Submitted:
11 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Genetic and Clinical Landscape of Prom1-Associated Retinal Diseases
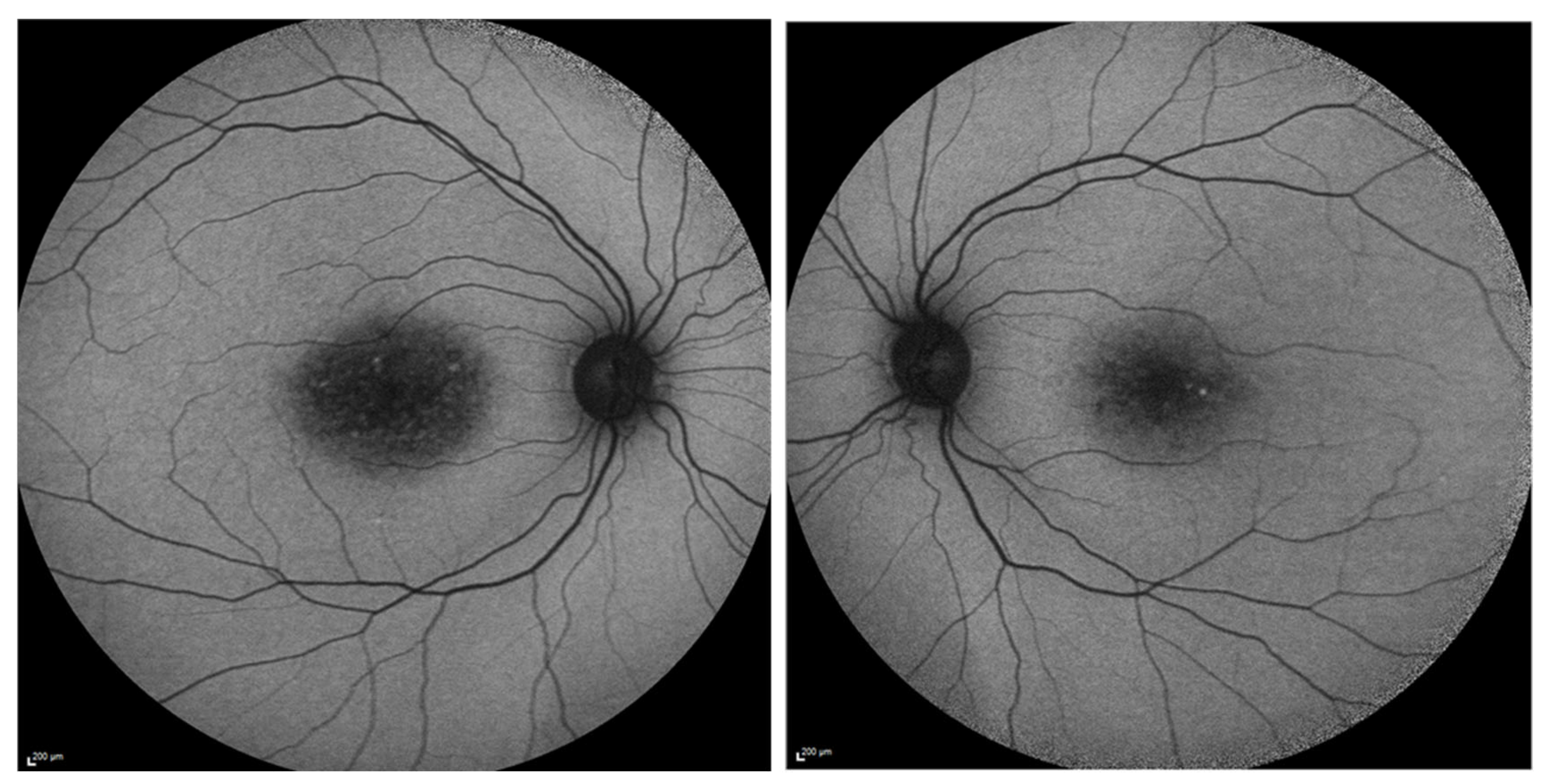
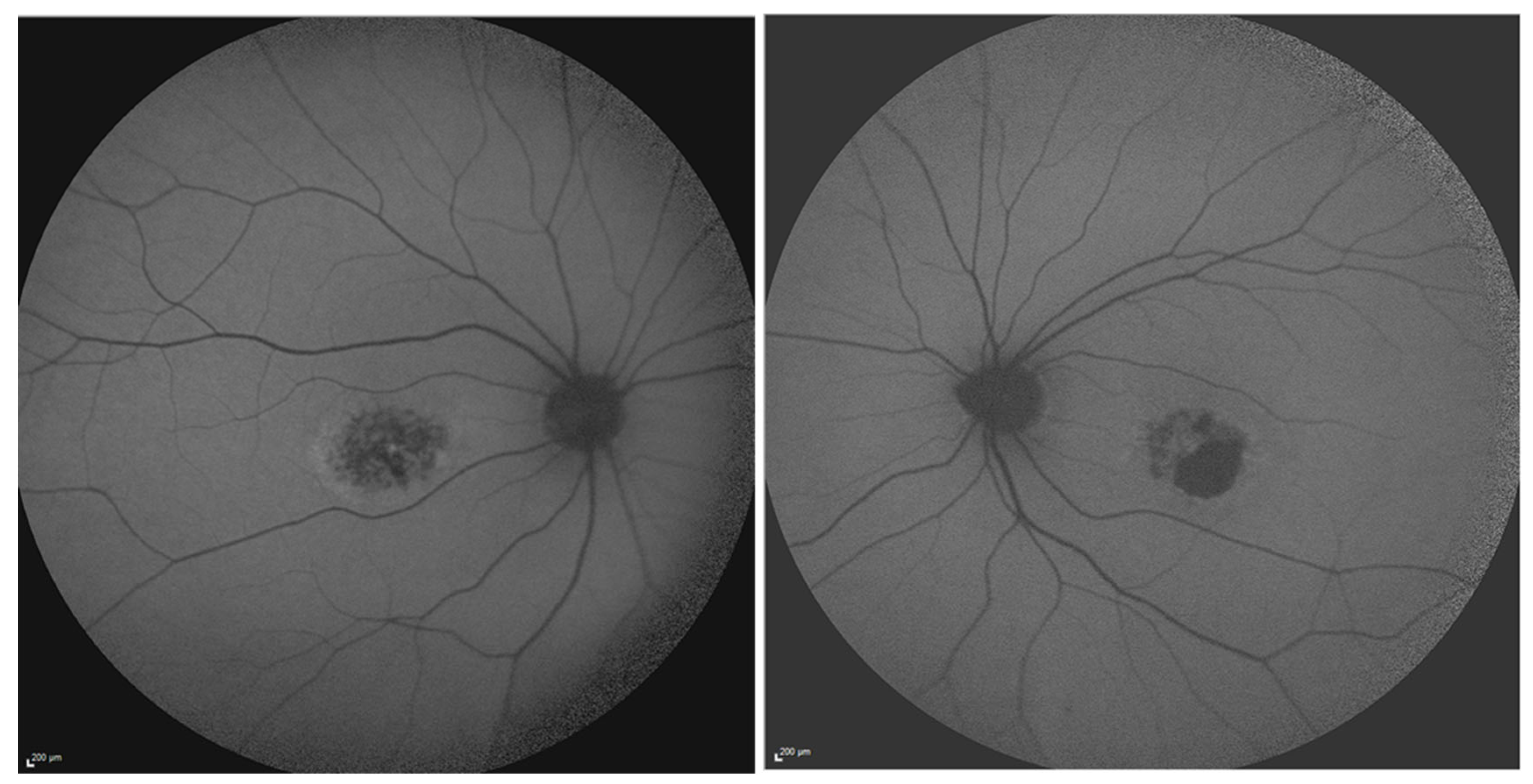
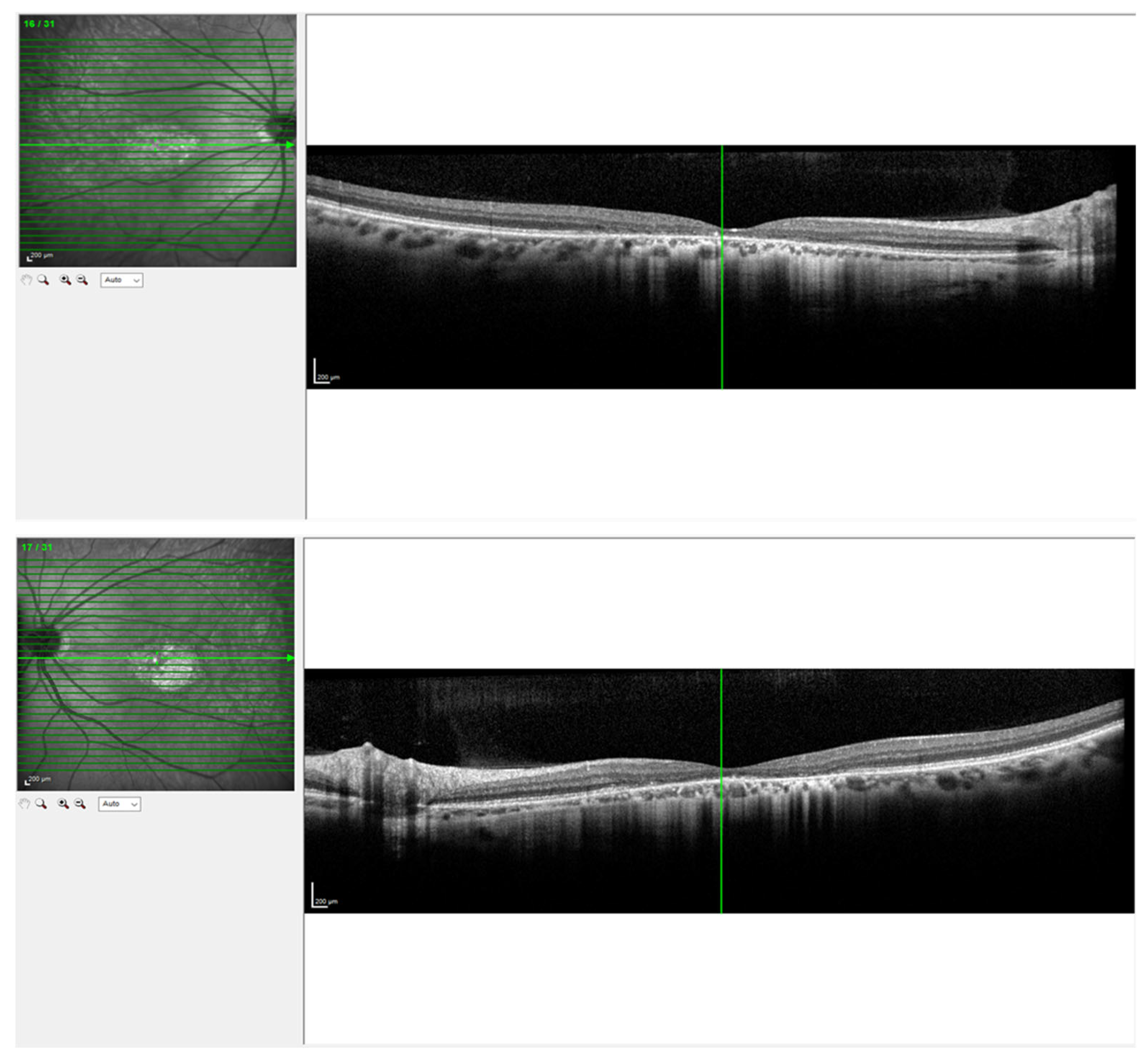
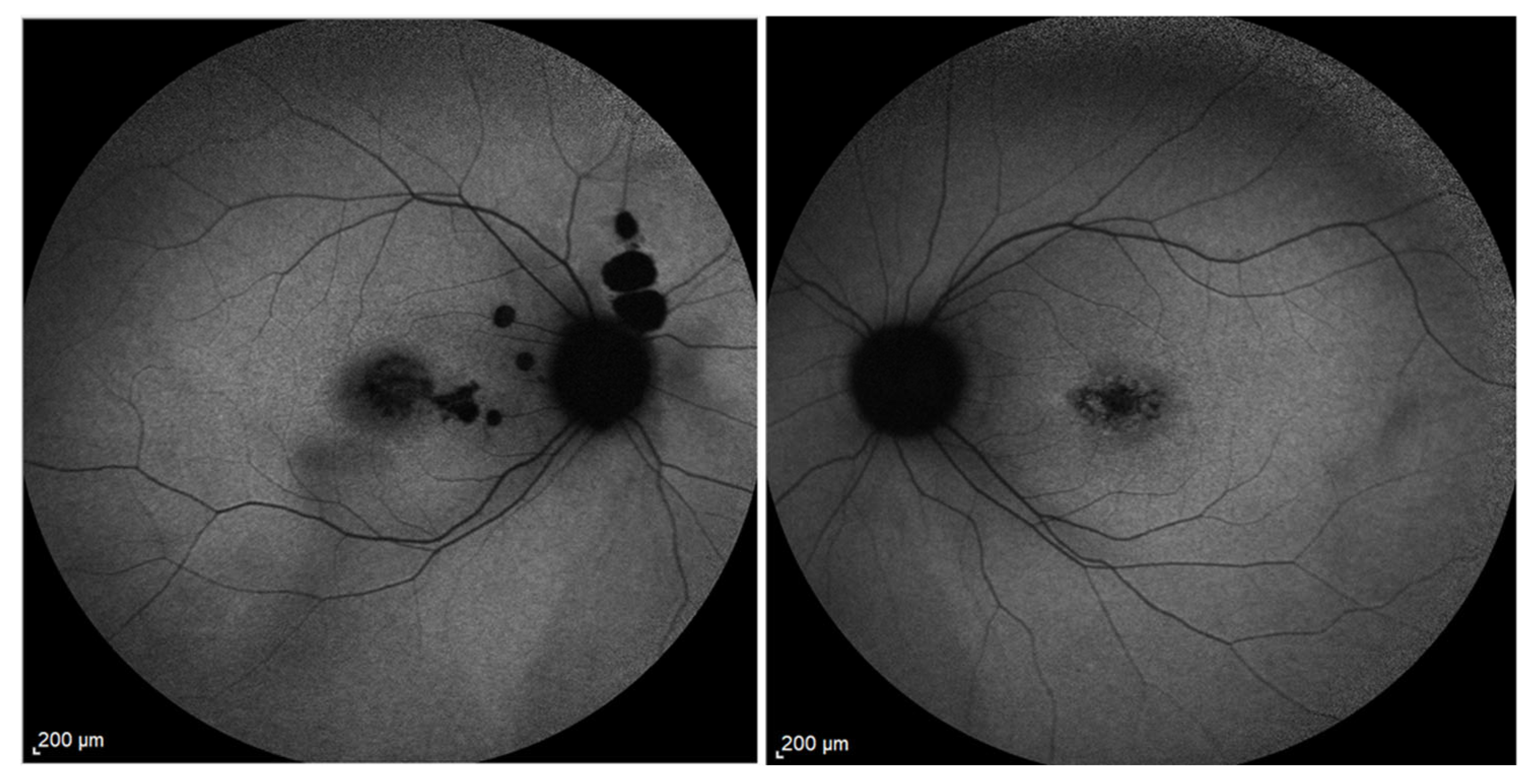
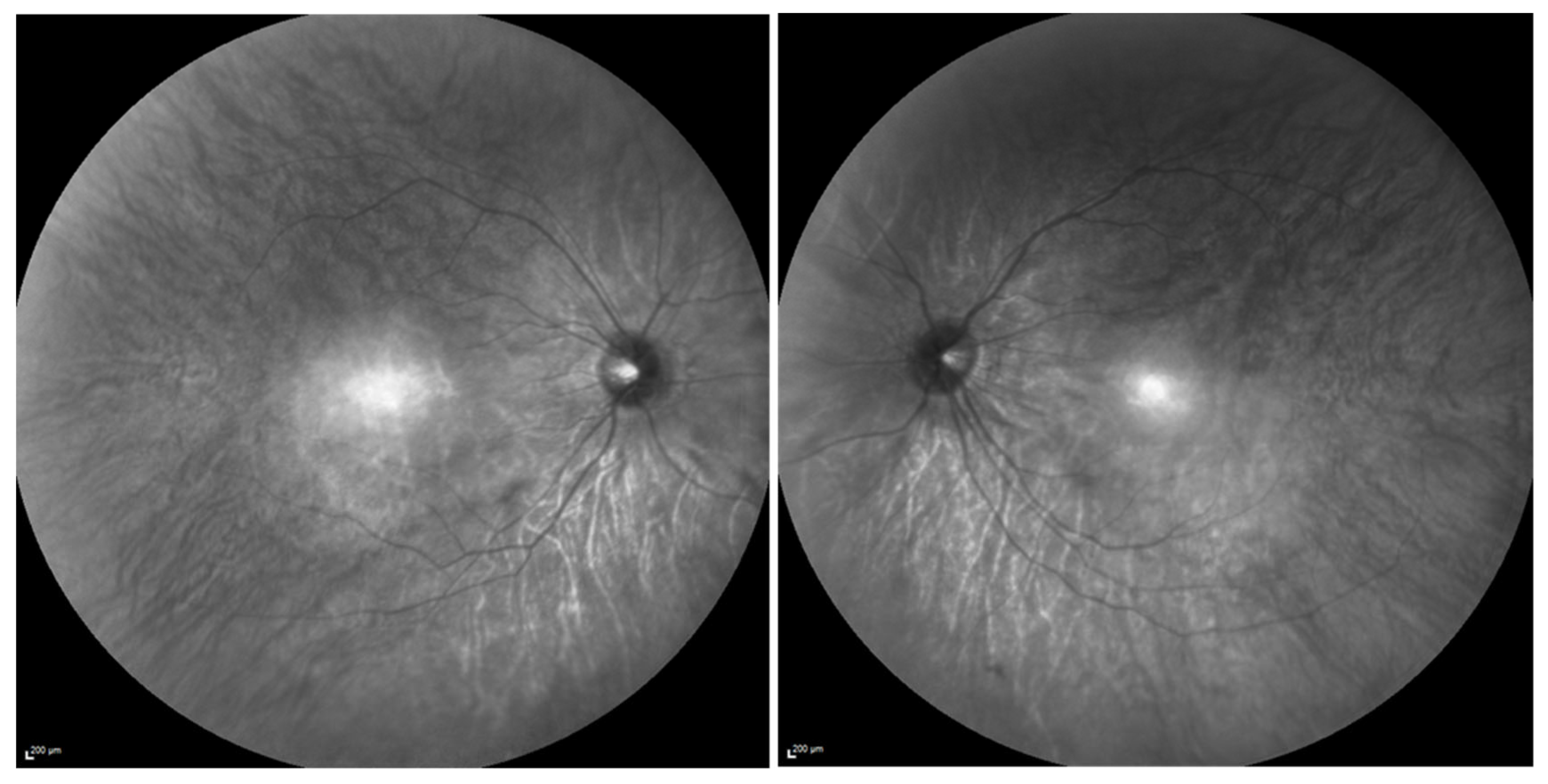
3. Prom1’s Presence and Role in the RPE
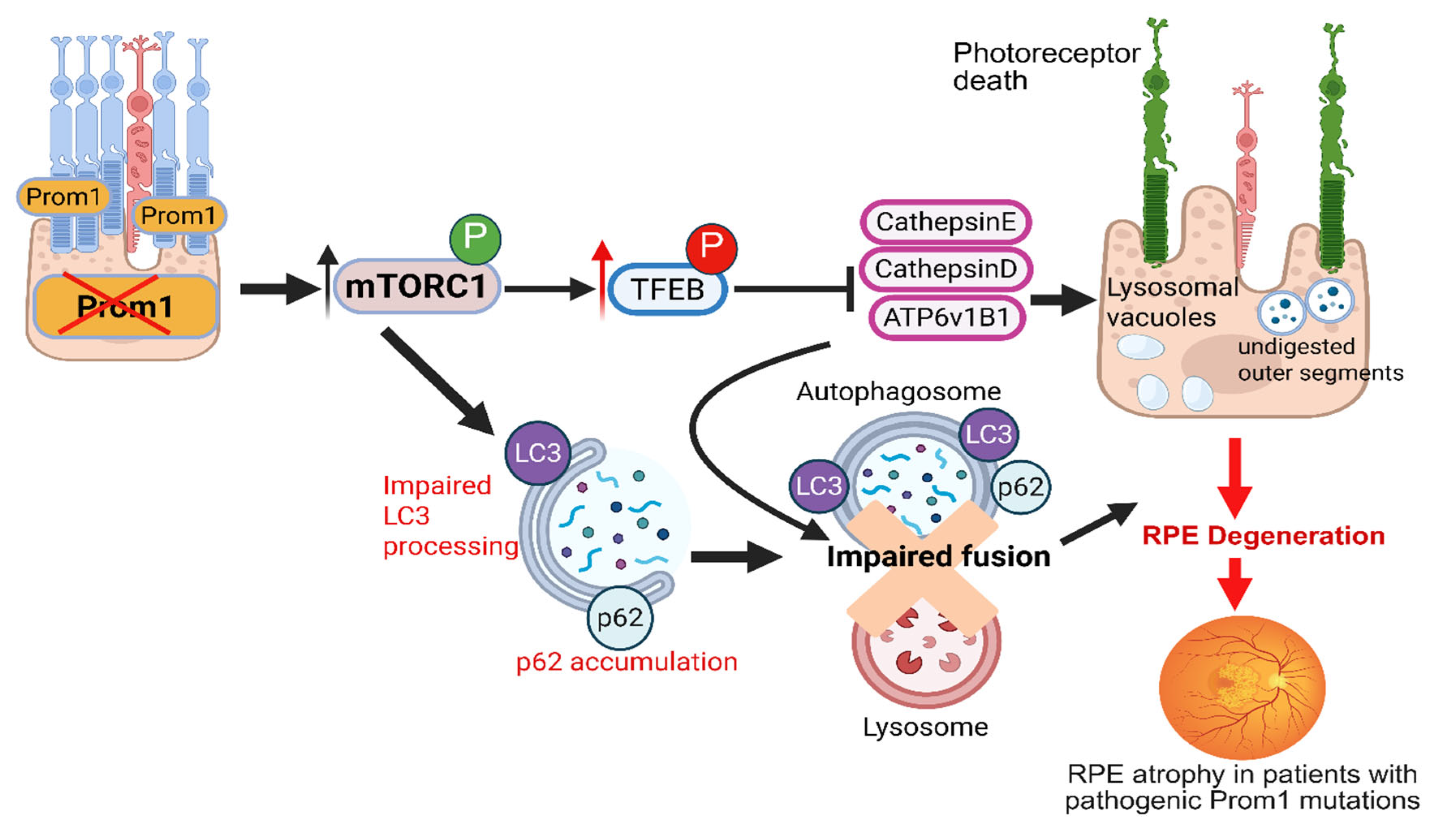
4. Role of Prom1 in RPE EMT
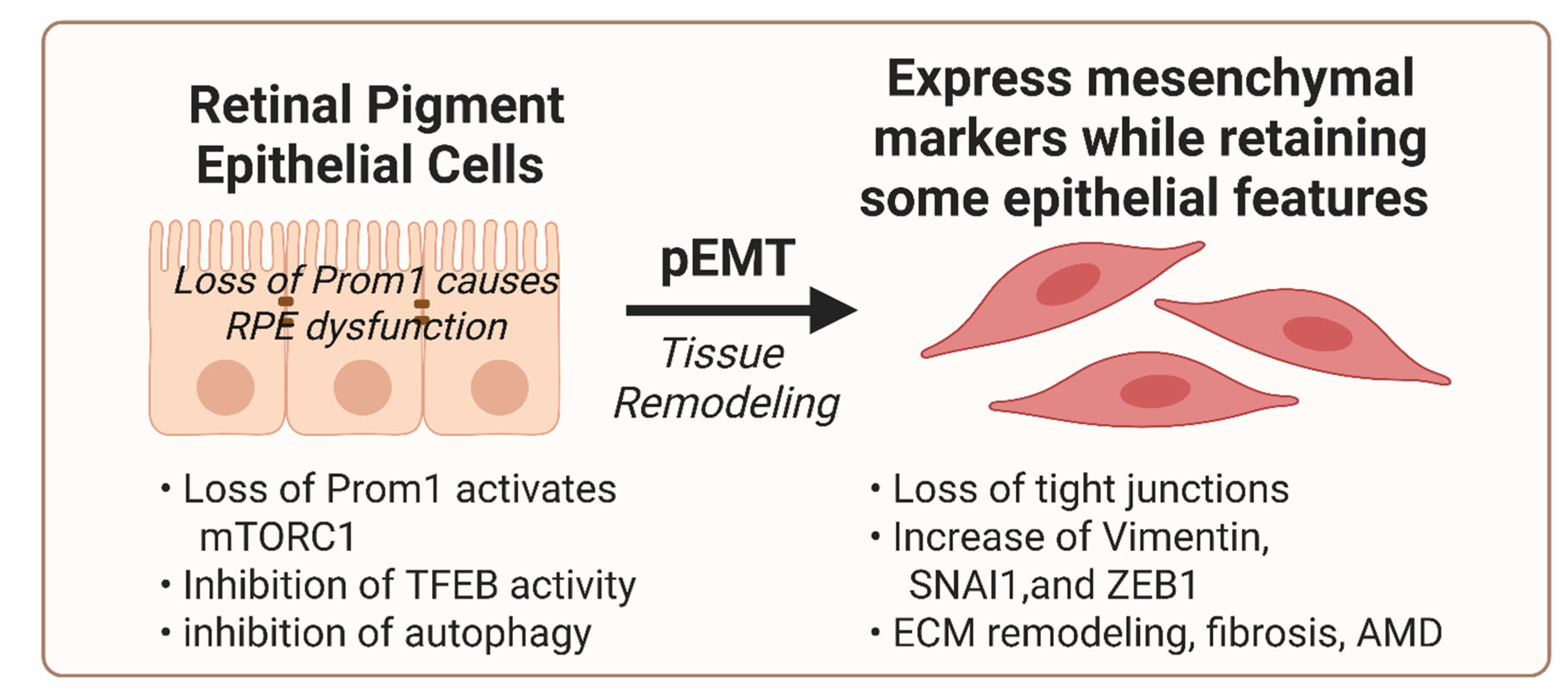
6. Future Directions and Therapeutic Implications
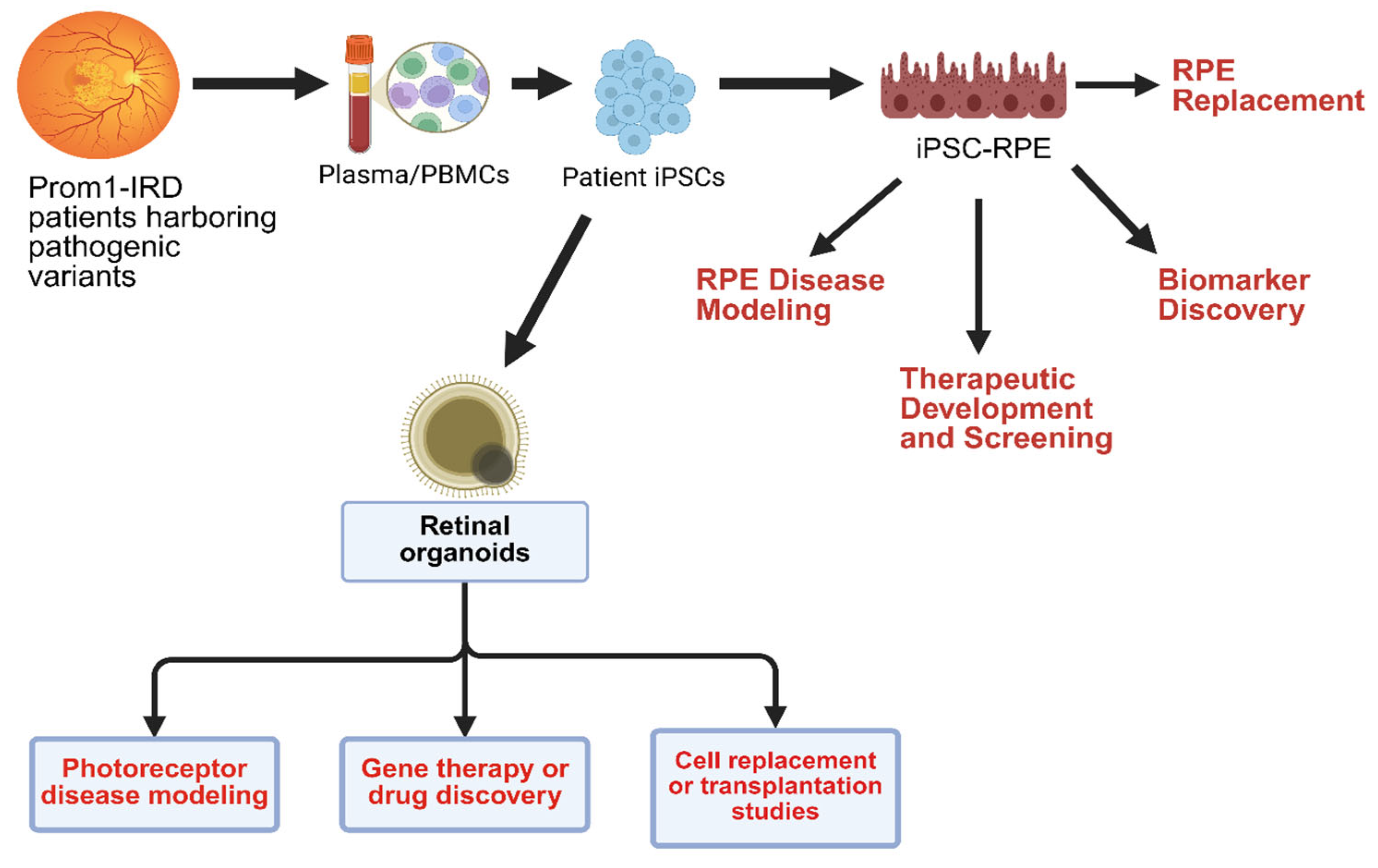
6.1. Gene Therapy
6.2. RPE-Targeted Therapies
6.3. Precision Modeling
6.4. Combined-Compartment Therapies
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| IRDs | Inherited Retinal Dystrophies |
| aAMD | Atrophic Age-related Macular Degeneration |
| RPE | Retinal Pigment Epithelium |
| mTORC1 | Mammalian Target of Rapamycin Complex 1 |
| TFEB | Transcription Factor EB |
| Prom1 | Prominin-1 (CD133) |
| AD | Autosomal Dominant |
| AR | Autosomal Recessive |
| STGD4 | Stargardt disease 4 |
| EMT | Epithelial-Mesenchymal Transition |
| pEMT | partial EMT |
| ECM | Extracellular matrix |
| AAV | Adeno-associated virus |
| POS | Photoreceptor Outer Segment |
| LCA | Leber congenital amaurosis |
| CRD | Cone-rod dystrophies |
| RP | Retinitis Pigmentosa |
| BEST1 | Bestrophin-1 |
| CHM | Choroideremia |
| LOF | Loss-of-function |
| FAF | Fundus autofluorescence |
| SD-OCT | Spectral Domain Optical Coherence Tomography |
| iPSC | induced pluripotent stem cell |
| PBMC | Peripheral Blood Mononuclear Cell |
| MMP | Matrix Metalloproteinases |
| TIMP | Tissue Inhibitors of Metalloproteinases |
| MerTK | Mer proto-oncogene Tyrosine-Kinase |
| FAK | Focal Adhesion Kinase |
| ROS | Reactive Oxygen Species |
| NLRP3 | NOD-like Receptor Protein 3 |
| TGF-beta | Transforming Growth Factor-beta |
References
- Rosa, J.G.S.; Disner, G.R.; Pinto, F.J.; Lima, C.; Lopes-Ferreira, M. Revisiting Retinal Degeneration Hallmarks: Insights from Molecular Markers and Therapy Perspectives. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Collaborators, G.B.D.G.A. Global burden of vision impairment due to age-related macular degeneration, 1990-2021, with forecasts to 2050: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Glob Health 2025, 13, e1175–e1190. [Google Scholar] [CrossRef] [PubMed]
- Manley, A.; Meshkat, B.I.; Jablonski, M.M.; Hollingsworth, T.J. Cellular and Molecular Mechanisms of Pathogenesis Underlying Inherited Retinal Dystrophies. Biomolecules 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Georgiou, M.; Fujinami, K.; Michaelides, M. Inherited retinal diseases: Therapeutics, clinical trials and end points-A review. Clin Exp Ophthalmol 2021, 49, 270–288. [Google Scholar] [CrossRef] [PubMed]
- Guziewicz, K.E.; Sinha, D.; Gomez, N.M.; Zorych, K.; Dutrow, E.V.; Dhingra, A.; Mullins, R.F.; Stone, E.M.; Gamm, D.M.; Boesze-Battaglia, K.; et al. Bestrophinopathy: An RPE-photoreceptor interface disease. Prog Retin Eye Res 2017, 58, 70–88. [Google Scholar] [CrossRef] [PubMed]
- Cehajic Kapetanovic, J.; Barnard, A.R.; MacLaren, R.E. Molecular Therapies for Choroideremia. Genes (Basel) 2019, 10. [Google Scholar] [CrossRef]
- Cideciyan, A.V. Leber congenital amaurosis due to RPE65 mutations and its treatment with gene therapy. Prog Retin Eye Res 2010, 29, 398–427. [Google Scholar] [CrossRef]
- Duncan, J.L.; Pierce, E.A.; Laster, A.M.; Daiger, S.P.; Birch, D.G.; Ash, J.D.; Iannaccone, A.; Flannery, J.G.; Sahel, J.A.; Zack, D.J.; et al. Inherited Retinal Degenerations: Current Landscape and Knowledge Gaps. Transl Vis Sci Technol 2018, 7, 6. [Google Scholar] [CrossRef] [PubMed]
- Datta, S.; Cano, M.; Ebrahimi, K.; Wang, L.; Handa, J.T. The impact of oxidative stress and inflammation on RPE degeneration in non-neovascular AMD. Prog Retin Eye Res 2017, 60, 201–218. [Google Scholar] [CrossRef]
- Lakkaraju, A.; Umapathy, A.; Tan, L.X.; Daniele, L.; Philp, N.J.; Boesze-Battaglia, K.; Williams, D.S. The cell biology of the retinal pigment epithelium. Prog Retin Eye Res 2020, 100846. [Google Scholar] [CrossRef]
- McClements, M.E.; MacLaren, R.E. Gene therapy for retinal disease. Transl Res 2013, 161, 241–254. [Google Scholar] [CrossRef]
- Maw, M.A.; Corbeil, D.; Koch, J.; Hellwig, A.; Wilson-Wheeler, J.C.; Bridges, R.J.; Kumaramanickavel, G.; John, S.; Nancarrow, D.; Roper, K.; et al. A frameshift mutation in prominin (mouse)-like 1 causes human retinal degeneration. Hum Mol Genet 2000, 9, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Corbeil, D.; Roper, K.; Fargeas, C.A.; Joester, A.; Huttner, W.B. Prominin: a story of cholesterol, plasma membrane protrusions and human pathology. Traffic 2001, 2, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Thamm, K.; Simaite, D.; Karbanova, J.; Bermudez, V.; Reichert, D.; Morgenstern, A.; Bornhauser, M.; Huttner, W.B.; Wilsch-Brauninger, M.; Corbeil, D. Prominin-1 (CD133) modulates the architecture and dynamics of microvilli. Traffic 2019, 20, 39–60. [Google Scholar] [CrossRef]
- Moya-Molina, M.; Dorgau, B.; Flood, E.; Letteboer, S.J.F.; Lorentzen, E.; Coxhead, J.; Smith, G.; Roepman, R.; Nagaraja Grellscheid, S.; Armstrong, L.; et al. Deciphering the impact of PROM1 alternative splicing on human photoreceptor development and maturation. Cell Death Dis 2024, 15, 721. [Google Scholar] [CrossRef] [PubMed]
- Del Pozo-Valero, M.; Martin-Merida, I.; Jimenez-Rolando, B.; Arteche, A.; Avila-Fernandez, A.; Blanco-Kelly, F.; Riveiro-Alvarez, R.; Van Cauwenbergh, C.; De Baere, E.; Rivolta, C.; et al. Expanded Phenotypic Spectrum of Retinopathies Associated with Autosomal Recessive and Dominant Mutations in PROM1. Am J Ophthalmol 2019, 207, 204–214. [Google Scholar] [CrossRef]
- Steinberg, R.H.; Fisher, S.K.; Anderson, D.H. Disc morphogenesis in vertebrate photoreceptors. J Comp Neurol 1980, 190, 501–508. [Google Scholar] [CrossRef]
- Zacchigna, S.; Oh, H.; Wilsch-Brauninger, M.; Missol-Kolka, E.; Jaszai, J.; Jansen, S.; Tanimoto, N.; Tonagel, F.; Seeliger, M.; Huttner, W.B.; et al. Loss of the cholesterol-binding protein prominin-1/CD133 causes disk dysmorphogenesis and photoreceptor degeneration. J Neurosci 2009, 29, 2297–2308. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.S.; Liang, J.; Gao, M.; Sun, J.R.; Liu, Y.; Chen, J.Q.; Zhao, X.H.; Wang, Y.M.; Chen, Y.H.; Wang, Y.W.; et al. Deletion of prominin-1 in mice results in disrupted photoreceptor outer segment protein homeostasis. Int J Ophthalmol 2021, 14, 1334–1344. [Google Scholar] [CrossRef]
- Yanardag, S.; Rhodes, S.; Saravanan, T.; Guan, T.; Ramamurthy, V. Prominin 1 is crucial for the early development of photoreceptor outer segments. Sci Rep 2024, 14, 10498. [Google Scholar] [CrossRef]
- Yang, Z.; Chen, Y.; Lillo, C.; Chien, J.; Yu, Z.; Michaelides, M.; Klein, M.; Howes, K.A.; Li, Y.; Kaminoh, Y.; et al. Mutant prominin 1 found in patients with macular degeneration disrupts photoreceptor disk morphogenesis in mice. J Clin Invest 2008, 118, 2908–2916. [Google Scholar] [CrossRef] [PubMed]
- Carr, B.J.; Yang, L.L.; Moritz, O.L. Prominin-1 and Photoreceptor Cadherin Localization in Xenopus laevis: Protein-Protein Relationships and Function. Adv Exp Med Biol 2019, 1185, 483–487. [Google Scholar] [CrossRef]
- Dellett, M.; Sasai, N.; Nishide, K.; Becker, S.; Papadaki, V.; Limb, G.A.; Moore, A.T.; Kondo, T.; Ohnuma, S. Genetic background and light-dependent progression of photoreceptor cell degeneration in Prominin-1 knockout mice. Invest Ophthalmol Vis Sci 2014, 56, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Grosspotzl, M.; Riedl, R.; Schliessleder, G.; Hu, Z.J.; Michaelides, M.; Sadda, S.; Birch, D.; Charbel Issa, P.; Wedrich, A.; Seidel, G.; et al. Progression of PROM1-Associated Retinal Degeneration as Determined by Spectral-Domain Optical Coherence Tomography Over a 24-Month Period. Am J Ophthalmol 2024, 259, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Kong, M.D.; Golebka, J.; Anderson, V.R.; Bao, C.; Bailey, J.A.; Exinor, A.; Demirkol, A.; Tsang, S.H. Divergent Manifestations in Biallelic Versus Monoallelic Variants of RP1-, BEST1-, and PROM1-Associated Retinal Disorders. Int J Mol Sci 2025, 26. [Google Scholar] [CrossRef]
- Hwang, S.; Kang, S.W.; Jang, J.H.; Kim, S.J. Genetic and clinical characteristics of PROM1-related retinal degeneration in Korean. Sci Rep 2023, 13, 21877. [Google Scholar] [CrossRef]
- Imani, S.; Cheng, J.; Shasaltaneh, M.D.; Wei, C.; Yang, L.; Fu, S.; Zou, H.; Khan, M.A.; Zhang, X.; Chen, H.; et al. Genetic identification and molecular modeling characterization reveal a novel PROM1 mutation in Stargardt4-like macular dystrophy. Oncotarget 2018, 9, 122–141. [Google Scholar] [CrossRef]
- Michaelides, M.; Gaillard, M.C.; Escher, P.; Tiab, L.; Bedell, M.; Borruat, F.X.; Barthelmes, D.; Carmona, R.; Zhang, K.; White, E.; et al. The PROM1 mutation p.R373C causes an autosomal dominant bull's eye maculopathy associated with rod, rod-cone, and macular dystrophy. Invest Ophthalmol Vis Sci 2010, 51, 4771–4780. [Google Scholar] [CrossRef]
- Zhang, Q.; Zulfiqar, F.; Xiao, X.; Riazuddin, S.A.; Ahmad, Z.; Caruso, R.; MacDonald, I.; Sieving, P.; Riazuddin, S.; Hejtmancik, J.F. Severe retinitis pigmentosa mapped to 4p15 and associated with a novel mutation in the PROM1 gene. Hum Genet 2007, 122, 293–299. [Google Scholar] [CrossRef]
- Puertas-Neyra, K.; Coco-Martin, R.M.; Hernandez-Rodriguez, L.A.; Gobelli, D.; Garcia-Ferrer, Y.; Palma-Vecino, R.; Telleria, J.J.; Simarro, M.; de la Fuente, M.A.; Fernandez-Bueno, I. Clinical exome analysis and targeted gene repair of the c.1354dupT variant in iPSC lines from patients with PROM1-related retinopathies exhibiting diverse phenotypes. Stem Cell Res Ther 2024, 15, 192. [Google Scholar] [CrossRef] [PubMed]
- Cehajic-Kapetanovic, J.; Birtel, J.; McClements, M.E.; Shanks, M.E.; Clouston, P.; Downes, S.M.; Charbel Issa, P.; MacLaren, R.E. Clinical and Molecular Characterization of PROM1-Related Retinal Degeneration. JAMA Netw Open 2019, 2, e195752. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Yin, J.; Huo, W.; Chaum, E. Loss of Prom1 impairs autophagy and promotes epithelial-mesenchymal transition in mouse retinal pigment epithelial cells. J Cell Physiol 2023. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Yang, T.S.; Nabit, B.P.; Krystofiak, E.S.; Rex, T.S.; Chaum, E. Prominin-1 Knockdown Causes RPE Degeneration in a Mouse Model. Cells 2024, 13. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Yin, J.; Winborn, C.S.; Zhang, Q.; Yue, J.; Chaum, E. Prominin-1 Is a Novel Regulator of Autophagy in the Human Retinal Pigment Epithelium. Invest Ophthalmol Vis Sci 2017, 58, 2366–2387. [Google Scholar] [CrossRef]
- Huo, W.; Yin, J.; Ghose, P.; Schafer, J.C.; Chaum, E.; Bhattacharya, S. Prominin-1 Regulates Retinal Pigment Epithelium Homeostasis: Transcriptomic Insights into Degenerative Mechanisms. Int J Mol Sci 2025, 26. [Google Scholar] [CrossRef]
- D'Esposito, F.; Gagliano, C.; Vallone, S.; Cappellani, F.; Gagliano, G.; Randazzo, V.; Tognetto, D.; Esposito, G.; Zeppieri, M. Clinical and Molecular Findings in PROM1-Associated Inherited Retinal Dystrophies. Genes (Basel) 2025, 16. [Google Scholar] [CrossRef]
- Lee, I.J.; Abbey, C.; Leys, M. Clinical Characterization of Autosomal Dominant and Autosomal Recessive PROM1 Mutation With a Report of Novel Mutation. Ophthalmic Surg Lasers Imaging Retina 2022, 53, 422–428. [Google Scholar] [CrossRef]
- de Castro-Miro, M.; Pomares, E.; Lores-Motta, L.; Tonda, R.; Dopazo, J.; Marfany, G.; Gonzalez-Duarte, R. Combined genetic and high-throughput strategies for molecular diagnosis of inherited retinal dystrophies. PLoS One 2014, 9, e88410. [Google Scholar] [CrossRef] [PubMed]
- Peter, V.G.; Kaminska, K.; Santos, C.; Quinodoz, M.; Cancellieri, F.; Cisarova, K.; Pescini Gobert, R.; Rodrigues, R.; Custodio, S.; Paris, L.P.; et al. The first genetic landscape of inherited retinal dystrophies in Portuguese patients identifies recurrent homozygous mutations as a frequent cause of pathogenesis. PNAS Nexus 2023, 2, pgad043. [Google Scholar] [CrossRef]
- Ricca, A.M.; Han, I.C.; Hoffmann, J.; Stone, E.M.; Sohn, E.H. Macular Atrophy and Phenotypic Variability in Autosomal Dominant Stargardt-Like Macular Dystrophy Due to Prom1 Mutation. Retina 2023, 43, 1165–1173. [Google Scholar] [CrossRef]
- Wu, C.; Zhang, L.; Guo, Q.; Li, Y.; Qiu, R.; Yao, S.; Lei, B. The pathogenicity of a novel frame-shift variant c.2321delC of PROM1 in an autosomal recessive cone-rod dystrophy pedigree may be associated with augment of autophagy. Exp Eye Res 2025, 257, 110453. [Google Scholar] [CrossRef]
- Carss, K.J.; Arno, G.; Erwood, M.; Stephens, J.; Sanchis-Juan, A.; Hull, S.; Megy, K.; Grozeva, D.; Dewhurst, E.; Malka, S.; et al. Comprehensive Rare Variant Analysis via Whole-Genome Sequencing to Determine the Molecular Pathology of Inherited Retinal Disease. Am J Hum Genet 2017, 100, 75–90. [Google Scholar] [CrossRef]
- Kim, J.M.; Lee, C.; Lee, G.I.; Kim, N.K.D.; Ki, C.S.; Park, W.Y.; Kim, B.J.; Kim, S.J. Identification of the PROM1 Mutation p.R373C in a Korean Patient With Autosomal Dominant Stargardt-like Macular Dystrophy. Ann Lab Med 2017, 37, 536–539. [Google Scholar] [CrossRef]
- Eidinger, O.; Leibu, R.; Newman, H.; Rizel, L.; Perlman, I.; Ben-Yosef, T. An intronic deletion in the PROM1 gene leads to autosomal recessive cone-rod dystrophy. Mol Vis 2015, 21, 1295–1306. [Google Scholar]
- Eisenberger, T.; Neuhaus, C.; Khan, A.O.; Decker, C.; Preising, M.N.; Friedburg, C.; Bieg, A.; Gliem, M.; Charbel Issa, P.; Holz, F.G.; et al. Increasing the yield in targeted next-generation sequencing by implicating CNV analysis, non-coding exons and the overall variant load: the example of retinal dystrophies. PLoS One 2013, 8, e78496. [Google Scholar] [CrossRef]
- Lee, W.; Paavo, M.; Zernant, J.; Stong, N.; Laurente, Z.; Bearelly, S.; Nagasaki, T.; Tsang, S.H.; Goldstein, D.B.; Allikmets, R. Modification of the PROM1 disease phenotype by a mutation in ABCA4. Ophthalmic Genet 2019, 40, 369–375. [Google Scholar] [CrossRef]
- Ragi, S.D.; Lima de Carvalho, J.R., Jr.; Tanaka, A.J.; Park, K.S.; Mahajan, V.B.; Maumenee, I.H.; Tsang, S.H. Compound heterozygous novel frameshift variants in the PROM1 gene result in Leber congenital amaurosis. Cold Spring Harb Mol Case Stud 2019, 5. [Google Scholar] [CrossRef]
- Permanyer, J.; Navarro, R.; Friedman, J.; Pomares, E.; Castro-Navarro, J.; Marfany, G.; Swaroop, A.; Gonzalez-Duarte, R. Autosomal recessive retinitis pigmentosa with early macular affectation caused by premature truncation in PROM1. Invest Ophthalmol Vis Sci 2010, 51, 2656–2663. [Google Scholar] [CrossRef] [PubMed]
- Salles, M.V.; Motta, F.L.; Dias da Silva, E.; Varela Lima Teixeira, P.; Antunes Costa, K.; Filippelli-Silva, R.; Martin, R.; Pesquero, J.B.; Ferraz Sallum, J.M. PROM1 gene variations in Brazilian patients with macular dystrophy. Ophthalmic Genet 2017, 38, 39–42. [Google Scholar] [CrossRef]
- Fujinami, K.; Oishi, A.; Yang, L.; Arno, G.; Pontikos, N.; Yoshitake, K.; Fujinami-Yokokawa, Y.; Liu, X.; Hayashi, T.; Katagiri, S.; et al. Clinical and genetic characteristics of 10 Japanese patients with PROM1-associated retinal disorder: A report of the phenotype spectrum and a literature review in the Japanese population. Am J Med Genet C Semin Med Genet 2020, 184, 656–674. [Google Scholar] [CrossRef] [PubMed]
- Boulanger-Scemama, E.; El Shamieh, S.; Demontant, V.; Condroyer, C.; Antonio, A.; Michiels, C.; Boyard, F.; Saraiva, J.P.; Letexier, M.; Souied, E.; et al. Next-generation sequencing applied to a large French cone and cone-rod dystrophy cohort: mutation spectrum and new genotype-phenotype correlation. Orphanet J Rare Dis 2015, 10, 85. [Google Scholar] [CrossRef] [PubMed]
- Weigmann, A.; Corbeil, D.; Hellwig, A.; Huttner, W.B. Prominin, a novel microvilli-specific polytopic membrane protein of the apical surface of epithelial cells, is targeted to plasmalemmal protrusions of non-epithelial cells. Proc Natl Acad Sci U S A 1997, 94, 12425–12430. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Yin, J.; Huo, W.; Chaum, E. Modeling of mitochondrial bioenergetics and autophagy impairment in MELAS-mutant iPSC-derived retinal pigment epithelial cells. Stem Cell Res Ther 2022, 13, 260. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Guo, F.; Wang, Y.; Chen, G.; Song, X.; Luo, B.Y.; Zheng, D.; Gao, G.; Yin, W.; Zhang, S.; et al. Exogenous Photoreceptor-Specific N-Glycosylated PROM1 Rescues Retinal Degeneration in Patient and Mouse Models. Mol Ther 2026. [Google Scholar] [CrossRef] [PubMed]
- Carr, B.J.; Skitsko, D.; Kriese, L.M.; Song, J.; Li, Z.; Ju, M.J.; Moritz, O.L. prominin-1-null Xenopus laevis develop subretinal drusenoid-like deposits, cone-rod dystrophy and RPE atrophy. J Cell Sci 2024, 137. [Google Scholar] [CrossRef] [PubMed]
- Carr, B.J.; Skitsko, D.; Kriese, L.M.; Song, J.; Li, Z.; Ju, M.J.; Moritz, O.L. Prominin-1 null Xenopus laevis develop subretinal drusenoid-like deposits, cone-rod dystrophy, and RPE atrophy. J Cell Sci 2024. [Google Scholar] [CrossRef]
- Zhou, M.; Geathers, J.S.; Grillo, S.L.; Weber, S.R.; Wang, W.; Zhao, Y.; Sundstrom, J.M. Role of Epithelial-Mesenchymal Transition in Retinal Pigment Epithelium Dysfunction. Front Cell Dev Biol 2020, 8, 501. [Google Scholar] [CrossRef]
- Li, J.; Nagasaka, Y.; Shen, H.; Zhou, X.; Ma, J.; Trevisan-Silva, D.; Sherman, N.E.; Ambati, J.; Gelfand, B.D.; Guo, L.W. TMEM97 governs partial epithelial-mesenchymal transition of retinal pigment epithelial cells via the CTNND2-ADAM10 axis. Mol Ther Nucleic Acids 2025, 36, 102460. [Google Scholar] [CrossRef]
- Feng, H.; Zhao, X.; Guo, Q.; Feng, Y.; Ma, M.; Guo, W.; Dong, X.; Deng, C.; Li, C.; Song, X.; et al. Autophagy resists EMT process to maintain retinal pigment epithelium homeostasis. Int J Biol Sci 2019, 15, 507–521. [Google Scholar] [CrossRef]
- Chowdhury, O.; Bammidi, S.; Gautam, P.; Babu, V.S.; Liu, H.; Shang, P.; Xin, Y.; Mahally, E.; Nemani, M.; Koontz, V.; et al. Activated mTOR Signaling in the RPE Drives EMT, Autophagy, and Metabolic Disruption, Resulting in AMD-Like Pathology in Mice. Aging Cell 2025, e70018. [Google Scholar] [CrossRef]
- Wang, N.; Wang, Y.; Zhang, L.; Yang, W.; Fu, S. Molecular Mechanisms of Epithelial-Mesenchymal Transition in Retinal Pigment Epithelial Cells: Implications for Age-Related Macular Degeneration (AMD) Progression. Biomolecules 2025, 15. [Google Scholar] [CrossRef] [PubMed]
- Blasiak, J.; Pawlowska, E.; Helotera, H.; Ionov, M.; Derwich, M.; Kaarniranta, K. Potential of autophagy in subretinal fibrosis in neovascular age-related macular degeneration. Cell Mol Biol Lett 2025, 30, 54. [Google Scholar] [CrossRef] [PubMed]
- Guha, S.; Baltazar, G.C.; Coffey, E.E.; Tu, L.A.; Lim, J.C.; Beckel, J.M.; Patel, S.; Eysteinsson, T.; Lu, W.; O'Brien-Jenkins, A.; et al. Lysosomal alkalinization, lipid oxidation, and reduced phagosome clearance triggered by activation of the P2X7 receptor. FASEB J 2013, 27, 4500–4509. [Google Scholar] [CrossRef]
- Higashijima, F.; Hasegawa, M.; Yoshimoto, T.; Kobayashi, Y.; Wakuta, M.; Kimura, K. Molecular mechanisms of TGFbeta-mediated EMT of retinal pigment epithelium in subretinal fibrosis of age-related macular degeneration. Front Ophthalmol (Lausanne) 2022, 2, 1060087. [Google Scholar] [CrossRef]
- Yang, H.S.; Jo, Y.H.; Kim, D.; Choi, S.; Moon, B.G.; Lee, Y.; Kim, M.; Koo, D.; Kim, J. Yes-Associated Protein (YAP) Is Associated With Disease Severity and Visual Prognosis in Epiretinal Membrane. Invest Ophthalmol Vis Sci 2025, 66, 39. [Google Scholar] [CrossRef]
- Shu, D.Y.; Butcher, E.; Saint-Geniez, M. EMT and EndMT: Emerging Roles in Age-Related Macular Degeneration. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef]
- Kaarniranta, K.; Blasiak, J.; Liton, P.; Boulton, M.; Klionsky, D.J.; Sinha, D. Autophagy in age-related macular degeneration. Autophagy 2023, 19, 388–400. [Google Scholar] [CrossRef] [PubMed]
- Falcao, A.S.; Pedro, M.L.; Tenreiro, S.; Seabra, M.C. Targeting Lysosomal Dysfunction and Oxidative Stress in Age-Related Macular Degeneration. Antioxidants (Basel) 2025, 14. [Google Scholar] [CrossRef] [PubMed]
- Brandstetter, C.; Mohr, L.K.; Latz, E.; Holz, F.G.; Krohne, T.U. Light induces NLRP3 inflammasome activation in retinal pigment epithelial cells via lipofuscin-mediated photooxidative damage. J Mol Med (Berl) 2015, 93, 905–916. [Google Scholar] [CrossRef] [PubMed]
| Prom1 variant (s) | Predicted amino acid change | Genotype | Variant Effect(s) | Phenotype | Reference |
| c.1557C>A | p.Tyr519* | Compound Heterozygous | Nonsense | Retinal dystrophy | ClinVar RCV000504778.6 [42] |
| c.1177_1178delAT | p.Ile393Argfs*21 | Compound Heterozygous | Frameshift | Retinal dystrophy | ClinVar RCV000504956 [31] |
| c.1354_1355insT (c.1354dupT) (LOF) |
p.Tr452Leufs*13 | Heterozygous and Homozygous | Frameshift | AR, CORD, RP, and STGD4 | PMID: 38956727 [30] |
| c.22del | p.Leu8fs* | Heterozygous | Frameshift (LOF) |
Severe retinal dystrophy | PMID: 31199449 [31] |
| c.436C>T | p.Arg146Ter | Heterozygous (dominant) or Homozygous (recessive) | Nonsense | AR, Retinitis Pigmentosa | ClinVar: RCV000987427.1 [29,42] |
| c.199C>T | p.Gln67* | Homozygous | Nonsense | Retinal dystrophy | PMID: 31199449 [31] |
| c.1142-1G>A | Splice acceptor site | Homozygous | Aberrant splicing | STGD4, Retinitis pigmentosa. CORD, macular dystrophy | ClinVar: RCV002497313.2 PMID: 31199449 [31] |
| c.1117C>T | p.Arg373Cys [AD] | Heterozygous | Missense-dominant negative | Macular and peripheral RPE degeneration | PMID: 38072963 PMID: 28840994 [26,43] |
| c.1901C>T c.2020C>T |
p>Gln637* [AD/AR] p.Arg684* [AD/AR] |
Heterozygous Homozygous |
Nonsense- Truncating-LOF Nonsense- Truncating-LOF |
AD- Bull’s eye maculopathy AR-panretinal dystrophy |
PMID: 35951719 [37] |
| c.642T>A | p.Tyr214* [AD] | Heterozygous | Nonsense- Premature stop-LOF | Retinal dystrophy Best retinal disease |
PMID: 26702251, 24265693. [44,45] Lee_2021 (abstract) |
| c.2110C>T | p.Arg704Cys [AR] | Heterozygous | Missense | Retinal dystrophy | ClinVar: RCV001058099.8 |
| c.303+2T>C | Splice donor | Heterozygous | Splicing abnormality, exon 4 skip- null function | Macular dystrophy; early-onset rod-cone dystrophy |
PMID: 40724865 [25] |
| c.718G>A | p.Gly240Arg | Heterozygous | Missense | AD Bull’s eye macular dystrophy | ClinVar: RCV00036867.2, RCV000262406.5 |
| c.400C>T | p.Arg134Cys | Heterozygous | Missense | AD Stargardt-like macular dystrophy | PMID: 31576780 [46] |
| c.1877_1878del | p.Ile626fs | Heterozygous | Frameshift deletion- loss of protein | Leber’s congenital amaurosis/ macular atrophy | PMID: 31836589 [47] |
| c.139del | p.His47fs | Heterozygous | Frameshift deletion- loss of protein | Leber’s Congenital Amaurosis/ Macular Atrophy | PMID: 31836589 [47] |
| c.734T>C | p.L245P | Heterozygous | Missense | Stargardt4-like macular Dystrophy | PMID: 29416601 [27] |
| c.1726C>T | p.Q576X | Homozygous | Missense | AR RP with macular degeneration RPE atrophy | PMID: 17605048 [29,44] |
| c.1841delG | p.G614Efs12X | Homozygous | Frameshift- truncated non-functional protein | AR RP with macular degeneration | PMID: 10587575 [12,44] |
| c.869delG | p.S290IfsX | Homozygous | Frameshift – truncated protein-LOF | AR RP with macular degeneration | PMID: 20042663 [48] |
| c.2321delC | p.A774Vfs*2 | Heterozygous | Frameshift | AR cone-rod dystrophy | PMID: 40414337 [41] |
| c.2485G>A | p.D829N | Heterozygous | Missense | AD Macular Dystrophy | PMID: 28095140 [49] |
| c.334T>C | p.C112R | Heterozygous | Missense | AD Macular Dystrophy | PMID: 32820593 [50] |
| c.2327A>T | p.D776V | Homozygous | Missense | AR Macular Dystrophy | PMID: 28095140 [49] |
| c.7dup | p.L3Pfs28* | Compound heterozygous | Frameshift | AD Stargardt-like macular dystrophy | PMID: 26103963 [51] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




