Submitted:
12 March 2026
Posted:
13 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- Better diagnosis and treatment planning: Spiro digital EEG monitoring allows for highly accurate detection and management of neurological disorders.
- Reduction of health economics costs: Improving diagnostic accuracy reduces misdiagnosis and subsequent incorrect treatment, which in turn lowers drug-related side effects and the corresponding costs.
- Better access to health care: Digital EEG systems provide easy remote data retrieval and analysis, thereby enhancing health service accessibility for those living in rural or less served parts of Oman.
- Improved patient outcome: Early and accurate diagnosis enables the appropriate intervention when required, resulting in better clinical outcomes and quality of life.
2. Literature Review
2.1. Automated and Machine-Learning-Based RBD Detection
2.2. Deep Learning and Automated Sleep Staging
2.3. Clinical Reviews and Standardization Efforts
2.4. Summary of Reviewed Studies (2020–2025)
2.5. Critical Analysis and Research Gaps
3. Materials and Methods
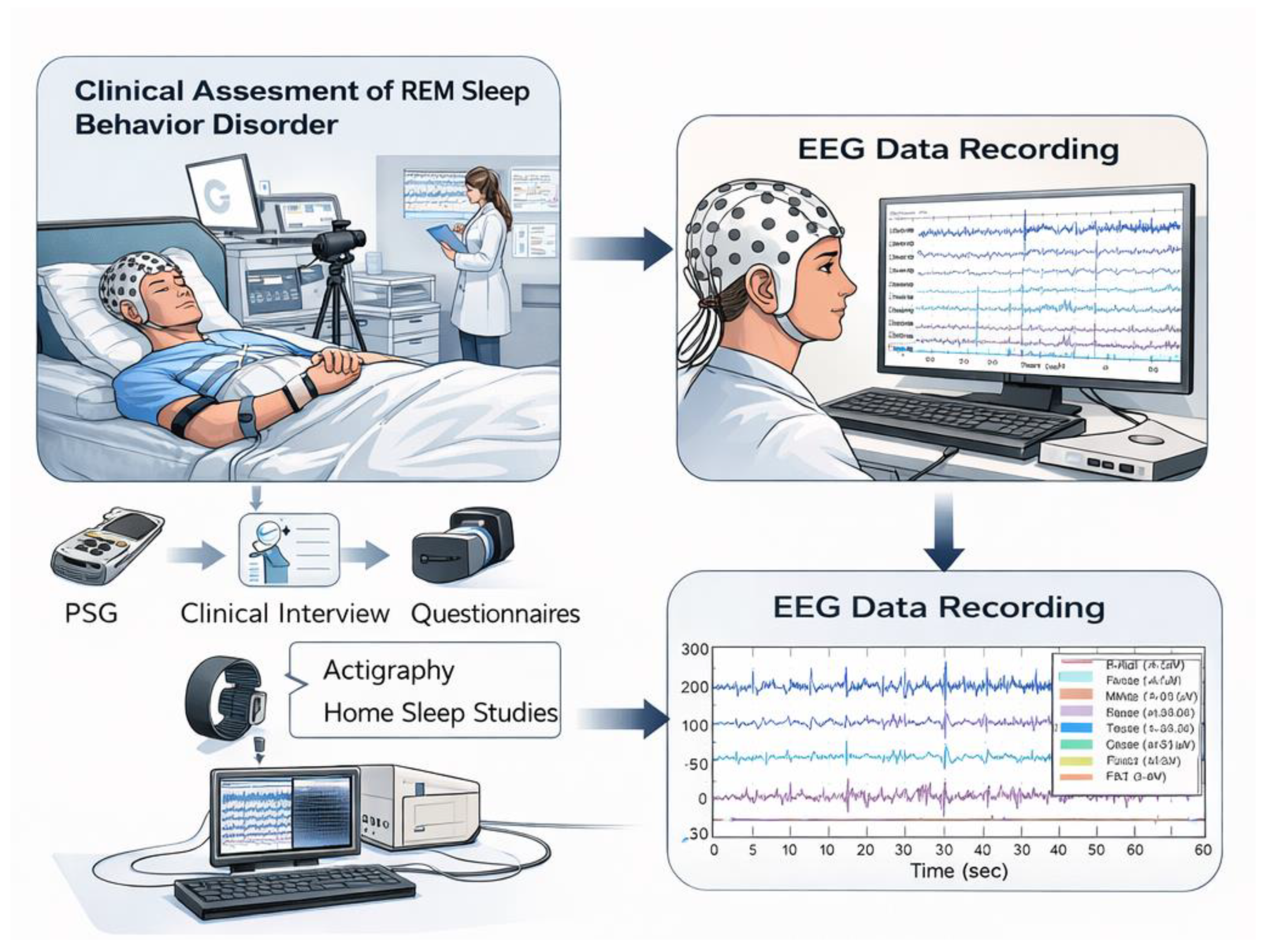
3.1. Clinical Assessment of REM Sleep Behavior Disorder
3.2. Processing of Cyclic Alternating Pattern (CAP) EEG Data
3.3. EEG Data Acquisition
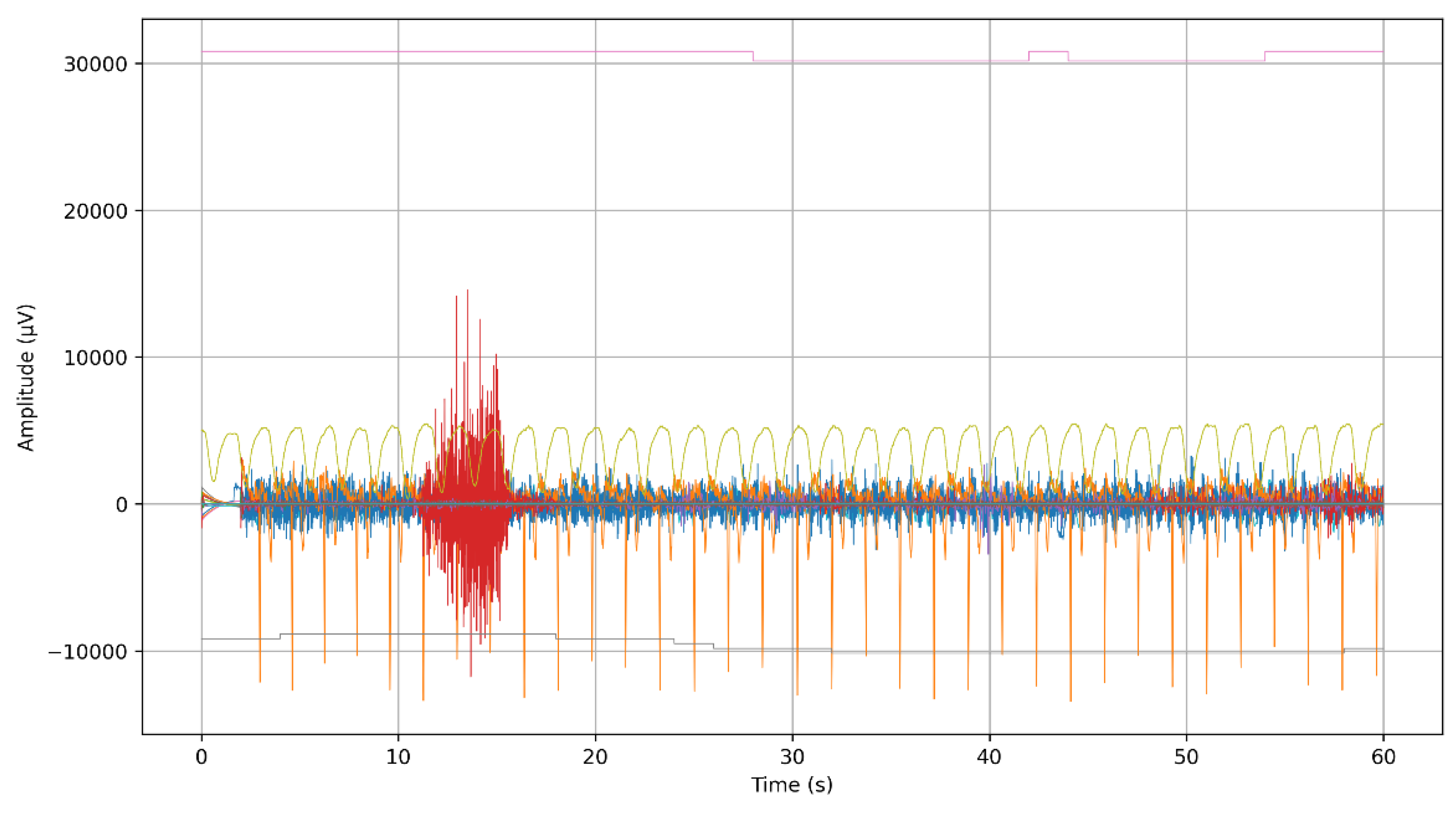
3.4. Signal Preprocessing
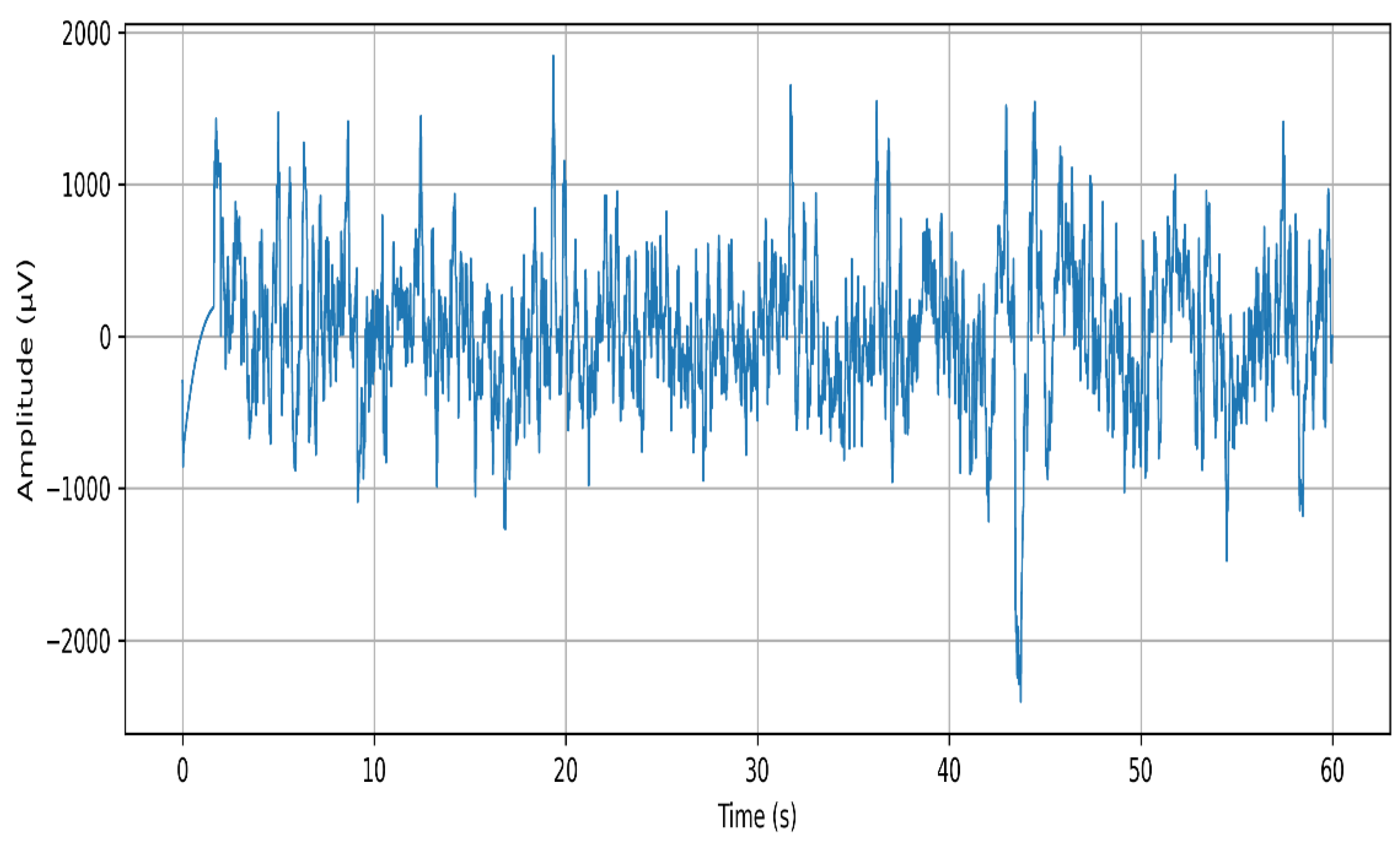
3.4.1. Band-Pass Filtering
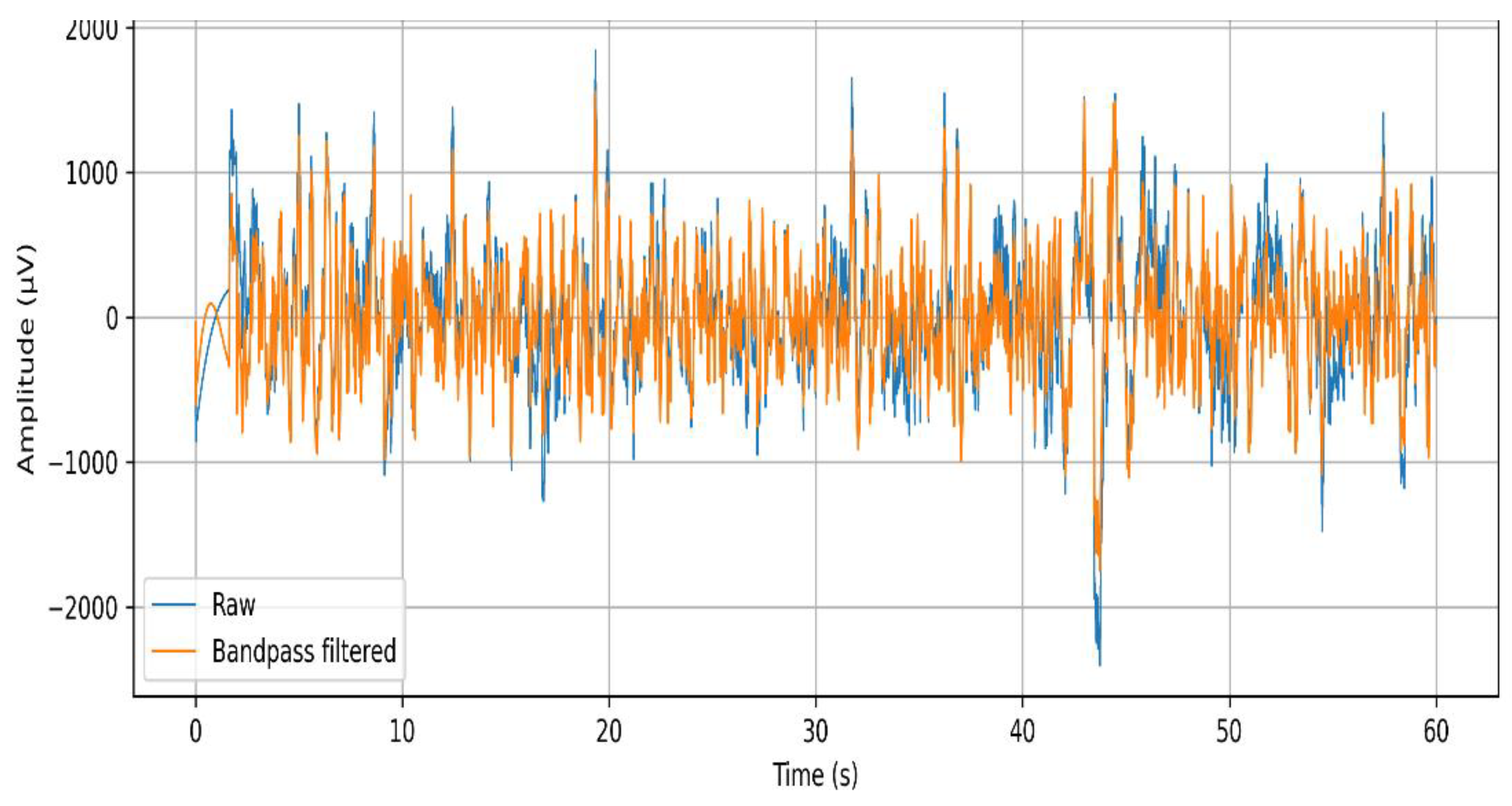
3.4.2. Notch Filtering
3.4.3. Epoch Segmentation and Artifact Rejection
- Amplitude thresholding (|x[n]| > 100 μV)
- Variance-based abnormality detection
- Visual inspection
3.5. Channel Selection and REM Stage Justification
3.6. Power Spectral Density Estimation
- is the Hamming window,
- is the segment length,
- is the number of overlapping segments,
- is the normalization factor.
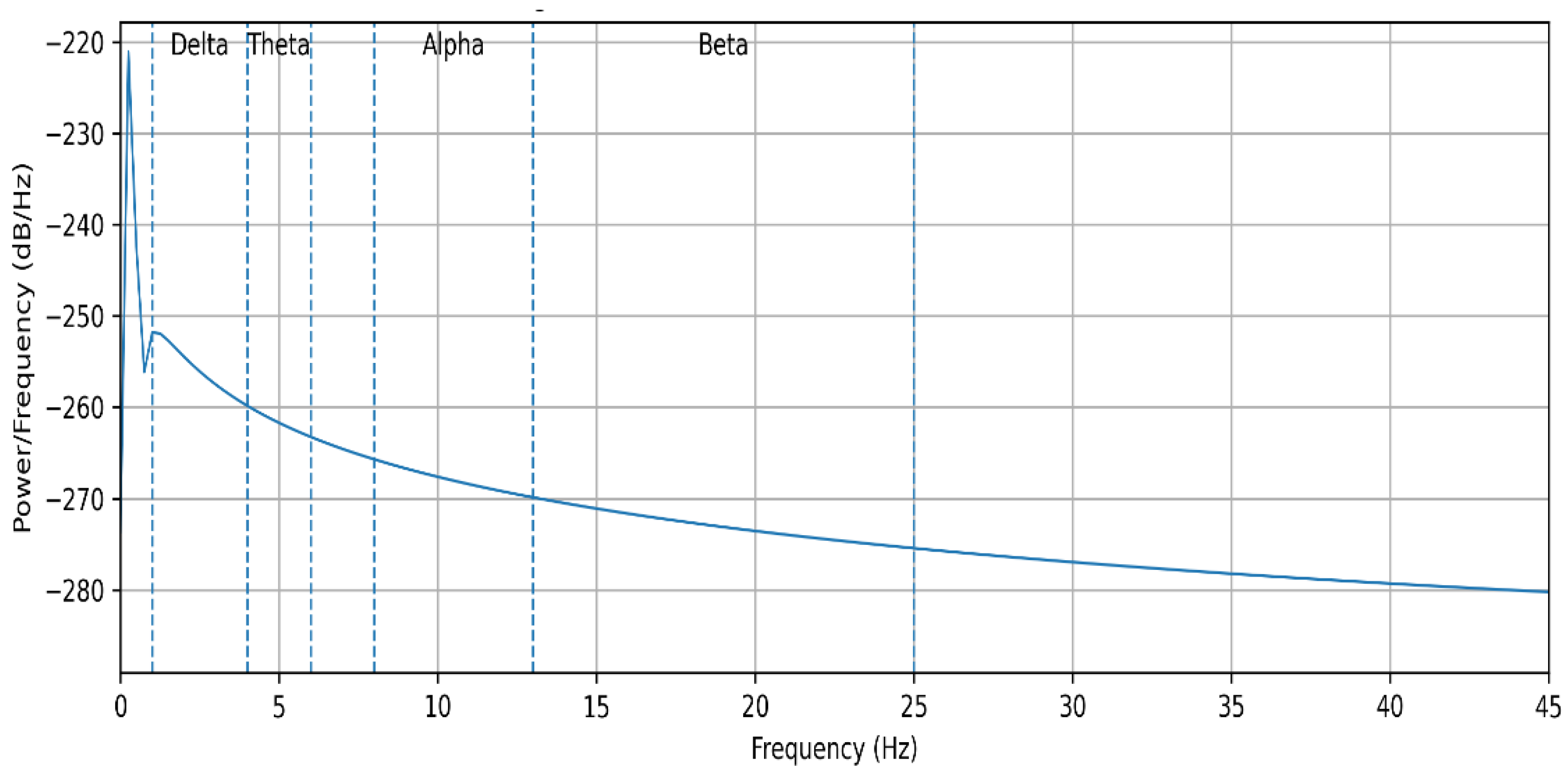
3.7. Spectral Feature Extraction
| Band | Frequency Range (Hz) | Clinical Relevance |
| Delta | 1–4 | Deep sleep activity |
| Theta | 4–6 | REM and transitional sleep |
| Alpha | 8–13 | Relaxed wakefulness |
| Beta | 13–25 | Cortical activation |
3.8. Normalization and Disorder Detection Algorithm
- Elevated beta activity during REM → RBD
- Altered theta–alpha ratios → Insomnia
- Spectral instability and delta suppression → Sleep-disordered breathing
4. Results
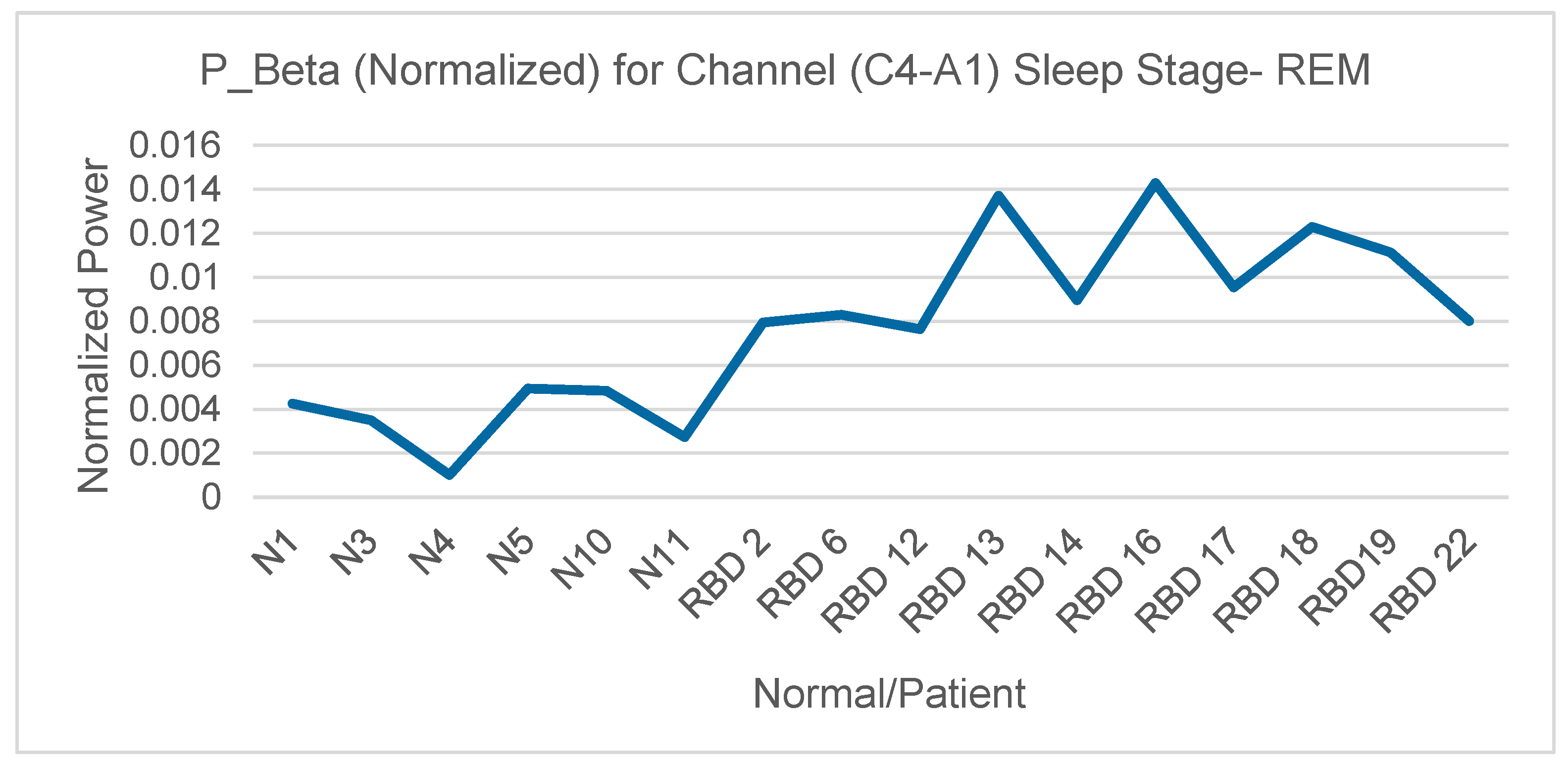
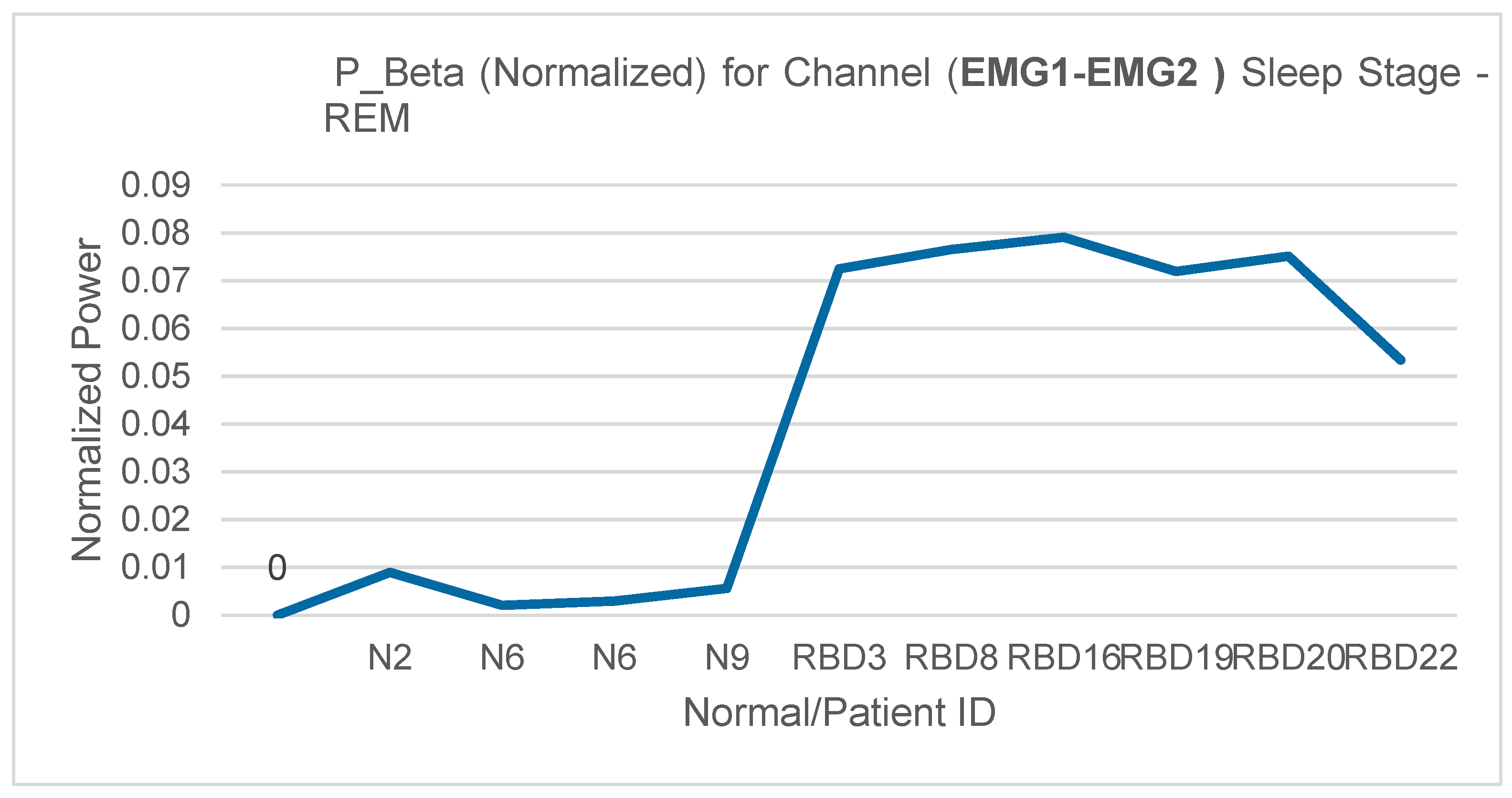
5. Conclusion
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
References
- Al-Dhahir, R.S.; Osorio, R.S.; O’Rourke, M.A. Machine learning for the diagnosis of neurological disorders using electroencephalography. J. Neural Eng. 2021, 18, 036006. [CrossRef]
- O’Rourke, M.A.; Osorio, R.S.; Bronzino, J.A. Artificial intelligence in medical imaging: Current status and future directions. IEEE J. Biomed. Health Inform. 2022, 26, 1178–1189. [CrossRef]
- Kostopoulos, D.K.; Georgopoulos, A.P. Machine learning in the diagnosis and management of neurological disorders. Nat. Rev. Neurol. 2021, 17, 221–234. [CrossRef]
- Chaudhary, R.A.; Osorio, R.S. Deep learning in neuroimaging: Current perspectives and future directions. NeuroImage 2020, 222, 116871. [CrossRef]
- Tzallas, A.T.; Tsipouras, M.G.; Fotiadis, D.I. Epileptic seizure detection in EEGs using time–frequency analysis. IEEE Trans. Inf. Technol. Biomed. 2009, 13, 703–710. [CrossRef]
- Bronzino, J.A.; Osorio, R.S. Artificial intelligence in neurology: Current status and future directions. Lancet Neurol. 2021, 20, 687–699. [CrossRef]
- Gómez, J.C.; del Río, M.A. Deep learning in neurological disorders: Current status and future directions. J. Clin. Med. 2021, 10, 1026. [CrossRef]
- Cooray, N.; Angelini, E.; Mace, M.; et al. Detection of REM sleep behaviour disorder by automated polysomnography analysis. Clin. Neurophysiol. 2019, 130, 505–514. [CrossRef]
- Cooray, N.; Beniczky, S.; Linne, M.; et al. Screening for REM sleep behaviour disorder with a minimal set of sensors. Clin. Neurophysiol. 2021, 132, 904–913. [CrossRef]
- Röthenbacher, A.; Cesari, M.; Doppler, C.E.J.; et al. RBDtector: An open-source software to detect REM sleep without atonia according to visual scoring criteria. Sci. Rep. 2022, 12, 25163. [CrossRef]
- Perslev, M.; Darkner, S.; Kempfner, L.; et al. U-Sleep: Resilient high-frequency sleep staging. npj Digit. Med. 2021, 4, 72. [CrossRef]
- Högl, B.; Stefani, A.; Videnovic, A. Idiopathic REM sleep behaviour disorder and neurodegeneration—An update. Nat. Rev. Neurol. 2018, 14, 40–55. [CrossRef]
- Postuma, R.B.; Arnulf, I.; Högl, B.; et al. Risk and predictors of dementia and parkinsonism in idiopathic REM sleep behaviour disorder. Brain 2019, 142, 744–759. [CrossRef]
- Frauscher, B.; Iranzo, A.; Gaig, C.; et al. Quantification of electromyographic activity during REM sleep in REM sleep behaviour disorder. Sleep 2008, 31, 1149–1159. [CrossRef]
- Sasai-Sakuma, T.; Inoue, Y.; et al. Quantitative assessment of isolated REM sleep without atonia. Sleep 2014, 37, 297–306. [CrossRef]
- Puligheddu, M.; Figorilli, M.; et al. Quantification of REM sleep without atonia: A review of scoring methods. Sleep Med. Rev. 2023, 67, 101708. [CrossRef]
- Jo, H.G.; Park, J.Y.; Lee, C.K.; An, S.K.; Yoo, S.K. Genetic fuzzy classifier for sleep stage identification. Comput. Biol. Med. 2010, 40, 629–634. [CrossRef]
- Bigan, C.; Woolfson, M.S. Time–frequency analysis of short segments of biomedical data. IEE Proc. Sci. Meas. Technol. 2000, 147, 368–373. [CrossRef]
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; et al. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000, 101, e215–e220. [CrossRef]
- Ebrahimi, F.; Mikaeili, M.; Estrada, E.; Nazeran, H. Automatic sleep stage classification using EEG and neural networks. In Proc. IEEE EMBS; 2008, 1151–1154. [CrossRef]
- Garcia-Molina, G.; Bellesi, M.; Pastoor, S.; et al. Online single-channel EEG-based automatic sleep staging. In Eng. Psychol. Cogn. Ergon.; Springer, 2013, 333–342. [CrossRef]
- Iranzo, A.; Santamaría, J.; Tolosa, E. The clinical and pathophysiological relevance of REM sleep behavior disorder. Sleep Med. Rev. 2009, 13, 385–401. [CrossRef]
- Schenck, C.H.; Boeve, B.F.; Mahowald, M.W. Delayed emergence of parkinsonism or dementia in idiopathic RBD. Neurology 2003, 60, 1134–1140. [CrossRef]
- Ferri, R.; Manconi, M.; Plazzi, G.; et al. Quantitative EMG analysis during REM sleep in RBD. J. Sleep Res. 2008, 17, 89–100. [CrossRef]
- Sors, A.; Bonnet, S.; Mirek, E.; Vercueil, L.; Payen, J.F. CNN-based sleep stage scoring from single-channel EEG. Biomed. Signal Process. Control 2018, 42, 107–114. [CrossRef]
- Phan, H.; Andreotti, F.; Cooray, N.; et al. Joint classification and prediction framework for sleep staging. IEEE Trans. Biomed. Eng. 2019, 66, 1285–1296. [CrossRef]
- Stephansen, J.B.; Olesen, A.N.; Olsen, M.; et al. Neural network analysis enables efficient diagnosis of narcolepsy. Nat. Commun. 2018, 9, 5229. [CrossRef]
- Kempfner, J.; Jennum, P.; Sørensen, H.B.D. Automatic detection of REM sleep behaviour disorder based on EMG. Clin. Neurophysiol. 2014, 125, 2328–2336. [CrossRef]
- Cesari, M.; Stefani, A.; Figorilli, M.; et al. Video-polysomnographic criteria for RBD. Sleep Med. 2018, 52, 22–29. [CrossRef]
- Xu, Y.; Yang, J.; Ming, W.; Wang, S.; Sawan, M. Probabilistic real-time EEG-based detection with reduced latency. Expert Syst. Appl. 2024, 236, 121359. [CrossRef]
- Sors, A.; Bonnet, S.; Mirek, S.; Vercueil, L.; Payen, J.F. A convolutional neural network for sleep stage scoring from raw EEG signals. Physiol. Meas. 2018, 39, 124005. [CrossRef]
- Yildirim, O.; Baloglu, U.B.; Acharya, U.R. A deep learning model for automated sleep stage classification using PSG signals. Int. J. Environ. Res. Public Health 2019, 16, 599. [CrossRef]
- Phan, H.; Tran, M.C.; Koch, P.; Mertins, A. Towards more accurate automatic sleep staging via deep transfer learning. IEEE Trans. Biomed. Eng. 2021, 68, 1787–1798. [CrossRef]
- Aljalal, M.; Alshammari, T.; Alshammari, A.; Alturki, F. Sleep stage classification using EEG signal analysis based on empirical mode decomposition and deep learning. Sensors 2021, 21, 722. [CrossRef]
- Li, Y.; Pan, W.; Wang, F.; Wang, J. A hybrid deep learning model for sleep stage classification based on EEG signals. IEEE Access 2020, 8, 118772–118781. [CrossRef]
- Zhu, T.; Luo, W.; Yu, F.; Zhang, C. Sleep stage classification based on multi-scale feature fusion using EEG signals. Biomed. Signal Process. Control 2021, 70, 102968. [CrossRef]
- Boashash, B. Time–Frequency Signal Analysis and Processing: A Comprehensive Reference; Academic Press: Oxford, UK, 2015. [CrossRef]
- Mallat, S. A Wavelet Tour of Signal Processing: The Sparse Way, 3rd ed.; Academic Press: Burlington, MA, USA, 2009. [CrossRef]
- Cohen, L. Time–Frequency Analysis; Prentice Hall: Englewood Cliffs, NJ, USA, 1995.
- Boashash, B.; Khan, N.A.; Ben-Jabeur, T. Time–frequency features for pattern recognition using high-resolution distributions. IEEE Signal Process. Mag. 2015, 32, 79–93. [CrossRef]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M.; Gertych, A.; Tan, R.S. Deep convolutional neural network for the automated detection and diagnosis of seizure using EEG signals. Comput. Biol. Med. 2018, 100, 270–278. [CrossRef]
- Dauwels, J.; Vialatte, F.; Musha, T.; Cichocki, A. A comparative study of synchrony measures for the early diagnosis of Alzheimer’s disease based on EEG. NeuroImage 2010, 49, 668–693. [CrossRef]
- Sharma, M.; Pachori, R.B.; Acharya, U.R. Automated sleep stage classification based on iterative filtering of EEG signals. Neural Comput. Appl. 2019, 31, 6759–6773. [CrossRef]
- Okano, K.; Kaczmarzyk, J.R.; Dave, N.; et al. Sleep quality, duration, and consistency are associated with better academic performance in college students. NPJ Sci. Learn. 2019, 4, 16. [CrossRef]
- Mindell, J.A.; Meltzer, L.J.; Carskadon, M.A.; et al. Developmental aspects of sleep hygiene: Findings from the 2004 National Sleep Foundation Sleep in America Poll. Sleep Med. 2009, 10, 771–779. [CrossRef]
- Berry, R.B.; Brooks, R.; Gamaldo, C.; et al. AASM scoring manual updates for 2017 (Version 2.4). J. Clin. Sleep Med. 2017, 13, 665–666. [CrossRef]
- Rechichi, I.; Iadarola, A.; Zibetti, M.; et al. Assessing REM sleep behaviour disorder: From machine learning classification to the definition of a continuous dissociation index. Int. J. Environ. Res. Public Health 2022, 19, 248. [CrossRef]

| Study (Year) | Source | Dataset / Sample | Method / Sensors | Key Findings |
|---|---|---|---|---|
| Cooray et al., 2021 | Clinical Neurophysiology | Minimal-sensor screening cohort (RBD vs controls) | Minimal sensors (EOG + EMG; optional ECG) + classical ML (e.g., Random Forest) | Demonstrated strong screening capability using a reduced sensor set; suitable for scalable screening. |
| Perslev et al., 2021 | npj Digital Medicine | 15,660 PSG recordings | Deep learning–based automated sleep staging (U-Sleep) | Performance comparable to expert scorers; robust and generalizable across large datasets. |
| Röthenbacher et al., 2022 | Scientific Reports | RSWA/RBD detection cohort (software validation) | SINBAR-based RSWA scoring (RBDtector, open-source) | Reported high diagnostic performance for RSWA detection; supports reproducible RBD/RWA screening. |
| Postuma et al., 2019 | Brain | Longitudinal idiopathic RBD cohort | Clinical follow-up with neurodegenerative outcome tracking | Confirmed iRBD as a strong prodromal marker for Parkinsonism/dementia with quantified risk predictors. |
| Puligheddu et al., 2023 | Sleep Medicine Reviews | Review of RSWA quantification literature | Comparative scoring and methodology review | Emphasized the need for harmonization and standardization of RSWA scoring methods. |
| Xu et al., 2024 | Expert Systems with Applications | EEG-based datasets | Probabilistic real-time detection framework | Reduced detection latency with high predictive performance; supports real-time clinical decision workflows. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




