1. Introduction
To evolve complex life requires a genetic code, which requires a genetic adapter. Without a code supported by an adapter molecule, the potential to evolve enduring and replicated complexity based on pre-life metabolic systems remained limited [
1,
2,
3,
4]. Life on Earth evolved around tRNA, tRNAomes, AARSomes, first proteins, ribosomes and the genetic code [
5,
6,
7,
8,
9,
10]. The purpose of this review is to concentrate on early tRNAome and AARSome networks to describe evolution of the first code on Earth.
For evolution of the code, tRNAs must diversify to tRNAomes. Most tRNAs are type I, initially, with a 5 nt V loop (V for variable). In Archaea, longer type II V arms (initially 14 nt) are utilized by tRNA
Leu (5 tRNA
Leu) and tRNA
Ser (4 tRNA
Ser). The type I V loop was processed from the primitive type II V arm by a 9 nt internal deletion [
11]. Leucine and serine occupy 6-codon sectors of the code, so their longer V arms were used in place of the anticodon stem-loop-stem as a major determinant for cognate AARS recognition. Arginine is also found in a 6-codon sector of the code (5 tRNA
Arg). Arginine utilizes significant anticodon loop unwinding to expose additional bases for recognition. It is not likely that the strategy utilized for arginine could support three amino acids in 6-codon boxes. Anticodon loop unwinding indicates allosteric effects of cognate tRNA-AARS binding [
11,
12,
13,
14]. Complex life on Earth evolved around tRNA, tRNAomes and AARSomes.
AARSomes diverged from class II to class I enzymes [
15,
16,
17,
18]. GlyRS-IIA (class II; subclass A) appears to be closest to the founding AARS. It follows that glycine was the founding amino acid in the code [
19,
20]. All class II enzymes were derived from GlyRS-IIA as the root sequence. A primitive ValRS-IA (class I; subclass A) was derived from GlyRS-IIA by appending an N-terminal extension, which redirected to the class I AARS fold. Early folds of class II and class I AARS were directed by Zn binding. All class I AARS appear derived from a primordial ValRS-IA as the root enzyme. AARS enzymes are analyzed for: 1) tRNA contacts; 2) tRNA deformation (allostery); 3) modifications of the anticodon loop; 4) amino acid identity (chemical features); and 5) fidelity (i.e., editing). These characteristics appear to be most central to establishment of the first code. At early stages, code innovation was more important than fidelity. At late stages, fidelity mechanisms froze the code.
We utilize the ancient Archaeon
Pyrococcus furiosus as a reference species that may be close to the last universal common (cellular) ancestor (LUCA) for translation functions [
21]. The
P. furiosus tRNAome is tightly clustered around the primordial tRNA sequence. Similarly, the AARSome appears to be diverged in an orderly manner from the primitive GlyRS-IIA root sequence. Of course, tRNAomes and AARSomes must diverge from root sequences to maintain cognate translational discrimination and accuracy.
2. Materials and Methods
P. furiosus was the reference species chosen to be similar to LUCA for translation functions [
21]. In the future, selecting a more advantageous reference species that is closer to LUCA may be possible. The idea behind the choice of
P. furiosus was to anchor to a system lacking huge divergence from the first code. The root sequence for tRNA evolution has been determined and essentially matches a typical tRNA from
P. furiosus [
22]. Defining or estimating root sequences is fundamental to understand early evolution and the pre-life to life transition. It appears that a primitive GlyRS-IIA diversified to all class II AARS. A primitive GlyRS-IIA apparently diverged to a primitive ValRS-IA by attachment of an N-terminal segment that redirected the protein fold [
15]. All class I AARS appear to diverge from a primitive ValRS-IA.
Sequence similarity of class II and class I AARS has been demonstrated. For instance, the sequence similarity of
Methanobacterium bryantii IleRS-IA (a class I AARS) and
Methanococcoides burtonii GlyRS-IIA (a class II AARS) was determined with an e-value of 4x10
-12 for a substantial in-phase alignment [
15,
18]. The e-value represents about 1 chance in 2.5x10
11 of the alignment resulting from a random occurrence. Many more examples of class II versus class I AARS homology can readily be obtained. These data are inconsistent with some other published models for class II and class I AARS evolution [
23,
24,
25,
26].
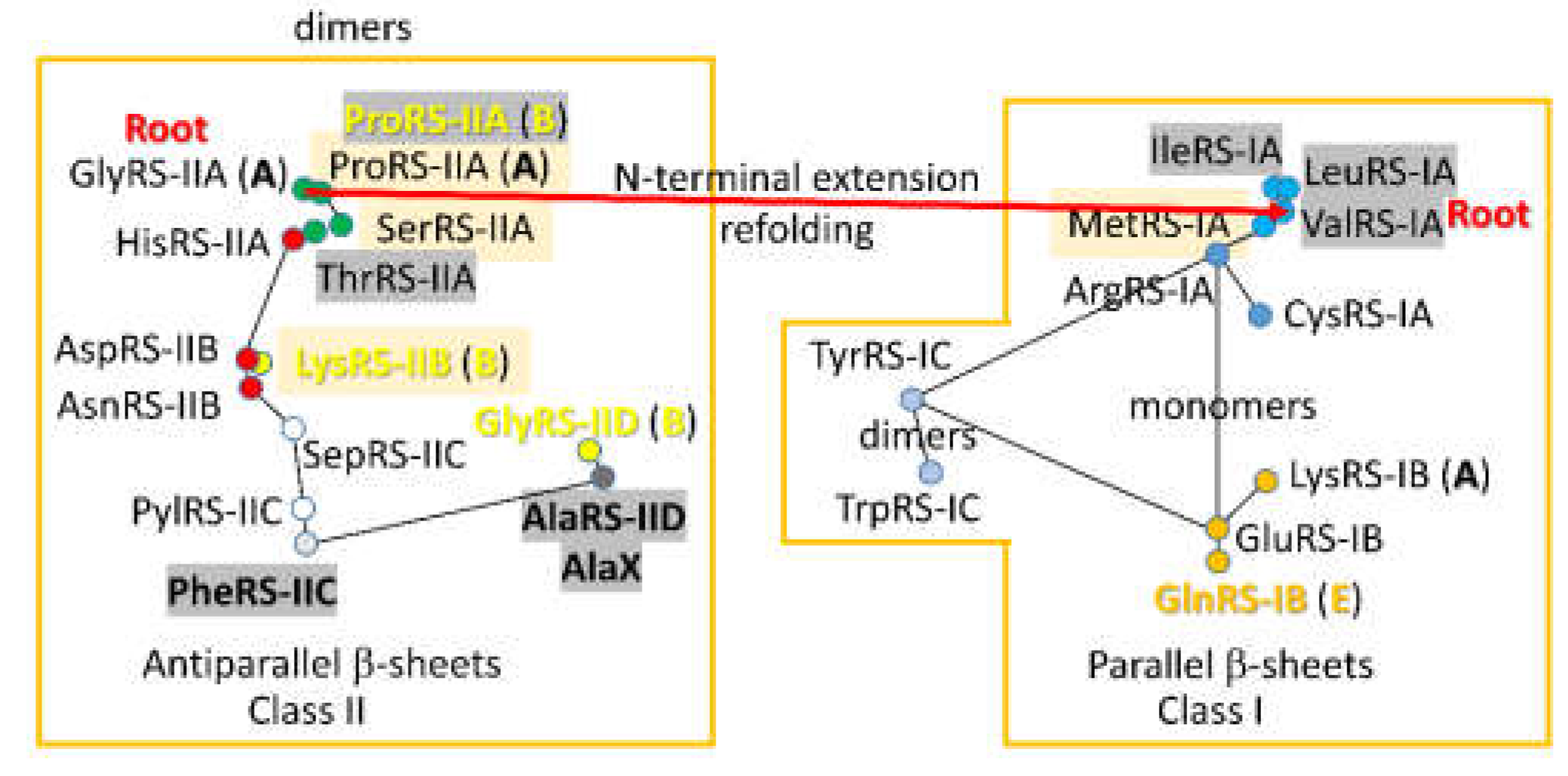
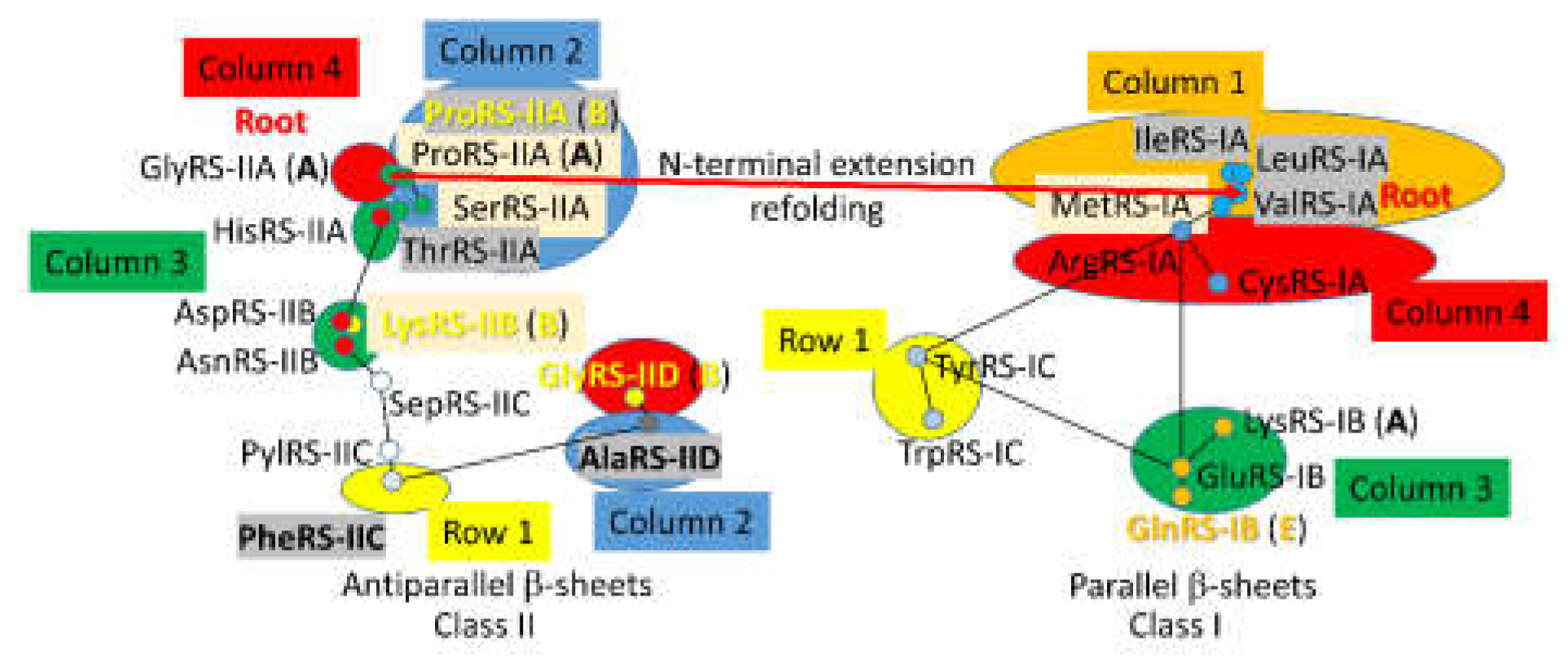
A 2-dimensional network for
P. furiosus AARS enzymes was previously published [
18]. Phyre 2 scoring of structural and sequence similarity was used to draw the maps for class II and class I AARS separately. Because high homology scores in Phyre 2 were assigned to closely related enzymes, reciprocal scores were used to draw the map. At the time the map was constructed, there was no objective mechanism to incorporate the sequence similarity of GlyRS-IIA, IleRS-IA and ValRS-IA.
Fidelity assays were generally done in a small number of reference organisms, so results may not universally apply. This caution extends to structural studies also.
ChimeraX was used to generate molecular graphics [
27,
28,
29]. AARS structures were selected that were close in structure to the first enzymes.
The Modomics database was used to identify anticodon loop modifications [
30].
P. furiosus modification data was obtained from reference [
31].
tRNA structures were drawn and colored according to internal homologies, based on the three 31 nt minihelix tRNA evolution theorem, as previously described [
15,
22,
32]. Historical numbering of tRNAs can be confusing, particularly within the D loop and V loop. Here, we number the D loop D
1 to D
17 and the V loop V
1 to V
n (V loop of n bases; initially, V
1→V
5 for type I, V
1→V
14 for type II; V
1→V
5 for type I align to V
1→V
5 for type II).
3. AARS Enzymes at the Base of Code Evolution
3.1. The AARS Mechanism
The AARS enzyme reaction is complex [
33,
34,
35]. Within the aminoacylating active site of the AARS, the amino acid carboxy terminus reacts with ATP to form an AMP adduct (aa-C=O, -O–AMP) releasing pyrophosphate. The tRNA 73-NCCA (N is the discriminator base) end displaces AMP to bind the aa-C=O, -O-tRNA, releasing AMP. Class II AARS attach the 76A ribose 3’-O to the cognate amino acid. Class I AARS attach the 76A ribose 2’-O to the cognate amino acid. ATP, the cognate amino acid and the cognate tRNA are substrates. Because the reaction progresses in two steps, the order of substrate additions may be important for AARS enzymes, affecting what reaction intermediate analogues or AARS-tRNA conformations can be visualized in crystal or cryo-electron microscopy structures. Pyrophosphate, AMP and aa-tRNA are products. In structures, non-reactive aa-AMP analogues were sometimes used to mimic a reaction intermediate.
3.2. GlyRS-IIA
A primordial GlyRS-IIA appears to be the founding AARS. All class II and class I AARS enzymes appear to be derived from this root.
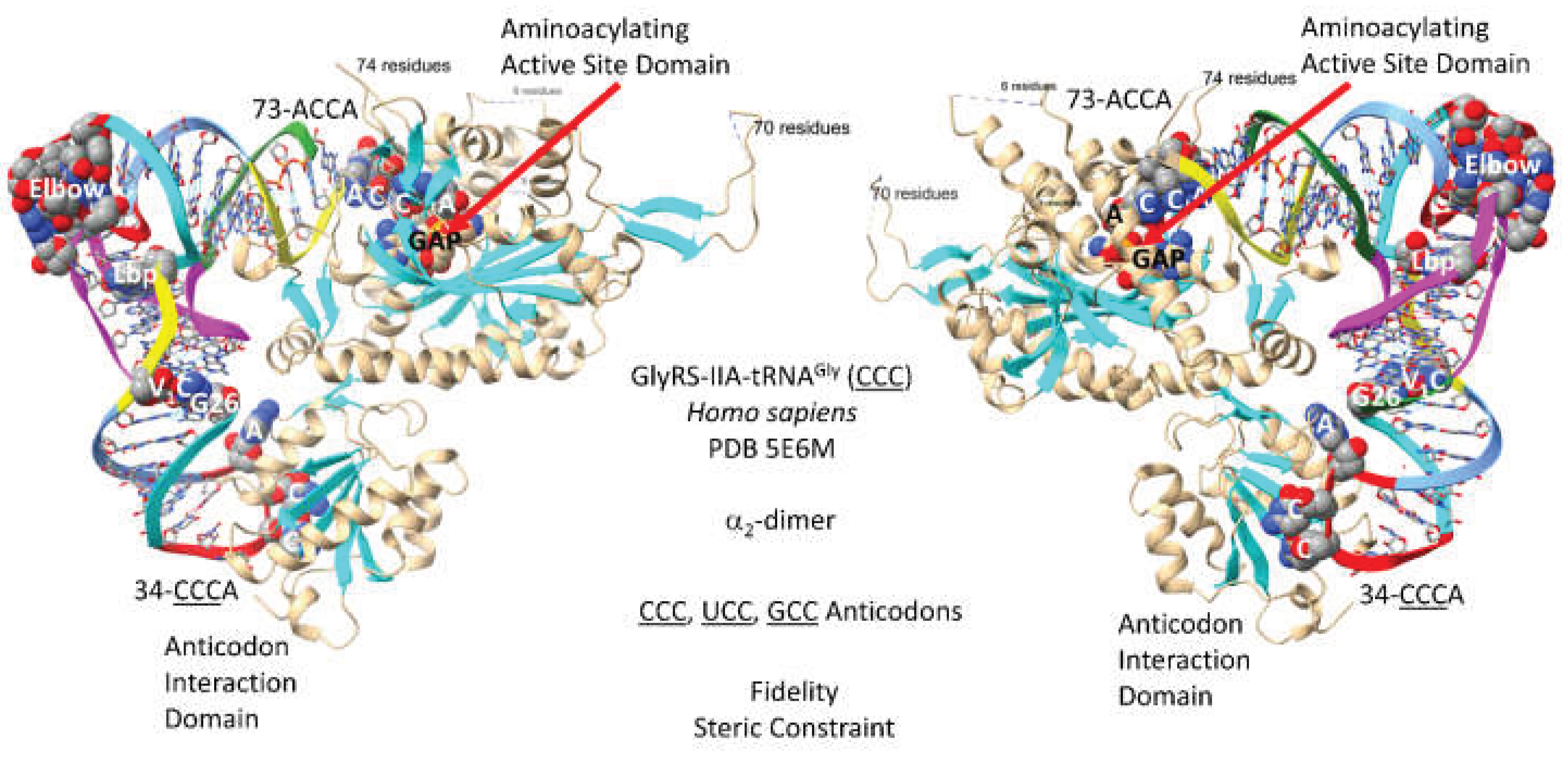
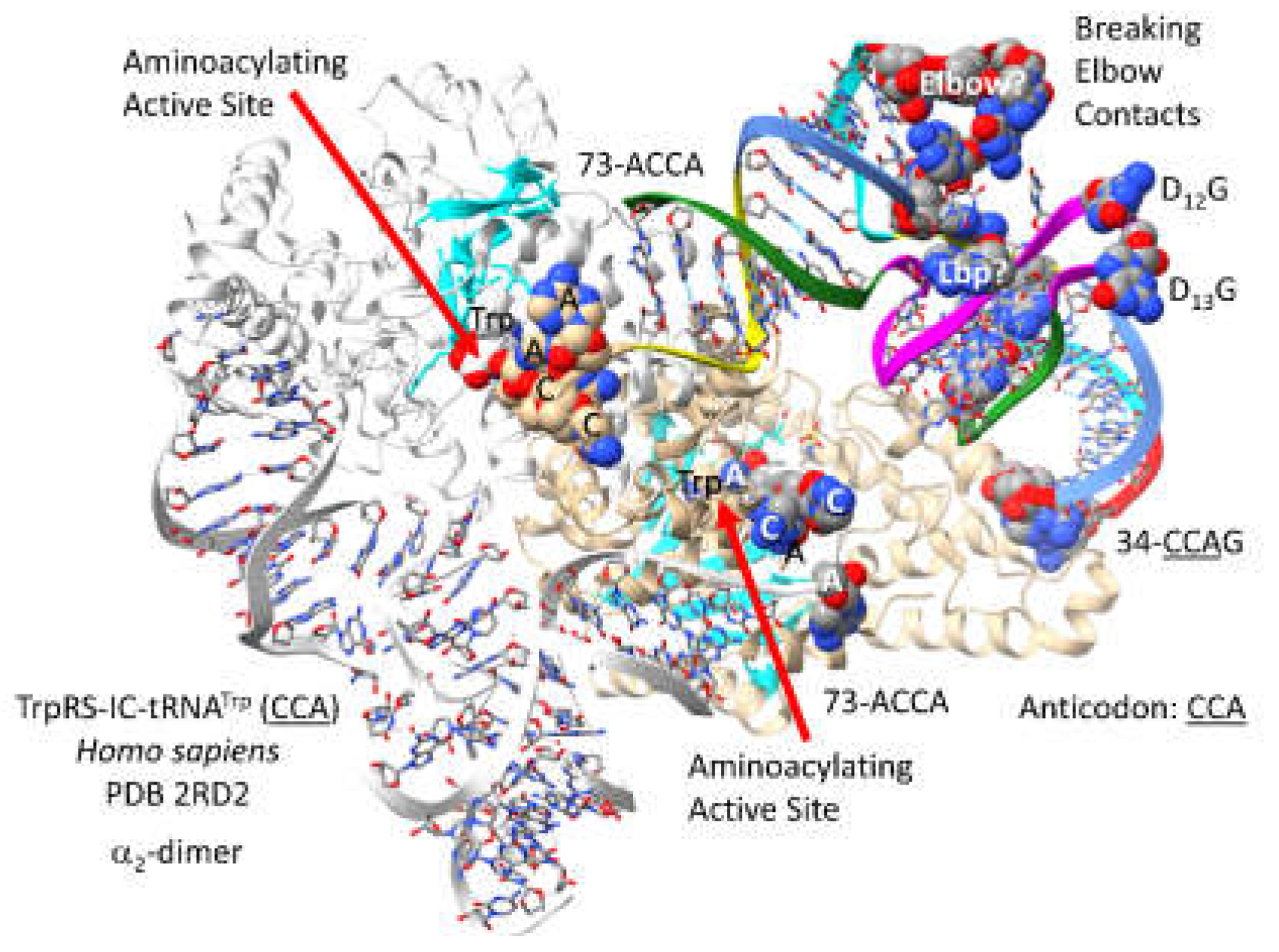
Figure 1 shows a GlyRS-IIA-tRNA
Gly (
CCC) structure from
Homo sapiens [
36]. Human GlyRS-IIA is similar in structure and sequence to archaeal GlyRS-IIA. GlyRS-IIA is an
2-dimer, but the image is of only a single GlyRS-IIA-tRNA
Gly (
CCC). The image was selected to demonstrate primary tRNA
Gly contacts to the anticodon loop and the tRNA 73-ACCA-76 3’-end. As a class II AARS, GlyRS-IIA has its aminoacylating active site on a surface of antiparallel -sheets. The GAP reaction intermediate analogue and the 73-ACCA-76 sequence identify the aminoacylating active site. tRNA
Gly utilizes 34-
CCC-36,
UCC and
GCC anticodons (anticodons are underlined for clarity), so the strongest interactions with GlyRS-IIA might be 35-
CCA-37, as indicated in the structure.
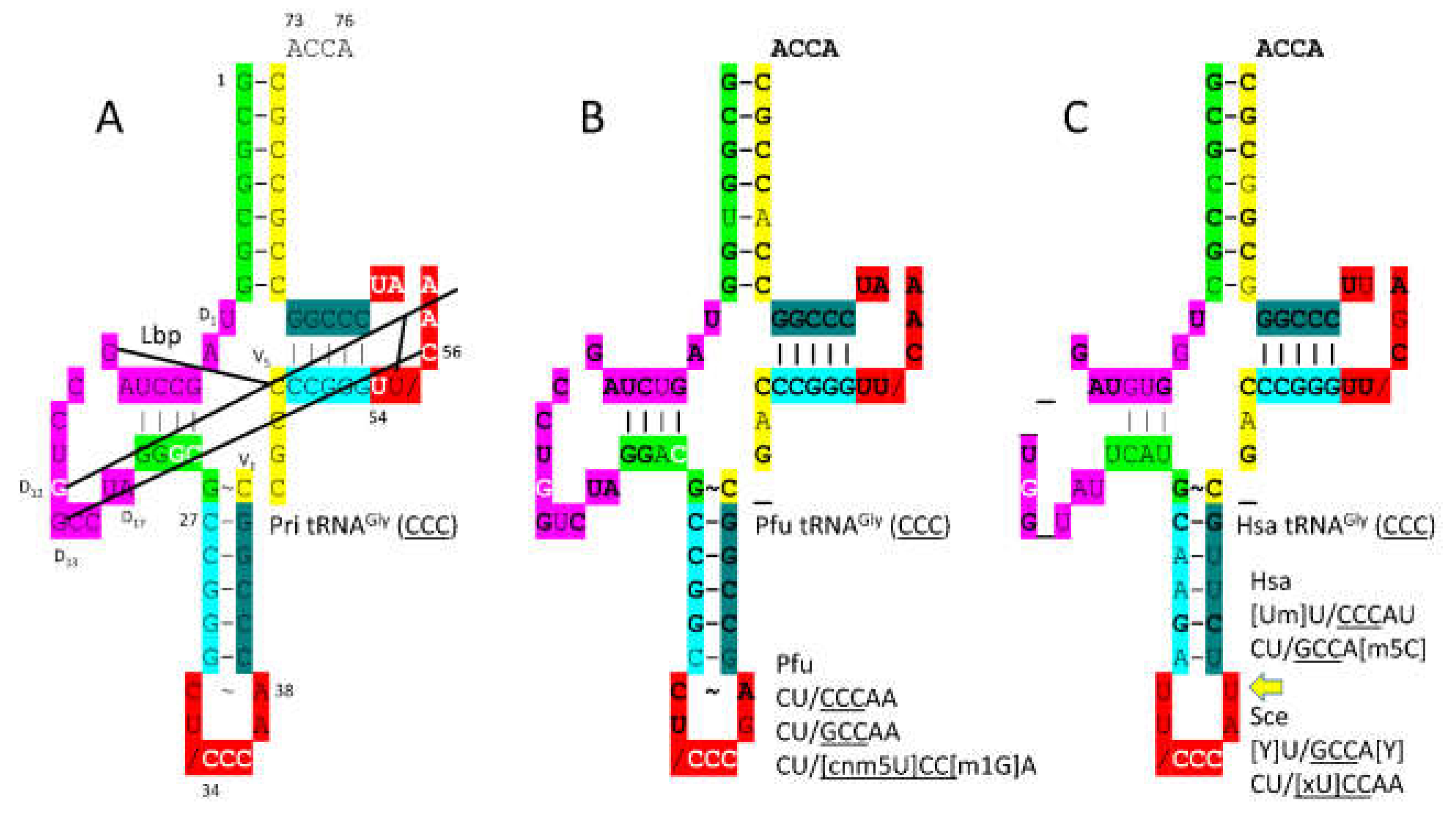
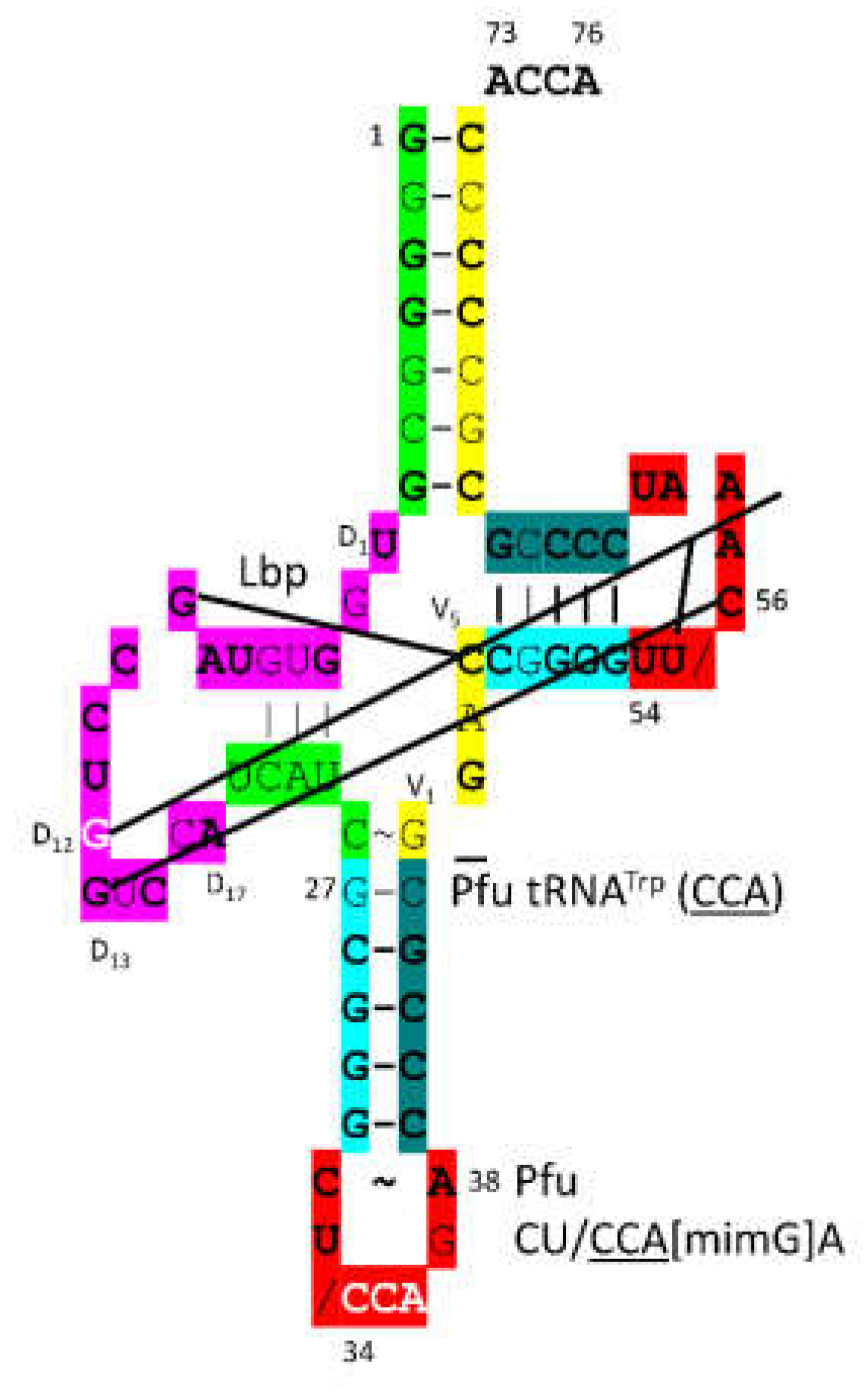
Figure 2 shows tRNA
Gly (
CCC).
Figure 2A is the primordial tRNA
Gly (
CCC).
Figure 2B is the
P. furiosus tRNA
Gly (
CCC).
Figure 2C is the human (Hsa) tRNA
Gly (
CCC), as in the structure in
Figure 1 [
30,
37,
38,
39]. Modifications to the anticodon loop are indicated and explained in the figure legend. Conserved bases compared to the primordial tRNA
Gly are bold in
Figure 2B
,C. As previously described, the primordial tRNA
Gly (
CCC) is a highly ordered sequence formed from GCG (5’-acceptor stem and 5’-acceptor stem remnant (5’-As*)), CGC (3’-acceptor stem and 3’-acceptor stem remnant (type I V loop)), and UAGCC (D loop) repeats and inverted repeats (stem-loop-stem; ~CCGGG_CU/
CCCAA_CCCGG; _ indicates separation of stem and loop; / indicates the U-turn; the anticodon is underlined) [
15,
22]. A few deviations from the perfectly ordered initial sequence are noted (
Figure 2A). These deviations, which pre-date LUCA, support the tRNA fold. D
12G (replacing D
12A in the third UAGCC repeat) intercalates between 57A and 58A and hydrogen bonds to 55U. D
13G forms a Watson-Crick pair with 56C. These are referred to as “elbow” contacts, where the D loop binds the T loop to stabilize the tRNA form [
15,
40]. The T loop is strongly selected to have the typical sequence UU/CAAAU to maintain the interaction with the D loop at the elbow.
In
P. furiosus, tRNA
Gly is the most similar tRNA to tRNA
Pri (Pri for primordial) [
21,
30]. The acceptor stem matches the primordial sequence in all but one bp. The primordial acceptor stem sequence is matched perfectly in some Archaea (i.e.,
Staphylothermus marinus tRNA
Gly (
GCC); GCGGCGG; a GCG repeat) [
42]. In tRNA
Gly (
CCC) of
P. furiosus, the D loop is intact (D
1-UAGUCUAGCCUGGUCUA-D
17;
Figure 2B) and very similar to tRNA
Pri (D
1-UAGCCUAGCCUGGCCUA-D
17;
Figure 2A) with only two base changes from the primordial sequence and no deleted bases relative to tRNA
Pri. The 5’-As* sequence GGACG varies in only a single base from the typical primordial sequence GGGCG. The anticodon stem matches tRNA
Pri in 4 of 5 bp. The V loop sequence C_GAC matches the primordial sequence CCGCC in 3 of 5 positions. The T stem-loop-stem matches tRNA
Pri at every position. As expected, human tRNA
Gly is more innovated from tRNA
Pri, but tRNA
Gly (
CCC) in humans is so similar that it might be monophyletic with tRNA
Gly (
CCC) in Archaea such as
P. furiosus.
In
P. furiosus, only the
34U anticodon loop for tRNA
Gly appears to be modified [
31]. The
34cnm5U modification is initiated by Elp3, which is a first enzyme as ancient as the genetic code. A
34U modified at the 5-carbon is expected to limit superwobbling. Unmodified
34U can read codon wobble 3-A,G,C and U. In mitochondria, superwobbling is utilized in 4-codon boxes to shrink the size of the tRNAome [
43,
44,
45]. A single unmodified
34U tRNA can read an entire 4-codon box. Because glycine is in a 4-codon box, unmodified
34U might be tolerated, but the
34cnm5U modification would limit reading to codon 3-A and 3-G.
Anticodon loop 32C-38A form a hydrogen bond [
41]. A 32Y-38Y (Y for pseudouridine), 32Um-38U, or 32C-38m5C interaction would be expected to change the dynamics of the loop.
Glycine is the smallest and most flexible amino acid. Steric hindrance of larger amino acids may be a mechanism by which GlyRS-IIA limits misincorporation of non-cognate amino acids. In more innovated Bacteria (i.e.,
Escherichia coli), GlyRS-IID substitutes for GlyRS-IIA. GlyRS-IIA is the more ancient enzyme and appears to be the root of both the class II and class I AARS lineages [
16,
18,
46].
The genetic code is hypothesized to have evolved initially to synthesize polyglycine, making tRNA
Gly the first tRNA, and a primitive GlyRS-IIA appears to be the founding AARS, as indicated by sequence. In pre-life, single-stranded RNA may have been stabilized by methylation at the 2’-O [
47]. This modification would render RNA resistant to ribozyme ribonucleases and base hydrolysis. If the 2’-O were modified in pre-life (i.e., by methylation), this might explain why GlyRS-IIA evolved initially to utilize the tRNA-76 ribose 3’-O to attach glycine.
3.3. ValRS-IA
The founding class I AARS appears to be a primitive version of ValRS-IA [
16,
17,
18,
46]. All class I AARS appear to radiate from this root. A primitive ValRS-IA was derived from a primitive GlyRS-IIA by attachment of an N-terminal sequence that redirected to the distinct class I fold. In ancient Archaea, ValRS-IA and IleRS-IA have two Zn motifs, one in the added N-terminal segment and one in the segment that is homologous to the N-terminal Zn motif in GlyRS-IIA. The added N-terminal Zn motif to form class I AARS generates the class I fold. It appears that early folding of class II and class I AARS was highly dependent on these Zn motifs. As evolution progressed, the Zn motifs were sometimes replaced by other folding determinants. Because AARS are first proteins (coevolved with the genetic code), early folding mechanisms dependent on Zn indicate early entry of cysteine into the code.
Glycine, alanine, aspartic acid and valine (GADV) are proposed to be the first encoded amino acids [
48,
49,
50,
51]. GADV (the four simplest amino acids) locate to the 4
th row of the code (tRNA-
36C). The genetic code appears to have sectored primarily by code columns. It is hypothesized that evolution in columns was commenced by filling the code with GADV on favored row 4 and, perhaps, expanding into other rows. It is hypothesized that earlier encoded amino acids occupied larger segments of the code that were then invaded by incoming amino acids. Amino acids that were added early, subsequently retreated to occupy the most favored sectors of the code (i.e., tRNA-
36C). Glycine locates to code column 4 (tRNA-
35C). Alanine locates to code column 2 (tRNA-
35G). Aspartic acid locates to code column 3 (tRNA-
35U). Valine locates to code column 1 (tRNA-
35A). It is hypothesized that row 4 (tRNA-
36C) and column 4 (tRNA-
35C) were the most favored in establishing the code. As the first encoded amino acid, glycine occupies the most favored row and column in the code. The genetic code is a highly ordered assembly.
Valine appears to be the founding amino acid for the assembly of column 1 of the code (tRNA-35A). Assembly of column 1 can be considered from the points of view of similar amino acids and homologous AARS enzymes. In column 1, V (row 4)→L (rows 1 and 2)→I (row 3)→M (row 3), as an order of assembly, appears reasonable. F appears to be a later addition to column 1, row 1, in disfavored row 1. According to closely homologous AARS enzymes, consider the following order of evolution: ValRS-IA→LeuRS-IA→IleRS-IA→MetRS-IA. Entry of phenylalanine and PheRS-IIC will be discussed below. Disfavored row 1 (tRNA-36A) appears to fill last and is a separate case. Sequence preference for rows appears to follow the order C (row 4; tRNA-36C)→G (row 2; tRNA-36G)→U (row 3; tRNA-36U)→→A (row 1; tRNA-36A). In row 3, it appears that Met invaded an Ile 4-codon sector, eliminating the UAU anticodon and inducing differential tRNA-34 modifications of CAU to discriminate Ile (GAU and agm2CAU) (agm2C for agmatidine) and Met (CAU (initiator) and CmAU (elongator)). Modification of the 2-carbon of C (agm2C) (Ile), slightly resembles G (Ile) and discriminates from Met (2-carbon C=O).
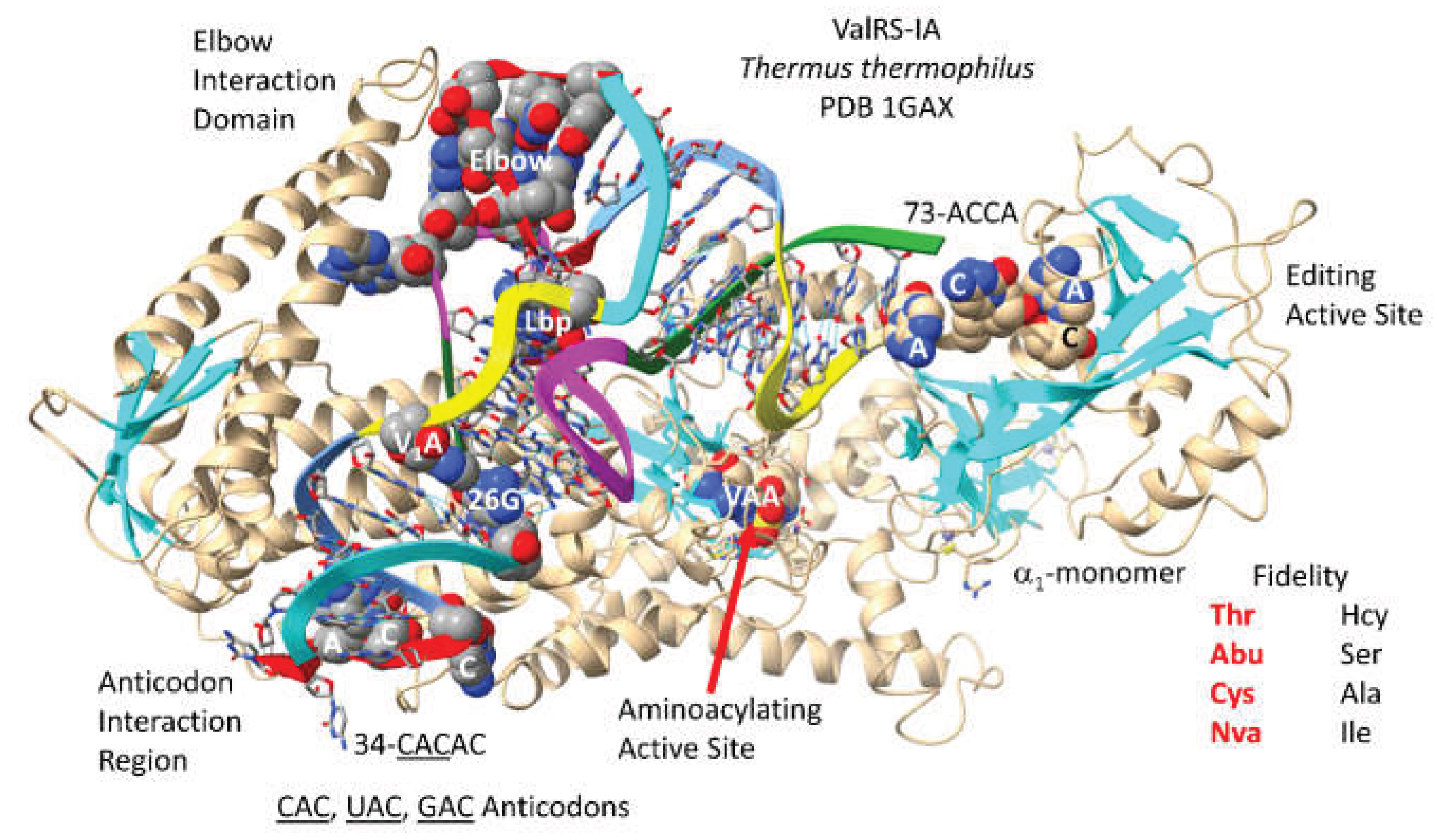
ValRS-IA of
Thermus thermophilus bound to tRNA
Val (
CAC) is shown in
Figure 3 [
52]. ValRS-IA functions as an
1-monomer. Because ValRS-IA is a class I AARS, the aminoacylating active site is at the C-terminal end of a set of parallel -sheets. The arrangement of parallel -sheets has been described as a Rossmann fold, but, undoubtedly, the aminoacylating active site arrangement of ValRS-IA and other class I AARS is unrelated to Rossmann fold proteins genetically. The aminoacylating active site can also be identified by the binding of VAA, a non-reactive Val-AMP analogue. 73-ACCA-76 locates to the separate editing active site that removes non-cognate amino acids from tRNA
Val. Non-cognate homocysteine, serine, alanine and isoleucine can be removed by the separate proofreading (editing) active site after attachment to tRNA
Val. Reactions within the ValRS-IA aminoacylating active site limit non-cognate threonine, -aminobutyric acid, cysteine and norvaline attachments to tRNA
Val. As a small, hydrophobic amino acid, valine has little chemical character, so editing reactions both before and after tRNA
Val attachment are important to limit inaccurate translation [
34,
35].
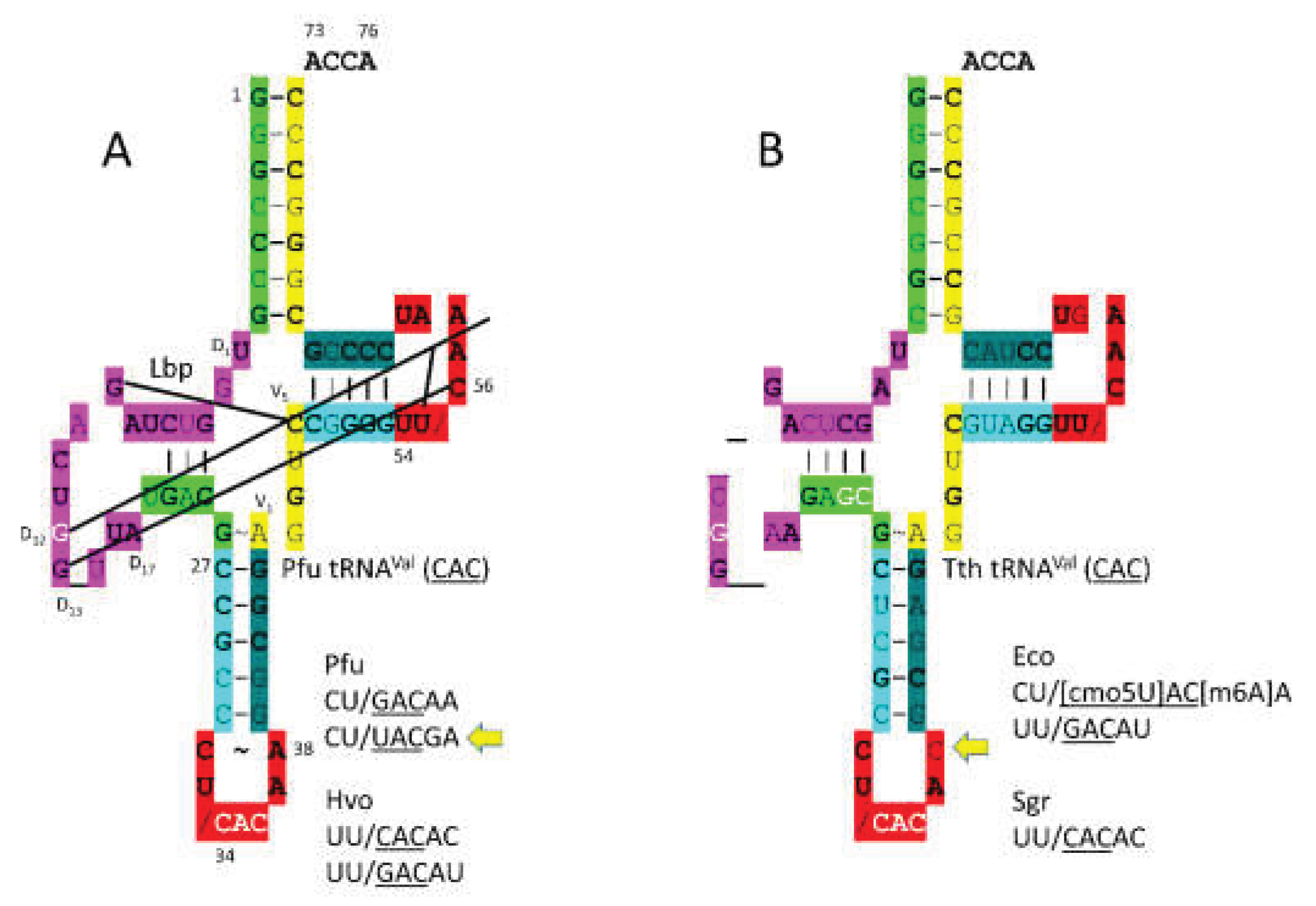
tRNA
Val is shown in
Figure 4. In
Figure 4A,
P. furiosus tRNA
Val (
CAC) is shown. The acceptor stem matches the primordial tRNA sequence in 4 of 7 bp. The D loop has the sequence D
1-UGGUCUAGACUGG_UUA-D
17 and matches the primordial sequence in all but 5 positions with a single base deleted from tRNA
Pri. The anticodon stem matches the primordial sequence in 3 of 5 bp. The T stem-loop-stem matches the primordial sequence in all but one stem bp. In
P. furiosus, tRNA
Val is similar to tRNA
Ala in sequence, indicating that Val and Ala may have entered the code at about the same time in evolution. GADV are proposed to have been the first 4 encoded amino acids [
48,
49,
50]. The
T. thermophilus tRNA
Val (
CAC) is more derived from the root sequence, as expected (
Figure 4B). Bacteria are more derived from LUCA than Archaea.
tRNA
Val utilizes
CAC,
UAC and
GAC anticodons. In
P. furiosus and
H. volcanii, none of these anticodon loops appears to be modified [
31,
53]. Unmodified
UAC would be predicted to read codon 3-A,G,C, and U by superwobbling [
43,
44,
45]. Because valine occupies a 4-codon sector, such promiscuity would be tolerated and might be selected. The tRNA
Val (
CAC) anticodon loop is substantially unwound by ValRS-IA, indicating allosteric effects of ValRS-IA-tRNA
Val binding. Allostery is likely important in selectively directing the tRNA
Val 3’-end to the aminoacylating or separate proofreading active site. Because ValRS-IA makes elbow contacts, these might leverage allosteric effects mediated through tRNA
Val. It appears that 35-
AC-36 and 38C make strongest contact to ValRS-IA, as might be expected. In
P. furiosus, 38A is present, rather than 38C, as in
E. coli. In
P. furiosus, tRNA
Val and tRNA
Ala are similar in sequence, consistent with the GADV hypothesis that indicates that valine and alanine were two of the first four encoded amino acids.
3.4. IleRS-IA
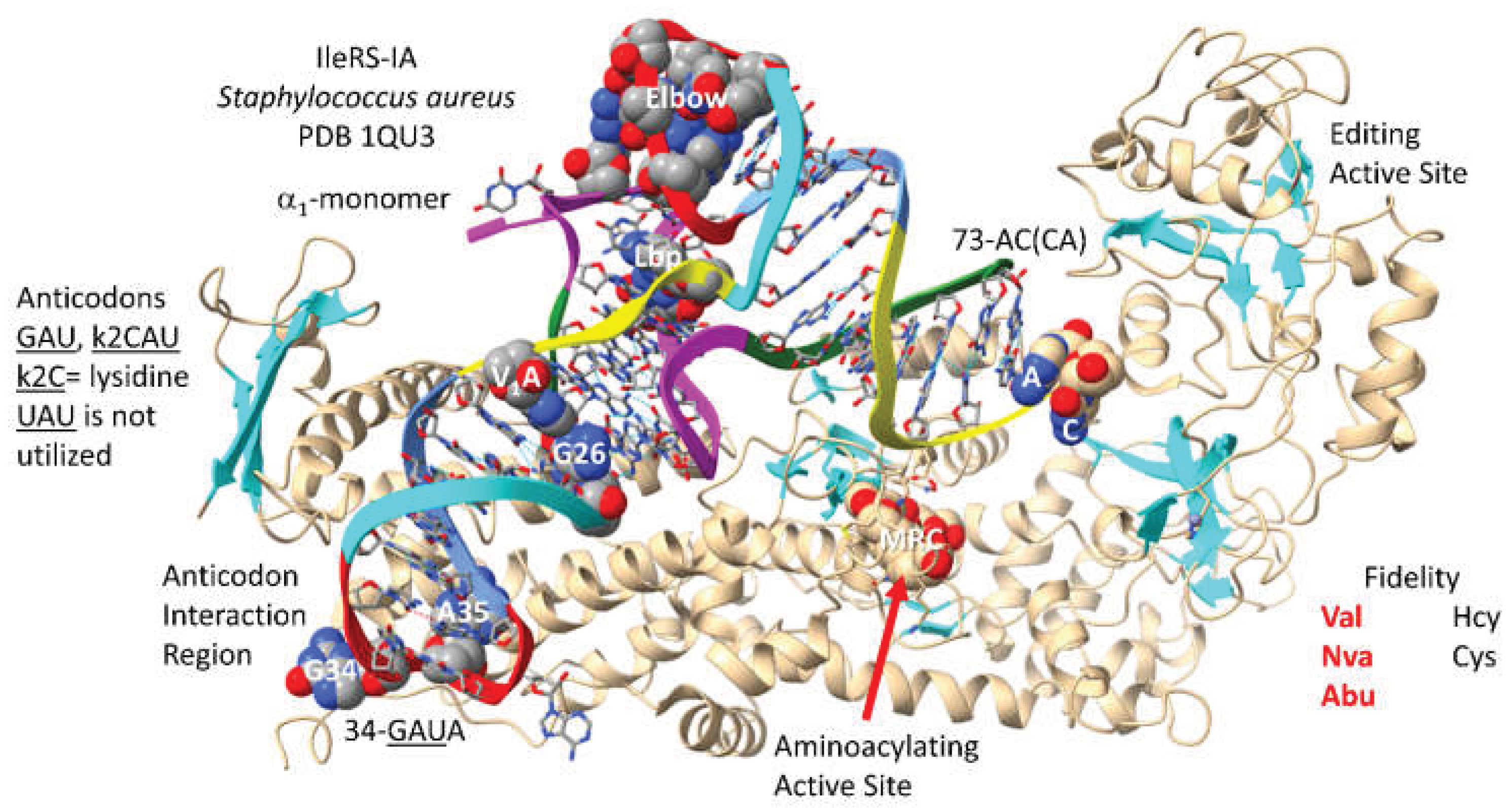
IleRS-IA-tRNA
Ile (
GUA) from Bacterium
Staphylococcus aureus is shown in
Figure 5 [
54]. By structure and sequence, IleRS-IA is closely related to ValRS-IA. Both enzymes function as
1-monomers. Compared to ValRS-IA, IleRS-IA appears to make no elbow contacts and unwinds the tRNA
Ile (
GAU and
k2CAU (
k2C for lysidine)) (in Archaea,
GAU and
agm2CAU (
agm2C for agmatidine)) anticodon loop to a somewhat lesser extent than ValRS-IA. Elbow contacts by an AARS may be used to leverage allosteric effects transmitted through a cognate tRNA to the aminoacylating or editing active sites. Apparently,
k2C and
agm2C can partly mimic
G for IleRS-IA binding. Minimally,
k2C and
agm2C are better
G mimics than 2-C=O, as in unmodified
C at the 2 carbon (i.e., Met anticodons). The
agm2C modification is added by a first protein (tRNA
Ile2 2-agmatinylcytidine synthase). As noted above, the
UAU anticodon is rarely encoded in Archaea. When anticodon
UAU is encoded, it is also modified to
agm2CAU to encode Ile. Also, MetRS-IA utilizes
CmAU (elongator) and
CAU (initiator) anticodons. Because Ile anticodons have tRNA-
36U, tRNA-37 is t6A or hn6A. It is hypothesized that modified tRNA-37A (i.e., t6A, hn6A) may have evolved to suppress wobbling at tRNA-
36U.
Because IleRS-IA is a class I AARS, the aminoacylating active site is at the C-terminal ends of a set of parallel -sheets. The reaction intermediate analogue MRC binds at this site. 73-AC(CA) is not fully resolved in the structure. IleRS-IA has a separate editing active site that can remove non-cognate homocysteine and cysteine from tRNAIle. The IleRS-IA aminoacylating active site limits incorporation of valine, norvaline and -aminobutyric acid. Similar to valine, isoleucine is a somewhat featureless amino acid that requires editing functions to suppress translation errors.
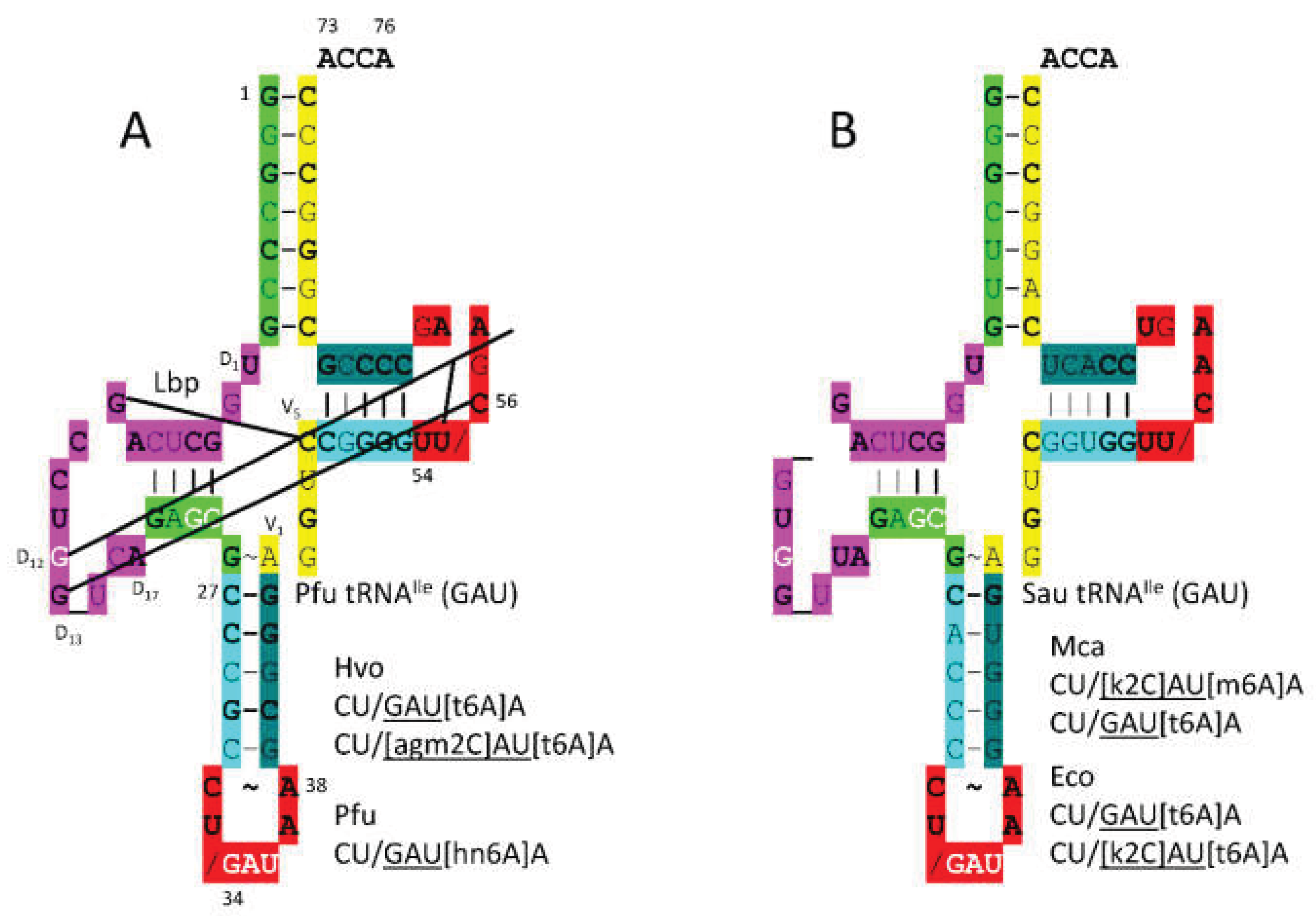
tRNA
Ile (
GAU) is shown in
Figure 6. In
P. furiosus, tRNA
Ile (
GAU) matches the primordial tRNA sequence in 4 bp within the acceptor stem. The D loop has the sequence D
1-UGGCUCAGCCUGG_UCA-D
17 matching the primordial tRNA in all but 6 positions with a single base deleted from tRNA
Pri. The 5’-As* sequence is GAGCG versus GGGCG in primordial tRNA [
15,
22]. The anticodon stem matches primordial tRNA in 3 bp. The T stem-loop-stem matches 4 of 5 bp in the stem and all but 2 bases in the loop compared to tRNA
Pri. In
S. aureus, the tRNA
Ile is more innovated.
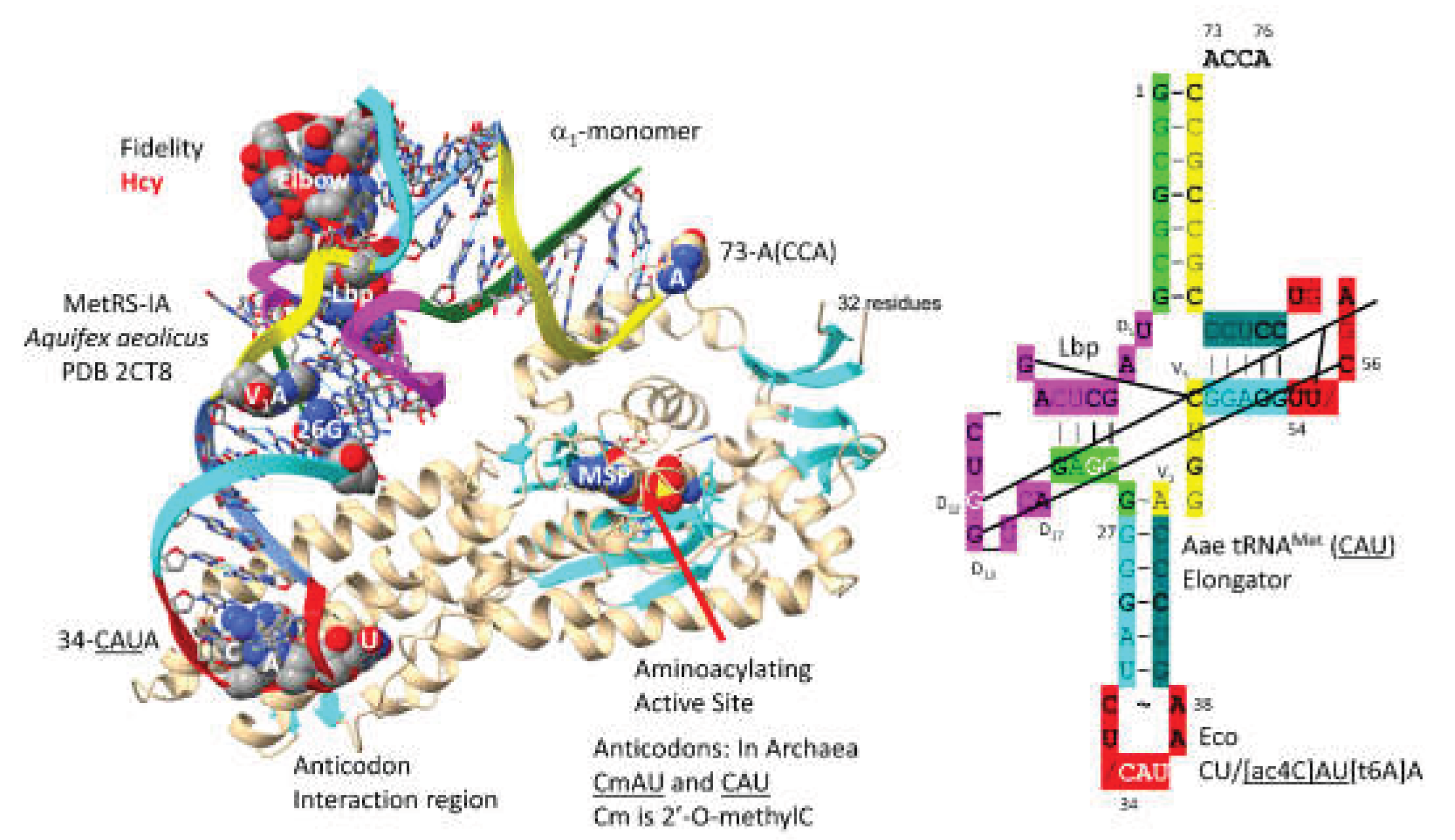
3.5. MetRS-IA
MetRS-IA-tRNA
Met (
CAU) from
Aquifex aeolicus is shown in
Figure 7 [
55]. In sequence and structure, MetRS-IA is very similar to IleRS-IA and ValRS-IA. MetRS-IA functions as an
1-monomer. There are no tRNA
Met (
CAU) elbow contacts. Unwinding of the anticodon loop to expose
CmAU or
CAU is slight, perhaps because only a single
CAU anticodon, lacking the
agm2C or
k2C modifications to encode isoleucine, is recognized. Because of deletion, MetRS-IA lacks an editing active site, but the aminoacylating active site of MetRS-IA limits incorporation of homocysteine. The aminoacylating active site is identified by MSP binding and a set of parallel -sheets. 73-A(CCA) is partly resolved in the structure, possibly indicating allosteric effects (i.e., of MSP analogue binding).
It is hypothesized that methionine entered the genetic code by invading a 4-codon isoleucine sector [
16]. The invasion resulted in the suppression of the
UAU anticodon that would cause ambiguity between methionine and isoleucine coding. Methionine adopted the
CmAU (elongator) and
CAU (initiator) anticodons (
Figure 8). In Archaea, to support a 3-codon box, isoleucine utilized the
agm2CAU and
GAU anticodons. In Bacteria, the
k2CAU and
GAU anticodons are utilized for isoleucine.
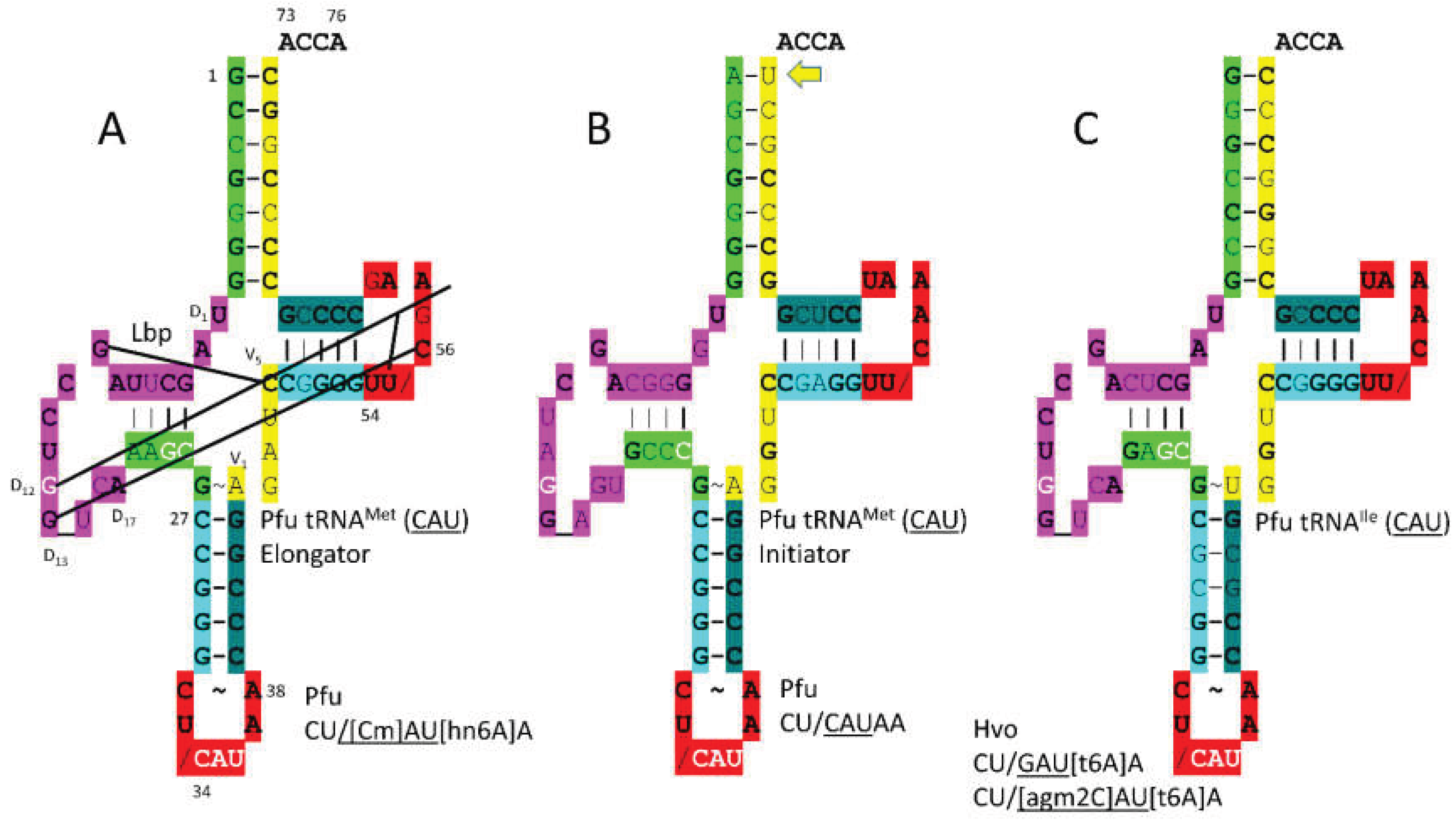
P. furiosus elongator tRNA
Met (
CmAU) is a close match to the primordial tRNA sequence (
Figure 8). The acceptor stem matches in 5 bp out of 7. The D loop sequence D
1-UAGCUUAGCCUGG_UCA-D
17 matches in all but 4 positions with a single base deleted from tRNA
Pri. The anticodon stem matches tRNA
Pri in 5 out of 5 bp. The T stem matches in 4 of 5 bp, and the T loop matches tRNA
Pri in all but 2 bases. The initiator tRNA
Met (
CAU) anticodon loop is unmodified. The 1A-72U pair in initiator tRNA
Met (
CAU) is unusual and more readily melted than 1G=72C (
Figure 8B), which is typical. The tRNA
Ile (
agm2CAU) sequence is similar to elongator tRNA
Met (
CmAU), indicating that tRNA
Met may have been derived from a primitive tRNA
Ile.
3.6. LeuRS-IA
tRNALeu is a type II tRNA with a longer V arm (initially, 14 nt; a 3’-acceptor stem ligated to a 5’-acceptor stem; initially, CCGCCGC_GCGGCGG). In Archaea, tRNALeu and tRNASer are type II tRNAs. Both leucine and serine are in 6-codon boxes. LeuRS-IA and SerRS-IIA utilize the longer tRNA V arms as major determinants for cognate tRNA charging rather than the anticodon loops, which LeuRS-IA and SerRS-IIA do not contact. Arginine is also in a 6-codon box, but tRNAArg is a type I tRNA. Rather than using a longer type II tRNAArg V arm, ArgRS-IA uses enhanced anticodon loop unwinding to expose bases for recognition. In Bacteria, tRNATyr is a type II tRNA, but type II tRNATyr (GUA) and its recognition by bacterial TyrRS-IC are bacterial innovations.
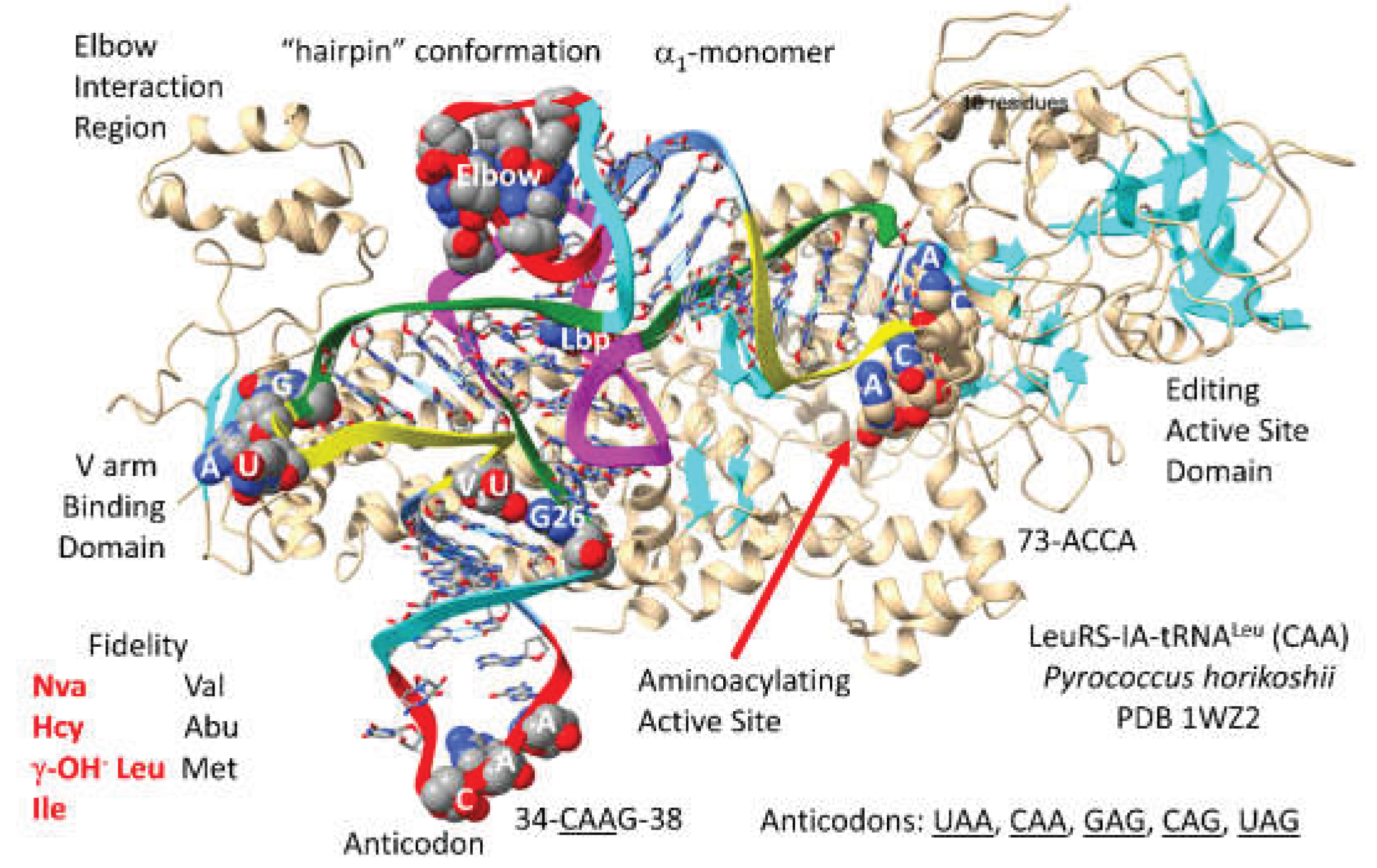
LeuRS-IA-tRNA
Leu (
CAA) of
Pyrococcus horikoshii is shown in
Figure 9 [
56,
57]. By structure and sequence, LeuRS-IA is closely related to ValRS-IA, IleRS-IA and MetRS-IA. LeuRS-IA functions as an
1-monomer. Archaeal and bacterial LeuRS-IA have different modes of contacting the type II V arm of their cognate tRNA
Leu. In Archaea (but not in Bacteria), LeuRS-IA contacts the end loop of the V arm at the typical sequence UAG [
11]. In both Archaea and Bacteria, the tRNA
Leu (
CAA) anticodon loop is not contacted by LeuRS-IA. Of all the AARS, only LeuRS-IA, SerRS-IIA and AlaRS-IID lack anticodon loop contacts for cognate tRNA recognition. A C-terminal region of archaeal LeuRS-IA contacts the elbow. The aminoacylating active site is identified by parallel -sheets and 73-ACCA-76. The tRNA
Leu (
CAA) 73-ACCA-76 is in the catalytic “hairpin” conformation for class I AARS enzymes, curving down into the aminoacylating active site. LeuRS-IA has a separate editing active site that removes non-cognate valine, -aminobutyrate and methionine from tRNA
Leu. The LeuRS-IA aminoacylating active site limits non-cognate norvaline, homocysteine, -OH
- leucine and isoleucine incorporation [
34]. Leucine is a hydrophobic amino acid with little chemical character and so requires editing to maintain translational accuracy.
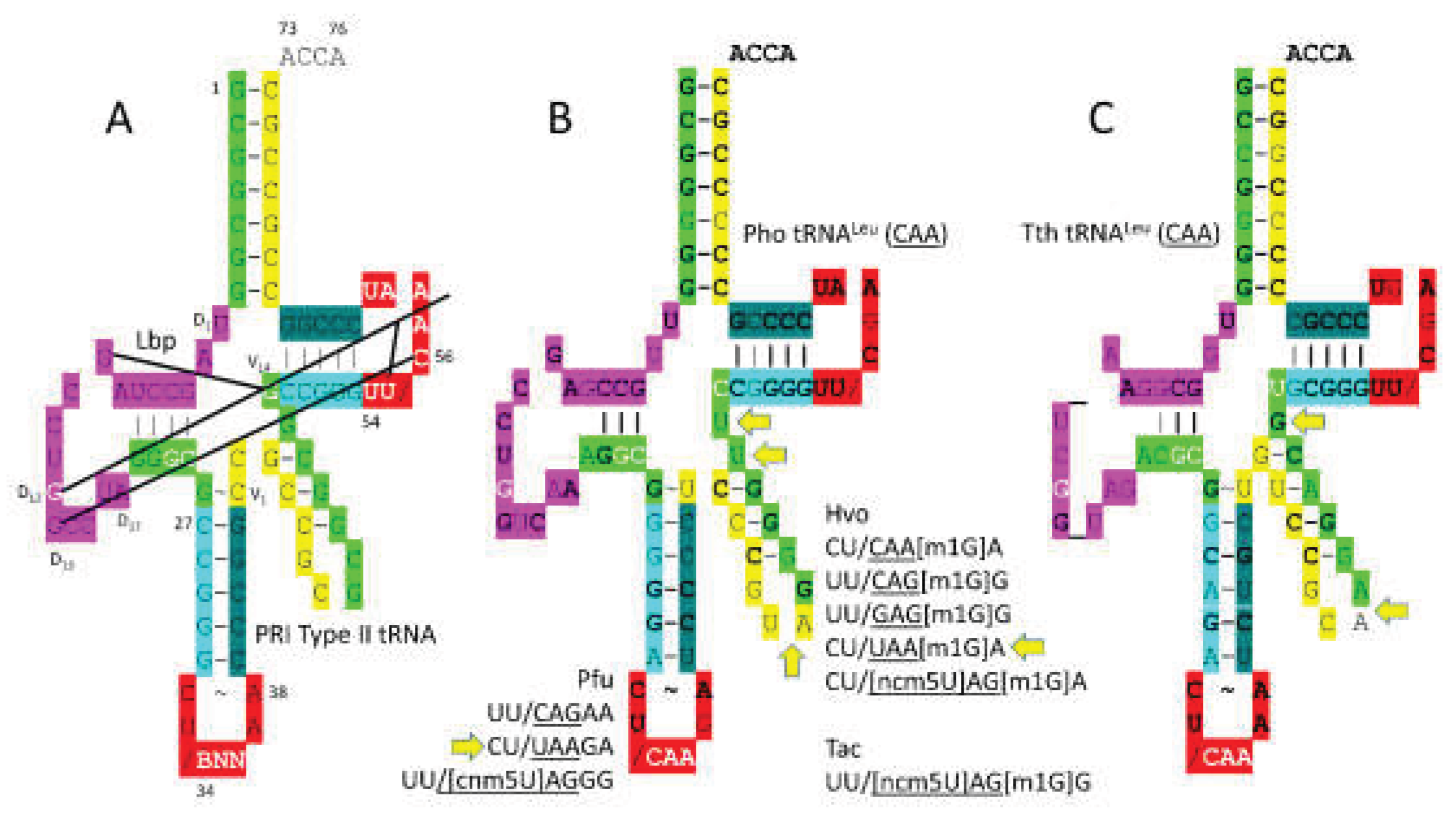
Figure 10 shows a comparison of a primordial type II tRNA (
Figure 10A) and archaeal tRNA
Leu (
CAA) (
Figure 10B). A bacterial tRNA
Leu is also shown (
Figure 10C). In
P. horikoshii, the tRNA
Leu (
CAA) type II V arm is 14 nt, as in type II tRNA
Pri. Most archaeal tRNA
Leu type II V arms are 14 nt in length, the primordial length. In Archaea, the tRNA
Leu type II V arm is a major determinant for cognate tRNA
Leu charging with leucine. The trajectory of the type II V arm is different than for tRNA
Ser, with two bases separating the 3’-V arm stem and the Levitt base for tRNA
Leu and with one base separating the 3’-V arm stem and the Levitt base for tRNA
Ser, in Archaea. For archaeal tRNA
Leu, the V arm end loop includes the UAG consensus to bind LeuRS-IA (
Figure 10B). The V arm end loop contact is not utilized by bacterial LeuRS-IA (
Figure 10C). tRNA
Leu utilizes
CAA,
UAA,
CAG,
GAG and
UAG anticodons. Because of superwobbling, unmodified
UAA anticodons, in principle, might utilize a phenylalanine UUU or UUC codon, substituting leucine for phenylalanine. We are not certain what limits miscoding, in this case. The anticodon
cnm5UAG is in a 4-codon box, but it has the 5-carbon
U modification that should suppress superwobbling.
The
P. horikoshii tRNA
Leu (
CAA) acceptor stem matches tRNA
Pri at 6 out of 7 bp (
Figure 10B). The D loop has the sequence D
1-UUGCCGAGCCUGGUCAA-D
17 matching the tRNA
Pri sequence in all but 4 positions and including no deletions compared to tRNA
Pri. The 5’-As* sequence is AGGCG matching typical tRNA
Pri GGGCG in all but one position. Two bases separate the type II 3’-V arm stem and the Levitt base. The V arm end loop has the sequence V
5-GUAG-V
8 that includes the V
6-UAG-V
8 consensus to bind LeuRS-IA (
Figure 9 and
Figure 10B). The T stem matches tRNA
Pri in all but one bp. The T loop matches in all but one base.
3.7. SerRS-IIA
At the base of code evolution, only tRNA
Leu and tRNA
Ser were selected to be type II tRNAs. The number of amino acids that are type II in an organism or domain is determined by the allowed trajectories of the V arm. In Archaea, the number of trajectories is two. In Bacteria, the number is three. In Archaea and Bacteria, the trajectories of type II V arms are different for tRNA
Leu and tRNA
Ser. SerRS-IIA is a very different AARS compared to LeuRS-IA. From sequence, however, it appears that type II tRNA
Ser may have been derived from type II tRNA
Leu. To maintain translational accuracy, the type II V arm of tRNA
Ser is recognized very differently than the type II V arm of tRNA
Leu. The type II tRNA
Ser V arm has a different trajectory from its tRNA body compared to the tRNA
Leu V arm [
11]. The trajectory of the type II V arm depends on the number of unpaired bases between the 3’-V arm stem and the Levitt reverse Watson-Crick base pair (i.e., D
8G=V
14C). In Archaea, for tRNA
Leu, the number is two unpaired bases (in Bacteria, the number is one unpaired base for the tRNA
Leu type II V arm). For tRNA
Ser, the number in Archaea is one unpaired base (in Bacteria, the number is zero unpaired bases for the tRNA
Ser type II V arm).
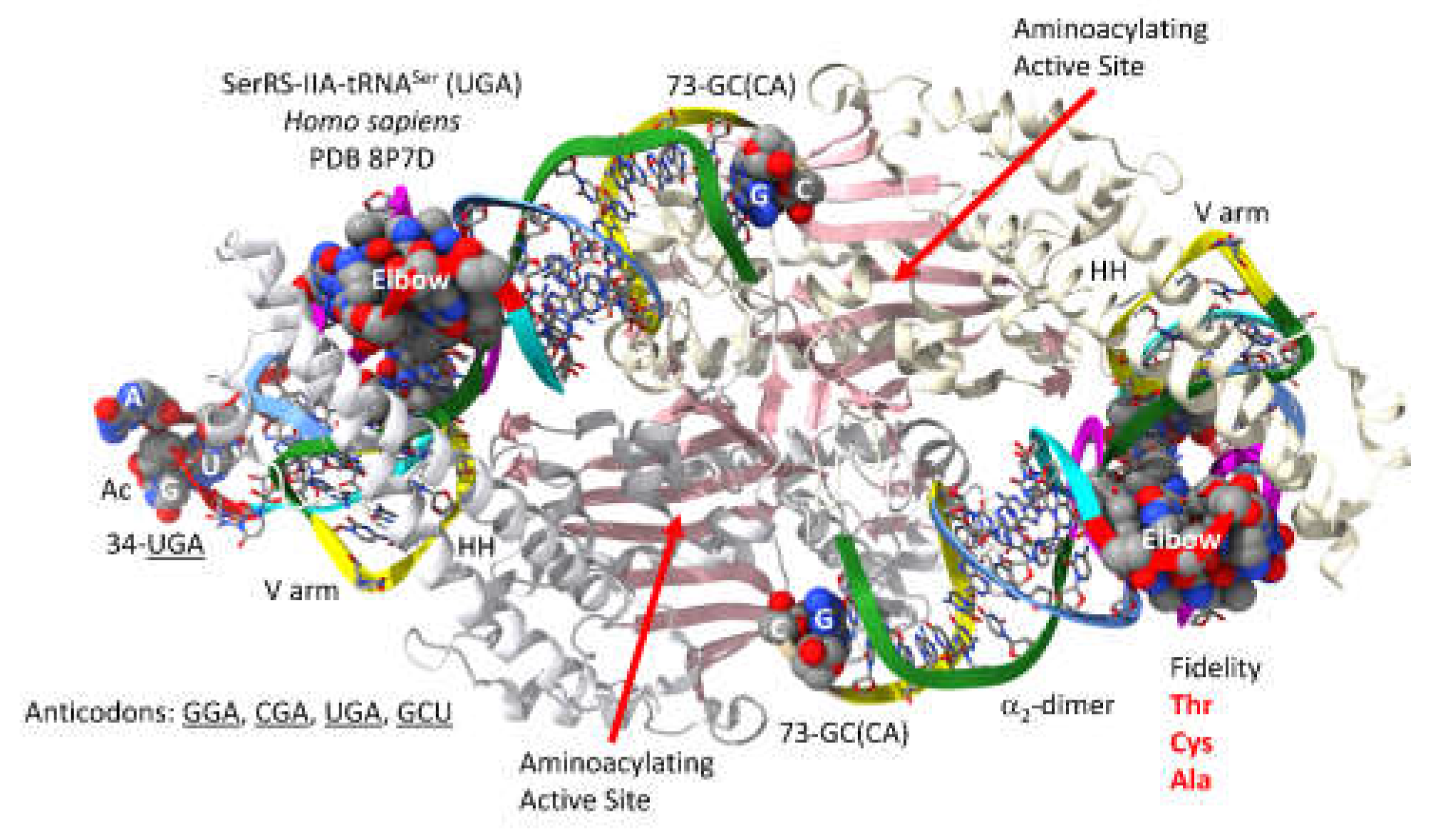
Human SerRS-IIA-tRNA
Ser (
UGA) is shown in
Figure 11 [
58]. SerRS-IIA has an N-terminal helix hairpin that lies across the type II V arm and interacts with the tRNA
Ser elbow. SerRS-IIA functions as an
2-dimer. The aminoacylating active site and the helix hairpin for a single tRNA
Ser are on separate -subunits. As noted above, no contacts are made by SerRS-IIA to the tRNA
Ser anticodon loop. The aminoacylating active site is on a surface of antiparallel -sheets. The SerRS-IIA aminoacylating active site limits non-cognate attachment of threonine, cysteine and alanine to tRNA
Ser.
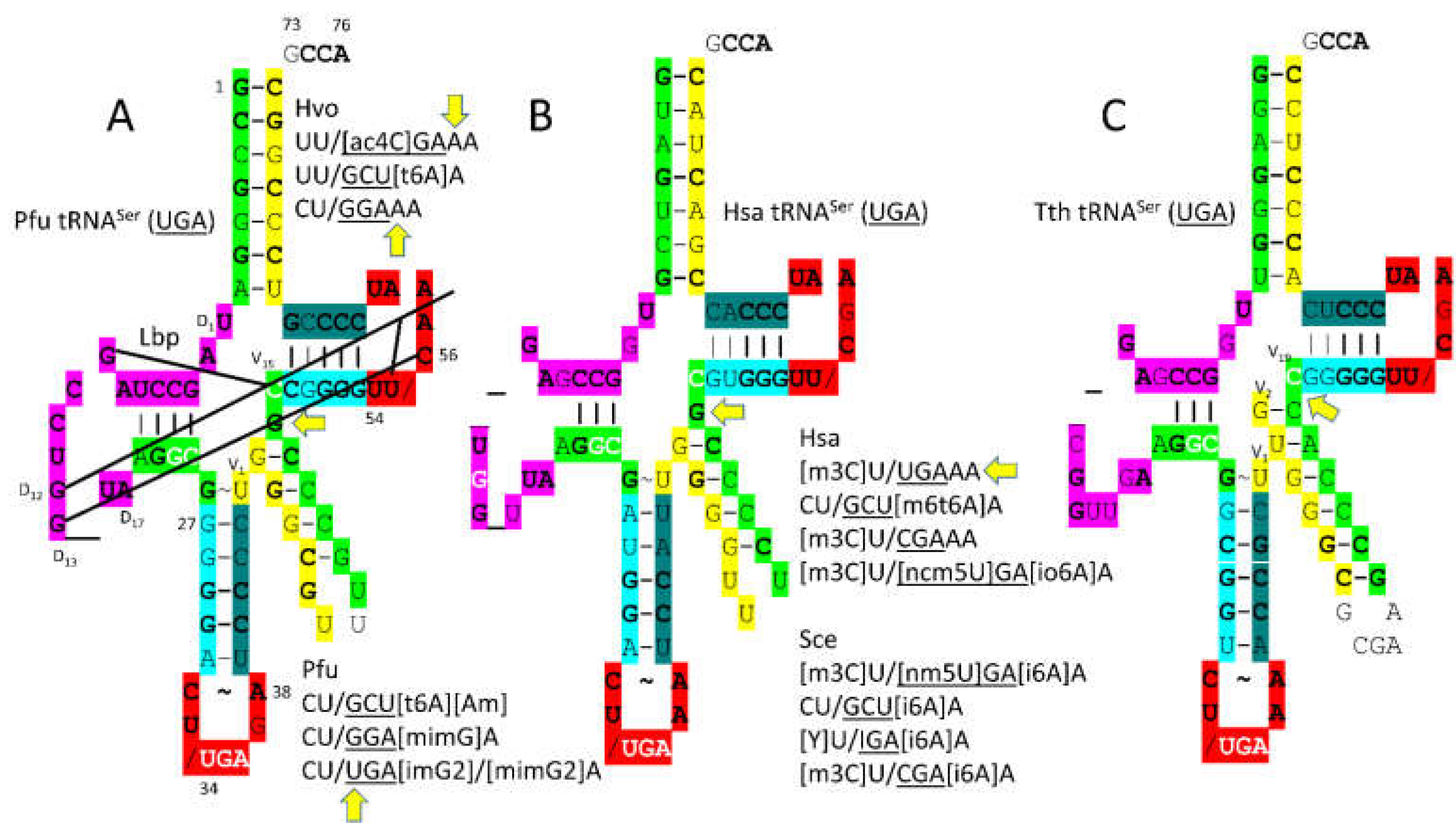
tRNA
Ser anticodon loop modifications are interesting and slightly unanticipated (
Figure 12). Generally, tRNA-
36U is associated with a modified tRNA-37A, as is observed. Generally, tRNA-
36A is associated with a modified tRNA-37G, but
GGA in
H. volcanii is followed by unmodified 37A. In
P. furiosus,
UGA is unmodified at
34U, but
UGA is in a 4-codon box, so implied superwobbling would not cause miscoding. In
P. furiosus tRNA
Ser (
UGA), the acceptor stem matches that of tRNA
Pri in 4 of 7 bp. In
H. sapiens, tRNA
Ser (
UGA) matches the tRNA
Pri acceptor stem in 5 of 7 bp (
Figure 12B). In the D loop, tRNA
Ser (
UGA) of
P. furiosus has two perfect UAGCC repeats, consistent with the three 31 nt tRNA evolution theorem. The D loop sequence is D
1-UAGCCUAGCCUGG__UA-D
17 matching the primordial tRNA sequence in all but two deleted positions. The 5’-As* sequence AGGCG matches tRNA
Pri in 4 of 5 positions. The human anticodon stem matches tRNA
Pri in 3 of 5 bp. The T stem-loop-stem of
P. furiosus matches tRNA
Pri in all but one stem bp.
In Bacteria, trajectories of tRNA type II V arms are different than in Archaea (compare
Figure 12A
,C). In Archaea, tRNA
Leu has two unpaired bases (
Figure 10B) and tRNA
Ser has one unpaired base (
Figure 12A) separating the 3’-V arm stem and the Levitt base. In Bacteria, tRNA
Tyr has 2 unpaired bases (see below), tRNA
Leu has one unpaired base (
Figure 10C) and tRNA
Ser has zero unpaired bases (
Figure 12C) separating the type II V arm 3’-stem and the Levitt base [
11]. Differences in type II V arm trajectories cause different modes of cognate AARS-tRNA recognition and are expected to limit horizontal type II tRNA gene transfers between Bacteria and Archaea.
Serine is the only amino acid that locates to two separate columns of the genetic code. It is hypothesized that serine jumped from column 2 to column 4 during code evolution. Being a type II tRNA lacking anticodon recognition by SerRS-IIA probably facilitated jumping. We suggest that jumping of serine in code evolution may correlate with introduction of cysteine into the code (see below).
3.8. ArgRS-IA
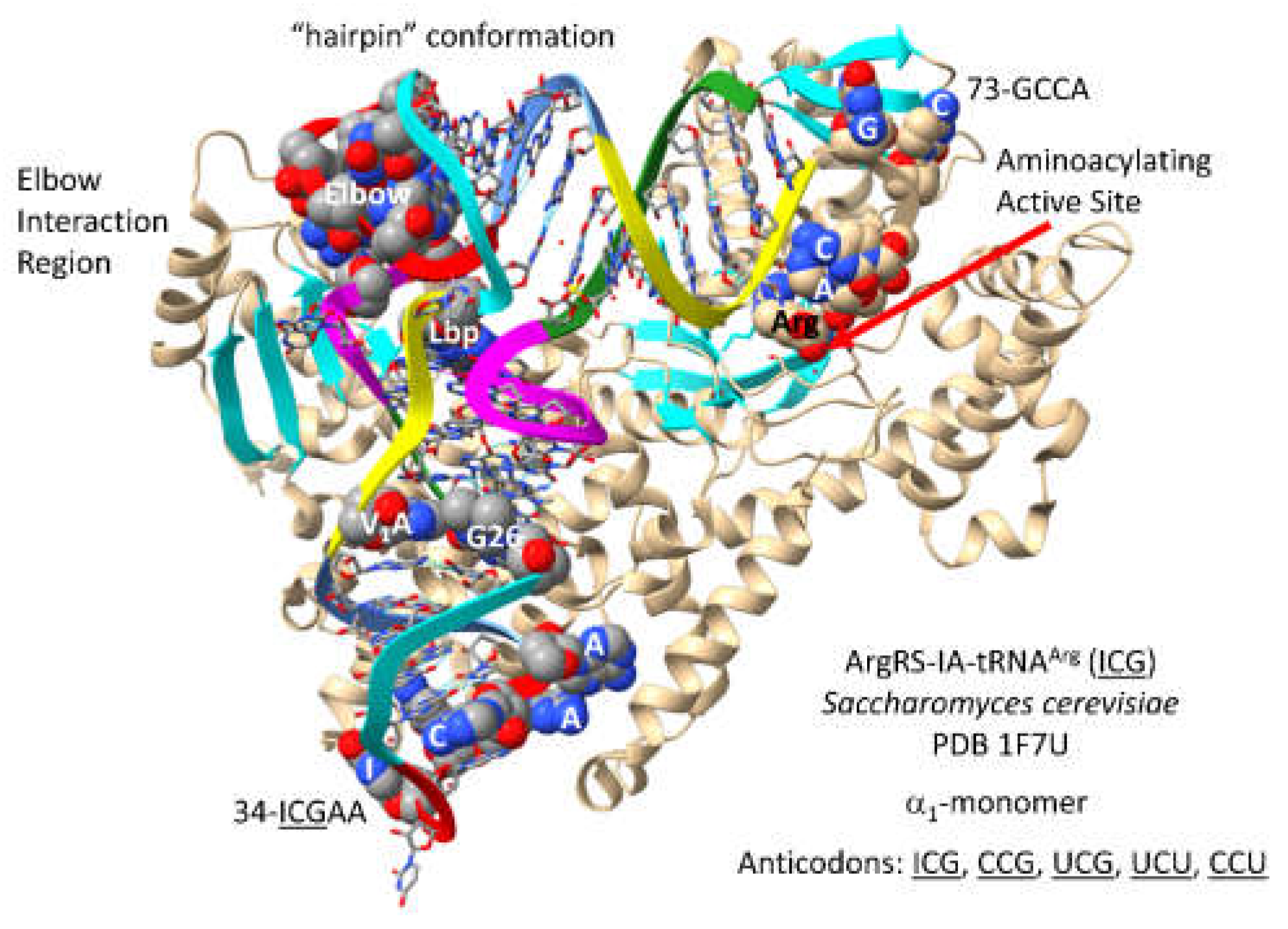
ArgRS-IA-tRNA
Arg (
ICG) of
S. cerevisiae is shown in
Figure 13 [
59]. As noted above, although arginine is in a 6-codon box, tRNA
Arg is a type I tRNA. Compared to LeuRS-IA and SerRS-IIA, ArgRS-IA utilizes the alternate strategy of increased unwinding of the type I tRNA
Arg anticodon loop to expose additional bases for cognate recognition. Three amino acids probably could not occupy 6-codon sectors in the code using the strategy that evolved for arginine, explaining why tRNA
Leu and tRNA
Ser evolved to substitute recognition of longer type II V arms, rather than utilizing anticodon loop determinant contacts. For tRNA
Arg, the 34-
ICGAA-38 sequence is substantially unwound.
35C, 37A and 38A appear to make strong contacts to ArgRS-IA. ArgRS-IA makes substantial elbow contacts that may help leverage anticodon loop opening through allosteric effects. 73-GCCA-76 is in the catalytic “hairpin” conformation for a class I AARS. Arginine binds at the aminoacylating active site. As expected for a class I AARS, parallel -sheets approach the aminoacylating active site.
In P. furiosus, the GCG anticodon would correspond most closely to the ICG anticodon in S. cerevisiae. Generally, when encoded ACG is modified to ICG by deamination, in Bacteria and Eukarya, the corresponding GCG anticodon is not utilized. Similarly, Archaea do not appear to utilize the 34A→I modification, but use the GCG anticodon instead. It is notable that 34A is not utilized in Archaea and, for the most part, in Bacteria (Bacteria utilize ICG to encode arginine). The lack of anticodon wobble base discrimination (i.e., pyrimidine versus purine, only) causes genetic code degeneracy.
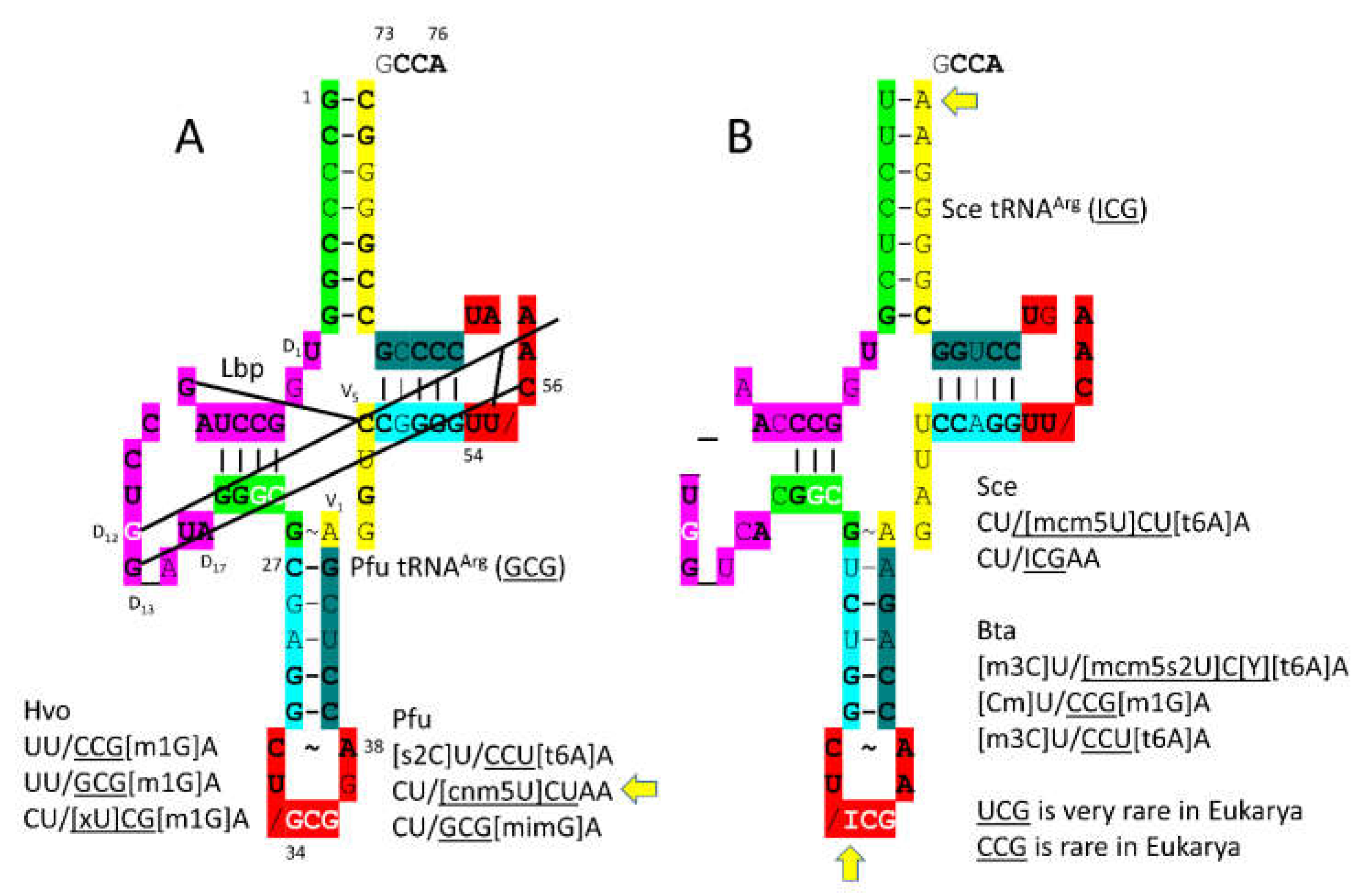
The
P. furiosus tRNA
Arg (
GCG) sequence is of interest (
Figure 14A). The acceptor stem matches tRNA
Pri in 5 of 7 bp. The D loop has the sequence D
1-UGGCCUAGCCUGG_AUA-D
17, which varies in only 3 positions from the primordial D loop sequence with a single base deleted relative to tRNA
Pri. The 5’-As* sequence is GGGCG, which matches the primordial tRNA sequence (GGGCG rearranged from GGCGG before LUCA). The V loop sequence AGGUC is typical. The T stem-loop-stem matches in all but one stem bp. The
S. cerevisiae tRNA
Arg (
ICG) sequence is more derived from the root sequence, as expected (
Figure 14B). In
P. furiosus, the
[cnm5U]CU anticodon is followed by an unmodified A, which is unexpected. Perhaps, the
cnm5U modification or another feature helps to compensate. Generally,
36U is followed by a modified 37A, as in
CCU[t6A]. CGA and CGG codons are rare in Eukarya. The corresponding
UCG and
CCG anticodons are also rare in Eukarya.
Arginine is an amino acid with significant discriminating characteristics. Arginine is positively charged and much stiffer than lysine. Also, arginine has significant hydrogen bonding potential. These characteristics discriminate arginine from lysine, which is much more flexible and has a more concentrated positive charge. We consider the idea that the first encoded positively charged amino acid may have been ornithine [
60]. Ornithine can be converted to arginine in two enzymatic steps, consistent with the notion that tRNA-linked chemistry may have contributed to the encoding of arginine and lysine. Ornithine can be converted to lysine in some Archaea and Bacteria [
61,
62,
63]. Consistent with this idea, tRNA
Arg and tRNA
Lys are similar in sequence in
P. furiosus.
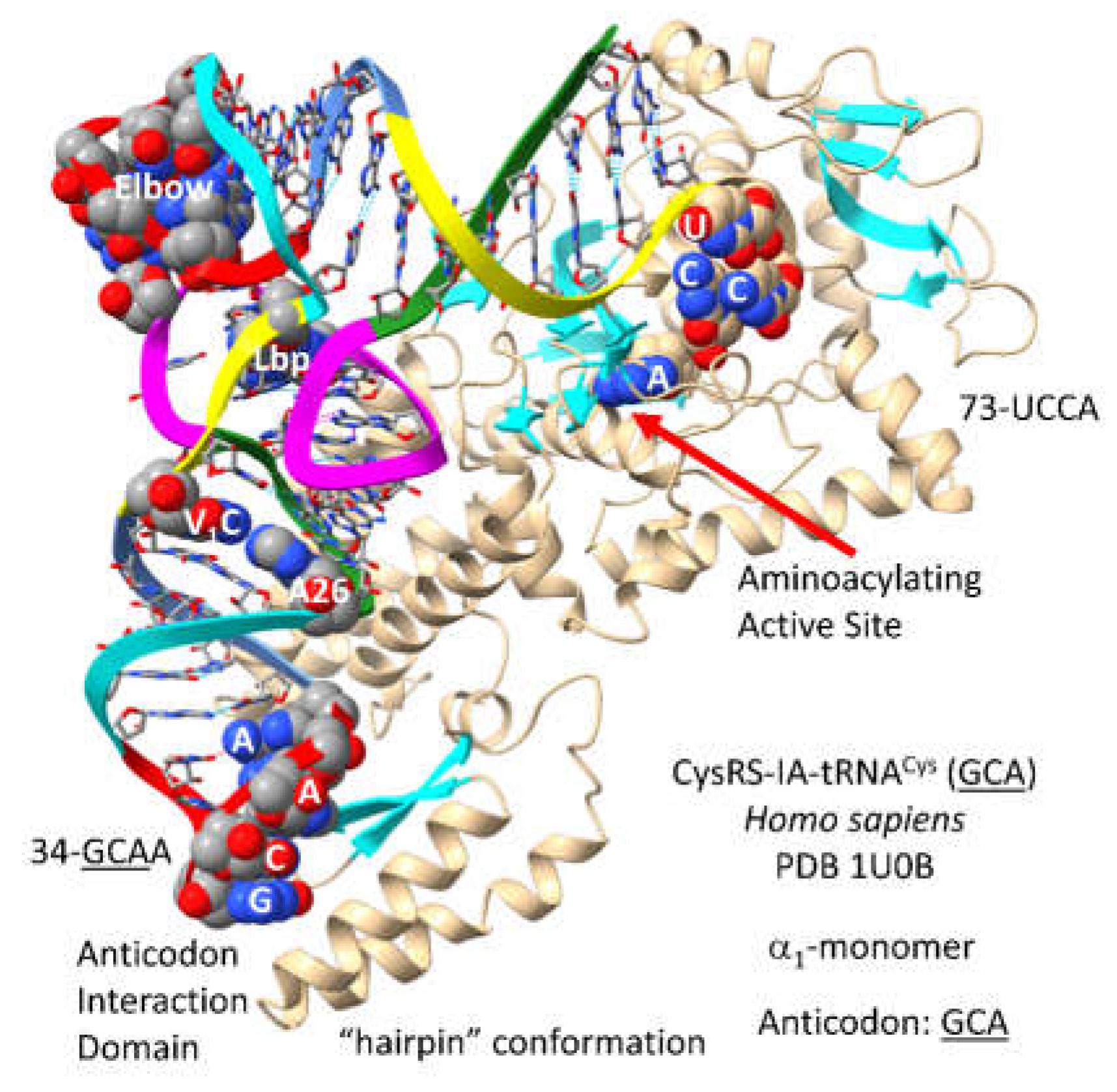
3.9. CysRS-IA
By sequence and structure, CysRS-IA (
Figure 15) is closely related to ArgRS-IA. Cysteine and arginine locate to column 4 of the genetic code, indicating evolution in code columns. Because CysRS-IA recognizes only the
GCA anticodon, 34-
GCA-36 can be recognized by the anticodon binding domain [
64]. In the structure, 73-UCCA-76 enters the CysRS-IA aminoacylating active site in the “hairpin” catalytic conformation. Discriminator 73U is rarely used. In
P. furiosus, 73U is only found in tRNA
Cys (1 tRNA
Cys) and tRNA
Thr (3 tRNA
Thr). The aminoacylating active site of CysRS-IA is at the C-terminal ends of a set of parallel -sheets, as expected. Cysteine is important for Zn binding. CysRS-IA utilizes Zn binding to bind and orient cysteine in its aminoacylating active site.
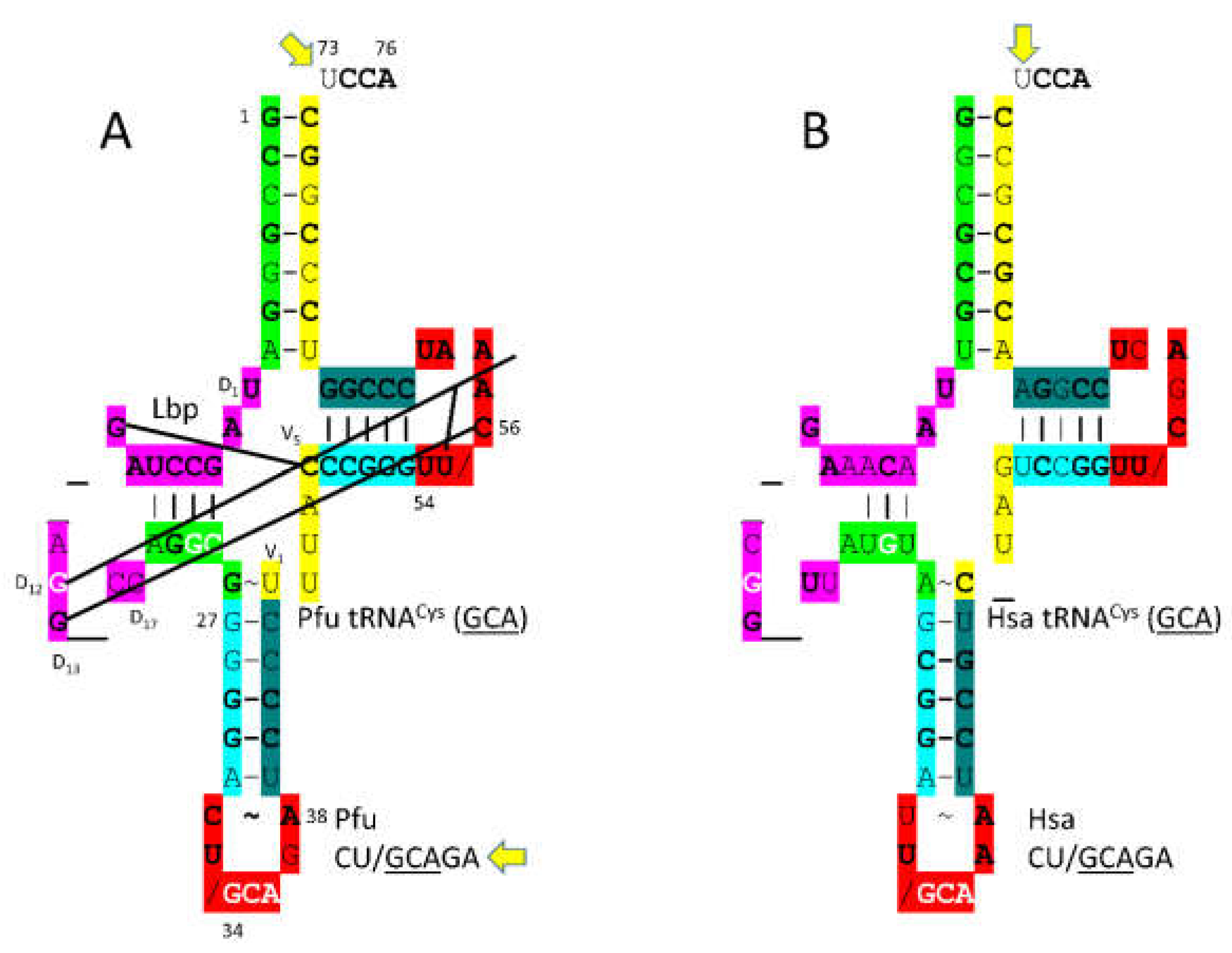
In
P. furiosus, tRNA
Cys (
GCA) is of interest (
Figure 16). The acceptor stem matches tRNA
Pri in 4 of 7 bp. The D loop sequence is D
1-UAGCCUAG__AGG__CC-D
17, matching the primordial tRNA sequence in the first 8 positions. The 5’-As* sequence is AGGCG, matching tRNA
Pri GGGCG in 4 of 5 positions. The anticodon stem matches tRNA
Pri in 2 bp. The T loop stem-loop-stem matches the primordial tRNA sequence exactly. Interestingly, for 34-
GCAG, the anticipated modified 37G is not present. The
P. furiosus tRNA
Cys (
GCA) is very similar to the human tRNA
Cys (
GCA), possibly indicating a monophyletic relationship between tRNA
Cys in Archaea and Eukarya.
Cysteine may have first entered the genetic code by tRNA-linked chemistry. There are two mechanisms by which Ser-tRNA
Cys might be converted to Cys-tRNA
Cys. pSer-tRNA
Cys can be converted to Cys-tRNA
Cys by pSer-tRNA
Cys→Cys-tRNA
Cys cysteine synthase (pSer for o-phosphoserine) [
65]. Serine can also be acetylated and then converted to cysteine with H
2S. It is hypothesized that serine jumping to column 4 of the genetic code from column 2 may have resulted from such a tRNA-linked mechanism. Cysteine ended up in column 4, row 1. Most row 1 amino acids (i.e., Phe, Tyr, Trp and Cys) appear to be among the last encoded. Cysteine, however, was important for Zn binding and protein folding (i.e., for AARS enzymes), indicating that cysteine must have entered the code earlier, before landing in its row 1 location [
15,
16,
17]. Serine may have occupied a larger sector of column 2 (i.e., rows 2 and 3). Serine or serine converted to cysteine may have jumped to row 4 (i.e., from column 2, row 3A (
GGU) to column 4, row 3A (
GCU)). Serine converted to cysteine could have shifted to column 4, row 1 (
GCU→
GCA), and CysRS-IA could have evolved from a primitive ArgRS-IA. In this manner, cysteine could have entered the code early with tRNA-linked synthesis but found its eventual position late.
GCU within a disrupted arginine sector would then have reverted to a serine anticodon. In column 2 of the code, Thr and Pro displaced Ser to its location in column 2, row 1. SerRS-IIA recognizes a type II tRNA
Ser, without anticodon recognition. A simple change in the tRNA
Ser anticodon might, therefore, be sufficient to achieve the jump from column 2 to column 4, but the change in the anticodon would not affect SerRS-IIA recognition. Serine split what was probably an enlarged arginine sector by jumping into column 4. The jumping of serine from column 2 to column 4 was some of the only chaos in generating the standard code.
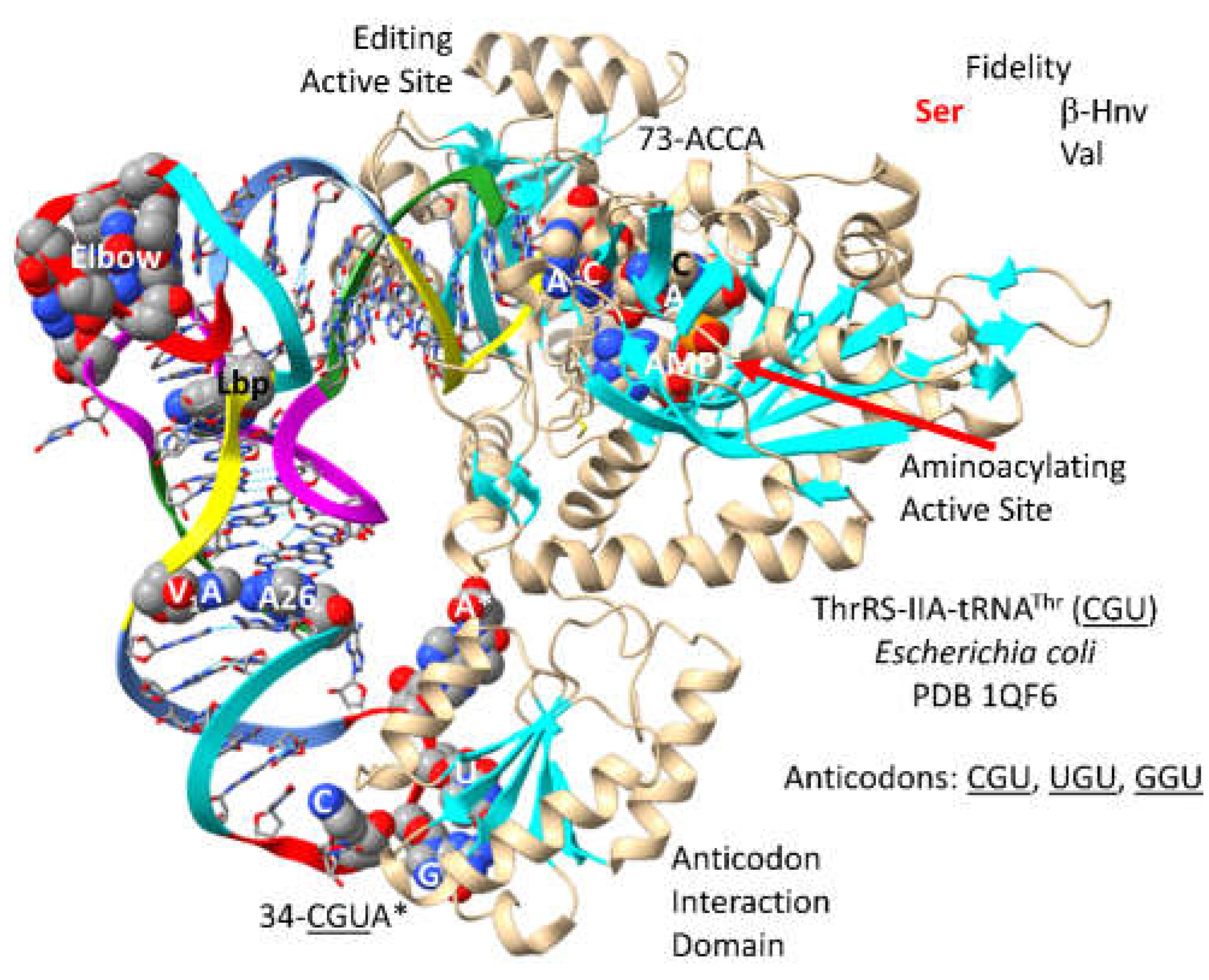
3.10. ThrRS-IIA
By structure and sequence, ThrRS-IIA (
Figure 17) is very similar to SerRS-IIA and GlyRS-IIA. As a class II AARS, ThrRS-IIA has its aminoacylating active site on a surface of antiparallel -sheets [
66]. 73-ACCA-76 penetrates the aminoacylating active site, where AMP binds. In
P. furiosus, the discriminator base is 73U rather than 73A, as in
E. coli. ThrRS-IIA has a separate editing active site that removes non-cognate -hydroxynorvaline and valine from tRNA
Thr. The aminoacylating active site of ThrRS-IIA limits non-cognate attachment of serine to tRNA
Thr. The anticodon binding region of ThrRS-IIA binds 35-
GU[m6t6A]-37 (in
E. coli).
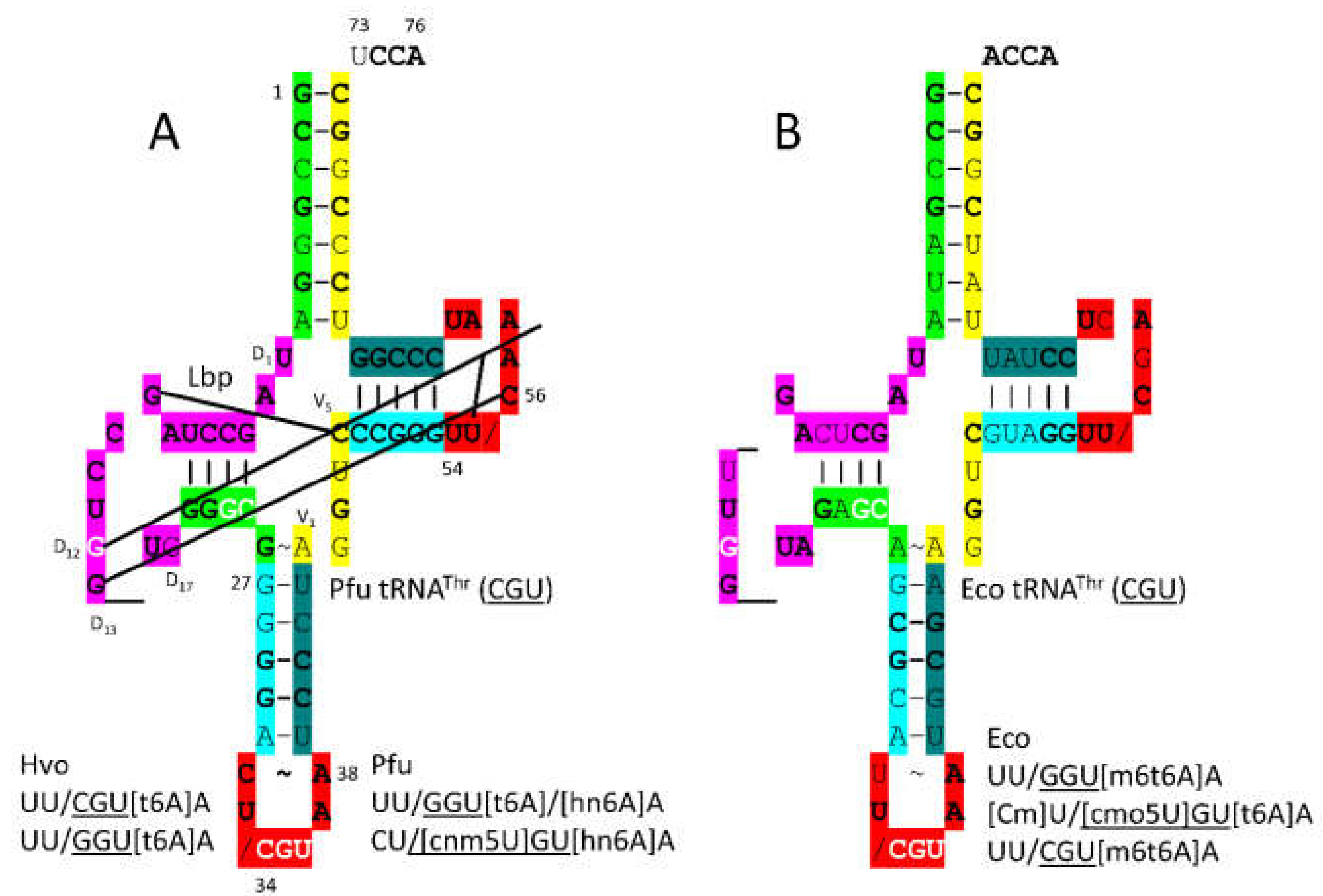
tRNA
Thr (
CGU) is shown in
Figure 18. In
P. furiosus, the acceptor stem matches the primordial tRNA sequence in 4 of 7 bp. The D loop has the sequence UAGCCUAGCCUGG__UG, which matches the primordial tRNA sequence in the first 13 positions exactly and in all but 3 positions, two of which are deletions. The 5’-As* sequence is GGGCG, which is typical. The anticodon stem matches tRNA
Pri in 2 bp. The V loop sequence AGGUC is typical. In
P. furiosus, the T stem-loop-stem matches the primordial tRNA sequence exactly. In Archaea and Bacteria,
36U is generally associated with a modified 37A (i.e., t6A or hn6A), as is observed. Modification of 37A may aid in accurate tRNA
Thr charging. Also, the modification of 37A may help to support the reading of
36U anticodons. In
P. furiosus, tRNA
Thr resembles tRNA
Ser in sequence, except for the V loop region (tRNA
Thr is type I; tRNA
Ser is type II) [
21].
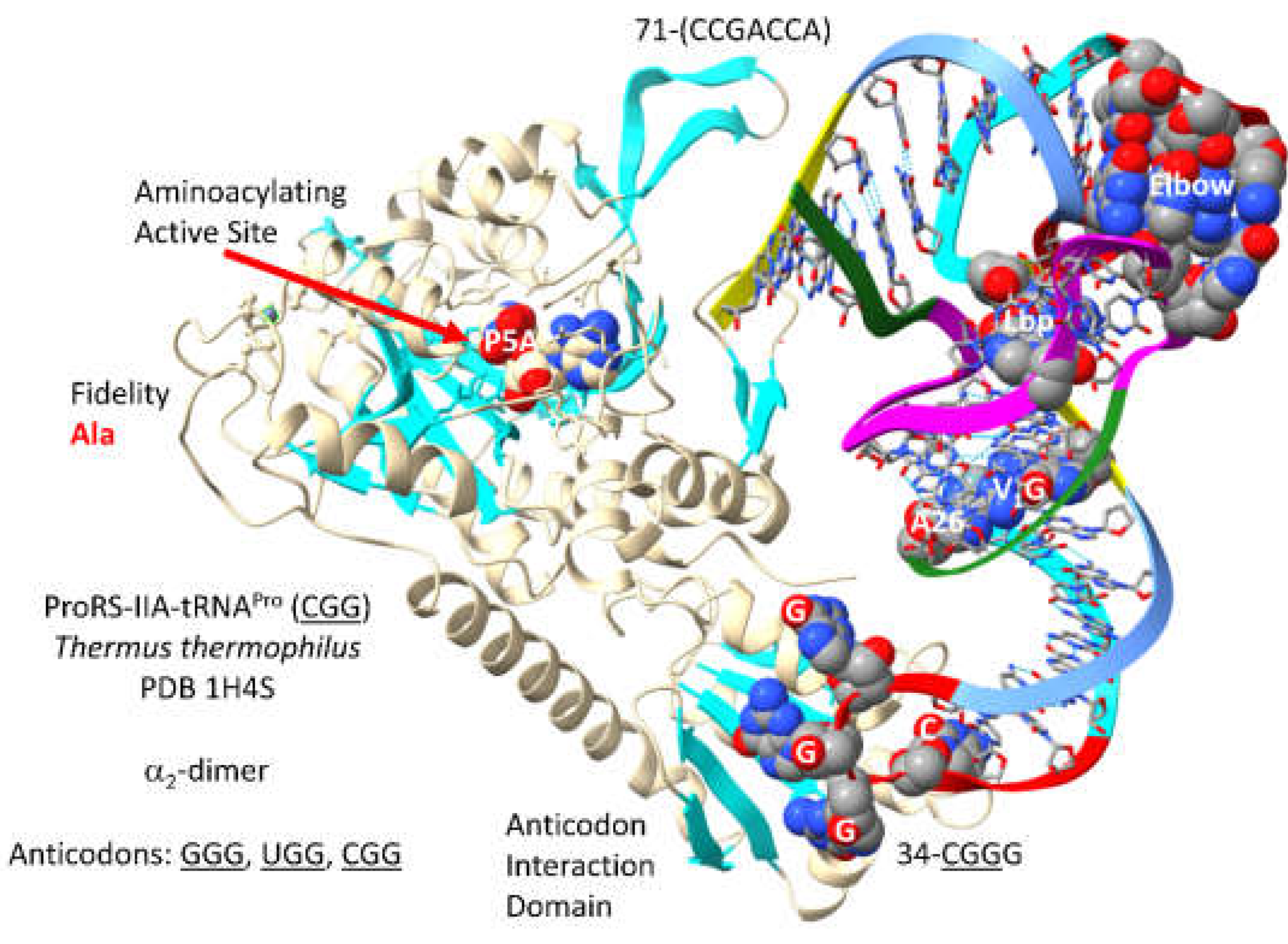
3.11. ProRS-IIA
ProRS-IIA-tRNA
Pro (
CGG) (
Figure 19) [
67] is closely related in sequence and structure to GlyRS-IIA, SerRS-IIA and ThrRS-IIA. The aminoacylating active site is on a surface of antiparallel -sheets. Reaction intermediate analogue P5A locates to the aminoacylating active site. In the structure 70-(CCGACCA)-76 is disordered. The anticodon loop 34-
CGGG-37 is substantially unwound indicating allosteric effects, which may also be indicated by disorder of the tRNA 3’-end. As expected, 35-
GGG-37 make the strongest ProRS-IIA binding contacts.
T. thermophilus and P. furiosus ProRS-IIA lack a separate editing active site that is, however, present in more derived Bacteria, such as E. coli. The aminoacylating active site of ProRS-IIA limits non-cognate alanine attachment to tRNAPro.
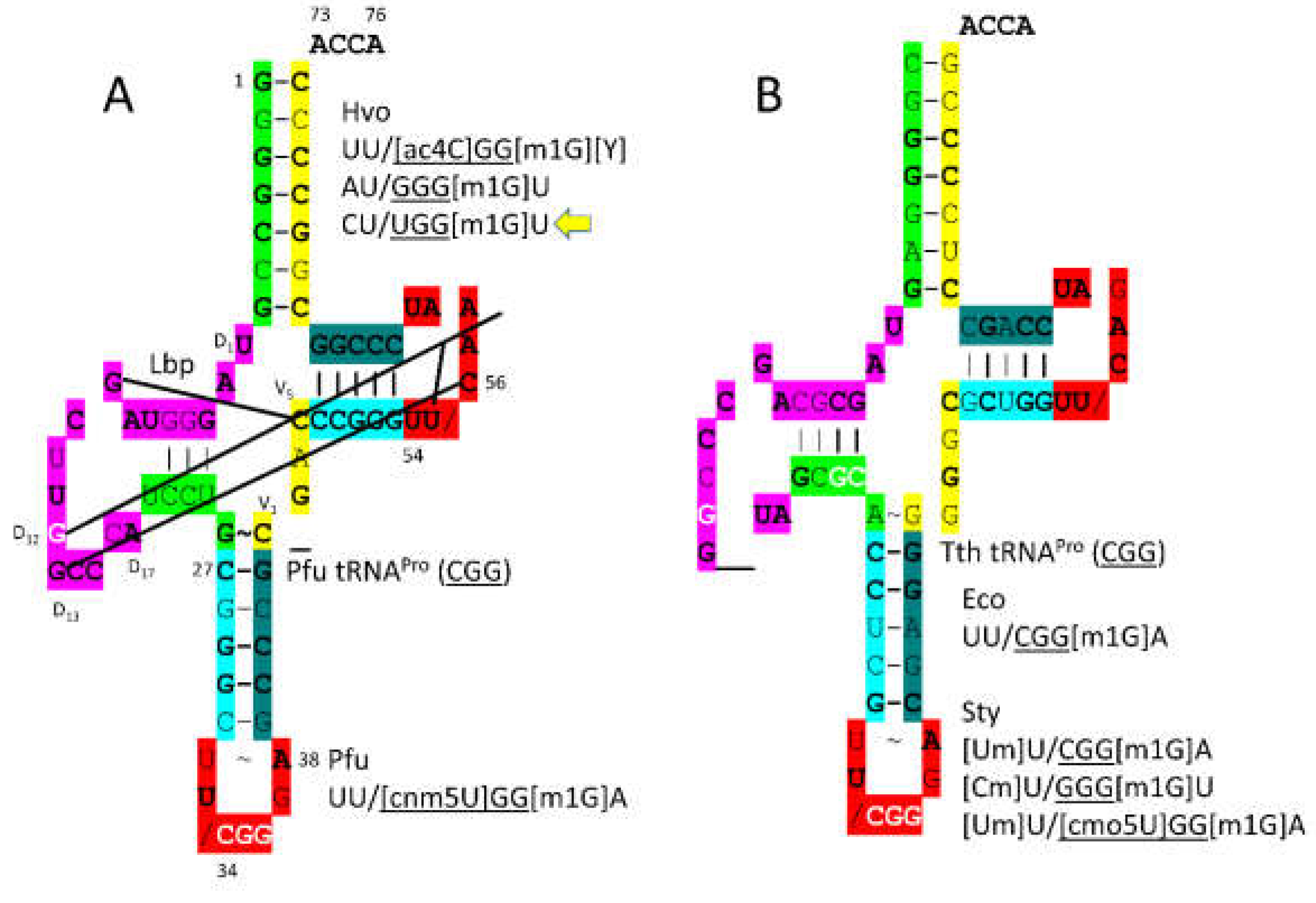
P. furiosus tRNA
Pro (
CGG) matches the acceptor stem of primordial tRNA in 5 of 7 positions (
Figure 20). tRNA
Pro (
CGG) has the D loop sequence D
1-UAGGGUAGCUUGGCCCA-D
17, which matches the primordial D loop except in 4 positions and has no deleted bases relative to tRNA
Pri. The anticodon stem matches the primordial sequence in 3 of 5 bp. The V loop sequence C_GAC matches the primordial sequence CCGCC in 3 positions. The T stem-loop-stem sequence matches the primordial tRNA sequence exactly. Proline is in a 4-codon box and, so, utilizes
CGG,
GGG and
UGG anticodons. Modifications are as expected, except that
H. volcanii 34U is unmodified. Because proline occupies a 4-codon box, superwobbling need not necessarily be suppressed. By contrast,
P. furiosus has the
34cnm5U modification.
3.12. AspRS-IIB
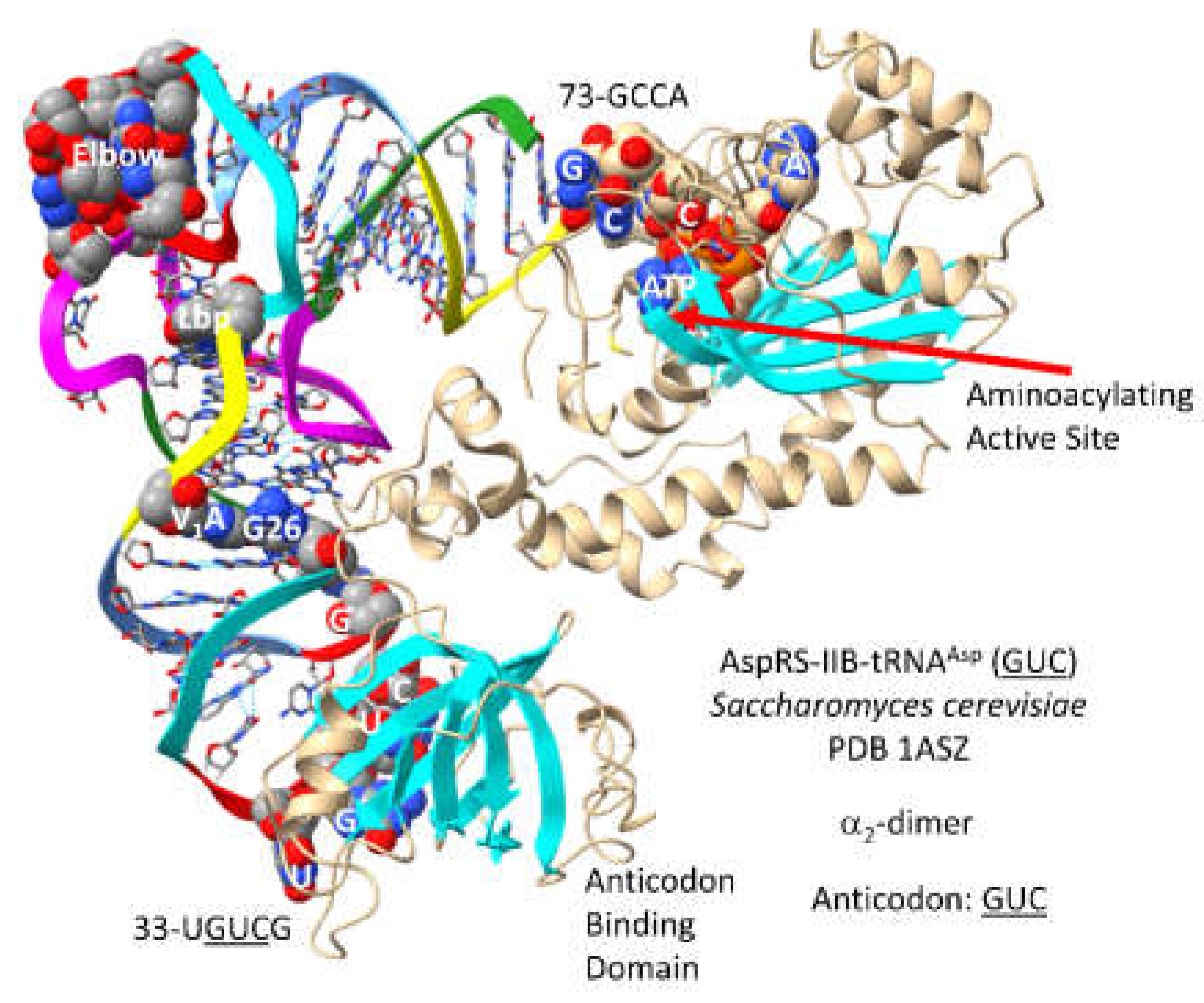
A primitive AspRS may be the founding AARS in column 3 of the code (tRNA-
35U). AspRS-IIB-tRNA
Asp (
GUC) of
S. cerevisiae is shown in
Figure 21 [
68]. Column 3 is the most innovated column, dividing into 2-codon sectors. For tRNA
Asp, only the
GUC anticodon is utilized. The anticodon loop is substantially unwound exposing 33-U
GUCG-38 to make AspRS-IIB contacts. Anticodon loop unwinding indicates allosteric effects communicated to the AspRS-IIB aminoacylating active site through tRNA
Asp (
GUC). 73-GCCA-76 enters the aminoacylating active site, where ATP binds. As expected, a surface of antiparallel -sheets is present at the aminoacylating active site.
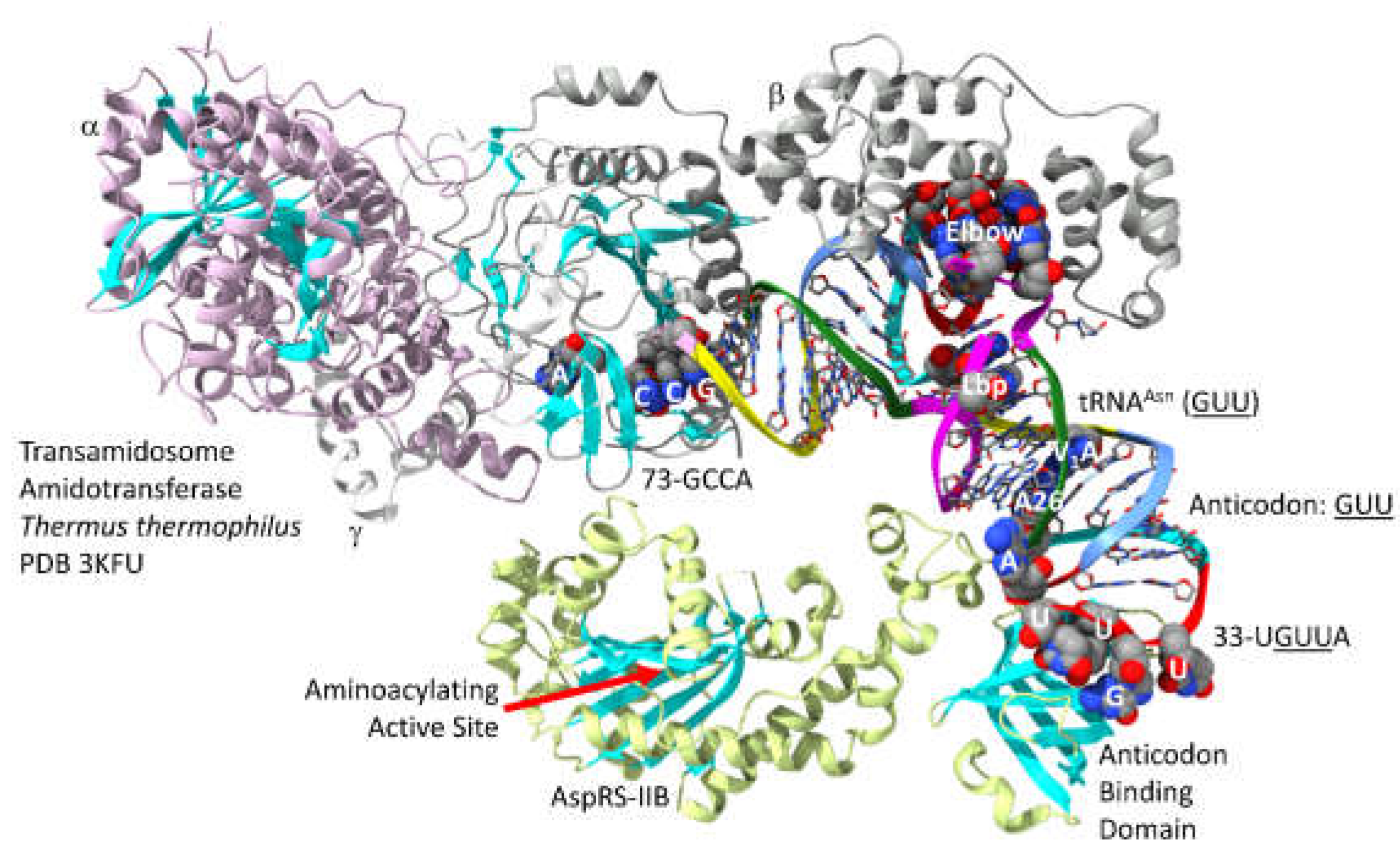
In
Figure 22, part of a
T. thermophilus transamidosome is shown [
69]. The image provides a partial approximation of the mechanism by which asparagine and glutamine may have first entered the genetic code [
70]. The -subunit of the amidotransferase that modifies Asp-tRNA
Asn to Asn-tRNA
Asn (
GUU) is homologous to an archaeal amidotransferase. Both asparagine and glutamine initially entered the code by tRNA-linked amidotransferase reactions. The tRNA
Asn (
GUU) anticodon loop is substantially unwound. 33-U
GUUA-37 interacts with the AspRS-IIB anticodon binding domain.
Asp and Glu are closely related negatively charged amino acids that locate to column 3, row 4. Asp has a shorter side chain than Glu and, so, generally forms better ion pair allosteric switches, particularly with Arg, which is stiffer than Lys. In
P. furiosus, tRNA
Asp, tRNA
Glu and tRNA
Gln are all closely related tRNAs by sequence [
21]. In
P. furiosus, tRNA
Asn is most similar to tRNA
Tyr. Deviation of tRNA
Asn from tRNA
Asp supports discrimination of chemically similar amino acids in coding.
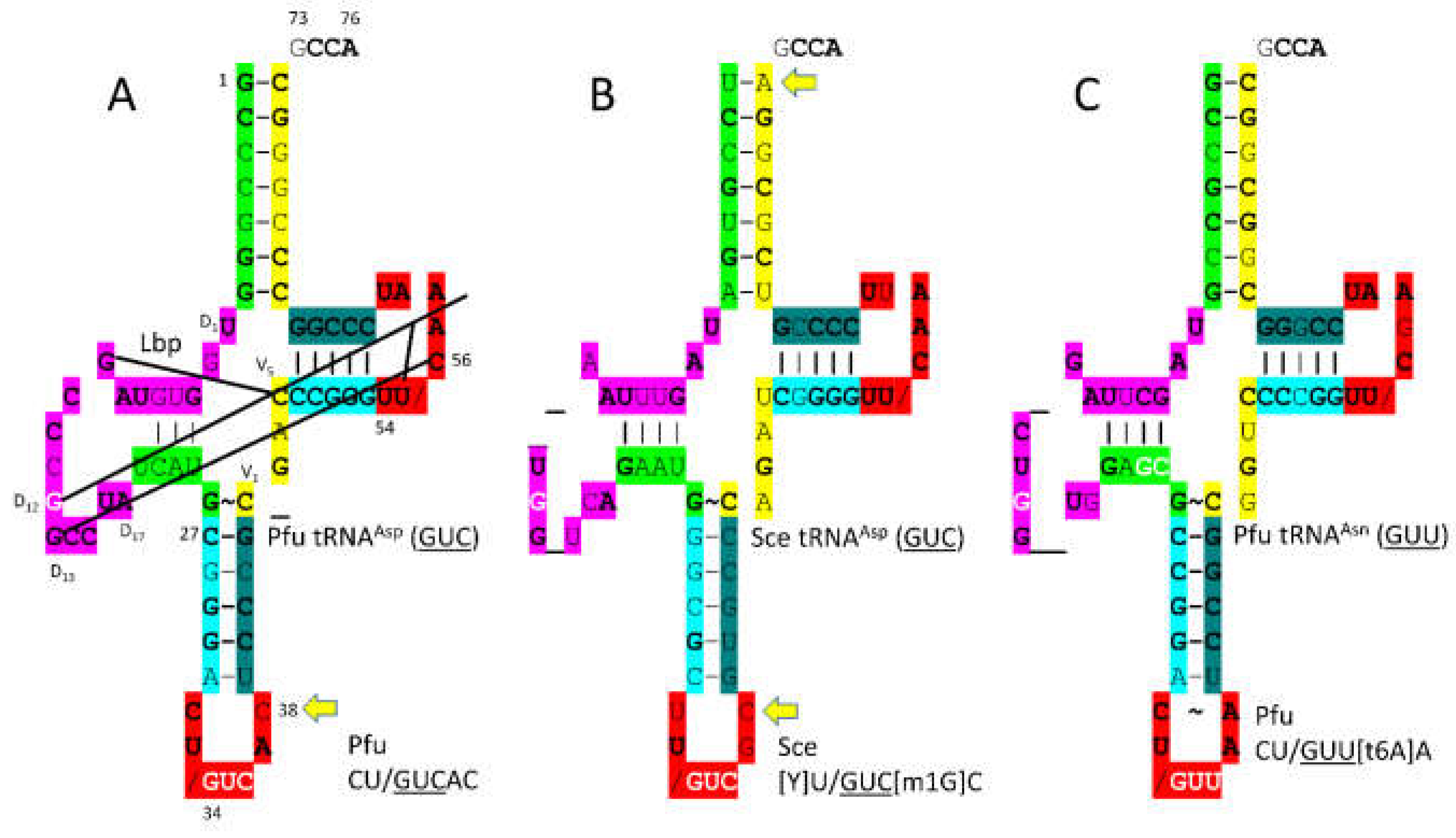
In
Figure 23, tRNA
Asp (
GUC) and tRNA
Asn (
GUU) are compared. It is hypothesized that tRNA
Asn (
GUU) evolved from tRNA
Asp (
GUC) and that AsnRS-IIB evolved from AspRS-IIB by duplication and divergence. The acceptor stem of
P. furiosus tRNA
Asp (
GUC) matches the primordial sequence at 4 of 7 bp. The D loop has the sequence D
1-UGGUGUAGCCCGGCCUA-D
17, which differs in 4 positions from the primordial tRNA but includes no deletions relative to tRNA
Pri. The D loop sequence D
6-UAGCCCGGCCUA-D
17 has only a single mismatch with tRNA
Pri. The anticodon stem matches tRNA
Pri in 3 bp. The T stem-loop-stem exactly matches the primordial tRNA sequence. The tRNA
Asp (
GUC) anticodon loop has a 32C-38C arrangement, which should alter the dynamics of the loop relative to 32C-38A, which is most common and primordial. The
P. furiosus tRNA
Asn (
GUU) matches the acceptor stem of the primordial tRNA in 5 of 7 bp. The D loop has the sequence D
1-UAGCUUAG_CUGG__UG-D
17, with 3 bases deleted from the primordial tRNA D loop but matching sequence in all but 5 positions. The 5’-As* sequence GAGCG matches the primordial sequence GGGCG in all but one base. The anticodon stem matches the primordial tRNA in 4 of 5 bp. The V loop sequence CGGUC matches tRNA
Pri CCGCC in 3 of 5 positions. The T stem-loop-stem matches in all but one stem bp and 1 loop base.
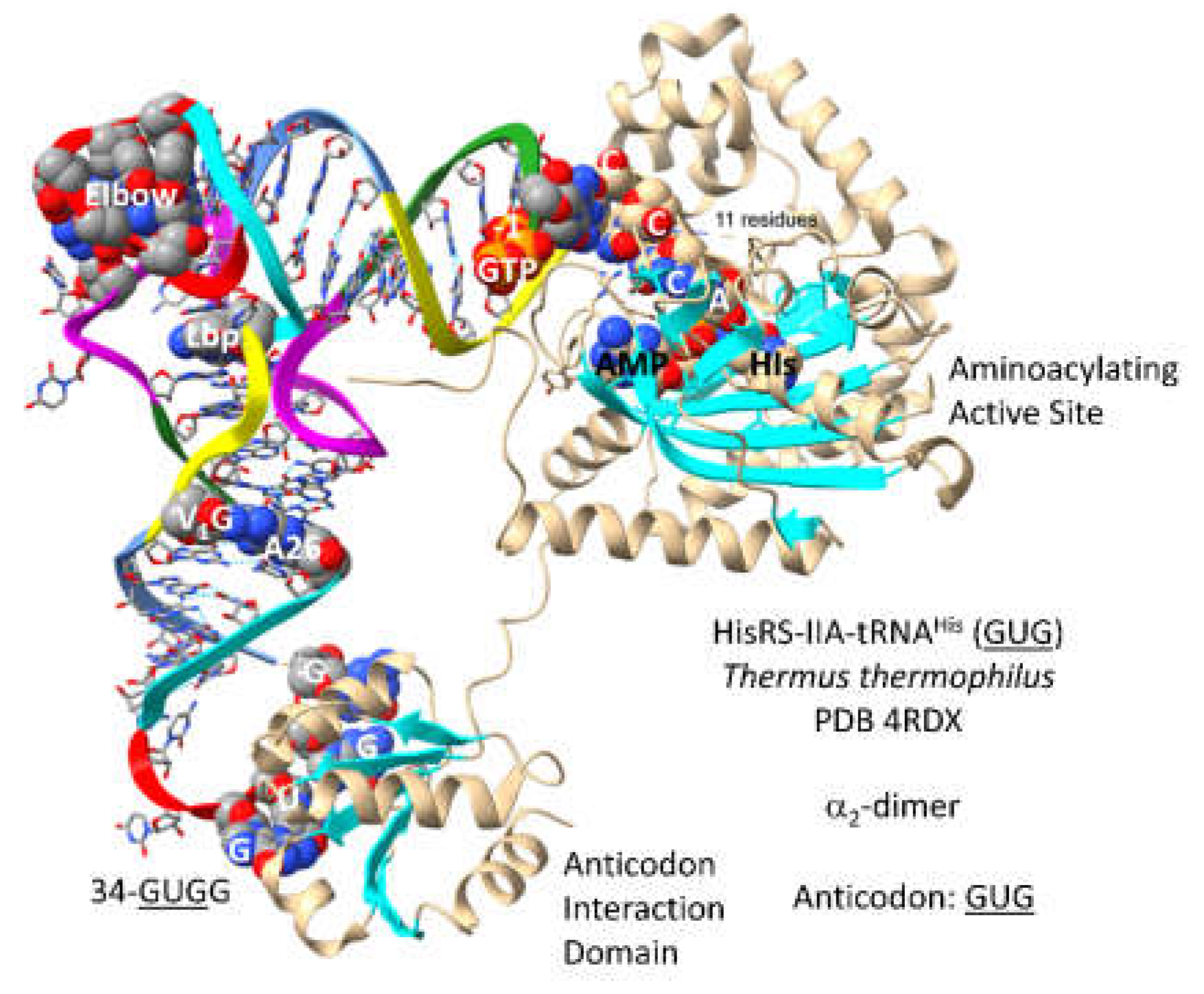
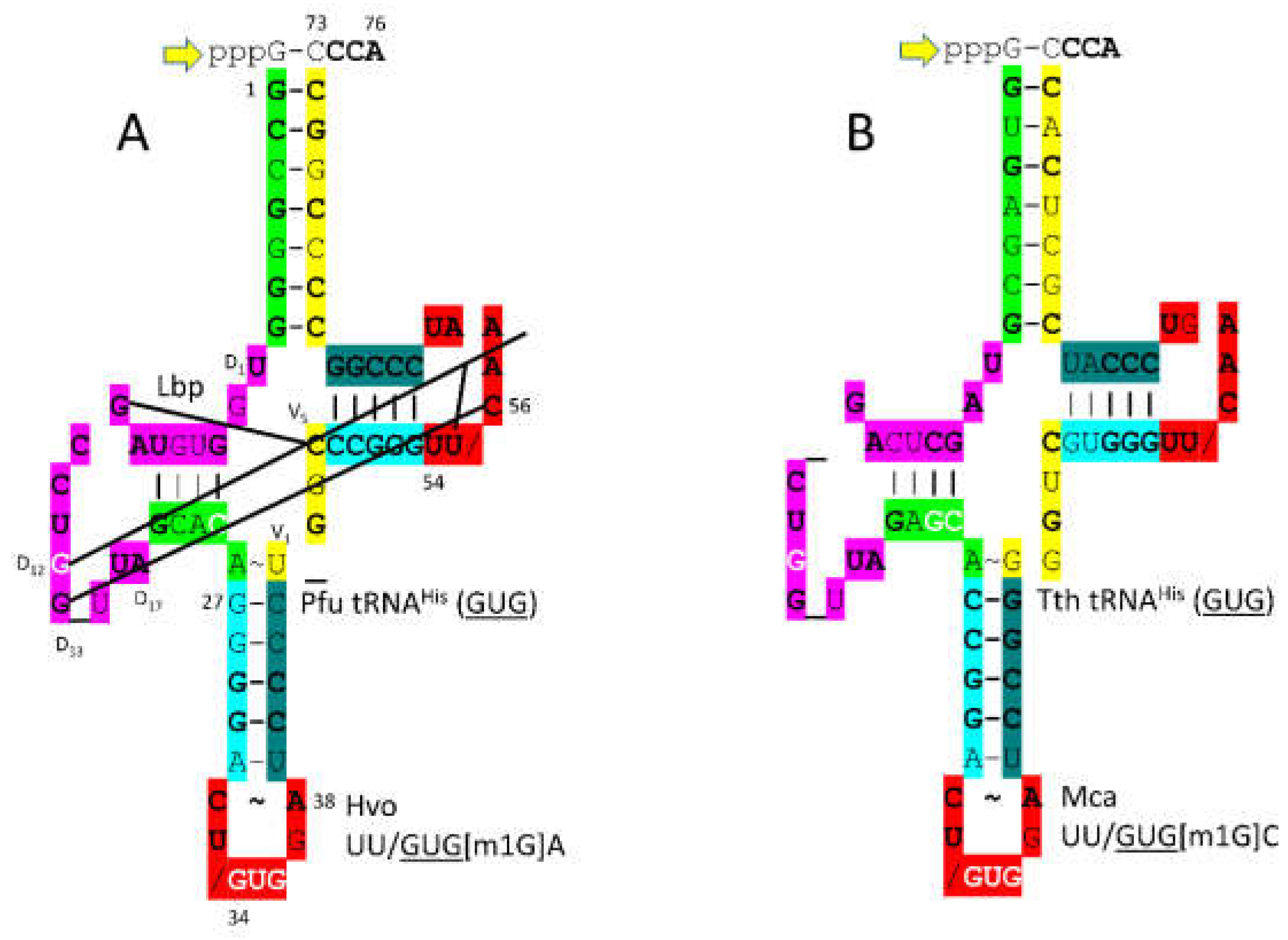
3.13. HisRS-IIA
Another column 3 amino acid is histidine. HisRS-IIA-tRNA
His (
GUG) from
T. thermophilus is shown in
Figure 24 [
71]. HisRS-IIA functions as an
2-dimer. As a class II AARS, HisRS-IIA has the aminoacylating active site on a surface of antiparallel -sheets. AMP and histidine bind in the aminoacylating active site, and 73-CCCA-76 enters the aminoacylating active site. On the ribosome, 74-CC-75 must pair with a GG sequence in the peptidyl site (P-site) of the peptidyl transferase center to orient the peptide-tRNA. Having the sequence 73-CCCA-76 in a tRNA, therefore, might cause problems with orienting the growing peptide chain during translation. To block 73C pairing with the ribosome G, tRNA
His (
GUG) is modified by addition of GTP at the -1 position. The enzyme that catalyzes this reaction is tRNA
His (-1) GTP transferase. This enzyme appears to be a first protein, as old as the genetic code. Also, the 73C=(-1)GTP base pair is a unique discriminator for cognate tRNA
His (
GUG) charging with histidine. As also for tRNA
Asp (
GUC) and tRNA
Asn (
GUU), the tRNA
His (
GUG) anticodon loop is unwound, exposing 34-
GUGG-37 to bind the HisRS-IIA anticodon binding domain. It is hypothesized that AspRS-IIB was originally AspRS-IIA, but diverged to suppress tRNA charging errors.
In
P. furiosus, the tRNA
His (
GUG) acceptor stem differs in only two bp from the primordial tRNA sequence (
Figure 25). the D loop of tRNA
His (
GUG) has the sequence D
1-UGGUGUAGCCUGG_UUA-D
17, differing in 5 positions from the primordial tRNA sequence with a single base deletion relative to tRNA
Pri. In
P. furiosus, the anticodon stem matches tRNA
Pri in 2 bp. In
T. thermophilus, the anticodon stem matches tRNA
Pri in 4 bp. In
P. furiosus, the T stem-loop-stem exactly matches the primordial tRNA sequence.
3.14. GluRS-IB
Column 3 of the genetic code is the most innovated column that encodes the most amino acids. It appears that column 3 may have sectored into 2-codon boxes initially by splitting Asp and Glu into a striped pattern of Asp in A rows (row 2A, 3A and 4A) and Glu in B rows (row 2B, 3B and 4B). A and B rows represent wobble tRNA-
34. tRNA-
34G is the anticodon base of the A row. At the base of code evolution, wobble tRNA-
34A is rarely or never used. tRNA-
34C or
34U is the anticodon base of the B row. Note that related amino acids and AARS enzymes Asp, Asn and His, charged to their cognate tRNAs by related enzymes AspRS-IIB, AsnRS-IIB and HisRS-IIA, locate to rows 4A, 3A and 2A. It is likely that AspRS was initially AspRS-IIA that evolved to AspRS-IIB to suppress translation errors. Glu, Lys and Gln locate to rows 4B, 3B and 2B. GluRS-IB and LysRS-IB (in Archaea) are closely related enzymes. GlnRS-IB was derived from GluRS-IB in Eukarya (~2.5 billion years ago) and then transferred to many prokaryotic species by horizontal gene transfers. At LUCA, GluRS-IB added glutamate to tRNA
Gln. Glu-tRNA
Gln was converted to Gln-tRNA
Gln by an amidotransferase. This is a similar tRNA-linked chemistry mechanism to that by which asparagine first entered the code [
70,
72,
73,
74].
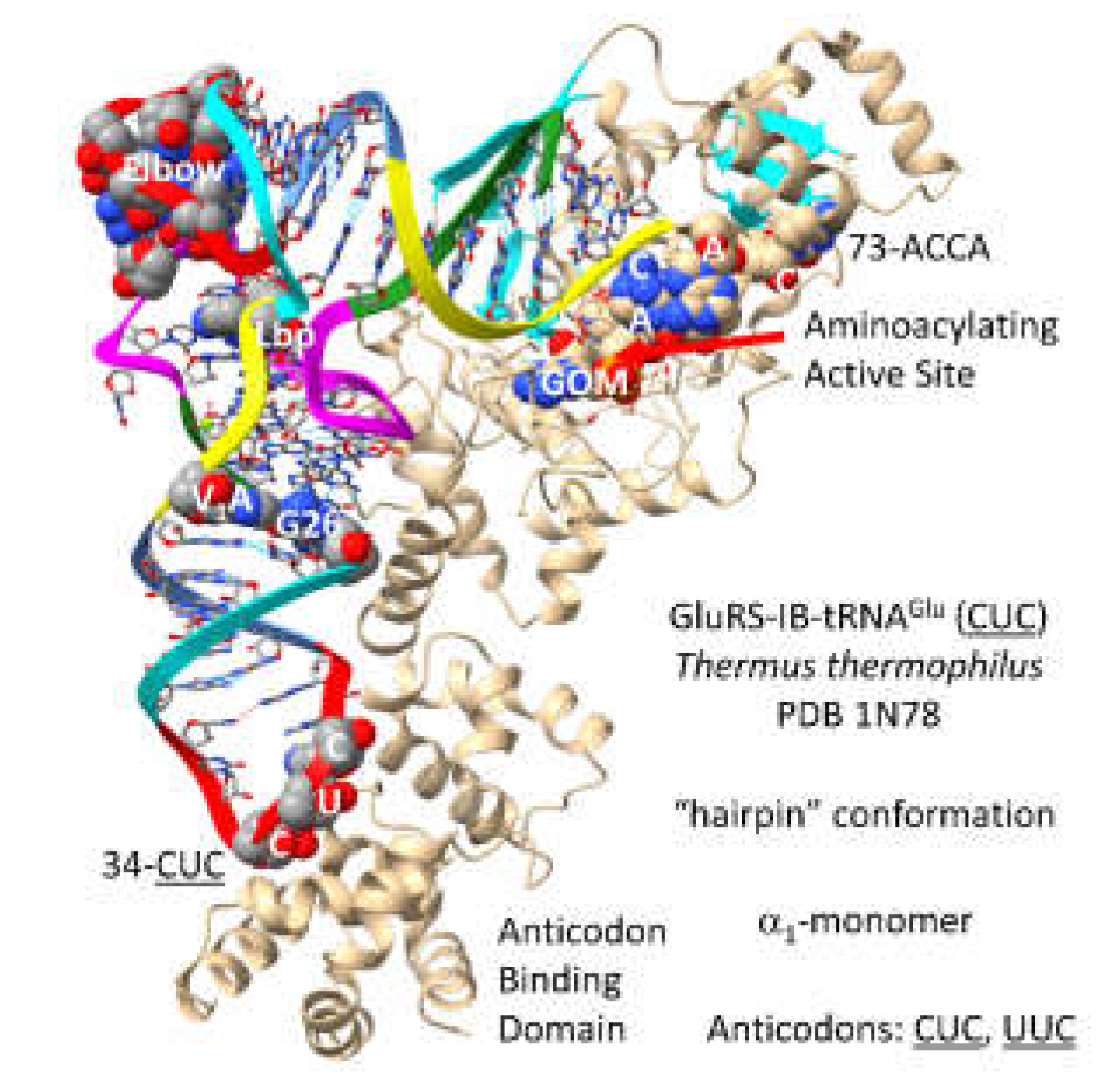
GluRS-IB (
Figure 26) [
75] may be derived from a primitive ArgRS-IA by duplication and repurposing. In contrast to AspRS-IIB, which is a class II AARS and an
2-dimer, GluRS-IB is a typical class I AARS that functions as an
1-monomer. The GluRS-IB aminoacylating active site is at the C-terminal ends of a set of parallel -sheets. 73-ACCA-76 penetrates to the aminoacylating active site in the catalytic hairpin conformation for class I AARS. The non-reactive GOM synthetic reaction intermediate binds here. In contrast to AspRS-IIB and AsnRS-IIB, the anticodon loop of tRNA
Glu is not substantially unwound. This difference and the difference of discriminator bases (73G (i.e., Asp) versus 73A (i.e., Glu)) may contribute to Asp versus Glu discrimination in cognate tRNA charging. 34-
CUC-36 binds the anticodon binding domain. Glutamate is a negatively charged amino acid with significant chemical character. No editing reactions are identified for GluRS-IB, consistent with the idea that glutamate is more readily discriminated by GluRS-IB than column 1 (Val, Leu, Ile, Met and Phe) and column 2 (Ala, Thr, Pro and Ser) amino acids that require cognate AARS enzymes that edit. Amino acids encoded in columns 3 and 4 have greater chemical character and less need of editing for error correction.
tRNA
Glu and tRNA
Gln are compared in Figure 27. In P. furiosus, tRNA
Glu (
CUC) (Figure 27A) is very close to the primordial tRNA sequence. The acceptor stem of tRNA
Glu (
CUC) varies by only 2 bp from tRNA
Pri. The D loop has the sequence D
1-UGGUGUAGCCCGGUCAA-D
17 differing from the primordial sequence in 6 positions but including no deletions relative to tRNA
Pri. By contrast, tRNA
Gln (
CUG) has two deletions from the primordial sequence in the D loop (Figure 27B). For tRNA
Glu (
CUC), the anticodon stem matches tRNA
Pri in 3 of 5 bp. For tRNA
Glu (
CUC) and tRNA
Gln (
CUG), the V loop has the sequence C_GAC, which matches the primordial sequence of CCGCC in 3 positions. Also, the V loop sequence C_GAC is found in tRNA
Asp (
GUC) (
Figure 23A), indicating that tRNA
Glu (
CUC) may be derived from tRNA
Asp, as might be expected. The T stem-loop-stem of tRNA
Glu (
CUC) is a perfect match to the primordial sequence. For tRNA
Gln (
CUG), the T stem-loop-stem sequence is slightly altered relative to tRNA
Pri. We note that tRNA
Gln (
CUG) has an unusual 1A=72U pair that is expected to separate more easily than 1G=72C in tRNA
Glu (
CUC) and many other P. furiosus tRNAs. Melting the 1A=72U pair in tRNA
Gln (
CUG) should contribute to discriminator function (i.e., in pre-life and until eukaryogenesis, for the Glu→Gln amidotransferase). In T. thermophilus, the tRNA
Glu (
CUC) is similar to P. furiosus but more derived from the root sequence, as expected. As mentioned above, tRNA
Asp, tRNA
Glu and tRNA
Gln are closely related sequences in P. furiosus.
Figure 27. tRNAGlu and tRNAGln. A) P. furiosus tRNAGlu (CUC). B) P. furiosus tRNAGln (CUG). C) T. thermophilus tRNAGlu (CUC). Sgr for Streptomyces griseus. xU indicates an unknown 5 carbon-U34 modification to suppress superwobbling. .
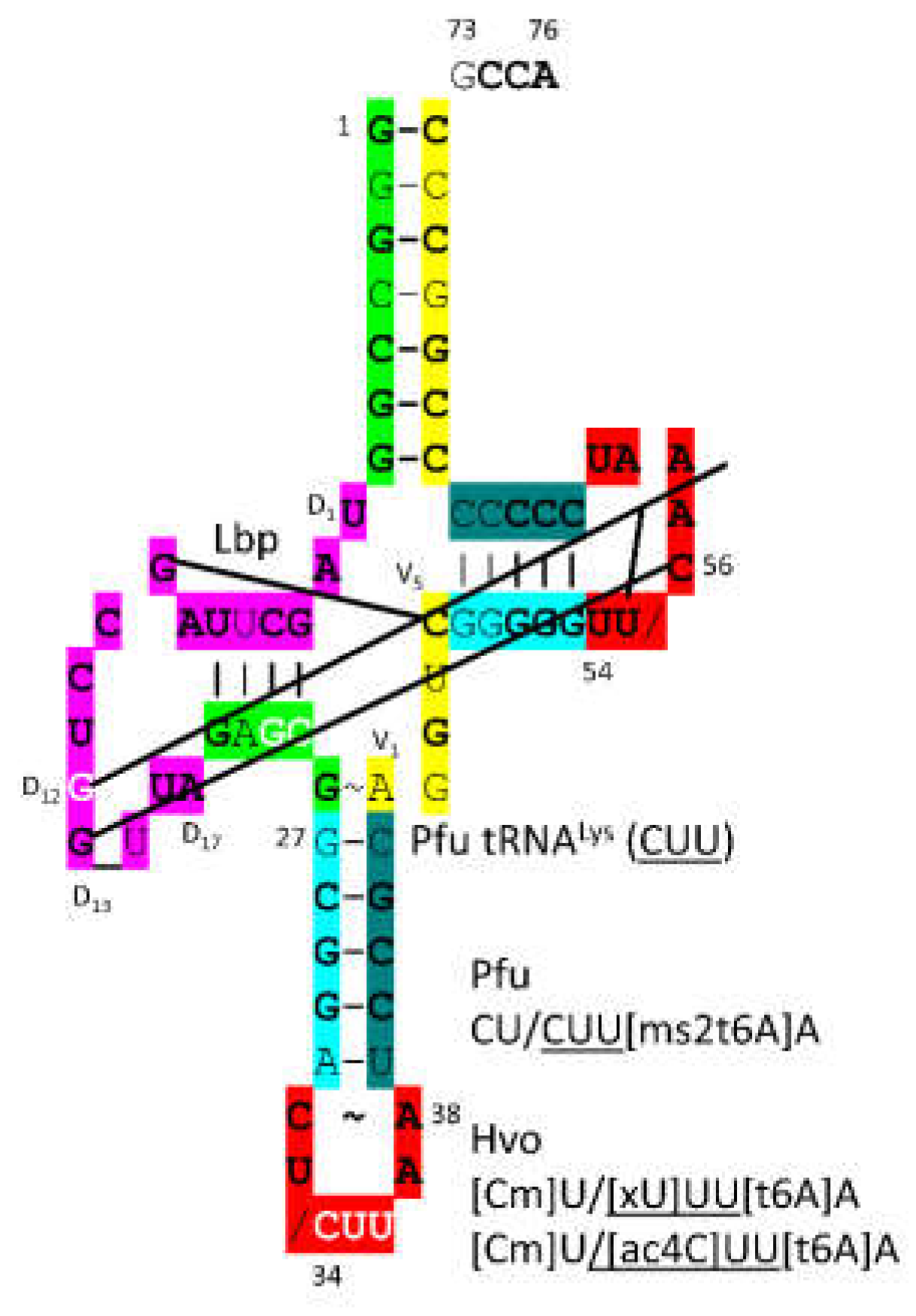
3.15. LysRS-IB
Currently, no suitable demonstration structure of archaeal LysRS-IB-tRNALys is available. Because of homology, we assume the structure would be similar to the image of GluRS-IB-tRNAGlu (CUC) (
Figure 26). LysRS-IB in Archaea appears to be the oldest LysRS. LysRS-IIB in Bacteria appears to be derived from AspRS-IIB, as a bacterial innovation. In Archaea, GluRS-IB, LysRS-IB (archaeal type) and GlnRS-IB (from Eukarya) are closely related AARS enzymes. In
Figure 28, a P. furiosus tRNA
Lys (
CUU) is shown. The acceptor stem sequence varies in only two bp from the primordial tRNA sequence. The D loop has the sequence D
1-UAGCUUAGCCUGG_UUA-D
17, differing in 3 positions from the primordial sequence, including a single base deletion relative to tRNA
Pri. The 5’-As* sequence GAGCG differs in only one position from the typical primordial sequence GGGCG. The anticodon stem matches tRNA
Pri in 3 of 5 bp. The type I V loop sequence AGGUC is typical. The T stem-loop-stem matches the primordial sequence in all but 2 stem bp. The modifications of the anticodon loop are as expected. In P. furiosus, tRNA
Lys is most similar to tRNA
Phe and somewhat similar to tRNA
Arg. Lysine and arginine are positively charged amino acids and may both be derived from ornithine by pre-life metabolism [
60].
3.16. AlaRS-IID
Alanine is proposed to be the founding amino acid for column 2 of the genetic code. It is hypothesized that AlaRS-IID may have replaced a now extinct AlaRS-IIA before LUCA, so there may be no sequence record of an earlier AlaRS-IIA. Homology comparing a IID and a IIA AARS is difficult to discern, so these are very different enzymes. The reason for the replacement may be to discriminate alanine, serine, threonine and proline. Column 2 of the genetic code includes SerRS-IIA, ThrRS-IIA and ProRS-IIA, indicating evolution in code columns. Column 2 of the code is divided into all 4-codon boxes.
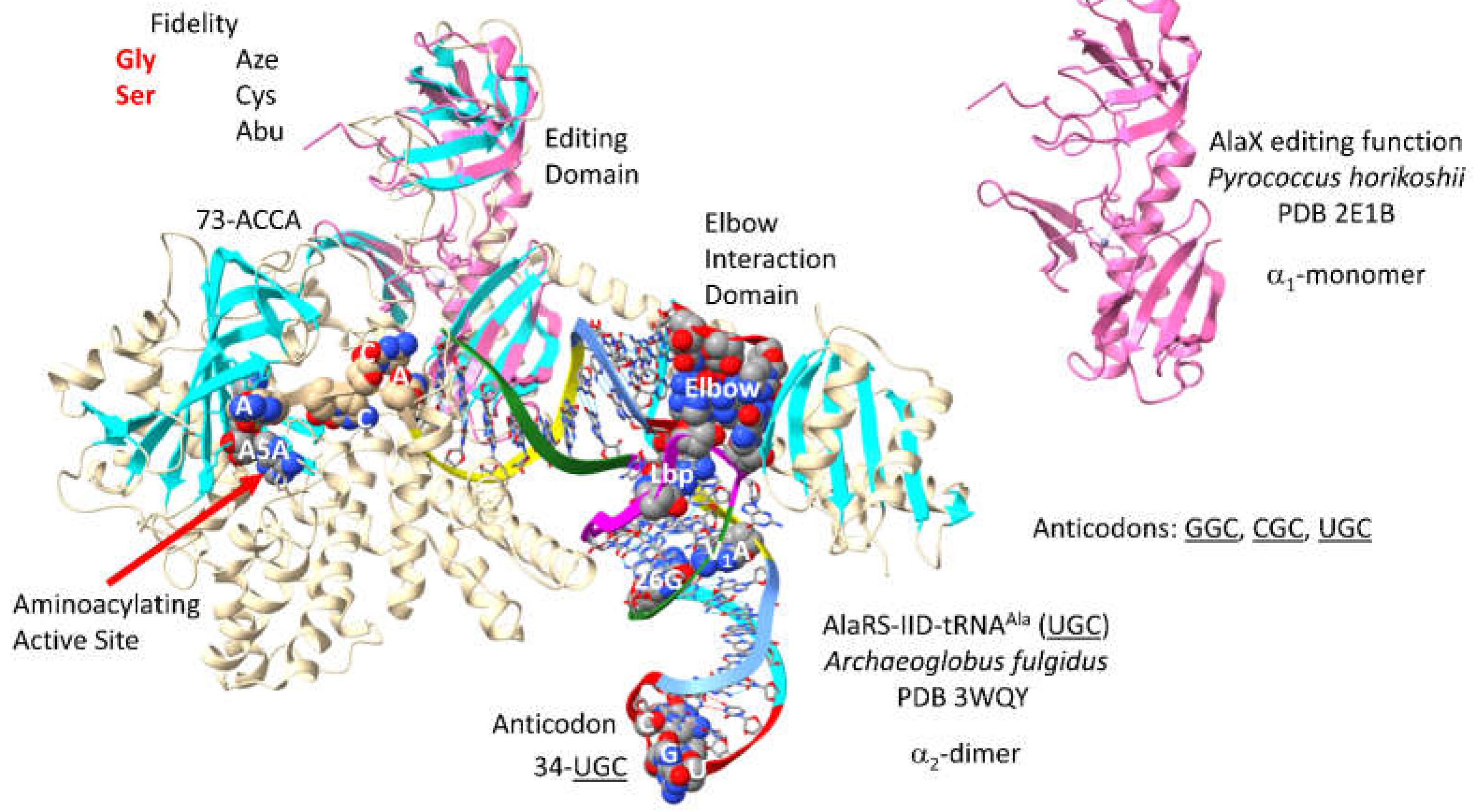
AlaRS-IID of Archaeon
Archaeoglobus fulgidus is shown in
Figure 29 [
76]. AlaRS-IID functions as an
2-dimer. The image is only half of the protein. Interestingly, although alanine locates to a 4-codon sector, AlaRS-IID makes no contacts to the tRNA
Ala anticodon loop. AlaRS-IID makes extensive elbow contacts, which may indicate tRNA
Ala distortion and allosteric effects of tRNA
Ala binding. 73-ACCA-76 penetrates the aminoacylating active site, which is also identified by a surface of antiparallel -sheets and A5A reaction intermediate analogue binding. AlaRS-IID includes a separate editing active site that removes non-cognate
azetidine-2-carboxylic acid, cysteine and -aminobutyrate from tRNA
Ala. The aminoacylating active site of AlaRS-IID limits non-cognate glycine and serine attachment to tRNA
Ala. In Archaea, a separate AlaX editing enzyme is also present that can remove non-cognate amino acids from tRNA
Ala [
77]. In the image shown, AlaX is light pink and was overlaid with the AlaRS-IID structure to locate the editing active site domain. AlaX may partly compensate for the lack of anticodon recognition by AlaRS-IID. Ala is a small hydrophobic amino acid with little chemical character, which may explain why AlaRS-IID has editing functions including the trans AlaX editing function.
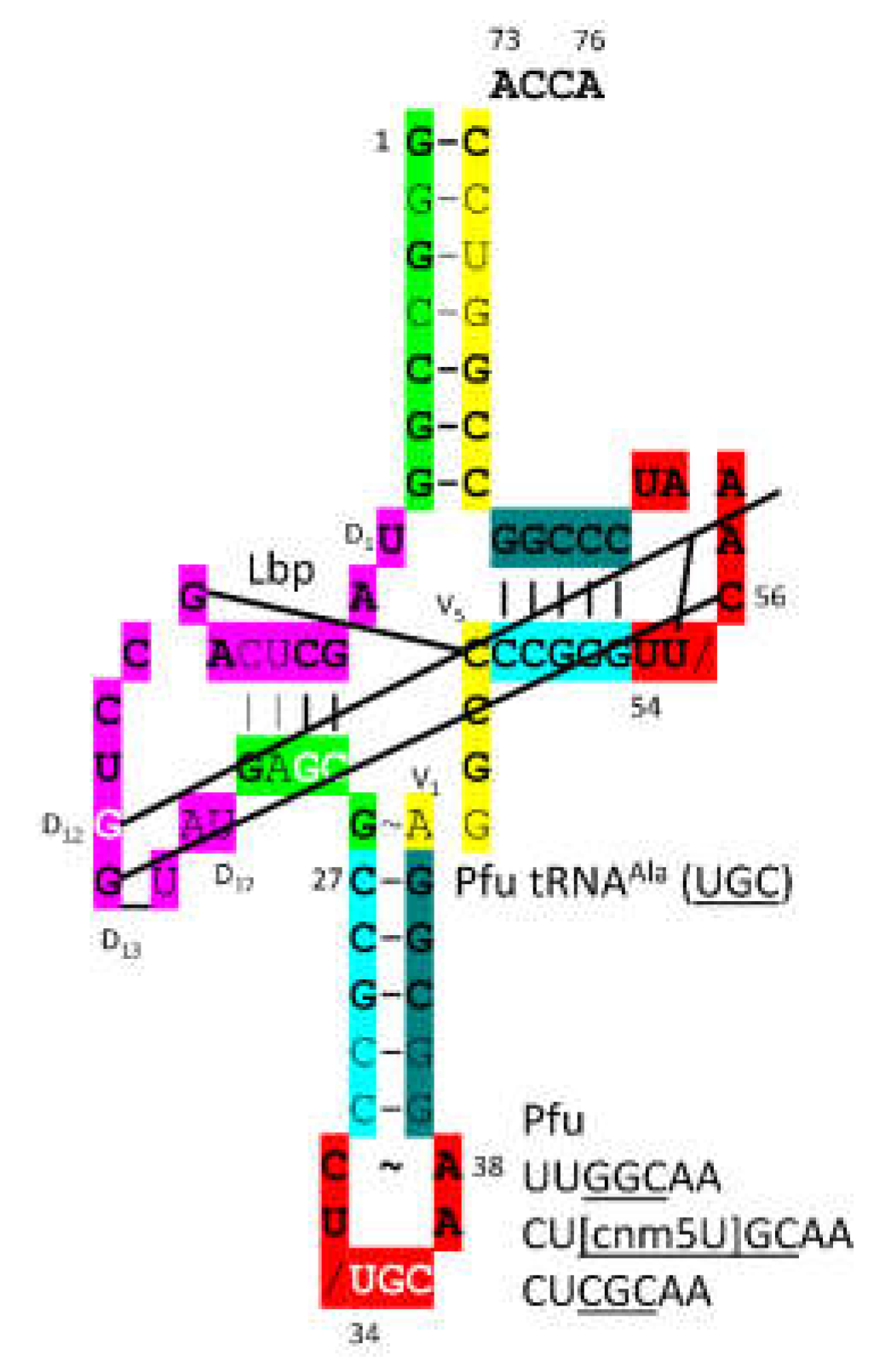
P. furiosus tRNA
Ala (
UGC) is shown in
Figure 30. The acceptor stem of tRNA
Ala (
UGC) varies in only two bp from the primordial tRNA sequence. The D loop has the sequence D
1-UAGCUCAGCCUGG_UAU-D
17, matching the primordial sequence in all but 6 positions with a single base deleted from tRNA
Pri. The 5’-As* sequence has the sequence GAGCG versus typical GGGCG. The anticodon stem matches tRNA
Pri in 3 of 5 bp. The V loop has the sequence AGGCC versus CCGCC for primordial tRNA and AGGUC for typical tRNA. The T stem-loop-stem matches the primordial tRNA.
P. furiosus tRNA
Ala has the appearance of an ancient tRNA, consistent with GADV being the first four amino acids in the code. As noted above,
P. furiosus tRNA
Ala is similar in sequence to tRNA
Val, consistent with alanine and valine being early additions to the code.
3.17. PheRS-IIC
It is hypothesized that aromatic amino acids (Phe, Tyr and Trp) entered the genetic code as some of the last amino acids added, in disfavored row 1 (tRNA-
36A) [
78]. It is suggested that row 1 (tRNA-
36A) was disfavored because, initially, both the tRNA-
34 and tRNA-
36 positions of the anticodon were wobble positions. During evolution of the code, tRNA-
34 remained a wobble position, but wobbling at tRNA-
36 was suppressed. Wobbling at tRNA-
36 was suppressed, in part, by modification of tRNA-37. Notably, if tRNA-
36U is present, generally, tRNA-37A is modified (i.e., t6A or hn6A). If tRNA-
36A is present, generally, tRNA-37G is modified (i.e., m1G). tRNA-37t6A may be more effective at suppressing tRNA-
36U wobbling compared to the efficacy of tRNA-37m1G at suppressing tRNA-
36A wobbling. In contrast to tRNA-
36, tRNA-
34 remained a wobble position. For one thing, modification of tRNA-33U cannot alter tRNA-
34 reading, because tRNA-33U is on the opposite side of the anticodon loop U-turn. At the base of code evolution, tRNA-33 is always U. Also, tRNA-
35 is a Watson-Crick position that cannot be modified in any way that interferes with coding. In evolution, tRNA-
34 wobbling could not be suppressed.
The following model is proposed for PheRS evolution. The initial PheRS may have been PheRS-IC derived distantly from a primitive ArgRS-IA or GluRS-IB. As TyrRS-IC and TrpRS-IC differentiated, there was insufficient discrimination between phenylalanine and tyrosine. PheRS-IC was then replaced by PheRS-IIC, before LUCA, leaving no sequence trace of PheRS-IC, except for TyrRS-IC and TrpRS-IC.
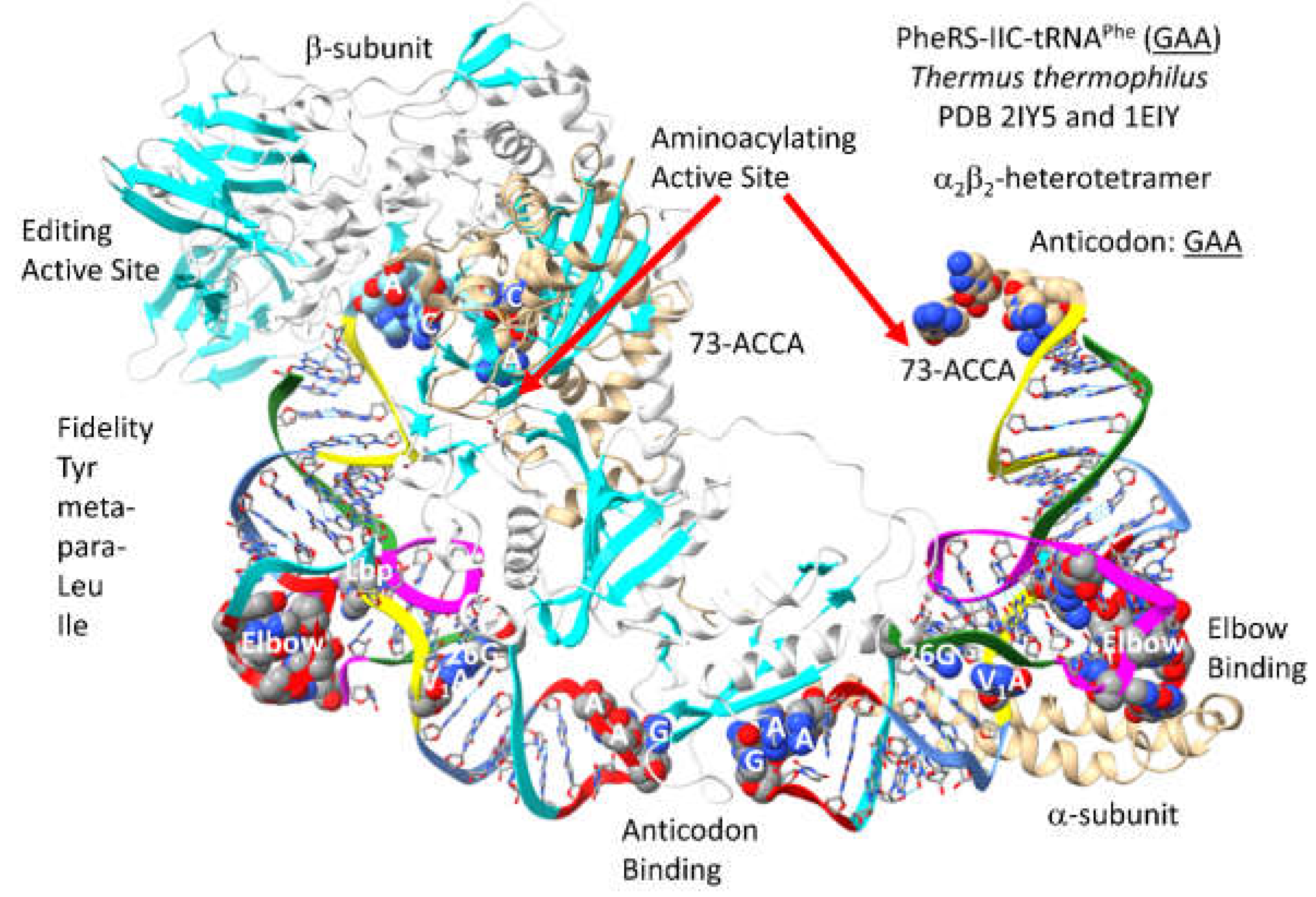
A detail of PheRS-IIC-tRNA
Phe (
GAA) from
T. thermophilus is shown in
Figure 31 [
79,
80]. PheRS-IIC in
T. thermophilus functions as an
2
2-dimer, which is also the archaeal form. Only one -unit is shown. To observe the relevant tRNA contacts, the two tRNA
Phe (
GAA) are visualized. The aminoacylating active site is a surface of antiparallel -sheets in the -subunit. 73-ACCA-76 penetrates to the aminoacylating active sites. The separate editing active site is within the -subunit. PheRS-IIC removes non-cognate tyrosine, meta- and para-substituted phenylalanine derivatives, leucine and isoleucine from tRNA
Phe (
GAA). An extrusion of the -subunit makes elbow contact. An extrusion of the -subunit makes anticodon contacts.
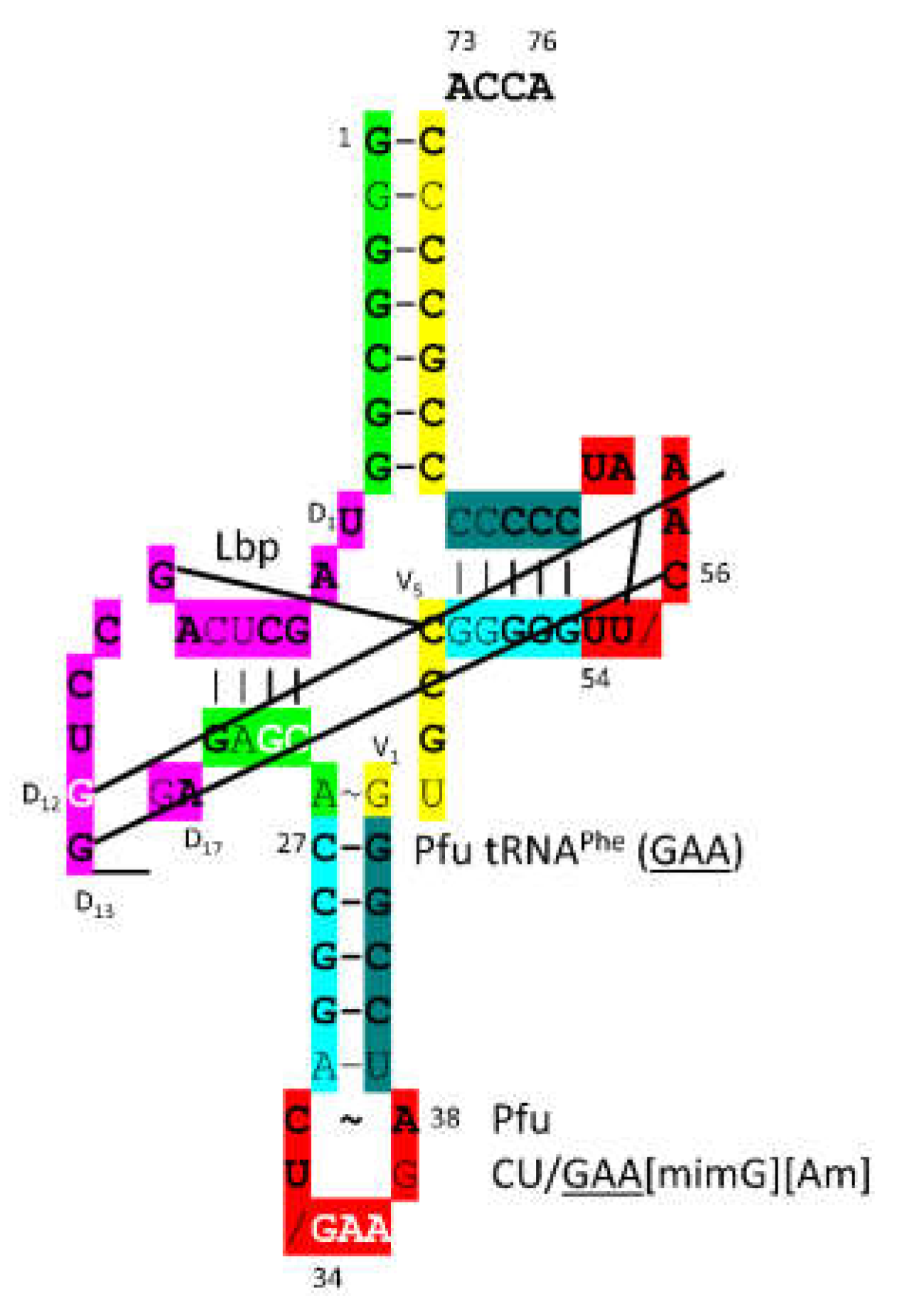
P. furiosus tRNA
Phe (
GAA) is shown in
Figure 32. The acceptor stem matches the primordial sequence in all but a single base pair. The D loop has the sequence D
1-UAGCUCAGCCUGG__GA-D
17, matching the primordial sequence in all but 5 positions with two bases deleted relative to tRNA
Pri. The 5’-As* sequence GAGCA matches the primordial sequence GGGCG in 3 positions. The anticodon stem matches the primordial sequence in 4 of 5 positions. The V loop sequence GUGCC matches primordial CCGCC in 3 positions. The T stem-loop-stem is very similar to the primordial sequence. Interestingly, tRNA
Phe (
GAA) appears to be a relatively early tRNA, although phenylalanine appears to be a later entry into the code. In
P. furiosus, tRNA
Phe (GAA) is closely related in sequence to tRNA
Lys (
UUU) and (
CUU).
3.18. TyrRS-IC
It is hypothesized that aromatic amino acids were a late addition to the genetic code along disfavored row 1 (tRNA-
36A). In evolution, TyrRS-IC and TrpRS-IC may be derived from a primitive ArgRS-IA or GluRS-IB. In contrast to most class I AARS, which are
1-monomers, TyrRS-IC and TrpRS-IC are obligate
2-dimers, with the anticodon binding and the aminoacylating active site for a single cognate tRNA in separate -subunits. TyrRS-IC-tRNA
Tyr (
GUA) from Archaeon
Methanocaldococcus jannaschii is shown in
Figure 33 [
81]. Aminoacylating active sites are at the C-terminal ends of a set of parallel -sheets. Tyrosine is bound at the aminoacylating active sites. 73-A(CCA)-76 is partly disordered in the structure. The anticodon 34-
GUA-36 contacts the anticodon interaction domain. One tRNA
Tyr (
GUA) is white and mostly obscured in the image.
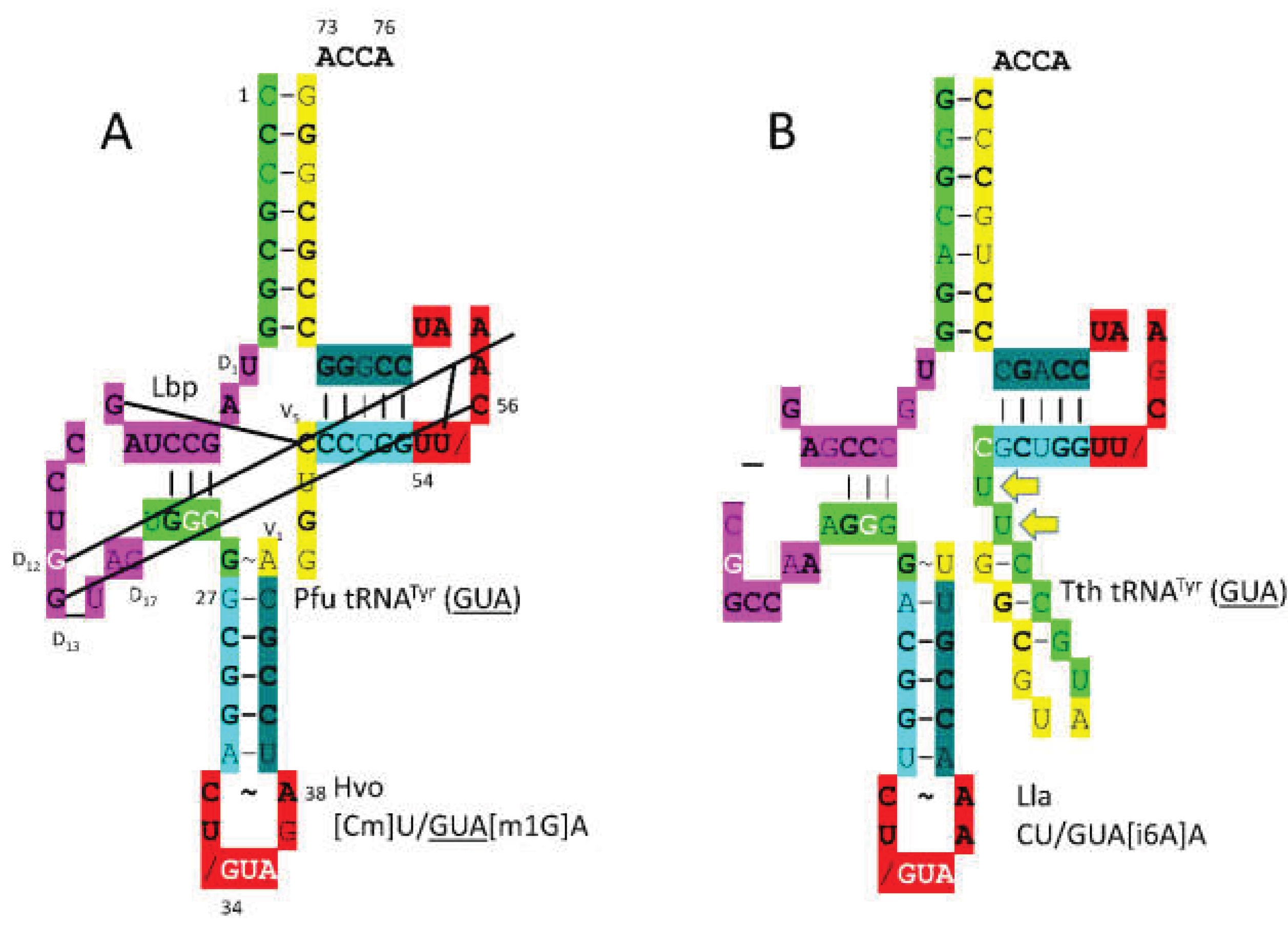
In
Figure 34A,
P. furiosus tRNA
Tyr (
GUA) is shown. The acceptor stem matches the primordial tRNA in all but 2 base pairs. The D loop sequence is D
1-UAGCCUAGCCUGG_UAG-D
17, matching the primordial sequence in all but 4 positions with a single base deleted relative to tRNA
Pri. Consistent with the three 31 nt minihelix tRNA evolution theorem, the D loop sequence begins with two perfect UAGCC repeats. The 5’-As* sequence is UGGCG, matching the typical primordial sequence GGGCG in all but a single base. The anticodon stem matches tRNA
Pri in 3 of 5 bp. The type I V loop is typical AGGUC. The T stem-loop-stem matches the primordial sequence in all but a single stem base pair. In Archaea, tRNA
Tyr (
GUA) is a type I tRNA. In Bacteria, by contrast, tRNA
Tyr (
GUA) is a type II tRNA [
11] (
Figure 34B). The difference appears to be a bacterial innovation.
3.19. TrpRS-IC
TrpRS-IC (
Figure 35) [
82] is a very similar enzyme to TyrRS-IC. In the TrpRS-IC-tRNA
Trp (
CCA) structure, 73-ACCA-76 enters the aminoacylating active site, where tryptophan binds. A set of parallel -sheets approach the aminoacylating active site. There are substantial allosteric effects on tRNA
Trp (
CCA) from TrpRS-IC binding. Elbow contacts between the D loop (D
12-GG-D
13) and the T loop (54-UU/CAA-58) are broken. The Levitt bp is also disrupted. Deformability of tRNA
Trp (
CCA) may contribute to cognate tryptophan charging. Tryptophan is in a 1-codon box in the code, which is generally not allowed. Tryptophan can be in a 1-codon box because Trp shares a 2-codon box with a stop codon (UGA), which does not utilize a tRNA but rather is recognized by a protein release factor binding to the UGA stop codon on the mRNA on the ribosome. Methionine is also in a 1-codon box that is shared with isoleucine (anticodon
CAU). In this case, different wobble
34C modifications explain how translational accuracy is maintained, and the
UAU Ile anticodon is generally not utilized.
P. furiosus tRNA
Trp (
CCA) is shown in
Figure 36. The acceptor stem matches the primordial tRNA sequence at 4 bp. The tRNA has a D loop with the sequence D
1-UGGUGUAGCCUGGUCCA-D
17, matching the primordial sequence in all but 5 positions and including no deletions from tRNA
Pri. The anticodon stem matches tRNA
Pri in 4 of 5 bp. The T stem-loop-stem matches the primordial tRNA sequence in all but one stem base pair. In
P. furiosus, tRNA
Trp (
CCA) is similar to tRNA
Pro (
GGG,
CGG and
UGG).
4. The Genetic Code
A model for evolution of the first genetic code is shown in
Figure 37 [
15,
16,
17,
18,
46]. Much of the data supporting this model is summarized in
Figure 38 and
Figure 39. The code is represented as a codon-anticodon table with a complexity of 32 assignments, rather than 64 assignments. Because of code degeneracy, 32 assignments (2x4x4) is the maximum complexity of the genetic code in tRNA, because a wobble position (tRNA-
34) has only purine versus pyrimidine resolution. The code is highly ordered. Most evolution is in code columns. The history of evolution of AARS enzymes (summarized below) relates a fairly straightforward story of evolution of the code. The genetic code is simpler and more ancient in Archaea. The code is more innovated in Bacteria and Eukarya [
43].
The model for evolution of the genetic code relies on the solution of tRNA evolution [
15,
22,
32]. For tRNA, the orderly mechanism of tRNA assembly and tRNA root sequences are known. tRNA evolved according to the three 31 nt minihelix tRNA evolution theorem. Based on sequence, this is much more of a theorem (a proven theory) than a conjecture or hypothesis or model. The original tRNA sequence was 100% RNA repeats (GCG, CGC and UAGCC) and inverted repeats (~CCGGG_CU/
GCCAA_CCCGG; _ separates stems and loop; / indicates the U-turn; the only sequence ambiguity is in the anticodon
GCC, which has since been scrambled in coding). Because the initial tRNA was so highly ordered, tRNA evolution was solved by inspection as a simple puzzle. ACCA-Gly was ligated at the tRNA 3’-end to synthesize polyglycine. There is no “chicken and egg” problem in evolution of the genetic code, because the code initially evolved to synthesize polyglycine and subsequently advanced to encode GADV polymers. The code did not need foresight of its evolving role in encoding RNA sequence-dependent proteins.
Evolution of the code makes best sense when viewed by code columns. The model for filling the code might be G→GADV→GADVLSER→GADVLSERNCQ→GADVLSERNCQPTIMHK→GADVLSERNCQPTIMHKFYW [
15,
16,
83]. At about the 11 amino acid stage, GADVLSERNCQ might be expected to support synthesis of the first proteins. Dividing the evolutionary history into code columns makes best sense (
Figure 37). NCQ, and possibly other amino acids, were added through tRNA-linked chemistry, giving insight into a major mechanism for RNA-linked pre-life metabolism.
4.1. Column 1
In column 1 (tRNA-
35A), valine may be the founding amino acid. It appears that valine (tRNA-
36C) goes to leucine goes to isoleucine goes to methionine. Phenylalanine is added last along disfavored row 1 (tRNA-
36A). In metabolism, valine can be converted to leucine in 5 steps. Thus, leucine may have been initially added to the code by tRNA-linked chemistry. Val-tRNA
Val may have been converted to Leu-tRNA
Leu in several steps, either supported by ribozymes or by first protein catalysts. We posit that tRNA-linked and RNA-linked chemistry were very ancient and were fundamental to evolution of pre-life metabolism and the genetic code. Notably, evolution of tRNA and divergence of tRNAomes is a story of RNA-amino acid and RNA-protein evolution [
47]. In the first code, tRNA
Val is type I and tRNA
Leu is type II, and type I tRNA was processed from a primitive type II tRNA. During early tRNAome assembly, tRNAs may have been mixed type I and type II, and the mixtures may have been sorted later by selection to construct the first code. In Archaea, only leucine and serine utilize type II tRNAs. In Bacteria, tyrosine, leucine and serine utilize type II tRNAs [
11]. The number of amino acids supported by type II tRNAs was limited by the number of allowed trajectory set points of the type II V arm.
In Archaea, type II tRNAs encoding leucine and serine were selected to substitute longer V arms for anticodon loop recognition by cognate AARS, because 5 tRNA
Leu and 4 tRNA
Ser were necessary [
11]. Isoleucine was the next amino acid to enter column 1. Neither valine nor leucine can be converted to isoleucine. Threonine (tRNA-
36U), which borders isoleucine (tRNA-
36U) in column 2 of the code, however, can be converted to isoleucine. It may be that Thr-tRNA
Thr evolved to Ile-tRNA
Ile via tRNA-linked chemistry. It is hypothesized that isoleucine briefly occupied a 4-codon sector of the code that was invaded by methionine. It appears that tRNA
Ile may have evolved to tRNA
Met. In
P. furiosus, tRNA
Ile and tRNA
Met are similar. It appears that methionine invaded a 4-codon isoleucine sector. At the base of the code evolution, the anticodon
UAU was eliminated. Without modification,
UAU would cause confusion between encoding isoleucine and methionine. Both initiator and elongator tRNA
Met (
CAU) evolved. The initiator tRNA
Met (
CAU) is unmodified at
34C. The elongator tRNA
Met (
CAU) utilizes the
34Cm modification. Phenylalanine was added to the code late (described below). In Archaea,
34agm2CAU (agm2C for agmatidine) encodes isoleucine. In Bacteria,
34k2CAU (k2C for lysidine) encodes isoleucine.
4.2. Column 2
Column 2 (tRNA-35G) of the code appears to have evolved from alanine to serine to proline and threonine. Serine appears to have jumped from column 2 to column 4 of the genetic code, perhaps, in part, to obtain a more favorable anticodon (tRNA-
35C appears to be favored over tRNA-
35G). Alanine can be converted to serine in several steps, so Ala-tRNA
Ala may have evolved to Ser-tRNA
Ser. If this is the case, however, type I tRNA
Ser was probably replaced from type II tRNA
Leu. From sequence, it appears that type II tRNA
Ser was derived from type II tRNA
Leu. Serine is a special case because serine is the only amino acid that appears to jump columns in establishment of the genetic code. Also, serine can be converted to cysteine by tRNA-linked chemistry [
65,
84,
85]. It is hypothesized that conversion of serine to cysteine may relate to the jumping of serine from column 2 to column 4 of the code. To establish the standard code, cysteine landed in column 4, disfavored row 1. Cysteine, however, must have entered the code earlier, perhaps linked to serine within an expanded serine sector. In proteins, cysteine is necessary for Zn binding, which was required for first protein folding. AARS enzymes are an example of first proteins, coevolved with the genetic code, whose folding depended on cysteine binding Zn. Cysteine may have occupied the disfavored row 1 (tRNA-
36A) late in evolution of the code. Serine may have displaced cysteine in row 3, column 4, where serine now resides. Serine appears to have jumped columns by invading and splitting an expanded arginine sector.
Because serine utilizes a type II tRNASer, and because SerRS-IIA lacks tRNASer anticodon recognition, these features may have facilitated serine or serine/cysteine jumping in evolution of the code. A change of Ser/Cys-tRNASer/Cys (GGU)→Ser/Cys-tRNASer/Cys (GCU) could account for serine jumping columns. GGU becomes a threonine anticodon, indicating that the threonine 4-codon sector (column 2, row 3) displaced serine from an expanded serine sector. Threonine and serine are chemically related amino acids. Proline also appears to have displaced serine to form a 4-codon sector (column 2, row 2).
It is hypothesized that the first AlaRS may have been an AlaRS-IIA from which column 2 SerRS-IIA, ThrRS-IIA and ProRS-IIA were derived. As the genetic code was built up, however, we posit that AlaRS-IIA was replaced by AlaRS-IID. The proposed replacement is analogous to the replacement of archaeal type GlyRS-IIA by GlyRS-IID in more derived Bacteria. If the AlaRS-IIA to AlaRS-IID replacement event was prior to LUCA, there may now be no sequence record of AlaRS-IIA. The AlaRS-IID innovation helped discriminate neutral amino acids, alanine, serine, threonine and proline. AlaRS-IID has editing functions, and AlaRS-IID has a separate editing domain. In ancient organisms, AlaRS-IID utilizes AlaX editing protein to support accuracy (
Figure 29). The AlaRS-IID aminoacylating active site also has editing functions. AlaX protein may partially compensate for the lack of AlaRS-IID anticodon loop recognition. Alanine is in a 4-codon box, but AlaRS-IID does not utilize the tRNA
Ala anticodon loop as a determinant for accurate charging.
4.3. Column 3
Column 3 is the most innovated column in the code, encoding the most amino acids. Notably, column 3 is broken into all 2-codon sectors. It is hypothesized that column 3 may have sectored by a slightly different mechanism compared to columns 1, 2 and 4. We suggest that early in code evolution both tRNA-34 and tRNA-36 were wobble positions, but only a single wobble position could be utilized at a time. According to this view, columns 1, 2 and 4 primarily utilized Watson-Crick 35 and wobble 36. Column 3 primarily utilized Watson-Crick 35 and wobble 34. tRNA-35 was always easiest to read because this is the central base in the anticodon. In a wobble position, only purine-pyrimidine discrimination is achieved, so only 2 possible code assignments are obtained. In such a scenario, the complexity of the evolving code would be 2x4 or 4x2 or 8 amino acids, depending on the wobble position (tRNA-34 or tRNA-36). Because of tRNA-linked chemistry adding NCQ, the limited code probably expanded to 11 amino acids (GADVLSER→GADVLSERNCQ). We posit that evolution of the genetic code was hung up at 8 or 11 amino acids until wobbling at tRNA-36 could be suppressed. Wobbling at tRNA-36 was suppressed, in part, by modifications at tRNA-37. tRNA-37G modifications (i.e., m1G) were used to read tRNA-36A. tRNA-37A modifications (i.e., t6A) were used to read tRNA-36U. It appears that wobbling at tRNA-36U was more readily suppressed than wobbling at tRNA-36A. Notably, 37t6A to suppress 36U is a more dramatic modification than 37m1G to suppress 36A. In evolution of the code, row 3 (tRNA-36U) of the genetic code appears to have been more favorable than row 1 (tRNA-36A).
Column 3 appears to have first encoded aspartic acid. The chemically related glutamic acid may then have invaded the B rows. This resulted in a Glu-Asp-Glu-Asp-Glu-Asp (column 3, row 4B-4A-3B-3A-2B-2A) pattern (
Figure 37). Asparagine displaced aspartic acid in column 3, row 3A. Histidine displaced aspartic acid in column 3, row 2A. Lysine displaced glutamate in column 3, row 3B. Glutamine displaced glutamate in column 3, row 2B. Stop codons and tyrosine were added late across disfavored row 1.
As soon as aspartic acid and glutamate entered the code, tRNA-linked chemistry generated asparagine and glutamine [
70,
73,
74,
86,
87]. Serine can be converted to cysteine by tRNA-linked chemistry [
65,
84,
85]. The 8 amino acid code (i.e., GADVLSER), therefore, rapidly evolved to an 11 amino acid code (GADVLSERNCQ), by tRNA-linked chemistry. The 11 amino acid code appears to be sufficient to generate first RNA sequence-dependent proteins.
4.4. Column 4
Column 4 (tRNA-
35C) appears to be the most favored code column. Glycine appears to occupy the most favored sector of the genetic code (tRNA-
35C, tRNA-
36C). It is hypothesized that glycine is the founding amino acid in the code. GADV, the four simplest and probably the four initial encoded amino acids, occupy the most favored row of the genetic code (tRNA-
36C) [
48,
49,
50,
51,
88,
89]. These observations are consistent with glycine being the first encoded amino acid [
19,
20]. In
P. furiosus, tRNA
Gly is the most similar to tRNA
Pri, indicating that glycine may have been the first encoded amino acid [
21]. GlyRS-IIA appears to be the root for all class II and class I AARS. Glycine is the smallest and the most flexible amino acid. It is very likely that glycine was the founding amino acid in evolution of the genetic code.
By contrast to glycine, arginine, which is also in column 4, is a complex amino acid. It is hypothesized that ornithine may have been the founding positively charged amino acid [
60]. Ornithine can be converted to arginine in two metabolic steps. In some Archaea and Bacteria, ornithine can be converted to lysine by the α-aminoadipate pathway [
90,
91,
92,
93,
94,
95]. Thus, arginine and lysine may have entered the genetic code through tRNA-linked reactions: Orn-tRNA
Orn→Arg-tRNA
Arg (column 4) and Orn-tRNA
Orn (
UCU,
CCU)(column 4)→Lys-tRNA
Lys (
UUU,
CUU) (column 3).
ArgRS-IA is the closest relative of CysRS-IA, indicating how cysteine may have evolved to its current placement in column 4, row 1A, of the code. In P. furiosus, only tRNAThr (column 2) and tRNACys (column 4) utilize discriminator base 73U. tRNAThr and tRNACys are similar in sequence in P. furiosus.
4.5. Disfavored Row 1
Disfavored row 1 of the genetic code appears to have sectored last. Phenylalanine, tyrosine and tryptophan are complex aromatic amino acids. It has been hypothesized that initially phenylalanine spread across row 1 utilizing a primitive PheRS-IC. From PheRS-IC, both TyrRS-IC and TrpRS-IC may have been derived. To suppress translation errors, it is hypothesized that PheRS-IC was replaced by PheRS-IIC before LUCA. There now appears to be no sequence trace of PheRS-IC. PheRS-IIC has a separate editing active site to suppress non-cognate charging with tyrosine and other amino acids. In Bacteria, tRNA
Tyr (
GUA) is a type II tRNA, but this is a bacterial innovation, perhaps, to suppress translation errors (i.e., enhancing discrimination of Phe and Tyr). Amino acids that differ only in a hydroxyl group are difficult for AARS enzymes to distinguish [
34].
Serine appears to have ended up in column 2, row 1, from its particular chaotic history in genetic code evolution that may have involved invasion of an expanded serine sector by threonine and proline. Cysteine may have ended up in column 4, row 1A, from the history of serine jumping from column 2 to column 4.
We conclude that a rational explanation can be provided for the placements of: 1) all amino acids; 2) class II and class I AARS enzymes; and 3) many tRNAs in evolution of the genetic code. When this project was started, this outcome was not anticipated.
4.6. Stop Codons and Evolution of Translational Fidelity
Stop codons locate to column 3, row 1B, and column 4, row 1B. We consider it likely that stop codons were a late addition to the code. The evolution of the genetic code can be viewed as the evolution of intellectual property initially to support polypeptide polymer synthesis as a pre-life chemistry emulsifier and then progressing to cognate coding with the inception of complex life. According to this view, initially, in pre-life, long protein polymers and innovation in amino acid additions were selected over fidelity. Translational fidelity became ever more important, however, as the genetic code evolved and the system developed intellectual property in tRNAomes, AARSomes and first proteins that was more strongly selected.
Initially, the code evolved to synthesize polyglycine and then GADV polymers as emulsifiers for metabolic reaction components and to coalesce the first protocells. As accurate coding became more strongly selected, the selective pressure was toward evolution of fidelity mechanisms, such as editing mechanisms in AARS aminoacylating active sites and separate proofreading domains. Stop codons and frame maintenance are also fidelity mechanisms.
Amino acids with little chemical character locate to the left half of the genetic code (Val, Met, Ile, Leu, Phe, Ala, Thr, Pro, and Ser) (columns 1 and 2). These amino acids are charged to cognate tRNAs by AARS enzymes that edit either within separate proofreading active sites, within the aminoacylating active site, or both. Amino acids from the right half of the code have more chemical character (Glu, Asp, Lys, Asn, Gln, His, Tyr, Gly, Arg, Trp and Cys). Cognate charging of right half amino acids (columns 3 and 4) generally does not require editing. Right half amino acids have more chemical character (i.e., charge, hydrogen bonding, metal binding (i.e., Cys)) that is used to support accurate charging of their cognate tRNA.
Initially in pre-life, stop codons were not as important as later in evolution, because making longer emulsifying polymers was more important than accurate stops. Also, because of RNA ligations for RNA replication, combinations of primitive protein reading frames were strongly selected to generate more complex first proteins and new functions. When sequences were fused out of frame, therefore, more complex proteins were initially synthesized using frame shifts, before translation frames evolved to be in phase. A primitive class I ValRS-IA evolved by ligation of an N-terminal encoding RNA to a primitive class II GlyRS-IIA encoding RNA. In the absence of hard stop codons, the initial ligation was not necessarily in phase. Initially, innovation was strongly valued over accuracy. Stop codons are read by protein release factors in mRNA. Protein release factors bind to stop codons in mRNA and effect the nascent protein release from the ribosome [
96]. No tRNA is associated with stop codons and translation termination. In suppressor strains, a tRNA anticodon mutates into a stop codon to add an amino acid, somewhat inefficiently, in place of a stop.
5. Radiation of AARSomes
Figure 37,
Figure 38 and
Figure 39 document the evolution of the first genetic code.
Figure 37 shows the ordered structure of the code, indicating relatedness of AARS enzymes [
15,
16,
17,
18,
46]. Most evolution is in code columns, as indicated in the model.
Figure 38 relates the relationships among all class II and class I AARS in the ancient Archaeon
P. furiosus. Some AARS missing in
P. furiosus were supplied from other species, as appropriate.
P. furiosus was selected because
P. furiosus has an ancient tRNAome that is similar to that of LUCA [
21]. It was assumed that the
P. furiosus AARSome would also be similar to LUCA.
Figure 38 was prepared as previously described using Phyre 2 homology scoring by structure and sequence [
18,
97]. Relatedness of GlyRS-IIA and ValRS-IA and IleRS-IA sequence has previously been demonstrated [
15,
16,
17,
18,
46].
Figure 39 shows how the structure of the genetic code relates to the apparent lineages of AARS enzymes by correlating the map in
Figure 37 to the pattern of AARS evolution shown in
Figure 38. In
Figure 39, AARS enzymes are assigned background colors that relate to the structure of the code shown in
Figure 37. These three figures summarize an ordered model for AARS and genetic code evolution.
A primitive GlyRS-IIA appears to be the root of both the class II lineage and the class I lineage. Based on sequence homology, it is hypothesized that a primitive ValRS-IA was derived from a primitive GlyRS-IIA by ligation of an N-terminal encoding RNA to a GlyRS-IIA encoding RNA. In pre-life, replication of RNAs required ribozyme ligases that generated long and complex RNAs and complex proteins very early in evolution. tRNA evolution required ligation and complementary replication to generate tRNA out of 31 nt minihelices. Attachment of the N-terminal encoding RNA to the primitive GlyRS-IIA encoding RNA altered the folding of the translated protein to a primitive ValRS-IA. Zn binding motifs were important in folding the first AARS enzymes.
AARS enzymes are first proteins, coevolved with the genetic code. Without full tRNAomes and AARSomes, there is no standard code. It is hypothesized that sequence-dependent proteins emerged at about the 11 amino acid stage of code evolution (i.e., GADVLSERNCQ) (R may initially have been O (ornithine) that radiated to R and K). The 11 amino acid stage provides sufficient chemical diversity (i.e., flexibility, hydrophobicity, hydrogen bonding, charge) to encode first proteins. Addition of amino acids to the code improved protein structure and function until 20 amino acids and stops were encoded. As the code froze, adding additional amino acids became more of a liability because of the threat posed to translational accuracy. There is a tension between innovation and error catastrophe. To prevent error catastrophe, fidelity mechanisms such as amino acid identity (chemical character) and editing by AARS froze the code. Early in code evolution, innovation was more strongly selected. Late in code evolution fidelity mechanisms evolved to protect the intellectual property that pre-biology and emerging biology had generated.
The model for AARS radiation (
Figure 37,
Figure 38 and
Figure 39) is a working model. More advanced network and evolutionary analyses will be necessary to confirm or improve the model. To enhance tRNAome networks, alignments of tRNAs must be optimized in the D loop region and V loop region.
6. Evolution of Complex Life
The pathway to evolve complex life on Earth, supported by a genetic adapter and genetic code, is mostly elucidated [
15,
32]. Once the genetic code arose, all features of complex life and biodiversity became possible. The solution is embedded in the sequence of tRNA
Pri and in the order of assembly of the genetic code. tRNA was formed from GCG, CGC and UAGCC repeats and inverted repeats ~CCGGG_CU/
GCCAA_CCCGG. tRNA was evolved from ligation of three 31 nt minihelices of mostly known sequence (GCGGCGG_UAGCC_UA
GCCUA_GCCUA_CCGCCGC and ~GCGGCGG_ CCGGG_CU/
GCCAA_CCCGG_CCGCCGC). ACCA-Gly was ligated to various RNAs including tRNAs to synthesize polyglycine. The genetic code evolved as described in this report. Primitive pre-mRNAs and pre-rRNAs were generated by similar processes of ligation and genetic recombination.
To evolve tRNA required a small number of catalytic functions (i.e., ribozymes). The process required a mechanism to generate RNA repeats and inverted repeats. Multiple functions were necessary including RNA ligase, RNA replicase (complementary replication), exo- and endo-nucleases, ribose 2’-O-methyltransferase (for RNA stability) and ACCA-Gly transferase. Complementary replication utilizing snap-back primers (i.e., 31 nt minihelices) was needed. With these ingredients and little else, it should be possible to recreate most of the origin of tRNA and the genetic code in a laboratory. Evolution of tRNA and the genetic code describe an RNA-amino acid and RNA-peptide world overlaid on primitive metabolism with coevolution of protocells to generate the first life on Earth.
7. Discussion
The genetic code coevolved with tRNA, tRNAomes, AARSomes, ribosomes and first proteins [
5,
6,
7,
8,
9,
10]. Evolution of AARSomes is evident in genetic code columns. In column 1, ValRS-IA, LeuRS-IA, IleRS-IA and MetRS-IA are closely related enzymes. In column 2, SerRS-IIA, ProRS-IIA and ThrRS-IIA are closely related enzymes. AlaRS-IID may have replaced a now extinct AlaRS-IIA before LUCA. Column 3 demonstrates a striped pattern of related AARS enzymes. AspRS-IIB, AsnRS-IIB and HisRS-IIA are closely related enzymes in rows 4A, 3A and 2A. GluRS-IB, LysRS-IB (in Archaea) and GlnRS-IB (a eukaryotic innovation) are closely related enzymes in rows 4B, 3B and 2B. A primitive GlyRS-IIA appears to be the founding AARS. tRNA
Gly appears to be the founding tRNA that is most similar to tRNA
Pri [
21]. Glycine appears to be the founding amino acid [
19,
20], and glycine occupies the most favored sector in the code (tRNA-
35C, tRNA-
36C). In column 4, ArgRS-IA and CysRS-IA are closely related enzymes. Row 1 of the genetic code appears to have sectored last. TrpRS-IC and TyrRS-IC are closely related enzymes. PheRS-IIC appears to be a late substitution perhaps for a PheRS-IC, from which TyrRS-IC and TrpRS-IC were derived, that is now extinct. Cysteine may have first entered the code through tRNA-linked chemistry within an expanded serine sector.
Coding evolved around tRNA and the tRNA anticodon. Coding should be viewed as arising first in the tRNA anticodon. In tRNA, the maximum number of coding assignments is limited to 32 by wobbling. tRNA cannot support 64 genetic code assignments, as could DNA and mRNA. Coding coevolved from tRNA anticodons into mRNA codons and then was cast into DNA for more stable information storage. Degeneracy of the code is a feature of tRNA and the tRNA anticodon. Wobbling at tRNA-
34 created code degeneracy. Suppression of wobbling at tRNA-
36 gives the history of genetic code establishment. Even with modification, no tRNA-
34A is utilized at the base of genetic code evolution. Elp3 and subsequent tRNA-
34U 5-carbon modifications suppressed superwobbling in order to evolve 2-codon sectors in the code (i.e., column 3) [
43,
44,
45]. tRNA-
34, tRNA-37 and other tRNA modifications were necessary to evolve the first code.
Because of the placement of the anticodon loop U-turn, wobbling at tRNA-
36 was suppressed, but wobbling at tRNA-
34 was not. Next to tRNA-
34 is tRNA-33U, which is on the opposite side of the anticodon U-turn. Because of the placement of the U-turn, modifying tRNA-33U would be unlikely to influence reading at tRNA-
34. Also, tRNA-33U is almost never substituted, indicating that a purine at that position might disrupt loop geometry. Modifications of tRNA-
35 cannot compensate because
35 is a Watson-Crick position for coding that cannot be specified in sequence or modified in a manner that affects coding. Apparently, modifications of tRNA-37 helped to suppress wobbling at tRNA-
36, particularly for tRNA-
36U (i.e., tRNA-37t6A) and tRNA-
36A (i.e., tRNA-37m1G) [
43]. To evolve the first code, these modifications may have been universal. As systems have evolved, some compensations for some modifications may have coevolved. Wobbling at tRNA-
34 (regulated) versus tRNA-
36 (suppressed) appears to explain why columns 1, 2 and 4 differ in their sectoring from column 3, which is the most innovated column.
8. Conclusions
The genetic code is simpler in Archaea than in Bacteria and Eukarya, indicating that the archaeal code is most similar to the LUCA code. The code in Archaea is highly ordered, and the order provides the history for first code establishment. tRNAomes are simpler in Archaea. Organisms with the simplest tRNAomes are the closest to LUCA. tRNAome and AARSome networks of ancient organisms describe the history of establishment of the first code.
tRNA evolved from RNA repeats and inverted repeats of known sequence. Three 31 nt minihelices were ligated and processed by orderly internal 9 nt deletion(s) into type I and type II tRNAs [
15,
22,
98]. Multiple RNAs were joined as replication intermediates, generating long functional RNAs such as tRNAs, pre-mRNAs and primitive rRNAs. tRNA evolution is a story of amino acid-RNA and protein-RNA linked chemistry [
47], so life evolved from a complex RNA-amino acid- RNA-protein-metabolism world, packaged in coevolved protocells. When coupled with coded protein synthesis, this evolving pre-life world generated remarkable complexity and fostered surprising innovation. The first proteins that coevolved with the genetic code are highly evolved, innovated and complex constructs many of which remain largely unaltered to present day. With the freezing of the first code, life as currently known emerged on Earth. The history of tRNA evolution is embedded in tRNA sequence, which can be read. The history of the evolution of the genetic code is embedded in code structure and interacting tRNAome, AARSome and first protein networks.
The core history of abiogenesis is the evolution of tRNA, which was recorded and preserved in tRNA sequence. The history of genetic code evolution was written into the standard genetic code structure and AARSome radiation. In terms of astrobiology, it is difficult to see how life could evolve separately on another planet or moon by a very different chemistry or different pathway. If there is another route to a suitable genetic adapter than tRNA, we are not certain what that might be. Life without a genetic adapter and genetic code has limited possibilities.
Figure 1.
GlyRS-IIA-tRNAGly (CCC) from H. sapiens. A primitive GlyRS-IIA appears to be the founding AARS. The image was selected to emphasize tRNA contacts. Protein is beige. -sheets are cyan. tRNA is colored according to the three 31 nt minihelix tRNA evolution theorem. Some GlyRS-IIA amino acids that were not imaged are noted. Lbp for Levitt base pair. The elbow is where the D loop binds the T loop. .
Figure 1.
GlyRS-IIA-tRNAGly (CCC) from H. sapiens. A primitive GlyRS-IIA appears to be the founding AARS. The image was selected to emphasize tRNA contacts. Protein is beige. -sheets are cyan. tRNA is colored according to the three 31 nt minihelix tRNA evolution theorem. Some GlyRS-IIA amino acids that were not imaged are noted. Lbp for Levitt base pair. The elbow is where the D loop binds the T loop. .
Figure 2.
tRNA
Gly (
CCC). A) A primordial tRNA
Gly (
CCC). B)
P. furiosus tRNA
Gly (
CCC). C) Human tRNA
Gly (
CCC). tRNAs are colored according to the three 31 nt minihelix tRNA evolution theorem [
15,
22]. In
Figure 2A, the Levitt base pair (D
8G=V
5C) (Lbp) and some elbow contacts are indicated. The Levitt base pair is a reverse Watson-Crick pair that forms two hydrogen bonds. D
12G intercalates between 57A and 58A and hydrogen bonds to 55U [
41]. D
13G forms a Watson-Crick pair with T loop 56C. Anticodon sequences are underlined or white bold. / indicates a U-turn. Modifications of the anticodon loop are indicated. Modomics notation is used for tRNA anticodon loop modifications [
30]. xU indicates an unknown 5-carbon U modification to suppress superwobbling. Yellow arrows indicate features that may be of interest. 32U-38U is expected to alter dynamics of the anticodon loop.
Figure 2.
tRNA
Gly (
CCC). A) A primordial tRNA
Gly (
CCC). B)
P. furiosus tRNA
Gly (
CCC). C) Human tRNA
Gly (
CCC). tRNAs are colored according to the three 31 nt minihelix tRNA evolution theorem [
15,
22]. In
Figure 2A, the Levitt base pair (D
8G=V
5C) (Lbp) and some elbow contacts are indicated. The Levitt base pair is a reverse Watson-Crick pair that forms two hydrogen bonds. D
12G intercalates between 57A and 58A and hydrogen bonds to 55U [
41]. D
13G forms a Watson-Crick pair with T loop 56C. Anticodon sequences are underlined or white bold. / indicates a U-turn. Modifications of the anticodon loop are indicated. Modomics notation is used for tRNA anticodon loop modifications [
30]. xU indicates an unknown 5-carbon U modification to suppress superwobbling. Yellow arrows indicate features that may be of interest. 32U-38U is expected to alter dynamics of the anticodon loop.
Figure 3.
ValRS-IA-tRNA
Val (
CAC) from
T. thermophilus. The image was colored as in
Figure 1. Non-cognate amino acids that are blocked from incorporation within the aminoacylating active site are indicated in red. Non-cognate amino acids that are removed from tRNA
Val after attachment within the separate proofreading (editing) active site are indicated in black. .
Figure 3.
ValRS-IA-tRNA
Val (
CAC) from
T. thermophilus. The image was colored as in
Figure 1. Non-cognate amino acids that are blocked from incorporation within the aminoacylating active site are indicated in red. Non-cognate amino acids that are removed from tRNA
Val after attachment within the separate proofreading (editing) active site are indicated in black. .
Figure 4.
tRNAVal (CAC). A) From P. furiosus (Pfu). The yellow arrow indicates an unmodified 34U. B) From T. thermophilus (Tth). The yellow arrow indicates a 32C-38C arrangement, which may affect loop dynamics. Hvo indicates Haloferax volcanii. Eco indicates E. coli. Sgr indicates Streptomyces griseus.
Figure 4.
tRNAVal (CAC). A) From P. furiosus (Pfu). The yellow arrow indicates an unmodified 34U. B) From T. thermophilus (Tth). The yellow arrow indicates a 32C-38C arrangement, which may affect loop dynamics. Hvo indicates Haloferax volcanii. Eco indicates E. coli. Sgr indicates Streptomyces griseus.
Figure 5.
IleRS-IA-tRNA
Ile (
GAU) from
S. aureus. IleRS-IA has a separate proofreading active site that removes non-cognate homocysteine and cysteine attached to tRNA
Ile (black text). Non-cognate valine, norvaline and -aminobutyrate are blocked from attachment to tRNA
Ile through reactions at the aminoacylating active site (red text) [
34,
35]. MRC binds the aminoacylating active site.
Figure 5.
IleRS-IA-tRNA
Ile (
GAU) from
S. aureus. IleRS-IA has a separate proofreading active site that removes non-cognate homocysteine and cysteine attached to tRNA
Ile (black text). Non-cognate valine, norvaline and -aminobutyrate are blocked from attachment to tRNA
Ile through reactions at the aminoacylating active site (red text) [
34,
35]. MRC binds the aminoacylating active site.
Figure 6.
tRNAIle (GAU). A) P. furiosus tRNAIle (GAU). B) S. aureus tRNAIle (GAU). Modifications to the anticodon loop are as expected. Mca for Mycoplasma capricolum. .
Figure 6.
tRNAIle (GAU). A) P. furiosus tRNAIle (GAU). B) S. aureus tRNAIle (GAU). Modifications to the anticodon loop are as expected. Mca for Mycoplasma capricolum. .
Figure 7.
MetRS-IA-tRNAMet (CAU) from Aquifex aeolicus.
Figure 7.
MetRS-IA-tRNAMet (CAU) from Aquifex aeolicus.
Figure 8.
tRNAMet (CmAU and CAU) and tRNAIle (agm2CAU). A) P. furiosus elongator tRNAMet (CmAU). B) P. furiosus initiator tRNAMet (CAU). C) P. furiosus tRNAIle (agm2CAU).
Figure 8.
tRNAMet (CmAU and CAU) and tRNAIle (agm2CAU). A) P. furiosus elongator tRNAMet (CmAU). B) P. furiosus initiator tRNAMet (CAU). C) P. furiosus tRNAIle (agm2CAU).
Figure 9.
LeuRS-IA-tRNALeu (CAA) of P. horikoshii. LeuRS-IA has a separate editing active site that removes non-cognate valine, -aminobutyric acid and methionine from tRNALeu (black text). The aminoacylating active site blocks norvaline, homocysteine, -hydroxy leucine and isoleucine incorporation (red text). In P. horikoshii, tRNALeu is a type II tRNA with a 14 nt V arm. .
Figure 9.
LeuRS-IA-tRNALeu (CAA) of P. horikoshii. LeuRS-IA has a separate editing active site that removes non-cognate valine, -aminobutyric acid and methionine from tRNALeu (black text). The aminoacylating active site blocks norvaline, homocysteine, -hydroxy leucine and isoleucine incorporation (red text). In P. horikoshii, tRNALeu is a type II tRNA with a 14 nt V arm. .
Figure 10.
Comparison of type II tRNAPri and tRNALeu. A) type II tRNAPri. In the anticodon, B indicates G, C or U, but not A. B) P. horikoshii tRNALeu (CAA). In Archaea, two bases separate the 3’-V arm stem from the Levitt base (V14C) giving the trajectory of the V arm. The V6-UAG-V8 consensus to bind LeuRS-IA is indicated. In principle, the CU/UAAGA anticodon loop could cause leucine substitution for phenylalanine by superwobbling. C) Bacterial T. thermophilus tRNALeu (CAA) has a different trajectory of the V arm and lacks the V arm end loop UAG consensus. The trajectory of the V arm is given by the number of unpaired bases (one) separating the 3’-V arm stem and the Levitt base V15U.
Figure 10.
Comparison of type II tRNAPri and tRNALeu. A) type II tRNAPri. In the anticodon, B indicates G, C or U, but not A. B) P. horikoshii tRNALeu (CAA). In Archaea, two bases separate the 3’-V arm stem from the Levitt base (V14C) giving the trajectory of the V arm. The V6-UAG-V8 consensus to bind LeuRS-IA is indicated. In principle, the CU/UAAGA anticodon loop could cause leucine substitution for phenylalanine by superwobbling. C) Bacterial T. thermophilus tRNALeu (CAA) has a different trajectory of the V arm and lacks the V arm end loop UAG consensus. The trajectory of the V arm is given by the number of unpaired bases (one) separating the 3’-V arm stem and the Levitt base V15U.
Figure 11.
SerRS-IIA-tRNASer (UGA) from H. sapiens. The full 2-dimer is shown. One -subunit is colored white; one is wheat. -sheets are light pink. HH indicates the N-terminal helix hairpin that binds the type II V arm stems and the elbow of tRNASer. .
Figure 11.
SerRS-IIA-tRNASer (UGA) from H. sapiens. The full 2-dimer is shown. One -subunit is colored white; one is wheat. -sheets are light pink. HH indicates the N-terminal helix hairpin that binds the type II V arm stems and the elbow of tRNASer. .
Figure 12.
tRNASer (UGA). A) P. furiosus tRNASer (UGA). In Archaea, one base separates the 3’-V stem and the Levitt base. B) H. sapiens tRNASer (UGA). C) T. thermophilus tRNASer (UGA). Zero bases separate the 3’-V arm stem and the Levitt base (V19C). .
Figure 12.
tRNASer (UGA). A) P. furiosus tRNASer (UGA). In Archaea, one base separates the 3’-V stem and the Levitt base. B) H. sapiens tRNASer (UGA). C) T. thermophilus tRNASer (UGA). Zero bases separate the 3’-V arm stem and the Levitt base (V19C). .
Figure 13.
ArgRS-IA-tRNAArg (ICG) of S. cerevisiae.
Figure 13.
ArgRS-IA-tRNAArg (ICG) of S. cerevisiae.
Figure 14.
tRNAArg. A) P. furiosus tRNAArg (GCG). B) S. cerevisiae tRNAArg (ICG). Yellow arrows indicate features of possible interest. Bta indicates Bos taurus.
Figure 14.
tRNAArg. A) P. furiosus tRNAArg (GCG). B) S. cerevisiae tRNAArg (ICG). Yellow arrows indicate features of possible interest. Bta indicates Bos taurus.
Figure 15.
CysRS-IA-tRNACys (GCA) from H. sapiens.
Figure 15.
CysRS-IA-tRNACys (GCA) from H. sapiens.
Figure 16.
tRNACys (GCA). A) P. furiosus tRNACys (GCA). B) H. sapiens tRNACys (GCA).
Figure 16.
tRNACys (GCA). A) P. furiosus tRNACys (GCA). B) H. sapiens tRNACys (GCA).
Figure 17.
ThrRS-IIA-tRNAThr (CGU) from E. coli. A* is 37m6t6A. .
Figure 17.
ThrRS-IIA-tRNAThr (CGU) from E. coli. A* is 37m6t6A. .
Figure 18.
tRNAThr (CGU). A) P. furiosus tRNAThr (CGU). B) E. coli tRNAThr (CGU).
Figure 18.
tRNAThr (CGU). A) P. furiosus tRNAThr (CGU). B) E. coli tRNAThr (CGU).
Figure 19.
ProRS-IIA-tRNAPro (CGG) from T. thermophilus. P5A is a reaction intermediate analogue that binds in the aminoacylating active site. .
Figure 19.
ProRS-IIA-tRNAPro (CGG) from T. thermophilus. P5A is a reaction intermediate analogue that binds in the aminoacylating active site. .
Figure 20.
tRNAPro (CGG). A) P. furiosus tRNAPro (CGG). B) T. thermophilus tRNAPro (CGG). Sty indicates Salmonella typhimurium.
Figure 20.
tRNAPro (CGG). A) P. furiosus tRNAPro (CGG). B) T. thermophilus tRNAPro (CGG). Sty indicates Salmonella typhimurium.
Figure 21.
AspRS-IIB-tRNAAsp (GUC) from S. cerevisiae. .
Figure 21.
AspRS-IIB-tRNAAsp (GUC) from S. cerevisiae. .
Figure 22.
tRNA-linked chemistry. A detail of the T. thermophilus transamidosome is shown.
Figure 22.
tRNA-linked chemistry. A detail of the T. thermophilus transamidosome is shown.
Figure 23.
tRNAAsp (GUC) and tRNAAsn (GUU). A) P. furiosus tRNAAsp (GUC). B) S. cerevisiae tRNAAsp (GUC). C) P. furiosus tRNAAsn (GUU). .
Figure 23.
tRNAAsp (GUC) and tRNAAsn (GUU). A) P. furiosus tRNAAsp (GUC). B) S. cerevisiae tRNAAsp (GUC). C) P. furiosus tRNAAsn (GUU). .
Figure 24.
HisRS-IIA-tRNAHis (GUG) from T. thermophilus. .
Figure 24.
HisRS-IIA-tRNAHis (GUG) from T. thermophilus. .
Figure 25.
tRNAHis (GUG). A) P. furiosus tRNAHis (GUG). B) T. thermophilus tRNAHis (GUG). The yellow arrows indicates the unique (-1)GTP=73C discriminators that also may suppress misalignment of the P-site peptide-tRNA on the ribosome.
Figure 25.
tRNAHis (GUG). A) P. furiosus tRNAHis (GUG). B) T. thermophilus tRNAHis (GUG). The yellow arrows indicates the unique (-1)GTP=73C discriminators that also may suppress misalignment of the P-site peptide-tRNA on the ribosome.
Figure 26.
GluRS-IB-tRNAGlu (CUC) from T. thermophilus.
Figure 26.
GluRS-IB-tRNAGlu (CUC) from T. thermophilus.
Figure 28.
P. furiosus tRNALys (CUU). xU is an unidentified 5 carbon-U34 modification to suppress superwobbling.
Figure 28.
P. furiosus tRNALys (CUU). xU is an unidentified 5 carbon-U34 modification to suppress superwobbling.
Figure 29.
AlaRS-IID-tRNAAla (UGC) of A. fulgidus. The AlaX protein of P. horikoshii (pink) is also shown and overlaid on the A. fulgidus structure to locate the editing active site. No contacts are made by AlaRS-IID to the tRNAAla anticodon loop. .
Figure 29.
AlaRS-IID-tRNAAla (UGC) of A. fulgidus. The AlaX protein of P. horikoshii (pink) is also shown and overlaid on the A. fulgidus structure to locate the editing active site. No contacts are made by AlaRS-IID to the tRNAAla anticodon loop. .
Figure 30.
P. furiosus tRNAAla (UGC). .
Figure 30.
P. furiosus tRNAAla (UGC). .
Figure 31.
PheRS-IIC-tRNAPhe (GAA) from T. thermophilus. Both tRNAPhe (GAA) are shown to indicate all relevant PheRS-IIC-tRNAPhe (GAA) contacts. .
Figure 31.
PheRS-IIC-tRNAPhe (GAA) from T. thermophilus. Both tRNAPhe (GAA) are shown to indicate all relevant PheRS-IIC-tRNAPhe (GAA) contacts. .
Figure 32.
P. furiosus tRNAPhe (GAA).
Figure 32.
P. furiosus tRNAPhe (GAA).
Figure 33.
TyrRS-IC-tRNATyr (GUA) of M. jannaschii.
Figure 33.
TyrRS-IC-tRNATyr (GUA) of M. jannaschii.
Figure 34.
tRNATyr (GUA). A) Archaeal P. furiosus tRNATyr (GUA) (type I). B) Bacterial T. thermophilus tRNATyr (GUA) (type II). Lla for Lactobacillus lactis. In Bacteria, type II tRNATyr has two unpaired bases separating the 3’-V stem from the Levitt base.
Figure 34.
tRNATyr (GUA). A) Archaeal P. furiosus tRNATyr (GUA) (type I). B) Bacterial T. thermophilus tRNATyr (GUA) (type II). Lla for Lactobacillus lactis. In Bacteria, type II tRNATyr has two unpaired bases separating the 3’-V stem from the Levitt base.
Figure 35.
TrpRS-IC-tRNATrp (CCA) from H. sapiens.
Figure 35.
TrpRS-IC-tRNATrp (CCA) from H. sapiens.
Figure 36.
P. furiosus tRNATrp (CCA).
Figure 36.
P. furiosus tRNATrp (CCA).
Figure 37.
A model for evolution of the first code. A codon-anticodon table is shown with a maximum complexity of 32 assignments, as in tRNA. Codons are shown in sectors marked 1
st, 2
nd and 3
rd. Anticodons (Ac) are indicated (i.e., 34-[
A/G]AA-36). Anticodons that are not utilized are shown with red letters. No tRNA matches stop codons (UAA, UAG, UGA). Blue
34U indicates a modification to limit superwobbling such as
34cnm5U. As indicated above, some exceptions have been noted, but wobble
5C-U modifications to suppress superwobbling may have been universal at the inception of the first code. 37m1G is associated with
36A. 37t6A is associated with
36U. Column 1, row 3B,
34C modifications (orange) discriminate Ile and Met. The genetic code evolved primarily in columns, as indicated in the model. In column 1, ValRS-IA, LeuRS-IA, IleRS-IA and MetRS-IA are closely related enzymes (yellow type). In column 2, SerRS-IIA, ProRS-IIA and ThrRS-IIA are closely related enzymes (red type). In column 3, AspRS-IIB, AsnRS-IIB and HisRS-IIA are closely related (green type), and GluRS-IB, LysRS-IB (in Archaea) and GlnRS-IB (a eukaryotic innovation) are closely related (orange type). In column 4, ArgRS-IA and CysRS-IA are closely related. In row 1, TyrRS-IC and TrpRS-IC are closely related. A similar figure was previously published and is republished here with permission [
15,
16].
Figure 37.
A model for evolution of the first code. A codon-anticodon table is shown with a maximum complexity of 32 assignments, as in tRNA. Codons are shown in sectors marked 1
st, 2
nd and 3
rd. Anticodons (Ac) are indicated (i.e., 34-[
A/G]AA-36). Anticodons that are not utilized are shown with red letters. No tRNA matches stop codons (UAA, UAG, UGA). Blue
34U indicates a modification to limit superwobbling such as
34cnm5U. As indicated above, some exceptions have been noted, but wobble
5C-U modifications to suppress superwobbling may have been universal at the inception of the first code. 37m1G is associated with
36A. 37t6A is associated with
36U. Column 1, row 3B,
34C modifications (orange) discriminate Ile and Met. The genetic code evolved primarily in columns, as indicated in the model. In column 1, ValRS-IA, LeuRS-IA, IleRS-IA and MetRS-IA are closely related enzymes (yellow type). In column 2, SerRS-IIA, ProRS-IIA and ThrRS-IIA are closely related enzymes (red type). In column 3, AspRS-IIB, AsnRS-IIB and HisRS-IIA are closely related (green type), and GluRS-IB, LysRS-IB (in Archaea) and GlnRS-IB (a eukaryotic innovation) are closely related (orange type). In column 4, ArgRS-IA and CysRS-IA are closely related. In row 1, TyrRS-IC and TrpRS-IC are closely related. A similar figure was previously published and is republished here with permission [
15,
16].
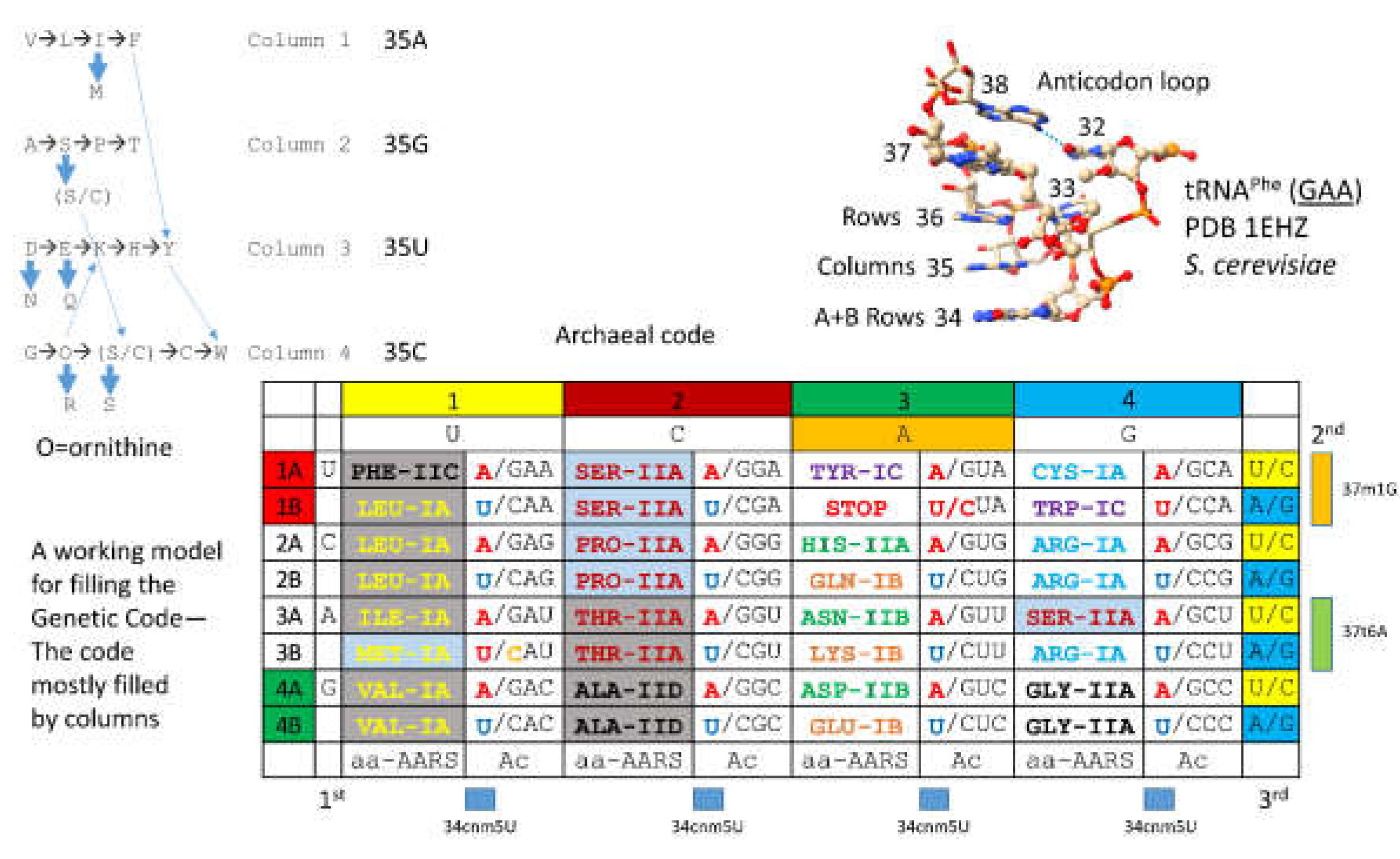
Figure 38.
Evolution of AARS enzymes. Phyre 2 homology scoring mostly to P. furiosus AARS sequences was used to draw the class II and class I AARS maps. GlyRS-IIA is homologous to ValRS-IA and IleRS-IA by sequence as indicated by the red arrow. AARS with separate editing active sites are shaded gray. AARS that have editing reactions only in their aminoacylating active sites are shaded pale yellow. Bacterial innovations are indicated (B). Archaeal-type AARS are indicated (A). GlnRS-IB was a eukaryotic innovation (E). GlyRS-IIA appears to be the root of all class II and class I AARS. A primitive ValRS-IA appears to be the root of all class I AARS. PheRS-IIC and AlaRS-IID are in bold because these enzymes may have replaced PheRS-IC and AlaRS-IIA before LUCA. Sep for o-phosphoserine. Pyl for pyrrolysine.
Figure 38.
Evolution of AARS enzymes. Phyre 2 homology scoring mostly to P. furiosus AARS sequences was used to draw the class II and class I AARS maps. GlyRS-IIA is homologous to ValRS-IA and IleRS-IA by sequence as indicated by the red arrow. AARS with separate editing active sites are shaded gray. AARS that have editing reactions only in their aminoacylating active sites are shaded pale yellow. Bacterial innovations are indicated (B). Archaeal-type AARS are indicated (A). GlnRS-IB was a eukaryotic innovation (E). GlyRS-IIA appears to be the root of all class II and class I AARS. A primitive ValRS-IA appears to be the root of all class I AARS. PheRS-IIC and AlaRS-IID are in bold because these enzymes may have replaced PheRS-IC and AlaRS-IIA before LUCA. Sep for o-phosphoserine. Pyl for pyrrolysine.
Figure 39.
Relationship of AARS enzymes and the genetic code. Column 1 amino acids and AARS are on an orange background. Column 2 amino acids and AARS are on a blue background. Column 3 amino acids and AARS are on a green background. Column 4 amino acids and AARS are on a red background. Row 1 amino acids and AARS are on a yellow background. Other indications are as in
Figure 38. .
Figure 39.
Relationship of AARS enzymes and the genetic code. Column 1 amino acids and AARS are on an orange background. Column 2 amino acids and AARS are on a blue background. Column 3 amino acids and AARS are on a green background. Column 4 amino acids and AARS are on a red background. Row 1 amino acids and AARS are on a yellow background. Other indications are as in
Figure 38. .