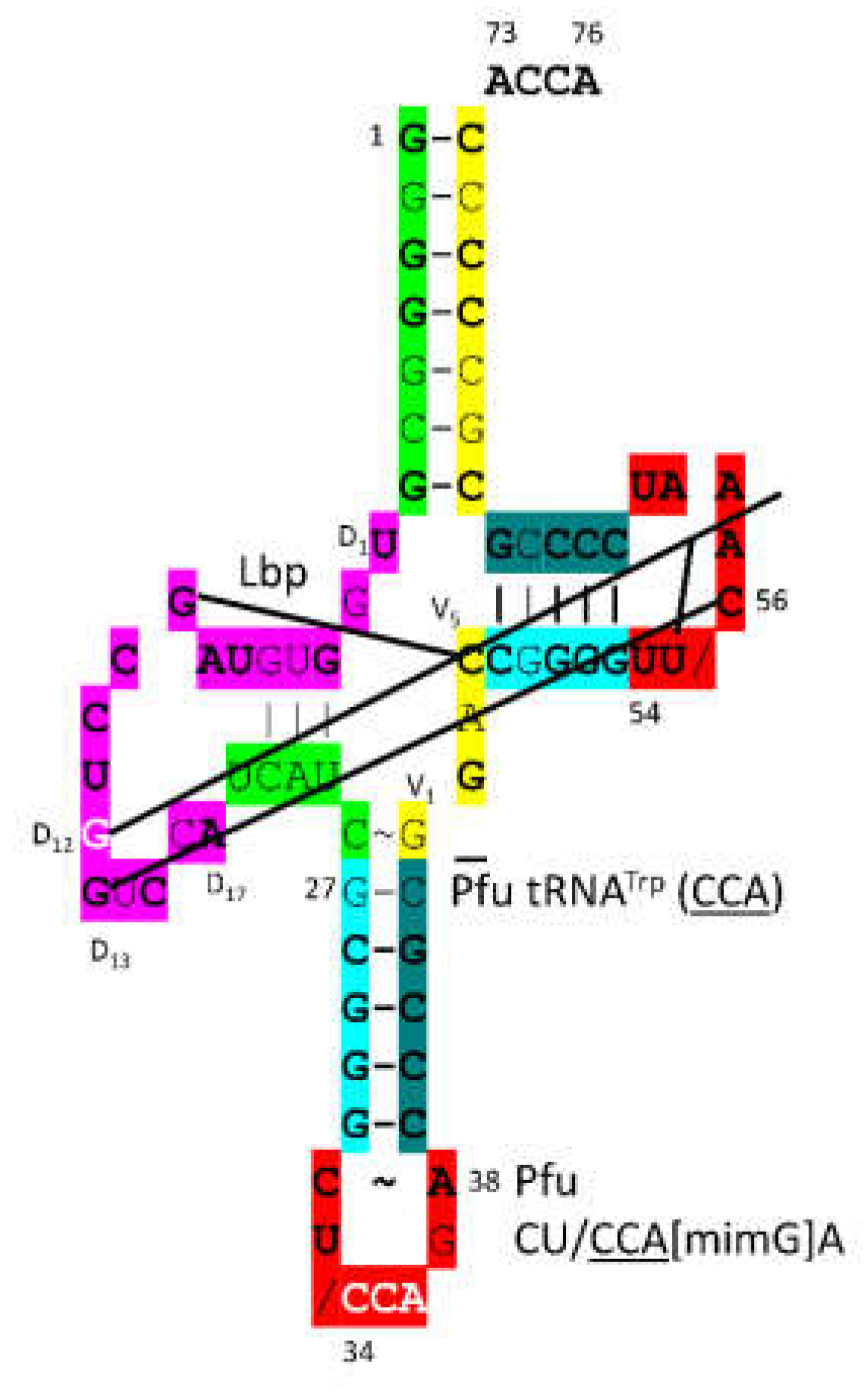
- Lei, L.; Burton, Z.F. Origin of Type II tRNA Variable Loops, Aminoacyl-tRNA Synthetase Allostery from Distal Determinants, and Diversification of Life. DNA 2024, 4, 252–275. [Google Scholar] [CrossRef]
- Li, R.; Macnamara, L.M.; Leuchter, J.D.; Alexander, R.W.; Cho, S.S. MD Simulations of tRNA and Aminoacyl-tRNA Synthetases: Dynamics, Folding, Binding, and Allostery. Int J Mol Sci 2015, 16, 15872–15902. [Google Scholar] [CrossRef]
- Han, Z.; Wang, X.; Wu, Z.; Li, C. Study of the Allosteric Mechanism of Human Mitochondrial Phenylalanyl-tRNA Synthetase by Transfer Entropy via an Improved Gaussian Network Model and Co-evolution Analyses. J Phys Chem Lett 2023, 14, 3452–3460. [Google Scholar] [CrossRef] [PubMed]
- Shao, Q.; Han, Z.; Cheng, J.; Wang, Q.; Gong, W.; Li, C. Allosteric Mechanism of Human Mitochondrial Phenylalanyl-tRNA Synthetase: An Atomistic MD Simulation and a Mutual Information-Based Network Study. J Phys Chem B 2021, 125, 7651–7661. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. Chemical Evolution of Life on Earth. Genes (Basel) 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. Evolution of the genetic code. Transcription 2021, 12, 28–53. [Google Scholar] [CrossRef]
- Lei, L.; Burton, Z.F. Evolution of Life on Earth: tRNA, Aminoacyl-tRNA Synthetases and the Genetic Code. Life (Basel) 2020, 10. [Google Scholar] [CrossRef]
- Kim, Y.; Opron, K.; Burton, Z.F. A tRNA- and Anticodon-Centric View of the Evolution of Aminoacyl-tRNA Synthetases, tRNAomes, and the Genetic Code. Life (Basel) 2019, 9. [Google Scholar] [CrossRef]
- Bernhardt, H.S.; Patrick, W.M. Genetic code evolution started with the incorporation of glycine, followed by other small hydrophilic amino acids. J Mol Evol 2014, 78, 307–309. [Google Scholar] [CrossRef]
- Bernhardt, H.S.; Tate, W.P. Evidence from glycine transfer RNA of a frozen accident at the dawn of the genetic code. Biol Direct 2008, 3, 53. [Google Scholar] [CrossRef]
- Pak, D.; Du, N.; Kim, Y.; Sun, Y.; Burton, Z.F. Rooted tRNAomes and evolution of the genetic code. Transcription 2018, 9, 137–151. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. The 3 31 Nucleotide Minihelix tRNA Evolution Theorem and the Origin of Life. Life (Basel) 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Rodriguez, L.; Erdogan, O.; Jimenez-Rodriguez, M.; Gonzalez-Rivera, K.; Williams, T.; Li, L.; Weinreb, V.; Collier, M.; Chandrasekaran, S.N.; Ambroggio, X.; et al. Functional Class I and II Amino Acid-activating Enzymes Can Be Coded by Opposite Strands of the Same Gene. J Biol Chem 2015, 290, 19710–19725. [Google Scholar] [CrossRef]
- Carter, C.W., Jr.; Li, L.; Weinreb, V.; Collier, M.; Gonzalez-Rivera, K.; Jimenez-Rodriguez, M.; Erdogan, O.; Kuhlman, B.; Ambroggio, X.; Williams, T.; et al. The Rodin-Ohno hypothesis that two enzyme superfamilies descended from one ancestral gene: an unlikely scenario for the origins of translation that will not be dismissed. Biol Direct 2014, 9, 11. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, S.N.; Yardimci, G.G.; Erdogan, O.; Roach, J.; Carter, C.W., Jr. Statistical evaluation of the Rodin-Ohno hypothesis: sense/antisense coding of ancestral class I and II aminoacyl-tRNA synthetases. Mol Biol Evol 2013, 30, 1588–1604. [Google Scholar] [CrossRef]
- Rodin, A.S.; Rodin, S.N.; Carter, C.W., Jr. On primordial sense-antisense coding. J Mol Evol 2009, 69, 555–567. [Google Scholar] [CrossRef]
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Couch, G.S.; Pearson, Z.J.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci 2023, 32, e4792. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci 2018, 27, 14–25. [Google Scholar] [CrossRef]
- Cappannini, A.; Ray, A.; Purta, E.; Mukherjee, S.; Boccaletto, P.; Moafinejad, S.N.; Lechner, A.; Barchet, C.; Klaholz, B.P.; Stefaniak, F.; et al. MODOMICS: a database of RNA modifications and related information. 2023 update. Nucleic Acids Res 2024, 52, D239–D244. [Google Scholar] [CrossRef]
- Wolff, P.; Villette, C.; Zumsteg, J.; Heintz, D.; Antoine, L.; Chane-Woon-Ming, B.; Droogmans, L.; Grosjean, H.; Westhof, E. Comparative patterns of modified nucleotides in individual tRNA species from a mesophilic and two thermophilic archaea. RNA 2020, 26, 1957–1975. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. A Recipe to Evolve Complex Life Chemically on Earth. Genes (Basel) 2025, 16. [Google Scholar] [CrossRef] [PubMed]
- Giege, R.; Eriani, G. The tRNA identity landscape for aminoacylation and beyond. Nucleic Acids Res 2023, 51, 1528–1570. [Google Scholar] [CrossRef] [PubMed]
- Tawfik, D.S.; Gruic-Sovulj, I. How evolution shapes enzyme selectivity - lessons from aminoacyl-tRNA synthetases and other amino acid utilizing enzymes. FEBS J 2020, 287, 1284–1305. [Google Scholar] [CrossRef]
- Giege, R.; Springer, M. Aminoacyl-tRNA Synthetases in the Bacterial World. EcoSal Plus 2016, 7. [Google Scholar] [CrossRef]
- Qin, X.; Deng, X.; Chen, L.; Xie, W. Crystal Structure of the Wild-Type Human GlyRS Bound with tRNA(Gly) in a Productive Conformation. J Mol Biol 2016, 428, 3603–3614. [Google Scholar] [CrossRef]
- Abe, T.; Inokuchi, H.; Yamada, Y.; Muto, A.; Iwasaki, Y.; Ikemura, T. tRNADB-CE: tRNA gene database well-timed in the era of big sequence data. Front Genet 2014, 5, 114. [Google Scholar] [CrossRef]
- Abe, T.; Ikemura, T.; Sugahara, J.; Kanai, A.; Ohara, Y.; Uehara, H.; Kinouchi, M.; Kanaya, S.; Yamada, Y.; Muto, A.; et al. tRNADB-CE 2011: tRNA gene database curated manually by experts. Nucleic Acids Res 2011, 39, D210–213. [Google Scholar] [CrossRef]
- Abe, T.; Ikemura, T.; Ohara, Y.; Uehara, H.; Kinouchi, M.; Kanaya, S.; Yamada, Y.; Muto, A.; Inokuchi, H. tRNADB-CE: tRNA gene database curated manually by experts. Nucleic Acids Res 2009, 37, D163–168. [Google Scholar] [CrossRef]
- Zhang, J.; Ferre-D'Amare, A.R. The tRNA Elbow in Structure, Recognition and Evolution. Life (Basel) 2016, 6. [Google Scholar] [CrossRef]
- Shi, H.; Moore, P.B. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: a classic structure revisited. RNA 2000, 6, 1091–1105. [Google Scholar] [CrossRef] [PubMed]
- Juhling, F.; Morl, M.; Hartmann, R.K.; Sprinzl, M.; Stadler, P.F.; Putz, J. tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res 2009, 37, D159–162. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Burton, Z.F. Superwobbling" and tRNA-34 Wobble and tRNA-37 Anticodon Loop Modifications in Evolution and Devolution of the Genetic Code. Life (Basel) 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Alkatib, S.; Scharff, L.B.; Rogalski, M.; Fleischmann, T.T.; Matthes, A.; Seeger, S.; Schottler, M.A.; Ruf, S.; Bock, R. The contributions of wobbling and superwobbling to the reading of the genetic code. PLoS Genet 2012, 8, e1003076. [Google Scholar] [CrossRef]
- Rogalski, M.; Karcher, D.; Bock, R. Superwobbling facilitates translation with reduced tRNA sets. Nat Struct Mol Biol 2008, 15, 192–198. [Google Scholar] [CrossRef]
- Pak, D.; Kim, Y.; Burton, Z.F. Aminoacyl-tRNA synthetase evolution and sectoring of the genetic code. Transcription 2018, 9, 205–224. [Google Scholar] [CrossRef]
- Muller, F.; Escobar, L.; Xu, F.; Wegrzyn, E.; Nainyte, M.; Amatov, T.; Chan, C.Y.; Pichler, A.; Carell, T. A prebiotically plausible scenario of an RNA-peptide world. Nature 2022, 605, 279–284. [Google Scholar] [CrossRef]
- Ikehara, K. Why Were [GADV]-amino Acids and GNC Codons Selected and How Was GNC Primeval Genetic Code Established? Genes (Basel) 2023, 14. [Google Scholar] [CrossRef]
- Ikehara, K. Evolutionary Steps in the Emergence of Life Deduced from the Bottom-Up Approach and GADV Hypothesis (Top-Down Approach). Life (Basel) 2016, 6. [Google Scholar] [CrossRef]
- Ikehara, K. [GADV]-protein world hypothesis on the origin of life. Orig Life Evol Biosph 2014, 44, 299–302. [Google Scholar] [CrossRef]
- Ikehara, K. Possible steps to the emergence of life: the [GADV]-protein world hypothesis. Chem Rec 2005, 5, 107–118. [Google Scholar] [CrossRef]
- Fukai, S.; Nureki, O.; Sekine, S.; Shimada, A.; Vassylyev, D.G.; Yokoyama, S. Mechanism of molecular interactions for tRNA(Val) recognition by valyl-tRNA synthetase. RNA 2003, 9, 100–111. [Google Scholar] [CrossRef] [PubMed]
- Sordyl, D.; Boileau, E.; Bernat, A.; Maiti, S.; Mukherjee, S.; Moafinejad, S.N.; Farsani, M.A.; Shavina, A.; Cappannini, A.; Agostini, G.; et al. MODOMICS: a database of RNA modifications and related information. 2025 update and 20th anniversary. Nucleic Acids Res 2025. [Google Scholar] [CrossRef] [PubMed]
- Silvian, L.F.; Wang, J.; Steitz, T.A. Insights into editing from an ile-tRNA synthetase structure with tRNAile and mupirocin. Science 1999, 285, 1074–1077. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, K.; Ogiso, Y.; Nakama, T.; Fukai, S.; Nureki, O. Structural basis for anticodon recognition by methionyl-tRNA synthetase. Nat Struct Mol Biol 2005, 12, 931–932. [Google Scholar] [CrossRef]
- Fukunaga, R.; Yokoyama, S. Aminoacylation complex structures of leucyl-tRNA synthetase and tRNALeu reveal two modes of discriminator-base recognition. Nat Struct Mol Biol 2005, 12, 915–922. [Google Scholar] [CrossRef]
- Fukunaga, R.; Yokoyama, S. Crystal structure of leucyl-tRNA synthetase from the archaeon Pyrococcus horikoshii reveals a novel editing domain orientation. J Mol Biol 2005, 346, 57–71. [Google Scholar] [CrossRef]
- Throll, P.; G.D., L.; Rico-Lastres, P.; Arnold, K.; Tengo, L.; Basu, S.; Kaiser, S.; Schneider, R.; Kowalinski, E. Structural basis of tRNA recognition by the m(3)C RNA methyltransferase METTL6 in complex with SerRS seryl-tRNA synthetase. Nat Struct Mol Biol 2024, 31, 1614–1624. [Google Scholar] [CrossRef]
- Delagoutte, B.; Keith, G.; Moras, D.; Cavarelli, J. Crystallization and preliminary X-ray crystallographic analysis of yeast arginyl-tRNA synthetase-yeast tRNAArg complexes. Acta Crystallogr D Biol Crystallogr 2000, 56, 492–494. [Google Scholar] [CrossRef]
- Longo, L.M.; Despotovic, D.; Weil-Ktorza, O.; Walker, M.J.; Jablonska, J.; Fridmann-Sirkis, Y.; Varani, G.; Metanis, N.; Tawfik, D.S. Primordial emergence of a nucleic acid-binding protein via phase separation and statistical ornithine-to-arginine conversion. Proc Natl Acad Sci U S A 2020, 117, 15731–15739. [Google Scholar] [CrossRef]
- Shi, W.; Yoshida, A.; Kosono, S.; Nishiyama, M. Evolution of lysine and arginine biosynthesis revealed by substrate specificity of lysine biosynthetic enzymes in Thermus thermophilus. FEBS J 2025. [Google Scholar] [CrossRef] [PubMed]
- Hashim, M.; Alam, I.; Ahmad, M.; Badruddeen; Akhtar, J.; Khan, M.I.; Islam, A.; Parveen, S. Comprehensive Review of L-Lysine: Chemistry, Occurrence, and Physiological Roles. Curr Protein Pept Sci 2025. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, J.; Wang, B.; Zhang, Y.; Li, H.; Liu, Y.; Yin, J.; He, D.; Luo, H.; Gan, F.; et al. Dissecting the Arginine and Lysine Biosynthetic Pathways and Their Relationship in Haloarchaeon Natrinema gari J7-2 via Endogenous CRISPR-Cas System-Based Genome Editing. Microbiol Spectr 2023, 11, e0028823. [Google Scholar] [CrossRef] [PubMed]
- Hauenstein, S.; Zhang, C.M.; Hou, Y.M.; Perona, J.J. Shape-selective RNA recognition by cysteinyl-tRNA synthetase. Nat Struct Mol Biol 2004, 11, 1134–1141. [Google Scholar] [CrossRef] [PubMed]
- Mukai, T.; Crnkovic, A.; Umehara, T.; Ivanova, N.N.; Kyrpides, N.C.; Soll, D. RNA-Dependent Cysteine Biosynthesis in Bacteria and Archaea. mBio 2017, 8. [Google Scholar] [CrossRef]
- Sankaranarayanan, R.; Dock-Bregeon, A.C.; Romby, P.; Caillet, J.; Springer, M.; Rees, B.; Ehresmann, C.; Ehresmann, B.; Moras, D. The structure of threonyl-tRNA synthetase-tRNA(Thr) complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell 1999, 97, 371–381. [Google Scholar] [CrossRef]
- Yaremchuk, A.; Cusack, S.; Tukalo, M. Crystal structure of a eukaryote/archaeon-like protyl-tRNA synthetase and its complex with tRNAPro(CGG). EMBO J 2000, 19, 4745–4758. [Google Scholar] [CrossRef]
- Cavarelli, J.; Eriani, G.; Rees, B.; Ruff, M.; Boeglin, M.; Mitschler, A.; Martin, F.; Gangloff, J.; Thierry, J.C.; Moras, D. The active site of yeast aspartyl-tRNA synthetase: structural and functional aspects of the aminoacylation reaction. EMBO J 1994, 13, 327–337. [Google Scholar] [CrossRef]
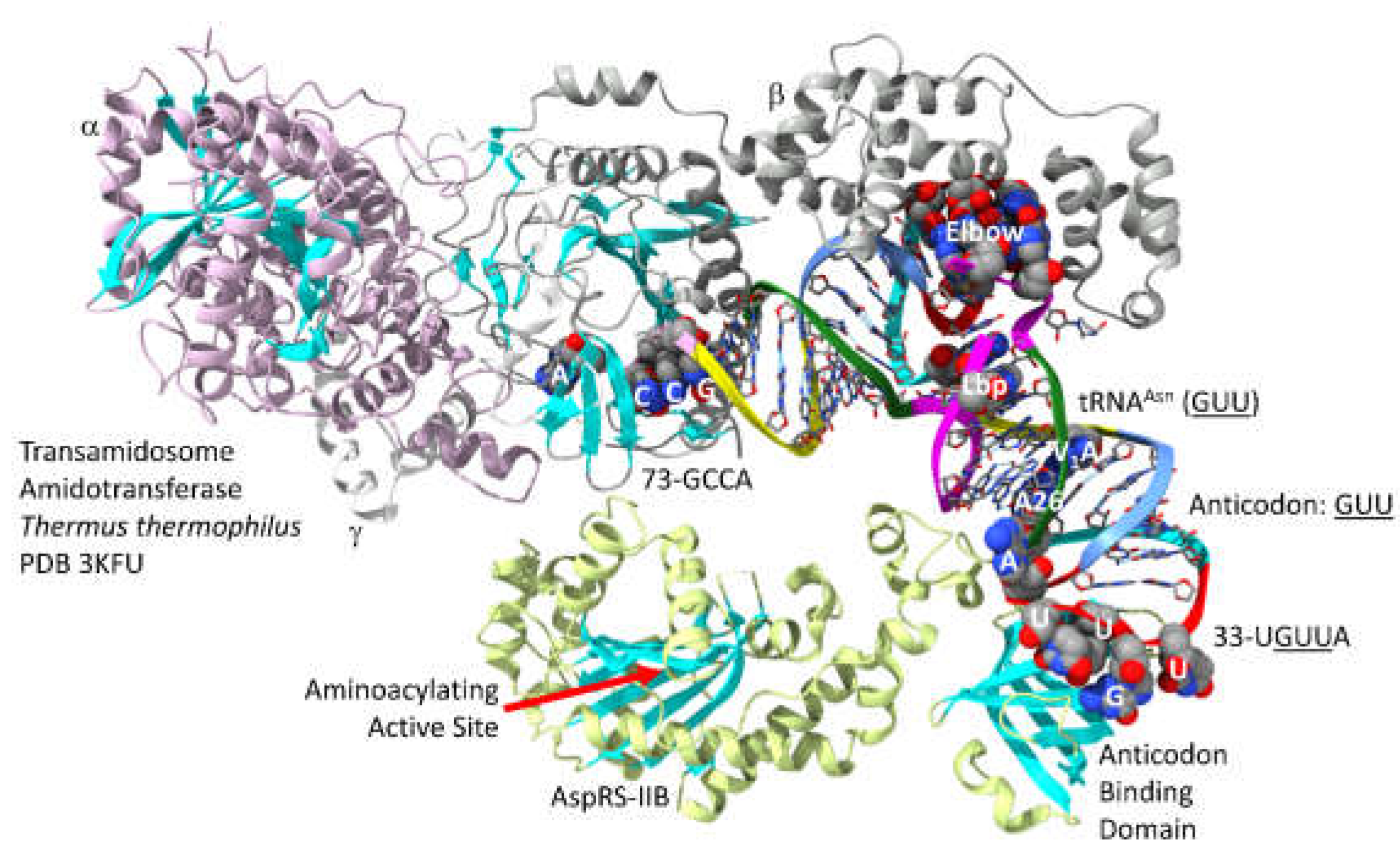
- Blaise, M.; Bailly, M.; Frechin, M.; Behrens, M.A.; Fischer, F.; Oliveira, C.L.; Becker, H.D.; Pedersen, J.S.; Thirup, S.; Kern, D. Crystal structure of a transfer-ribonucleoprotein particle that promotes asparagine formation. EMBO J 2010, 29, 3118–3129. [Google Scholar] [CrossRef]
- Rampias, T.; Sheppard, K.; Soll, D. The archaeal transamidosome for RNA-dependent glutamine biosynthesis. Nucleic Acids Res 2010, 38, 5774–5783. [Google Scholar] [CrossRef]
- Tian, Q.; Wang, C.; Liu, Y.; Xie, W. Structural basis for recognition of G-1-containing tRNA by histidyl-tRNA synthetase. Nucleic Acids Res 2015, 43, 2980–2990. [Google Scholar] [CrossRef] [PubMed]
- Di Giulio, M. The phylogenetic distribution of the glutaminyl-tRNA synthetase and Glu-tRNA(Gln) amidotransferase in the fundamental lineages would imply that the ancestor of archaea, that of eukaryotes and LUCA were progenotes. Biosystems 2020, 196, 104174. [Google Scholar] [CrossRef] [PubMed]
- Raczniak, G.; Becker, H.D.; Min, B.; Soll, D. A single amidotransferase forms asparaginyl-tRNA and glutaminyl-tRNA in Chlamydia trachomatis. J Biol Chem 2001, 276, 45862–45867. [Google Scholar] [CrossRef] [PubMed]
- Salazar, J.C.; Zuniga, R.; Raczniak, G.; Becker, H.; Soll, D.; Orellana, O. A dual-specific Glu-tRNA(Gln) and Asp-tRNA(Asn) amidotransferase is involved in decoding glutamine and asparagine codons in Acidithiobacillus ferrooxidans. FEBS Lett 2001, 500, 129–131. [Google Scholar] [CrossRef]
- Sekine, S.; Nureki, O.; Dubois, D.Y.; Bernier, S.; Chenevert, R.; Lapointe, J.; Vassylyev, D.G.; Yokoyama, S. ATP binding by glutamyl-tRNA synthetase is switched to the productive mode by tRNA binding. EMBO J 2003, 22, 676–688. [Google Scholar] [CrossRef]
- Naganuma, M.; Sekine, S.; Chong, Y.E.; Guo, M.; Yang, X.L.; Gamper, H.; Hou, Y.M.; Schimmel, P.; Yokoyama, S. The selective tRNA aminoacylation mechanism based on a single G*U pair. Nature 2014, 510, 507–511. [Google Scholar] [CrossRef]
- Fukunaga, R.; Yokoyama, S. Structure of the AlaX-M trans-editing enzyme from Pyrococcus horikoshii. Acta Crystallogr D Biol Crystallogr 2007, 63, 390–400. [Google Scholar] [CrossRef]
- Fournier, G.P.; Alm, E.J. Ancestral Reconstruction of a Pre-LUCA Aminoacyl-tRNA Synthetase Ancestor Supports the Late Addition of Trp to the Genetic Code. J Mol Evol 2015, 80, 171–185. [Google Scholar] [CrossRef]
- Moor, N.; Kotik-Kogan, O.; Tworowski, D.; Sukhanova, M.; Safro, M. The crystal structure of the ternary complex of phenylalanyl-tRNA synthetase with tRNAPhe and a phenylalanyl-adenylate analogue reveals a conformational switch of the CCA end. Biochemistry 2006, 45, 10572–10583. [Google Scholar] [CrossRef]
- Goldgur, Y.; Mosyak, L.; Reshetnikova, L.; Ankilova, V.; Lavrik, O.; Khodyreva, S.; Safro, M. The crystal structure of phenylalanyl-tRNA synthetase from thermus thermophilus complexed with cognate tRNAPhe. Structure 1997, 5, 59–68. [Google Scholar] [CrossRef]
- Kobayashi, T.; Nureki, O.; Ishitani, R.; Yaremchuk, A.; Tukalo, M.; Cusack, S.; Sakamoto, K.; Yokoyama, S. Structural basis for orthogonal tRNA specificities of tyrosyl-tRNA synthetases for genetic code expansion. Nat Struct Biol 2003, 10, 425–432. [Google Scholar] [CrossRef]
- Shen, N.; Guo, L.; Yang, B.; Jin, Y.; Ding, J. Structure of human tryptophanyl-tRNA synthetase in complex with tRNATrp reveals the molecular basis of tRNA recognition and specificity. Nucleic Acids Res 2006, 34, 3246–3258. [Google Scholar] [CrossRef]
- Wehbi, S.; Wheeler, A.; Morel, B.; Manepalli, N.; Minh, B.Q.; Lauretta, D.S.; Masel, J. Order of amino acid recruitment into the genetic code resolved by last universal common ancestor's protein domains. Proc Natl Acad Sci U S A 2024, 121, e2410311121. [Google Scholar] [CrossRef]
- Sun, F.J.; Caetano-Anolles, G. Transfer RNA and the origins of diversified life. Sci Prog 2008, 91, 265–284. [Google Scholar] [CrossRef] [PubMed]
- Hauenstein, S.I.; Perona, J.J. Redundant synthesis of cysteinyl-tRNACys in Methanosarcina mazei. J Biol Chem 2008, 283, 22007–22017. [Google Scholar] [CrossRef] [PubMed]
- Tumbula-Hansen, D.; Feng, L.; Toogood, H.; Stetter, K.O.; Soll, D. Evolutionary divergence of the archaeal aspartyl-tRNA synthetases into discriminating and nondiscriminating forms. J Biol Chem 2002, 277, 37184–37190. [Google Scholar] [CrossRef]
- Feng, L.; Stathopoulos, C.; Ahel, I.; Mitra, A.; Tumbula-Hansen, D.; Hartsch, T.; Soll, D. Aminoacyl-tRNA formation in the extreme thermophile Thermus thermophilus. Extremophiles 2002, 6, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Ikehara, K. Pseudo-replication of [GADV]-proteins and origin of life. Int J Mol Sci 2009, 10, 1525–1537. [Google Scholar] [CrossRef]
- Oba, T.; Fukushima, J.; Maruyama, M.; Iwamoto, R.; Ikehara, K. Catalytic activities of [GADV]-peptides. Formation and establishment of [GADV]-protein world for the emergence of life. Orig Life Evol Biosph 2005, 35, 447–460. [Google Scholar] [CrossRef]
- Liras, P.; Martin, J.F. Interconnected Set of Enzymes Provide Lysine Biosynthetic Intermediates and Ornithine Derivatives as Key Precursors for the Biosynthesis of Bioactive Secondary Metabolites. Antibiotics (Basel) 2023, 12. [Google Scholar] [CrossRef]
- Fazius, F.; Zaehle, C.; Brock, M. Lysine biosynthesis in microbes: relevance as drug target and prospects for beta-lactam antibiotics production. Appl Microbiol Biotechnol 2013, 97, 3763–3772. [Google Scholar] [CrossRef]
- Ouchi, T.; Tomita, T.; Horie, A.; Yoshida, A.; Takahashi, K.; Nishida, H.; Lassak, K.; Taka, H.; Mineki, R.; Fujimura, T.; et al. Lysine and arginine biosyntheses mediated by a common carrier protein in Sulfolobus. Nat Chem Biol 2013, 9, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Nishida, H.; Nishiyama, M. Evolution of lysine biosynthesis in the phylum deinococcus-thermus. Int J Evol Biol 2012, 2012, 745931. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, J.; Kobashi, N.; Nishiyama, M.; Yamane, H. Functional and evolutionary relationship between arginine biosynthesis and prokaryotic lysine biosynthesis through alpha-aminoadipate. J Bacteriol 2001, 183, 5067–5073. [Google Scholar] [CrossRef]
- Kosuge, T.; Hoshino, T. Lysine is synthesized through the alpha-aminoadipate pathway in Thermus thermophilus. FEMS Microbiol Lett 1998, 169, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Burroughs, A.M.; Aravind, L. The Origin and Evolution of Release Factors: Implications for Translation Termination, Ribosome Rescue, and Quality Control Pathways. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 2015, 10, 845–858. [Google Scholar] [CrossRef]
- Burton, Z.F. The 3-Minihelix tRNA Evolution Theorem. J Mol Evol 2020, 88, 234–242. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



