Submitted:
11 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Cloning and Transformation of BjYSL6.1, BjYSL6.4 in Schizosaccharomyces Pombe Vector pDES177N
2.3. Confocal Microscopy of GFP: BjYSL6.1-3’UTR and GFP: BjYSL6.4-3’UTR Constructs in S. pombe
2.4. Interaction Between BjYSL6 Proteins with BjNRAMP4.1 and BjHCF164
2.5. β-Galactosidase Assay Using Agarose Overlay
2.6. Agrobacterium Transformation
2.7. Localization of BjYSL6.1 and BjYSL6.4 in Brassica Determined by Confocal Microscopy
2.8. Bimolecular Fluorescence Complementation (BiFC) Assay to Study Interaction in Planta
2.9. GUS Staining of Transiently Transformed N. benthamiana Leaves
2.10. qRT-PCR PCR of BjYSL6 and BjNRAMP4.1 Gene
3. Results
3.1. BjYSL6.1 and BjYSL6.4 Are Homologs of Arabidopsis Thaliana YSL6 Protein AtYSL6
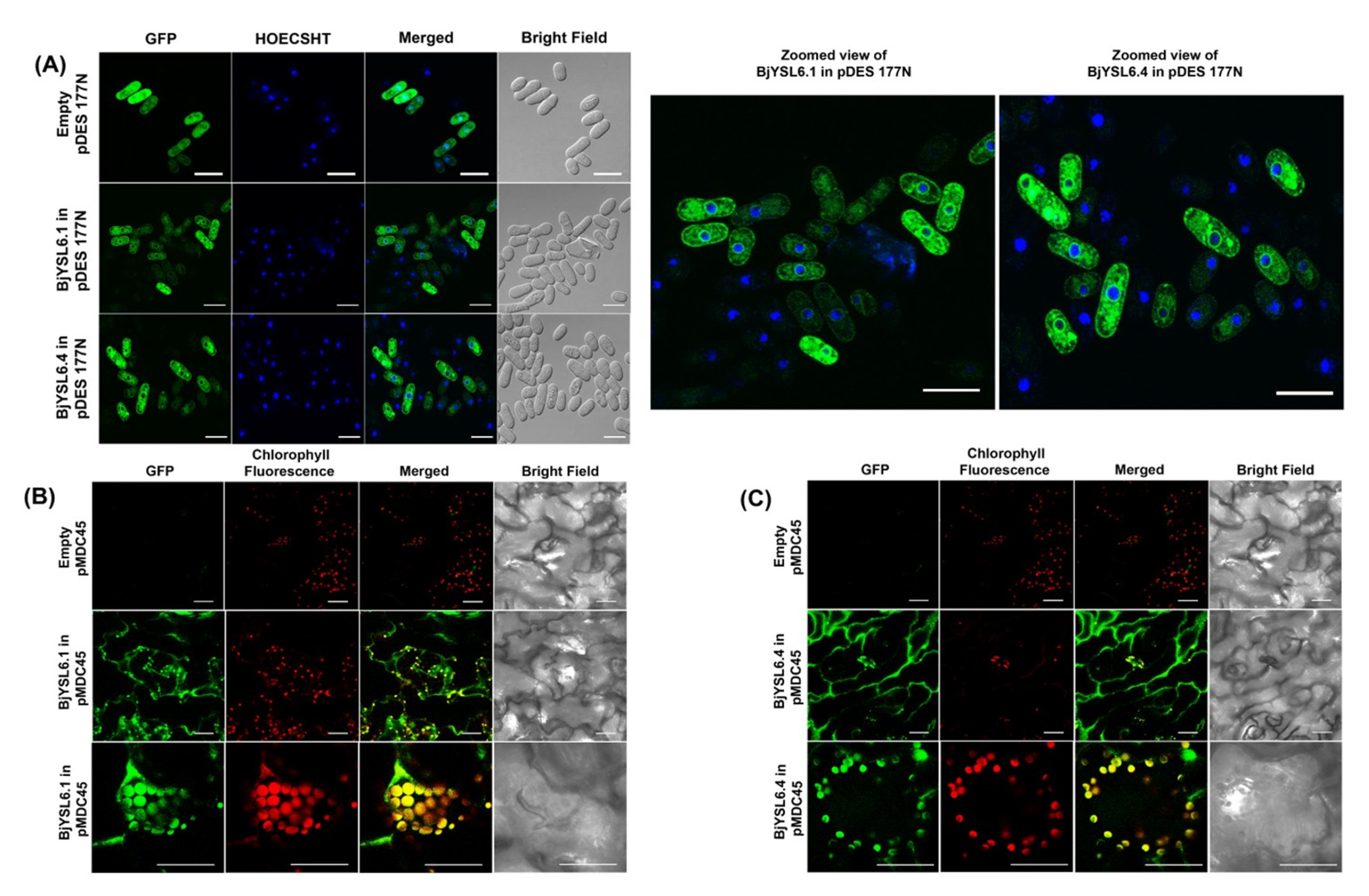
3.2. BjYSL6.1 and BjYSL6.4 Express in the Endomembranes of Schizosaccharomyces Pombe and Shoot Membrane, and Chloroplast of B. juncea
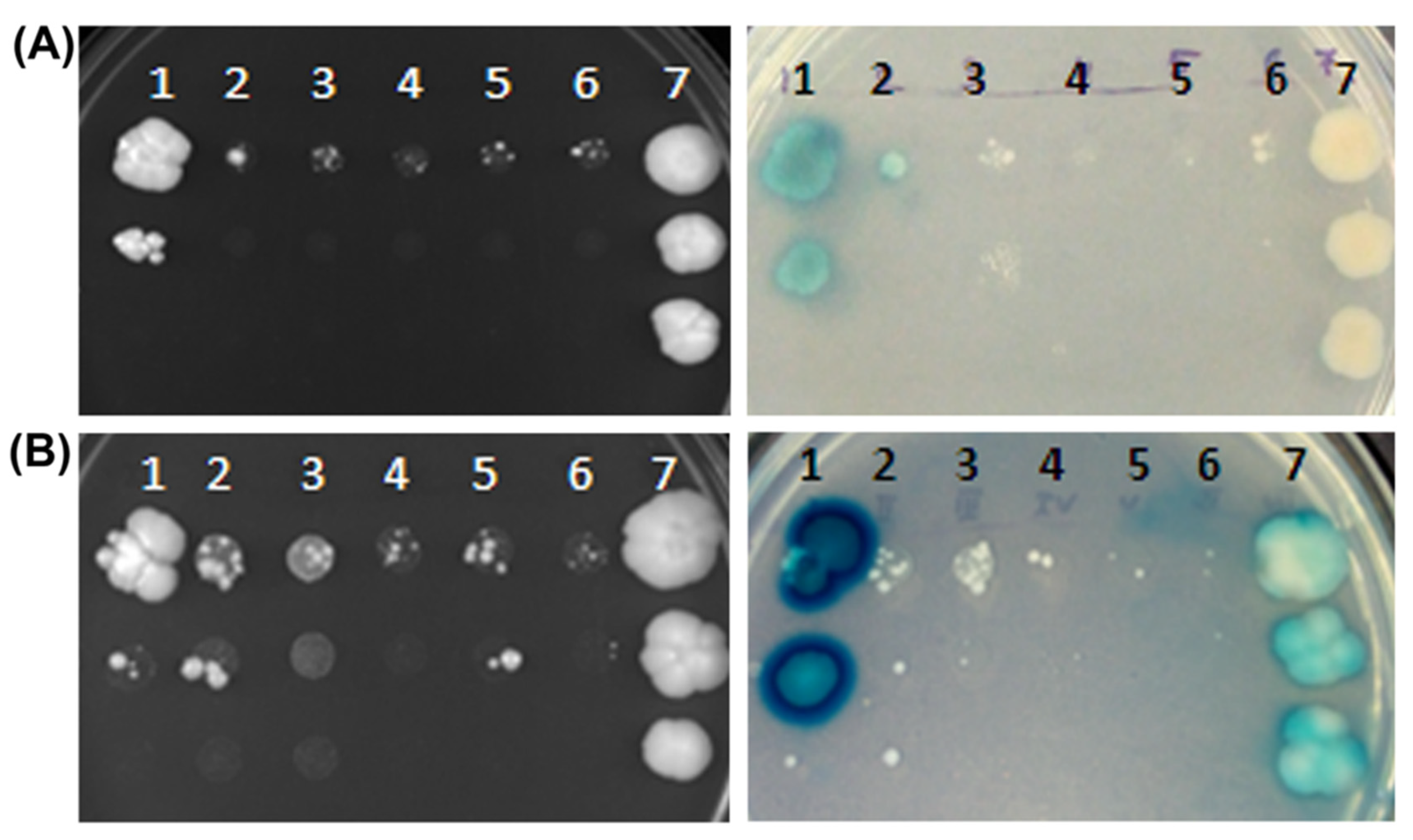
3.3. BjYSL6.1 and BjYSL6.4 Exhibit Distinct Interaction Patterns with BjNRAMP4.1 and BjHCF164 in Yeast Two-Hybrid Assays
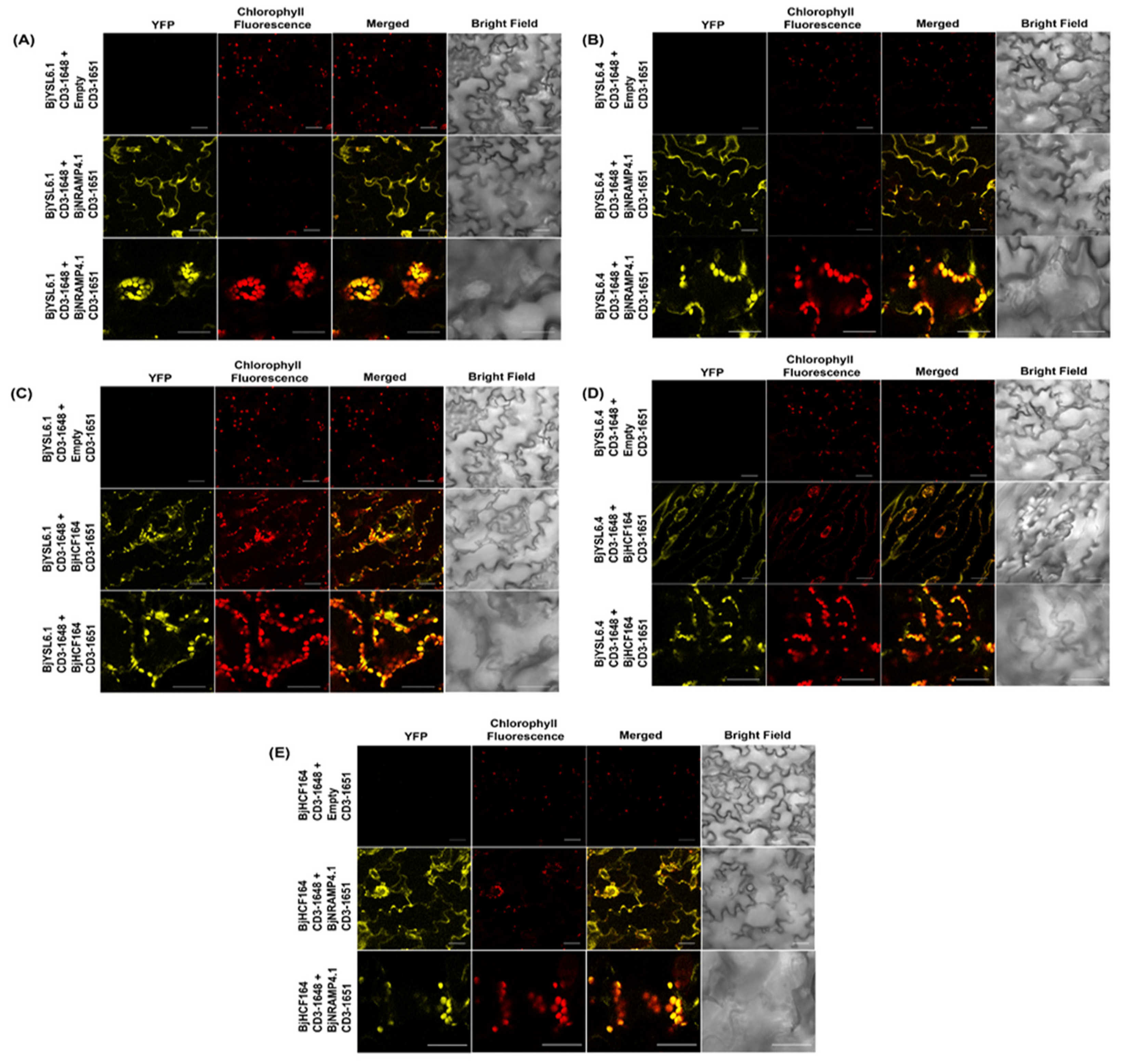
3.4. BjYSL6.1 and BjYSL6.4 Interact with BjNRAMP4.1 and BjHCF164 in Brassica juncea Leaf Albeit Differentially
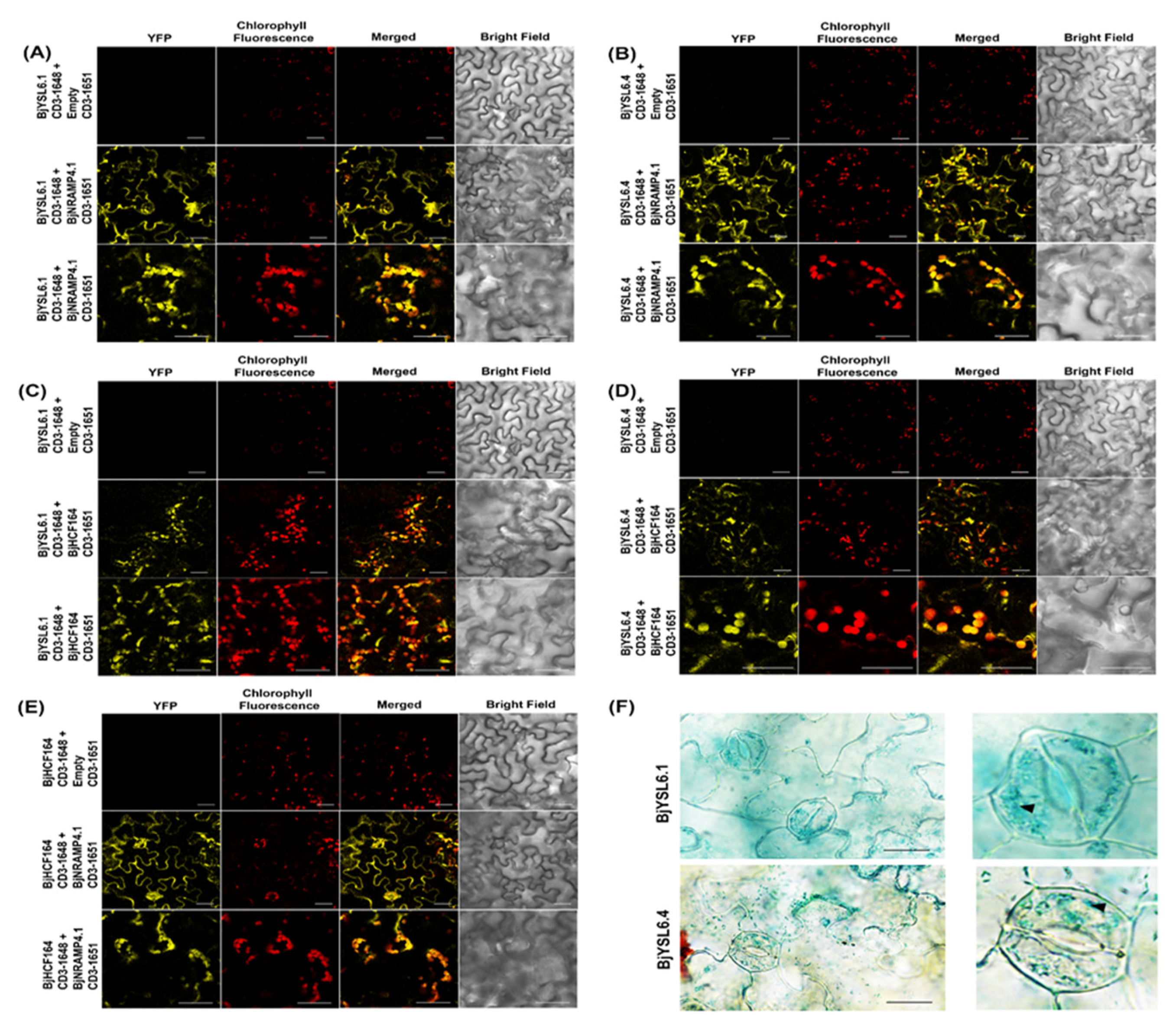
3.5. BjYSL6.1 and BjYSL6.4 Interact with BjNRAMP4.1 and BjHCF164 in Nicotiana benthamiana Leaves
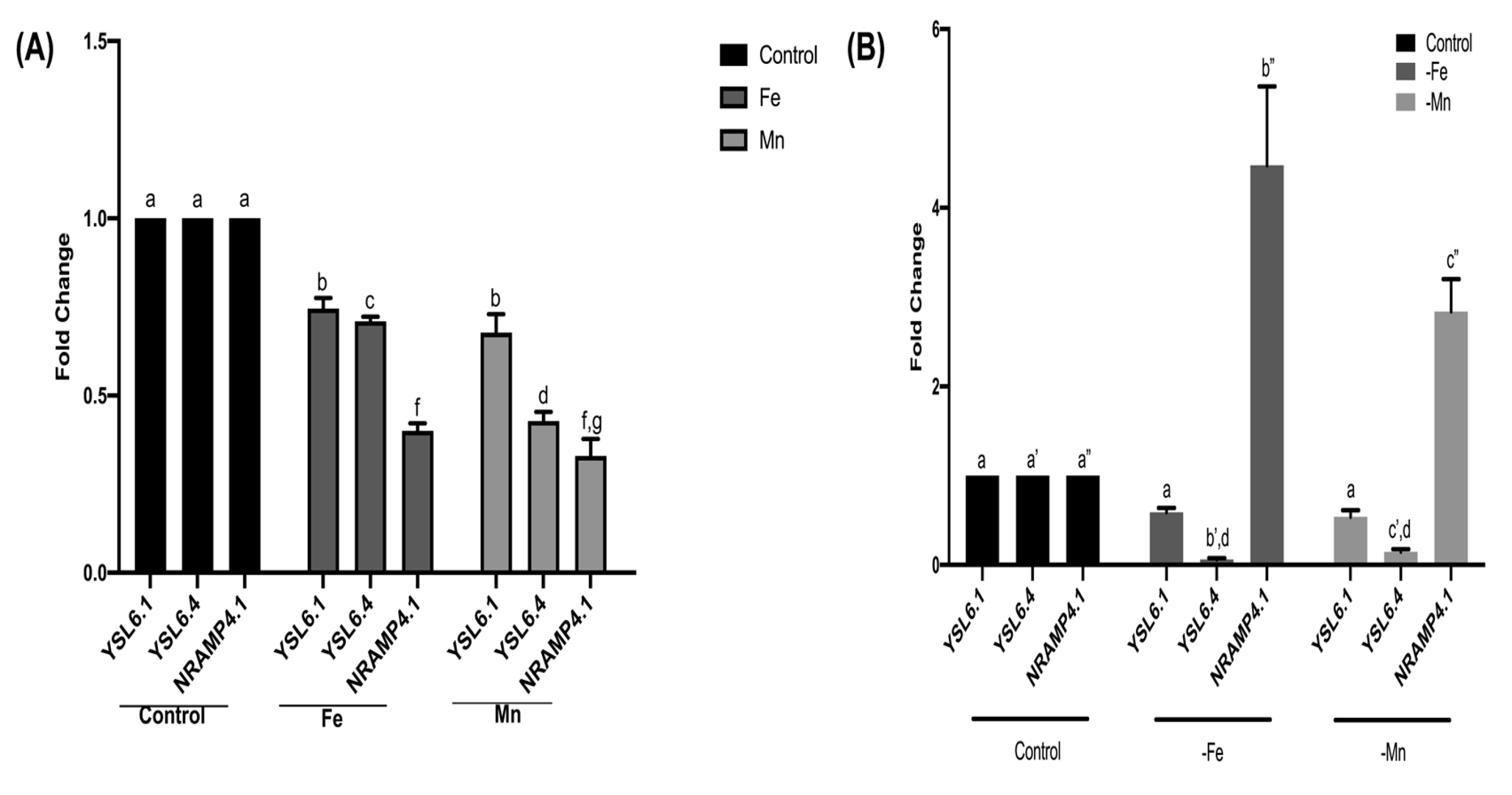
3.6. BjYSL6.1, BjYSL6.4, and BjNRAMP4.1 Exhibit Opposite Transcriptional Expression Under Metal Excess and Deficiency Conditions
4. Discussion
5. Conclusions
6. Limitations and Perspectives
Supplementary Materials
Acknowledgments
Conflicts of Interest
References
- Curie, C.; Panaviene, Z.; Loulergue, C.; Dellaporta, S.L.; Briat, J.-F.; Walker, E.L. Maize yellow stripe1 encodes a membrane protein directly involved in Fe (III) uptake. Nature 2001, 409, 346–349. [Google Scholar] [CrossRef]
- Roberts, L.A.; Pierson, A.J.; Panaviene, Z.; Walker, E.L. Yellow stripe1. Expanded roles for the maize iron-phytosiderophore transporter. Plant Physiology 2004, 135, 112–120. [Google Scholar] [CrossRef]
- Hopkins, B.G.; Jolley, V.D.; Brown, J.C. Plant utilization of iron solubilized by oat phytosiderophore. Journal of plant nutrition 1992, 15, 1599–1612. [Google Scholar] [CrossRef]
- Jolley, V.D.; Brown, J.C. Differential response of Fe-efficient corn and Fe-inefficient corn and oat to phytosiderophore released by Fe-efficient Coker 227 oat. Journal of plant nutrition 1991, 14, 45–58. [Google Scholar] [CrossRef]
- von Wirén, N.; Marschner, H.; Römheld, V. Uptake kinetics of iron-phytosiderophores in two maize genotypes differing in iron efficiency. Physiologia Plantarum 1995, 93, 611–616. [Google Scholar] [CrossRef]
- von Wirén, N.; Mori, S.; Marschner, H.; Romheld, V. Iron inefficiency in maize mutant ys1 (Zea mays L. cv Yellow-Stripe) is caused by a defect in uptake of iron phytosiderophores. Plant Physiology 1994, 106, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Eide, D.; Broderius, M.; Fett, J.; Guerinot, M.L. A novel iron-regulated metal transporter from plants identified by functional expression in yeast. Proceedings of the National Academy of Sciences of the United States of America 1996, 93, 5624–5628. [Google Scholar] [CrossRef]
- Murata, Y.; Ma, J.F.; Yamaji, N.; Ueno, D.; Nomoto, K.; Iwashita, T. A specific transporter for iron (III)–phytosiderophore in barley roots. The Plant journal 2006, 46, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Schaaf, G.; Ludewig, U.; Erenoglu, B.E.; Mori, S.; Kitahara, T.; von Wirén, N. ZmYS1 functions as a proton-coupled symporter for phytosiderophore-and nicotianamine-chelated metals. Journal of Biological Chemistry 2004, 279, 9091–9096. [Google Scholar] [CrossRef]
- Song, Z.; Li, S.; Li, Y.; Zhou, X.; Liu, X.; Yang, W.; Chen, R. Identification and characterization of yellow stripe-like genes in maize suggest their roles in the uptake and transport of zinc and iron. BMC Plant Biol 2024, 24, 3. [Google Scholar] [CrossRef]
- Dai, J.; Wang, N.; Xiong, H.; Qiu, W.; Nakanishi, H.; Kobayashi, T.; Nishizawa, N.K.; Zuo, Y. The Yellow Stripe-Like (YSL) Gene Functions in Internal Copper Transport in Peanut. Genes (Basel) 2018, 9. [Google Scholar] [CrossRef]
- Kumar, A.; Joon, R.; Singh, G.; Singh, J.; Pandey, A.K. The multifaceted role of YSL proteins: Iron transport and emerging functions in plant metal homeostasis. Biochim Biophys Acta Gen Subj 2025(1869), 130792. [CrossRef]
- Yuan, H.; Duan, Z.; Liang, G. Genome-wide identification and expression analysis of yellow stripe-like (YSL) genes in Nicotiana tabacum. Metallomics 2025, 17. [Google Scholar] [CrossRef] [PubMed]
- Finazzi, G.; Petroutsos, D.; Tomizioli, M.; Flori, S.; Sautron, E.; Villanova, V.; Rolland, N.; Seigneurin-Berny, D. Ions channels/transporters and chloroplast regulation. Cell Calcium 2015, 58, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Conte, S.S.; Chu, H.H.; Rodriguez, D.C.; Punshon, T.; Vasques, K.A.; Salt, D.E.; Walker, E.L. Arabidopsis thaliana Yellow Stripe1-Like4 and Yellow Stripe1-Like6 localize to internal cellular membranes and are involved in metal ion homeostasis. Front Plant Sci 2013, 4, 283. [Google Scholar] [CrossRef] [PubMed]
- Divol, F.; Couch, D.; Conéjéro, G.; Roschzttardtz, H.; Mari, S.; Curie, C. The Arabidopsis YELLOW STRIPE LIKE4 and 6 Transporters Control Iron Release from the. The Plant Cell 2013, 25, 1040–1055. [Google Scholar] [CrossRef] [PubMed]
- Gruenheid, S.; Pinner, E.; Desjardins, M.; Gros, P. Natural resistance to infection with intracellular pathogens: the Nramp1 protein is recruited to the membrane of the phagosome. The Journal of experimental medicine 1997, 185, 717–730. [Google Scholar] [CrossRef]
- Cailliatte, R.; Schikora, A.; Briat, J.-F.; Mari, S.; Curie, C. High-affinity manganese uptake by the metal transporter NRAMP1 is essential for Arabidopsis growth in low manganese conditions. The Plant Cell 2010, 22, 904–917. [Google Scholar] [CrossRef]
- Curie, C.; Alonso, J.M.; Jean, M.L.; Ecker, J.R.; Briat, J.-F. Involvement of NRAMP1 from Arabidopsis thaliana in iron transport. Biochemical Journal 2000, 347, 749–755. [Google Scholar] [CrossRef]
- Lanquar, V.; Ramos, M.S.; Lelièvre, F.; Barbier-Brygoo, H.; Krieger-Liszkay, A.; Krämer, U.; Thomine, S. Export of vacuolar manganese by AtNRAMP3 and AtNRAMP4 is required for optimal photosynthesis and growth under manganese deficiency. Plant physiology 2010, 152, 1986–1999. [Google Scholar] [CrossRef]
- Yang, M.; Zhang, W.; Dong, H.; Zhang, Y.; Lv, K.; Wang, D.; Lian, X. OsNRAMP3 is a vascular bundles-specific manganese transporter that is responsible for manganese distribution in rice. PLoS One 2013, 8, e83990. [Google Scholar] [CrossRef]
- Cailliatte, R.; Lapeyre, B.; Briat, J.-F.; Mari, S.; Curie, C. The NRAMP6 metal transporter contributes to cadmium toxicity. Biochemical Journal 2009, 422, 217–228. [Google Scholar] [CrossRef] [PubMed]
- Colangelo, E.P.; Guerinot, M.L. Put the metal to the petal: metal uptake and transport throughout plants. Current opinion in plant biology 2006, 9, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Gunshin, H.; Mackenzie, B.; Berger, U.V.; Gunshin, Y.; Romero, M.F.; Boron, W.F.; Nussberger, S.; Gollan, J.L.; Hediger, M.A. Cloning and characterization of a mammalian proton-coupled metal-ion transporter. Nature 1997, 388, 482–488. [Google Scholar] [CrossRef] [PubMed]
- Nevo, Y.; Nelson, N. The NRAMP family of metal-ion transporters. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research 2006, 1763, 609–620. [Google Scholar] [CrossRef]
- Sasaki, A.; Yamaji, N.; Yokosho, K.; Ma, J.F. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. The Plant Cell 2012, 24, 2155–2167. [Google Scholar] [CrossRef]
- Tiwari, M.; Sharma, D.; Dwivedi, S.; Singh, M.; Tripathi, R.D.; Trivedi, P.K. Expression in A rabidopsis and cellular localization reveal involvement of rice NRAMP, OsNRAMP 1, in arsenic transport and tolerance. Plant, Cell & Environment 2014, 37, 140–152. [Google Scholar] [CrossRef]
- Xia, J.; Yamaji, N.; Kasai, T.; Ma, J.F. Plasma membrane-localized transporter for aluminum in rice. Proceedings of the National Academy of Sciences 2010, 107, 18381–18385. [Google Scholar] [CrossRef]
- Marik, A.; Naiya, H.; Das, M.; Mukherjee, G.; Basu, S.; Saha, C.; Chowdhury, R.; Bhattacharyya, K.; Seal, A. Split-ubiquitin yeast two-hybrid interaction reveals a novel interaction between a natural resistance associated macrophage protein and a membrane bound thioredoxin in Brassica juncea. Plant molecular biology 2016, 92, 519–537. [Google Scholar] [CrossRef]
- Gabilly, S.T.; Dreyfuss, B.W.; Karamoko, M.; Corvest, V.; Kropat, J.; Page, M.D.; Merchant, S.S.; Hamel, P.P. CCS5, a thioredoxin-like protein involved in the assembly of plastid c-type cytochromes. Journal of Biological Chemistry 2010, 285, 29738–29749. [Google Scholar] [CrossRef]
- Lennartz, K.; Plücken, H.; Seidler, A.; Westhoff, P.; Bechtold, N.; Meierhoff, K. HCF164 encodes a thioredoxin-like protein involved in the biogenesis of the cytochrome b6f complex in Arabidopsis. The Plant Cell 2001, 13, 2539–2551. [Google Scholar] [CrossRef]
- Motohashi, K.; Hisabori, T. HCF164 receives reducing equivalents from stromal thioredoxin across the thylakoid membrane and mediates reduction of target proteins in the thylakoid lumen. The Journal of biological chemistry 2006, 281, 35039–35047. [Google Scholar] [CrossRef] [PubMed]
- Lemeille, S.; Willig, A.; Depège-Fargeix, N.; Delessert, C.; Bassi, R.; Rochaix, J.-D. Analysis of the chloroplast protein kinase Stt7 during state transitions. PLoS Biol 2009, 7, e1000045. [Google Scholar] [CrossRef]
- Pesaresi, P.; Hertle, A.; Pribil, M.; Kleine, T.; Wagner, R.; Strissel, H.; Ihnatowicz, A.; Bonardi, V.; Scharfenberg, M.; Schneider, A. Arabidopsis STN7 kinase provides a link between short-and long-term photosynthetic acclimation. The Plant Cell 2009, 21, 2402–2423. [Google Scholar] [CrossRef]
- Eisenhut, M.; Hoecker, N.; Schmidt, S.B.; Basgaran, R.M.; Flachbart, S.; Jahns, P.; Eser, T.; Geimer, S.; Husted, S.; Weber, A.P.M.; et al. The Plastid Envelope CHLOROPLAST MANGANESE TRANSPORTER1 Is Essential for Manganese Homeostasis in Arabidopsis. Molecular Plant 2018, 11, 955–969. [Google Scholar] [CrossRef] [PubMed]
- Schneider, A.; Steinberger, I.; Herdean, A.; Gandini, C.; Eisenhut, M.; Kurz, S.; Morper, A.; Hoecker, N.; Rühle, T.; Labs, M.; et al. The evolutionarily conserved protein PHOTOSYNTHESIS AFFECTED MUTANT71 is required for efficient manganese uptake at the thylakoid membrane in Arabidopsis. The Plant Cell 2016, 28, tpc.00812.02015. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Zhang, C.; Liu, C.; Jing, Y.; Wang, Y.; Jin, L.; Yang, L.; Fu, A.; Shi, J.; Zhao, F.; et al. Inner Envelope CHLOROPLAST MANGANESE TRANSPORTER 1 Supports Manganese Homeostasis and Phototrophic Growth in Arabidopsis. Molecular Plant 2018, 11, 943–954. [Google Scholar] [CrossRef]
- Duy, D.; Stübe, R.; Wanner, G.; Philippar, K. The Chloroplast Permease PIC1 Regulates Plant Growth and Development by Directing Homeostasis and Transport of Iron. Plant Physiology 2011, 155, 1709–1722. [Google Scholar] [CrossRef]
- Das, M.; Naiya, H.; Marik, A.; Mukherjee, G.; Seal, A. A protocol for functional study of genes in Brassica juncea by Agrobacterium-mediated transient expression: applicability in other Brassicaceae. Journal of Plant Biochemistry and Biotechnology 2020, 29, 368–379. [Google Scholar] [CrossRef]
- Lalonde, S.; Sero, A.; Pratelli, R.; Pilot, G.; Chen, J.; Sardi, M.I.; Parsa, S.A.; Kim, D.Y.; Acharya, B.R.; Stein, E.V.; et al. A membrane protein/signaling protein interaction network for Arabidopsis version AMPv2. Frontiers in physiology 2010, 1, 24. [Google Scholar] [CrossRef]
- Curtis, M.D.; Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant physiology 2003, 133, 462–469. [Google Scholar] [CrossRef]
- Xu, R.; Qingshun, L.Q. Protocol: Streamline cloning of genes into binary vectors in Agrobacterium via the Gateway® TOPO vector system. Plant Methods 2008, 4, 4. [Google Scholar] [CrossRef]
- Noël, L.D.; Cagna, G.; Stuttmann, J.; Wirthmüller, L.; Betsuyaku, S.; Witte, C.-P.; Bhat, R.; Pochon, N.; Colby, T.; Parker, J.E. Interaction between SGT1 and Cytosolic/Nuclear HSC70 Chaperones Regulates Arabidopsis Immune Responses. The Plant Cell 2007, 19, 4061–4076. [Google Scholar] [CrossRef]
- Blume, B.; Grierson, D. Expression of ACC oxidase promoter-GUS fusions in tomato and Nicotiana plumbaginifolia regulated by developmental and environmental stimuli. Plant J 1997, 12, 731–746. [Google Scholar] [CrossRef]
- Das, S.; Sen, M.; Saha, C.; Chakraborty, D.; Das, A.; Banerjee, M.; Seal, A. Isolation and expression analysis of partial sequences of heavy metal transporters from Brassica juncea by coupling high throughput cloning with a molecular fingerprinting technique. Planta 2011, 234, 139–156. [Google Scholar] [CrossRef]
- Thakur, A.; Bachhawat, A.K. Mutations in the N-terminal region of the Schizosaccharomyces pombe glutathione transporter pgt1+ allows functional expression in Saccharomyces cerevisiae. Yeast 2013, 30, 45–54. [Google Scholar] [CrossRef] [PubMed]
- Thaminy, S.; Miller, J.; Stagljar, I. The Split-Ubiquitin Membrane-Based Yeast Two-Hybrid System. Methods in molecular biology (Clifton, N.J.) 2004, 261, 297–312. [Google Scholar] [CrossRef]
- Sparkes, I.A.; Runions, J.; Kearns, A.; Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat Protoc 2006, 1, 2019–2025. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, S.B.; Eisenhut, M.; Schneider, A. Chloroplast Transition Metal Regulation for Efficient Photosynthesis. Trends in Plant Science 2020, 25, 817–828. [Google Scholar] [CrossRef]
- Pan, J.; Song, J.; Sharif, R.; Xu, X.; Li, S.; Chen, X. A mutation in the promoter of the yellow stripe-like transporter gene in cucumber results in a yellow cotyledon phenotype. Journal of Integrative Agriculture 2024, 23, 849–862. [Google Scholar] [CrossRef]
- Blum, H.; Beier, H.; Gross, H.J. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 1987, 8, 93–99. [Google Scholar] [CrossRef]
- Villarejo, A.; Burén, S.; Larsson, S.; Déjardin, A.; Monné, M.; Rudhe, C.; Karlsson, J.; Jansson, S.; Lerouge, P.; Rolland, N.; et al. Evidence for a protein transported through the secretory pathway en route to the higher plant chloroplast. Nature cell biology 2005, 7, 1224–1231. [Google Scholar] [CrossRef]
- Daher, Z.; Recorbet, G.; Valot, B.; Robert, F.; Balliau, T.; Potin, S.; Schoefs, B.; Dumas-Gaudot, E. Proteomic analysis of Medicago truncatula root plastids. Proteomics 2010, 10, 2123–2137. [Google Scholar] [CrossRef]
- Ferro, M.; Brugière, S.; Salvi, D.; Seigneurin-Berny, D.; Court, M.; Moyet, L.; Ramus, C.; Miras, S.; Mellal, M.; Le Gall, S.; et al. AT_CHLORO, a comprehensive chloroplast proteome database with subplastidial localization and curated information on envelope proteins. Mol Cell Proteomics 2010, 9, 1063–1084. [Google Scholar] [CrossRef] [PubMed]
- Bouchnak, I.; Brugière, S.; Moyet, L.; Le Gall, S.; Salvi, D.; Kuntz, M.; Tardif, M.; Rolland, N. Unraveling Hidden Components of the Chloroplast Envelope Proteome: Opportunities and Limits of Better MS Sensitivity*[S]. Molecular & Cellular Proteomics 2019, 18, 1285–1306. [Google Scholar] [CrossRef]
- Duan, Z.; Kong, F.; Zhang, L.; Li, W.; Zhang, J.; Peng, L. A bestrophin-like protein modulates the proton motive force across the thylakoid membrane in Arabidopsis. Journal of integrative plant biology 2016, 58, 848–858. [Google Scholar] [CrossRef] [PubMed]
- Kim, L.J.; Tsuyuki, K.M.; Hu, F.; Park, E.Y.; Zhang, J.; Iraheta, J.G.; Chia, J.C.; Huang, R.; Tucker, A.E.; Clyne, M.; et al. Ferroportin 3 is a dual-targeted mitochondrial/chloroplast iron exporter necessary for iron homeostasis in Arabidopsis. Plant J 2021, 107, 215–236. [Google Scholar] [CrossRef]
- Goodin, M.M.; Zaitlin, D.; Naidu, R.A.; Lommel, S.A. Nicotiana benthamiana: its history and future as a model for plant-pathogen interactions. Mol Plant Microbe Interact 2008, 21, 1015–1026. [Google Scholar] [CrossRef]
- Gandini, C.; Schmidt, S.; Husted, S.; Schneider, A.; Leister, D. The transporter SynPAM71 is located in the plasma membrane and thylakoids, and mediates manganese tolerance in Synechocystis PCC6803. New Phytologist 2017, 215. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



