Submitted:
11 March 2026
Posted:
11 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Ethical Statement
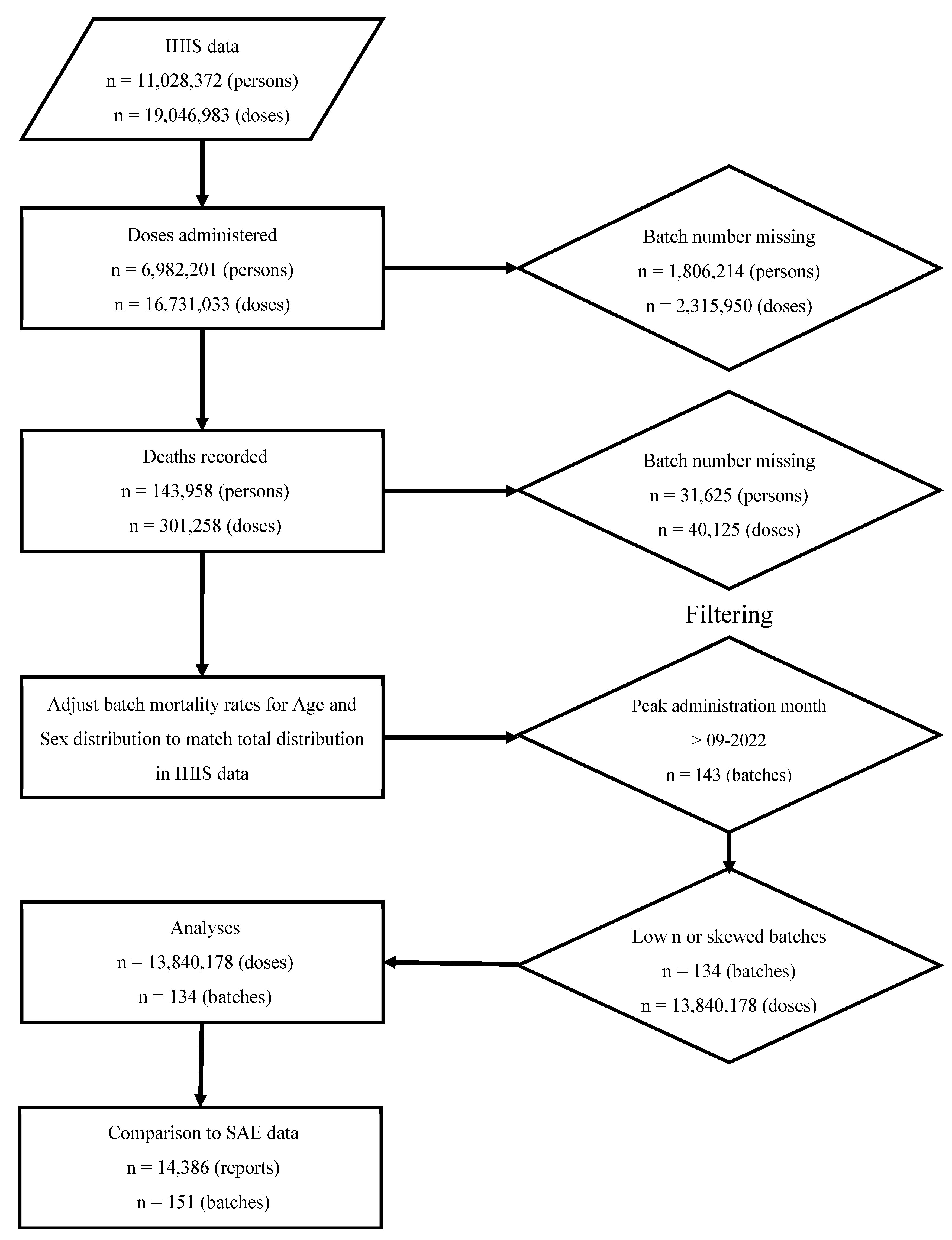
Data Set
Data Selection
Data Curation
Pfizer-BioNTech
Moderna
Astra Zeneca
Jansen
Data Analysis
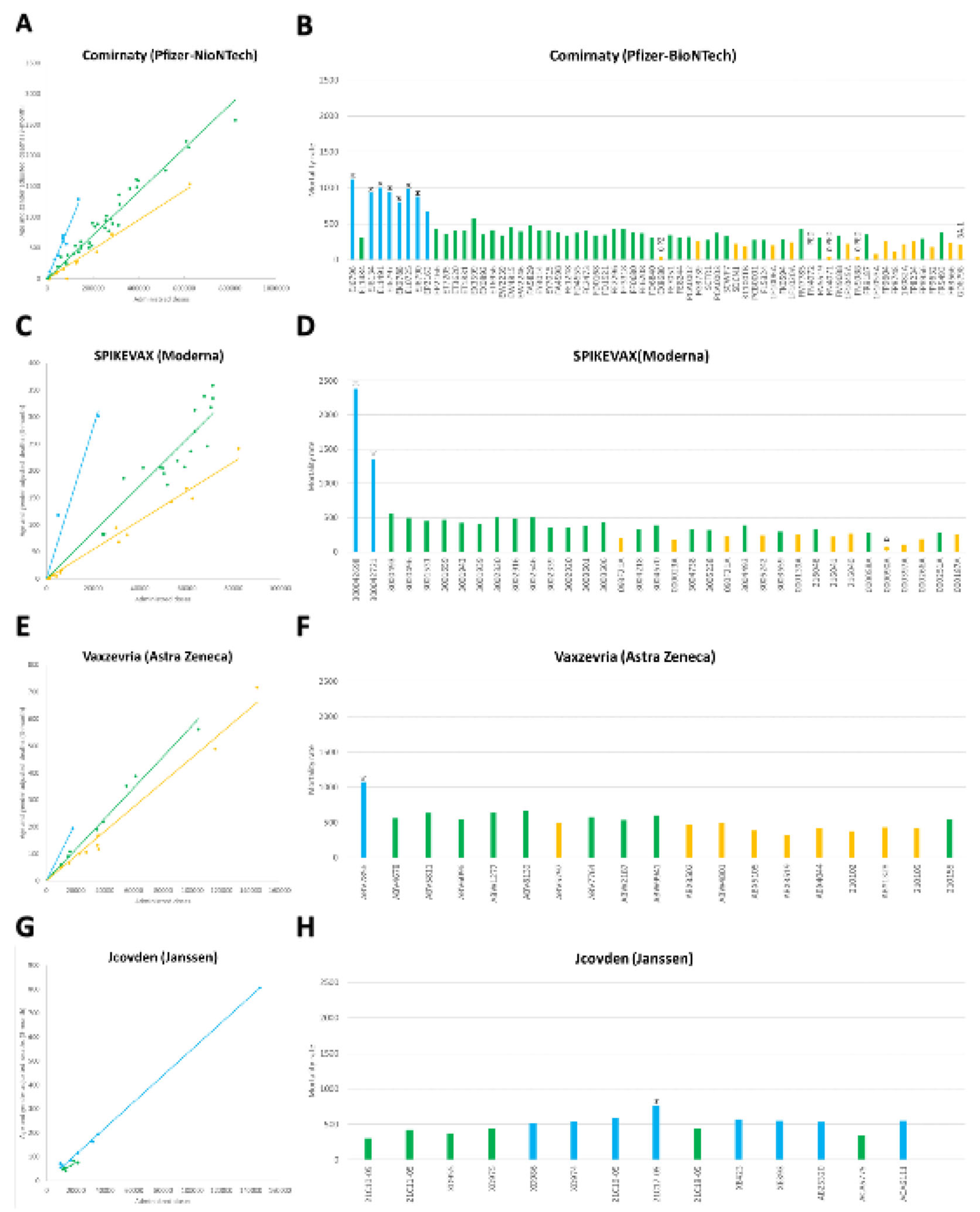
3. Results
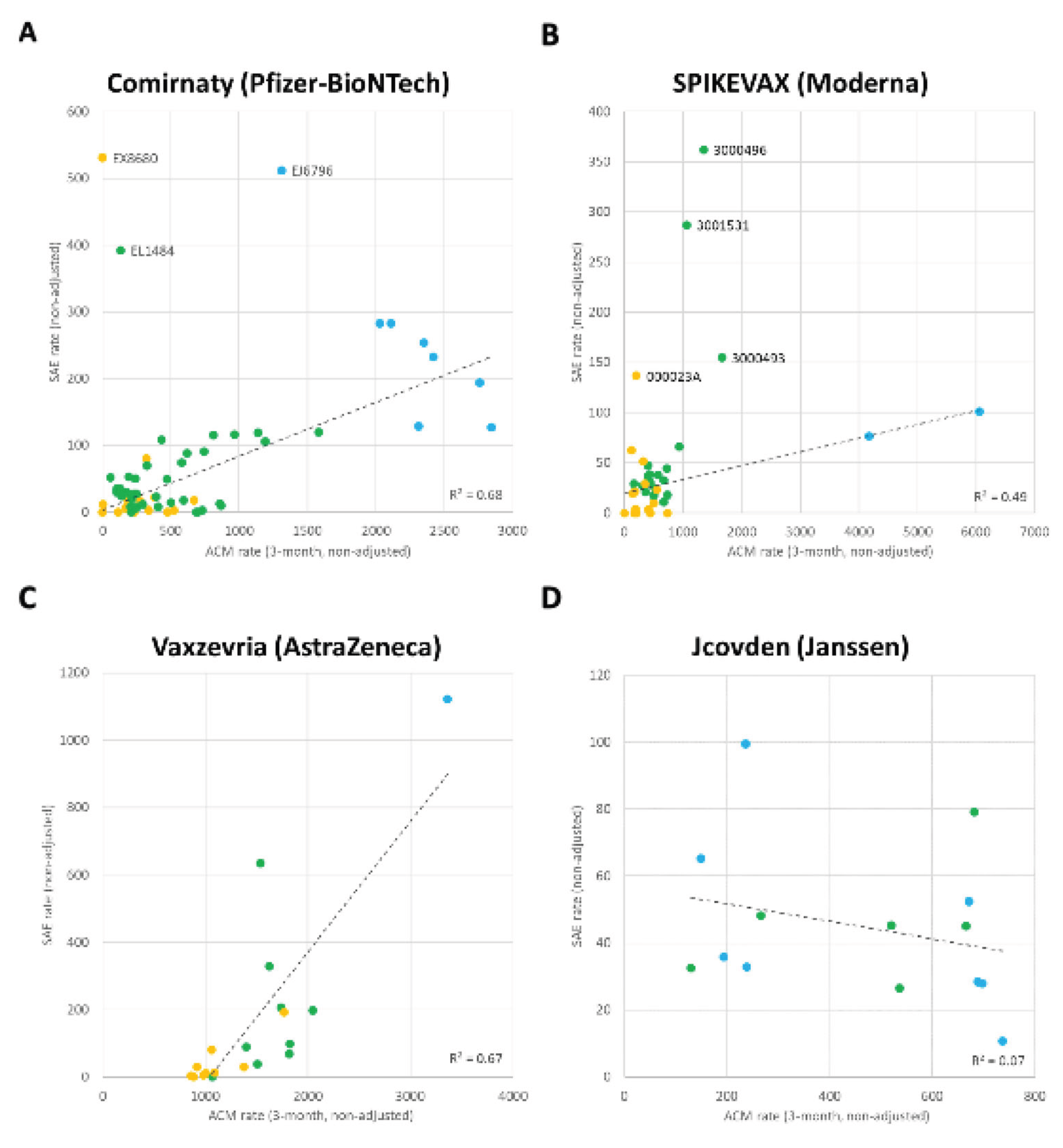
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wrapp, D; Wang, N; Corbett, KS; Goldsmith, JA; Hsieh, CL; Abiona, O; et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367(6483), 1260–3. [Google Scholar] [CrossRef]
- Semenzato, L; Le Vu, S; Botton, J; Bertrand, M; Jabagi, MJ; Drouin, J; et al. COVID-19 mRNA Vaccination and 4-Year All-Cause Mortality Among Adults Aged 18 to 59 Years in France. JAMA Netw Open. 2025, 8(12), e2546822. [Google Scholar] [CrossRef] [PubMed]
- Guo, W; Deguise, J; Tian, Y; Huang, PC; Goru, R; Yang, Q; et al. Profiling COVID-19 Vaccine Adverse Events by Statistical and Ontological Analysis of VAERS Case Reports. Front Pharmacol. 2022, 13, 870599. [Google Scholar] [CrossRef] [PubMed]
- Szebeni, J. Expanded Spectrum and Increased Incidence of Adverse Events Linked to COVID-19 Genetic Vaccines: New Concepts on Prophylactic Immuno-Gene Therapy, Iatrogenic Orphan Disease, and Platform-Inherent Challenges. Pharmaceutics 2025, 17(4). [Google Scholar] [CrossRef] [PubMed]
- Bartmann, C; Schmidt, V; Morz, M; Schwab, M; Rehn, M; Blau-Schneider, B; et al. Detection of spike protein in term placentas of COVID-19 vaccinated and/or SARS-CoV-2 infected women. PLoS One 2026, 21(3), e0344185. [Google Scholar] [CrossRef]
- Kuperwasser, C; El-Deiry, WS. COVID vaccination and post-infection cancer signals: Evaluating patterns and potential biological mechanisms. Oncotarget 2026, 17, 1–29. [Google Scholar] [CrossRef]
- Aarstad, J. Deaths among young people in England increased significantly in 10 of 11 weeks after COVID-19 vaccination and doubled in three. EXCLI J 2024, 23, 908–11. [Google Scholar]
- Okoro, EO; Ikoba, NA; Okoro, BE; Akpila, AS; Salihu, MO. Paradoxical increase in global COVID-19 deaths with vaccination coverage: World Health Organization estimates (2020-2023). Int J Risk Saf Med. 2025, 36(3), 103–12. [Google Scholar] [CrossRef]
- Imbrici, P; De Bellis, M; Liantonio, A; De Luca, A. Investigating the Benefit-Risk Profile of Drugs: From Spontaneous Reporting Systems to Real-World Data for Pharmacovigilance. Methods Mol Biol. 2025, 2834, 333–49. [Google Scholar]
- European Medicines Agency. EudraVigilance—European database of suspected adverse drug reaction reports. 2026 10/03/2026. Available online: https://www.adrreports.eu/en/.
- Postigo, R; Brosch, S; Slattery, J; van Haren, A; Dogne, JM; Kurz, X; et al. EudraVigilance Medicines Safety Database: Publicly Accessible Data for Research and Public Health Protection. Drug Saf. 2018, 41(7), 665–75. [Google Scholar] [CrossRef]
- Dubrall, D; Christ, P; Domgorgen, S; Schmid, M; Sachs, B. Factors associated with the completeness of information provided in adverse drug reaction reports of physicians, pharmacists and consumers from Germany. Sci Rep. 2025, 15(1), 23751. [Google Scholar] [CrossRef]
- Leitzen, S; Dubrall, D; Toni, I; Stingl, J; Christ, P; Koberle, U; et al. Adverse Drug Reactions in Children: Comparison of Reports Collected in a Pharmacovigilance Project Versus Spontaneously Collected ADR Reports. Paediatr Drugs 2023, 25(2), 203–15. [Google Scholar] [CrossRef]
- Dijkstra, L; Schink, T; Linder, R; Schwaninger, M; Pigeot, I; Wright, MN; et al. A discovery and verification approach to pharmacovigilance using electronic healthcare data. Front Pharmacol. 2024, 15, 1426323. [Google Scholar] [CrossRef]
- De Mattia, F; Chapsal, JM; Descamps, J; Halder, M; Jarrett, N; Kross, I; et al. The consistency approach for quality control of vaccines—a strategy to improve quality control and implement 3Rs. Biologicals 2011, 39(1), 59–65. [Google Scholar] [CrossRef] [PubMed]
- Bruce Yu, Y; Taraban, MB; Briggs, KT. All vials are not the same: Potential role of vaccine quality in vaccine adverse reactions. Vaccine 2021, 39(45), 6565–9. [Google Scholar] [CrossRef]
- Manniche, V; Schmeling, M; Gilthorpe, JD; Hansen, PR. Reports of Batch-Dependent Suspected Adverse Events of the BNT162b2 mRNA COVID-19 Vaccine: Comparison of Results from Denmark and Sweden. Medicina (Kaunas) 2024, 60(8). [Google Scholar] [CrossRef]
- Schmeling, M; Manniche, V; Hansen, PR. Batch-dependent safety of the BNT162b2 mRNA COVID-19 vaccine. Eur J Clin Invest. 2023, 53(12), e14102. [Google Scholar] [CrossRef] [PubMed]
- Jablonowski, K HB. Batch-dependent Safety of the BNT162b2 mRNA COVID-19 Vaccine in the United States. Science, Public Health Policy and the Law 2024, 2024;v5.2019-2024 v5.2019-2024. [Google Scholar]
- Furst, T; Sourek, P; Kratka, Z; Janosek, J. Batch-dependent safety of COVID-19 vaccines in the Czech Republic and comparison with data from Denmark. Eur J Clin Invest. 2024, 54(10), e14271. [Google Scholar] [CrossRef] [PubMed]
- IBM Corp. IBM SPSS Statistics for Windows. 2020, Version 26.0. [Google Scholar]
- Choi, MJ; Na, Y; Hyun, HJ; Nham, E; Yoon, JG; Seong, H; et al. Comparative safety analysis of mRNA and adenoviral vector COVID-19 vaccines: a nationwide cohort study using an emulated target trial approach. Clin Microbiol Infect. 2024, 30(5), 646–52. [Google Scholar] [CrossRef]
- Schmeling, M; Manniche, V; Hansen, PR. Safety of BNT162b2 mRNA COVID-19 Vaccine Batches: A Nationwide Cohort Study. Pharmacoepidemiol Drug Saf. 2025, 34(11), e70270. [Google Scholar] [CrossRef] [PubMed]
- Hviid, A; Svalgaard, IB. Safety of BNT162b2 mRNA COVID-19 Vaccine Batches: A Nationwide Cohort Study. Pharmacoepidemiol Drug Saf. 2025, 34(9), e70207. [Google Scholar] [CrossRef] [PubMed]
- Tinari, S. The EMA covid-19 data leak, and what it tells us about mRNA instability. BMJ. 2021, 372, n627. [Google Scholar] [CrossRef] [PubMed]
- European Medicines Agency. European public assessment report (EPAR) 2020. Available online: https://www.ema.europa.eu/en/documents/assessment-report/comirnaty-epar-public-assessment-report_en.pdf.
- Azzarone, B; Landolina, N; Mariotti, FR; Moretta, L; Maggi, E. Soluble SARS-CoV-2 Spike glycoprotein: considering some potential pathogenic effects. Front Immunol. 2025, 16, 1616106. [Google Scholar] [CrossRef]
- Brogna, C; Cristoni, S; Marino, G; Montano, L; Viduto, V; Fabrowski, M; et al. Detection of recombinant Spike protein in the blood of individuals vaccinated against SARS-CoV-2: Possible molecular mechanisms. Proteomics Clin Appl. 2023, 17(6), e2300048. [Google Scholar] [CrossRef]
- Speicher, DJ; Rose, J; McKernan, K. Quantification of residual plasmid DNA and SV40 promoter-enhancer sequences in Pfizer/BioNTech and Moderna modRNA COVID-19 vaccines from Ontario, Canada. Autoimmunity 2025, 58(1), 2551517. [Google Scholar] [CrossRef]
- US Food; Drug Administration. FDA Briefing Document—Vaccines and Related Biological Products Advisory Committee Meeting—October 26, 2021 2021. Available online: https://www.fda.gov/media/153447/download.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



