Submitted:
11 March 2026
Posted:
11 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Method
2.1. Generation of Transgenic Rice Plants Overexpressing PEPCK Gene
2.2. Tolerance Index Assessment of T1 Plants
2.3. Measurement of Salinity Tolerance by Leaf Disc Senescence Assay
2.4. Assessments of Antioxidants in PEPCK Transgenic Lines
2.4.1. Proline Estimation
2.4.2. Determination of H2O2 Content
2.4.3. Lipid Peroxidation Measurement (MDA Content)
2.4.4. Electrolytic Leakage (Membrane Permeability)
2.4.5. Relative Water Content
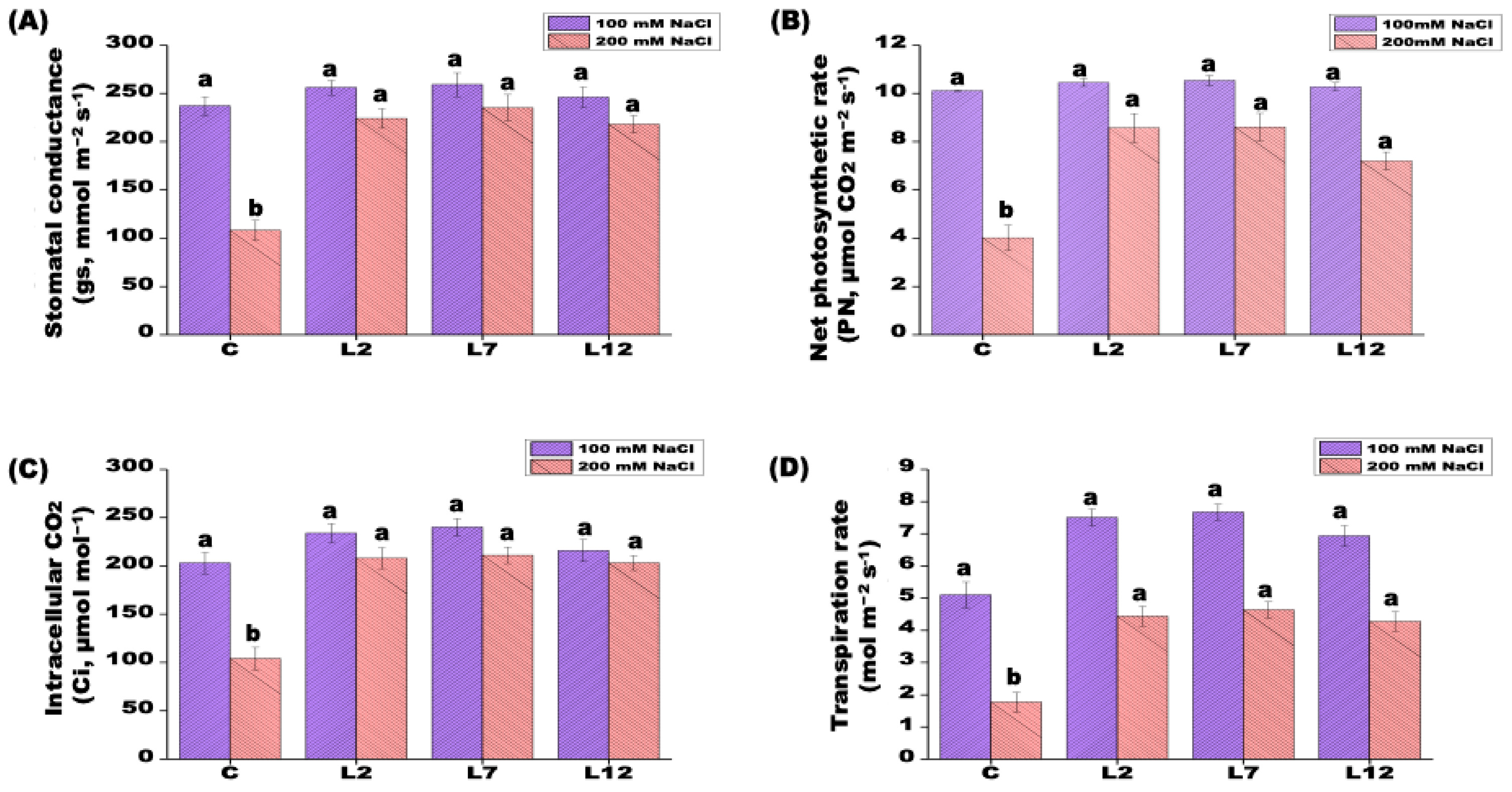
2.5. Quantification of Photosynthetic Parameters
2.6. The Agronomic Attributes of T1 Transgenic Plant
2.7. Determining the Endogenous Ion Content and Soluble Sugar and Hormones
2.8. Statistical Analysis
3. Results
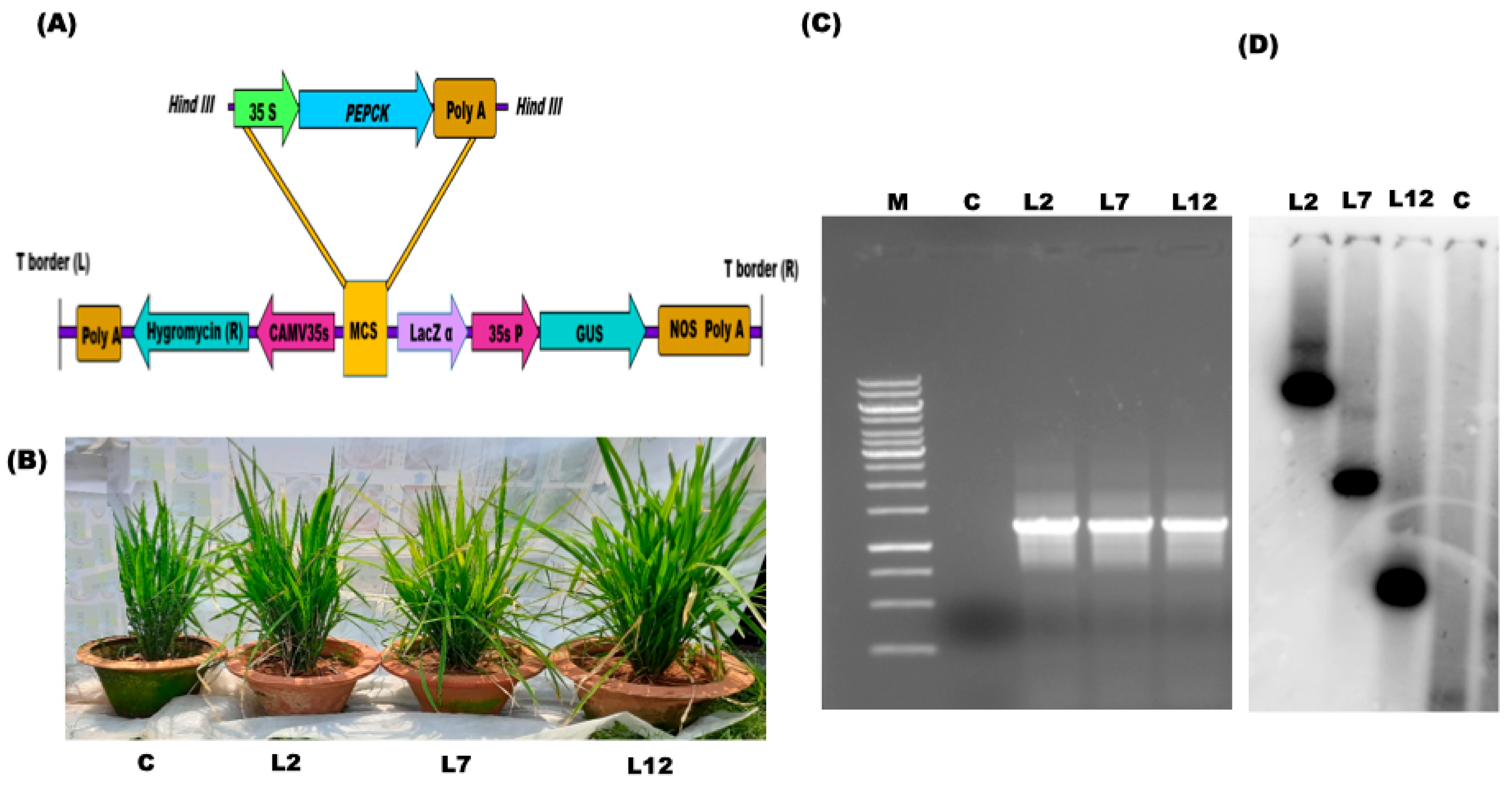
3.1. Molecular Assessment of the Transgenic PEPCK Plants
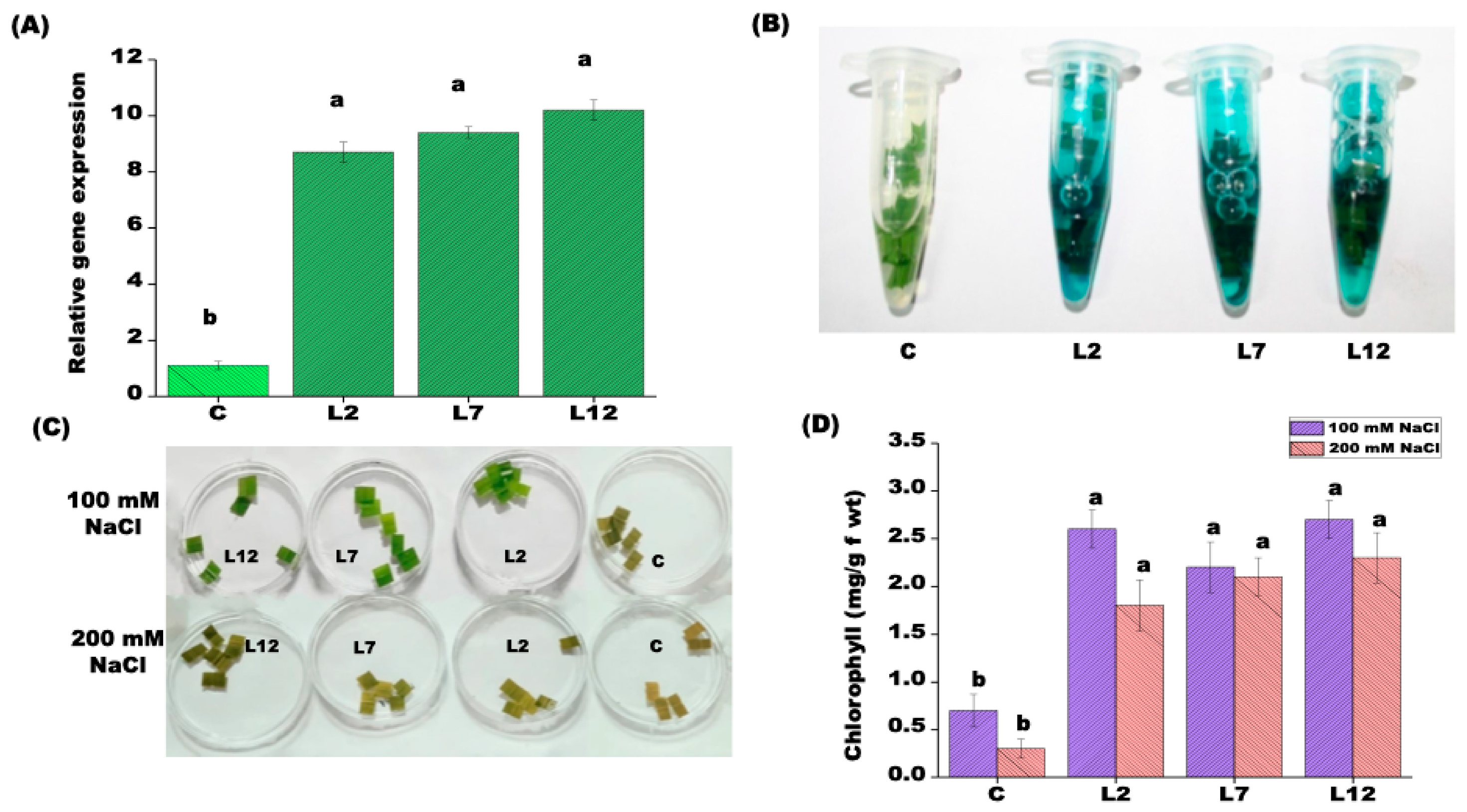
3.2. PEPCK Overexpression Confers Salt Tolerance
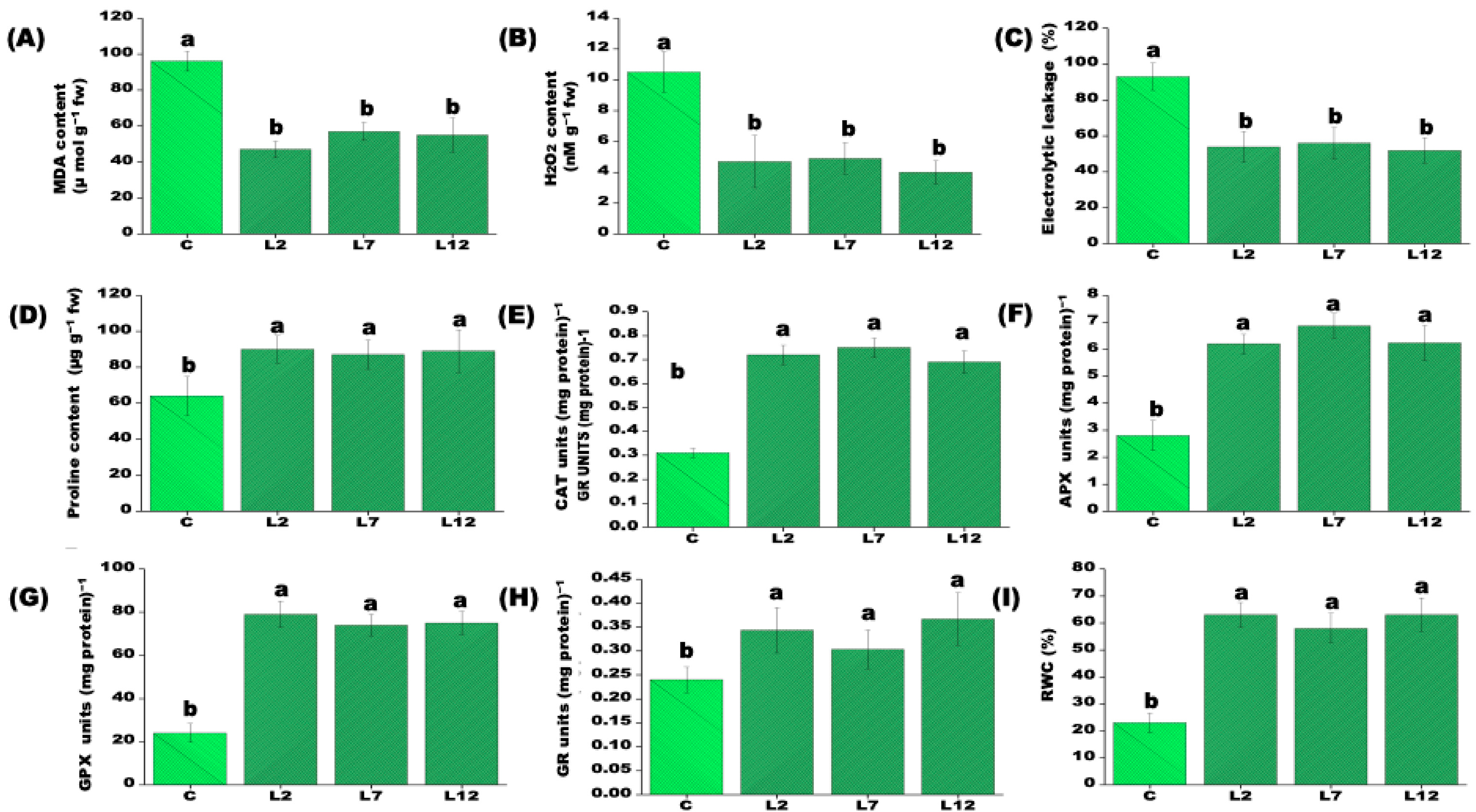
3.3. Biochemical and Physiological Responses of PEPCK Transgenic Plants Under Salt Stress
3.4. PEPCK Alleviates Agronomic Features in Rice Under Salt Stress
| WT (control) | Line 2 | Line 7 | Line 12 | |
| Segregation ratio (HygR:HygS [n]a | 0 | 2.62:1[178] | 2.72:1 [156] | 3.1:1[184] |
| Attributes | Control plants | NaCl (mM)-grown T1 PEPCK transgenic plants | ||||||
| L2 | L7 | L12 | ||||||
| 0 | 200 | 0 | 200 | 0 | 200 | 0 | 200 | |
| Plant height (cm) | 72±3.0 | 31±2.0 | 76± 2.6 | 68±2.6 | 71±3.0 | 66±3.6 | 78±2.0 | 71±3.0 |
| Root length (cm) | 26±1.0 | 12.3±1.1 | 28.3±0.5 | 23.1±0.7 | 23.8±0.7 | 23±0.5 | 29.6±0.7 | 25.1±1.0 |
| Root dry weight (g) | 2.5±0.4 | 1.3±0.2 | 2.8±0.1 | 1.9±0.2 | 3.1±0.3 | 2.5±0.4 | 2.6±0.3 | 2.3±0.1 |
| Leaf area (cm2/plant) | 93±3.6 | 39±3.0 | 93.3±2.5 | 84±3.0 | 97±2.0 | 92.3±2.5 | 100±2.6 | 87±2.0 |
| Total protein(mg g−1 fw) | 1.95±0.04 | 0.67±0.11 | 1.98±0.10 | 1.51±0.22 | 1.99±0.09 | 1.88±0.10 | 1.93±0.05 | 1.58±0.26 |
| Nitrogen (%) | 0.276±0.006 | 0.107±0.004 | 0.314±0.012 | 0.289±0.014 | 0.320±0.015 | 0.298±0.009 | 0.312±0.010 | 0.272±0.011 |
| Phosphorus (%) | 0.268±0.008 | 0.126±0.008 | 0.258±0.009 | 0.227±0.009 | 0.276±0.009 | 0.232±0.008 | 0.256±0.006 | 0.214±0.007 |
| Potassium (%) | 0.168±0.001 | 0.085±0.007 | 0.175±0.008 | 0.135±0.006 | 0.178±0.004 | 0.153±0.002 | 0.154±0.003 | 0.147±0.004 |
| Sodium (%) | 0.044±0.003 | 0.074±0.002 | 0.037±0.004 | 0.051±0.005 | 0.035±0.004 | 0.045±0.003 | 0.038±0.04 | 0.067±0.003 |
| PARAMETERS | Control plants | NaCl (mM)-grown T1 PEPCK transgenic plants | ||||||
| L2 | L7 | L12 | ||||||
| 0 | 200 | 0 | 200 | 0 | 200 | 0 | 200 | |
| Time required for flowering (days) | 86±3.6 | ND | 95±3.6 | 69.6±5.9 | 97±3.0 | 77±3.6 | 92±3.6 | 65.6±4.521 |
| No. of tillers/plant | 21.3±1.52 | ND | 24±0.17 | 16.6 ±0.57 | 26.6±0.57 | 20±0.17 | 22.6±1.15 | 14.3±0.57 |
| No. of panicle/plant | 25±1 | ND | 22.6±0.57 | 14±0.5 | 25.6±0.57 | 17±1.0 | 20±0.5 | 11.3±0.57 |
| No. of filled grain/panicle | 86±3.4 | ND | 89±3.0 | 80±3.0 | 94±3.0 | 83±3.0 | 84±3.6 | 72±3.6 |
| No. of chaffy grains/panicle | 10.3 ±0.57 | ND | 5.3±0.57 | 10.6±0.57 | 7.6±0.57 | 8.6±0.57 | 4.6±0.57 | 11±1.0 |
| Straw dry weight (g) | 55±3.0 | ND | 59±2.6 | 49±2.6 | 63±3.0 | 53.6±2.51 | 59.3±2.5 | 48.3±1.15 |
| 100 grain weight | 2.88±0.18 | ND | 2.96±0.30 | 2.27±0.13 | 3.24±0.15 | 2.63±0.06 | 2.52±0.13 | 2.25±0.06 |
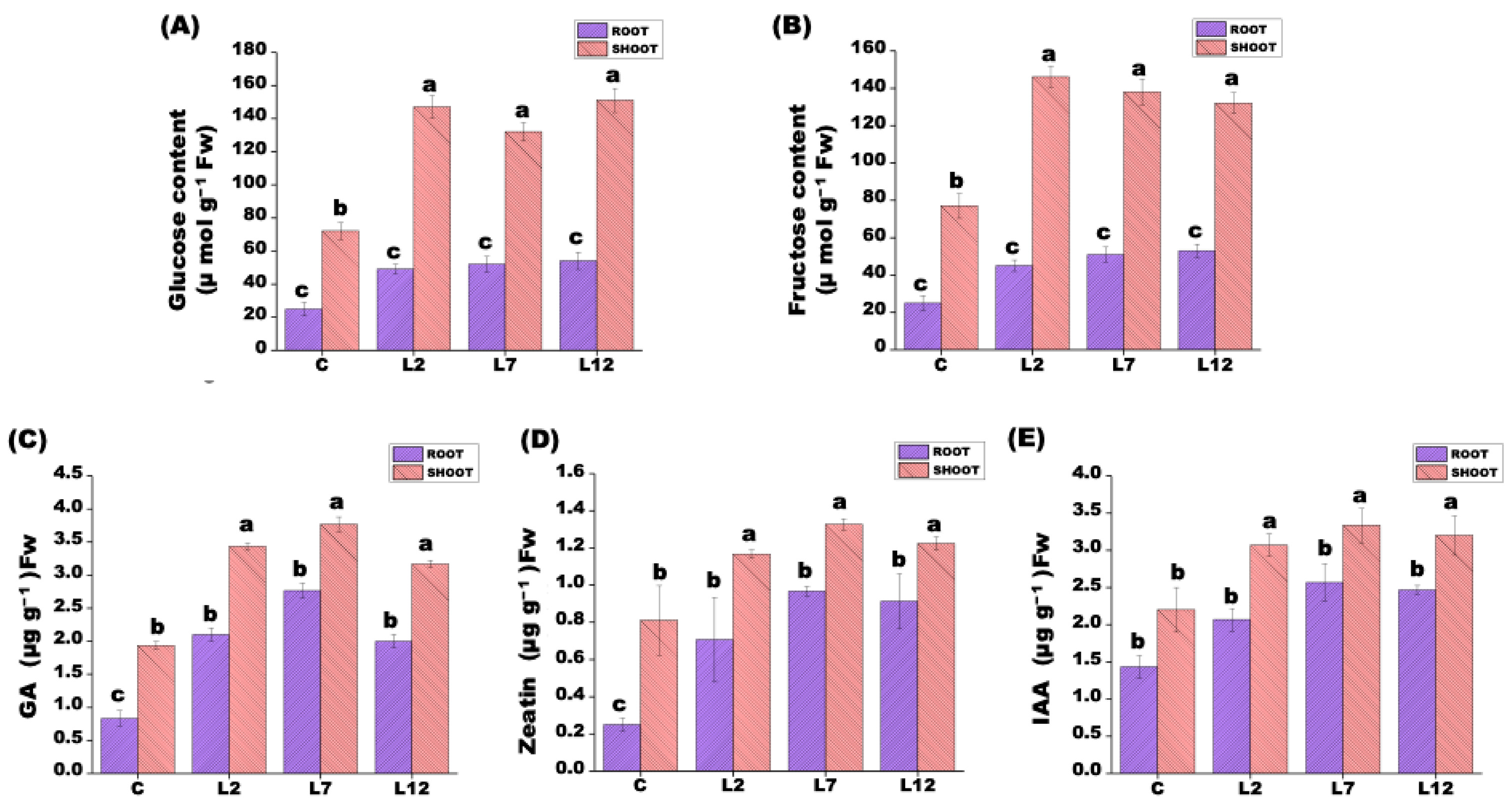
3.5. Carbohydrate Metabolism and Hormone Signaling Under Salt Stress
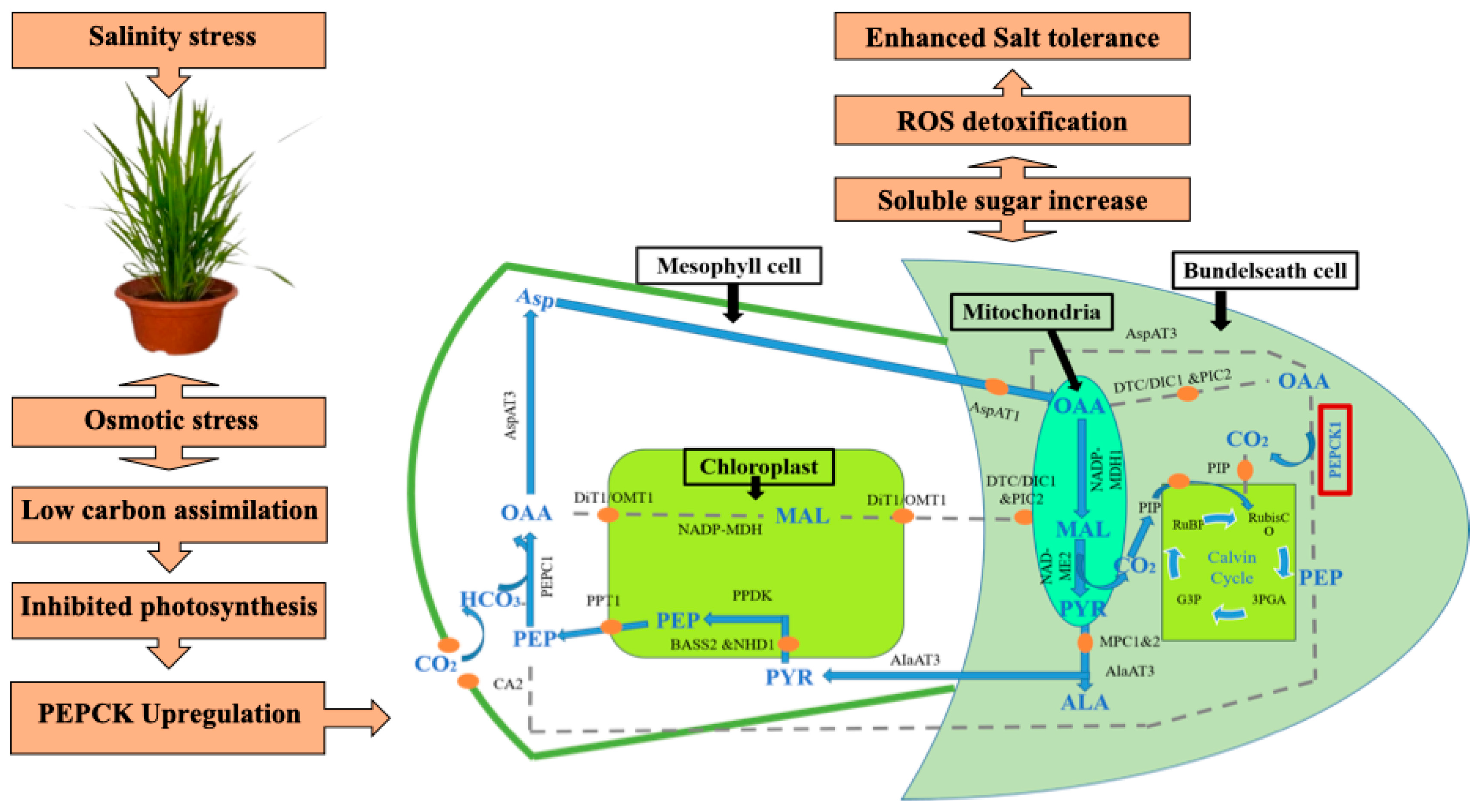
4. Discussion and Conclusion
Conflicts of Interest
References
- Li, Y.; Zhong, H.; Shan, Y.; Hang, Y.; Wang, D.; Zhou, Y.; Hubacek, K. Changes in global food consumption increase GHG emissions despite efficiency gains along global supply chains. In Nature Food; 2023. [Google Scholar] [CrossRef]
- Huang, S.; Huang, P.; Hareem, M.; Tahzeeb-ul-Hassan, M.; Younis, U.; Dawar, K.; Fahad, S.; Salmen, S.H.; Ansari, M.J.; Danish, S. Evaluating the hidden potential of deashed biochar in mitigating salinity stress for cultivation of fenugreek. Scientific Reports 2024, 14, 1, 141. [Google Scholar] [CrossRef]
- Shahid, S.A.; Zaman, M.; Heng, L. Soil Salinity: Historical Perspectives and a World Overview of the Problem. In Guideline for Salinity Assessment, Mitigation and Adaptation Using Nuclear and Related Techniques; Springer , 2018; pp. 43–53. [Google Scholar] [CrossRef]
- Mamasolieva, M.A.; L.A.G.; Hudoynazarov, I.A. Study the effect of a Biosolvent preparation on the amount of salts in saline soils and antioxidant enzymes of cotton. JSSWQ 2023, 15(2), 242–51. [Google Scholar]
- Jobe, T.O.; Zenzen, I.; Rahimzadeh, Karvansara, P.; Kopriva, S. Integration of sulfate assimilation with carbon and nitrogen metabolism in transition from C3 to C4 photosynthesis. J. Exp. Bot. 2019, 15(70), 4211–21. [Google Scholar] [CrossRef] [PubMed]
- Ermakova, M.; Arrivault, S.; Giuliani, R.; Danila, F.; Alonso-Cantabrana, H.; Vlad, D.; Ishihara, H.; Feil, R.; Guenther, M.; Borghi, G.L.; Covshoff, S.; Ludwig, M.; Cousins, A.B.; Langdale, J.A.; Kelly, S.; Lunn, J.E.; Stitt, M.; von Caemmerer, S.; Furbank, R.T. Installation of C4 photosynthetic pathway enzymes in rice using a single construct. Plant Biotechnol J. 2021, 19, 575–588. [Google Scholar] [CrossRef] [PubMed]
- Ermakova, M.; Danila, F.R.; Furbank, R.T.; von Caemmerer, S. On the road to C4 rice: advances and perspectives. T P J. 2020, 101(4), 940–50. [Google Scholar] [CrossRef]
- Shen, Z.; Dong, X.M.; Gao, Z.F.; Chao, Q.; Wang, B.C. Phylogenic and phosphorylation regulation difference of phosphoenolpyruvate carboxykinase of C3 and C4 plants. J plant physiol. 2017, 213, 16–22. [Google Scholar] [CrossRef]
- Prusty, S.; Sahoo, R.K. PEPCK Gene for Enhanced Photosynthesis and Salinity Stress Tolerance in Rice: A Review. Agricultural Reviews 2022. [Google Scholar] [CrossRef]
- Sahoo, R.K.; Tuteja, N. Development of Agrobacterium-mediated transformation technology for mature seed-derived callus tissues of indica rice cultivar IR64, GM Crops. Food 2012, 3, 123–128. [Google Scholar] [CrossRef]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2d ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, New York., 1989. [Google Scholar]
- Jayaraman, A.; Puranik, S.; Rai, N.K.; Vidapu, S.; Sahu, P.P.; Lata, C.; Prasad, M. cDNA-AFLP analysis reveals differential gene expression in response to salt stress in foxtail millet (Setaria italica L.). Mol Biotechnol. 2008, 40, 241–251. [Google Scholar] [CrossRef]
- Tuteja, N.; Sahoo, R.K.; Garg, B.; Tuteja, R. OsSUV3 dual helicase functions in salinity stress tolerance by maintaining photosynthesis and antioxidant machinery in rice (Oryza sativa L. cv. IR64). Plant J. 2013, 76, 115–127. [Google Scholar] [CrossRef]
- Garg, B.; Jaiswal, J.P.; Misra, S.; Tripathi, B.N; Prasad, M.A. A comprehensive study on dehydration-induced antioxidative responses during germination of Indian bread wheat (Triticum aestivum L. em Thell) cultivars collected from different agroclimatic zones. Physiol Mol Biol Plants 2012, 18, 217–228. [Google Scholar] [CrossRef]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of proline in water stress studies. Plant Soil. 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Jana, S.; Choudhuri, M.A. Glycolate metabolism of three submerged aquatic angiosperms during aging. Aquat. Bot. 1981, 12, 345–354. [Google Scholar] [CrossRef]
- De Vos, C.H.R.; Schat, H.; Vooijs, R.; Ernst. WHO Copper-induced damage to the permeability barrier in roots of Silene cucubalus. J. Plant Physiol. 1989, 135, 164–179. [Google Scholar] [CrossRef]
- Blum, A.; Ebercon, A. Cell membrane stability as a measure of drought and heat tolerance in wheat 1. Crop Science 1981, 21(1), 43–7. [Google Scholar] [CrossRef]
- Barrs, H.D.; Weatherley, P.E. A re-examination of the relative turgidity technique for estimating water deficit in leaves. Aust J Biol Sci. 1962, 15, 413–428. [Google Scholar] [CrossRef]
- Jackson, T.L. Soil chemical analysis; Prentice-Hall of India Private Limited: New Delhi, 1973. [Google Scholar]
- Gupta, P.K. Methods in environmental analysis water, soil and air. Agrobios, India 2004, 242–245. [Google Scholar]
- Chapman, H.D.; Pratt, P.F. Method and of analysis of soil, plant and water, 2nd edn; California University Agricultural Division, California, 1982; p. 170. [Google Scholar]
- Munns, R.; Wallace, P.A.; Teakle, N.L.; Colmer, T.D. Measuring soluble ion concentrations (Na+, K+, Cl−) in salt-treated plants. In Methods in molecular biology;Plant stress tolerance:methods and protocols; Sunkar, R, Ed.; Humana Press. Springer., 2010; vol 639, pp. 371–382. [Google Scholar] [CrossRef]
- Karkacier, M.; Erbas, M.; Uslu, M.K.; Aksu, M. Comparison of different extraction and detectionmethods for sugars using amino-bonded phase HPLC. J Chromatogr Sci. 2003, 41, 331–333. [Google Scholar] [CrossRef]
- Chen, J.; Du. X.M., G; Zhao, H.Y.; Zhou, X. Fluctuation in levels of endogenous plant hormones in ovules of normal and mutant cotton during flowering and their relation to fiber development. J. Plant Growth Regul. 1996, 15, 173–177. [Google Scholar] [CrossRef]
- Li, Y.; Tan, B.; Wang, D.; Mu, Y.; Li, G.; Zhang, Z.; Pan, Y.; Zhu, L. Proteomic analysis revealed different molecular mechanisms of response to PEG stress in drought-sensitive and drought-resistant sorghums. Int. J. Mol. Sci. 2022, 23, 2113297. [Google Scholar] [CrossRef]
- Feria, A.B.; Bosch, N.; Sánchez, A.; Nieto-Ingelmo, A.I.; Osa, C.; Echevarría, C.; Mauriño, S.G.; Monreal, J.A. Phosphoenolpyruvate carboxylase (PEPC) and PEPC-kinase (PEPC-k) isoenzymes in Arabidopsis thaliana: role in control and abiotic stress conditions. Planta 2016, 244, 901–913. [Google Scholar] [CrossRef] [PubMed]
- Hao, S.; Wang, Y.; Yan, Y.; Liu, Y.; Wang, J.; Chen, S. A review on plant responses to salt stress and their mechanisms of salt resistance. Horticulturae 2021, 7, 6–132. [Google Scholar] [CrossRef]
- Wang, J.; Nan, N.; Li, N.; Liu, Y.; Wang, T.J.; Hwang, I.; Liu, B.; Xu, Z.Y. A DNA methylation reader–chaperone regulator–transcription factor complex activates OsHKT1; 5 expression during salinity stress. Plant Cell. 2020, 32, 3535–58. [Google Scholar] [CrossRef] [PubMed]
- Tuteja, N.; Sahoo, R.K.; Huda, K.M.; Tula, S.; Tuteja, R. OsBAT1 augments salinity stress tolerance by enhancing detoxification of ROS and expression of stress-responsive genes in transgenic rice. Plant Mol Biol Rep. 2015, 33, 1192–209. [Google Scholar] [CrossRef]
- Sahoo, R.K.; Tuteja, R.; Gill, R.; Jiménez Bremont, J.F.; Gill, S.S.; Tuteja, N. Marker-free rice (Oryza sativa L. cv. IR 64) overexpressing PDH45 gene confers salinity tolerance by maintaining photosynthesis and antioxidant machinery. Antioxidants 2022, 11–770. [Google Scholar] [CrossRef]
- Abdala, K.A.; Liao, J.; Mohammed, E.A.; Wu, J.M. Transgenic rice (Oryza sativa L.) varieties exhibit superior salt stress resilience: a comparative study of physiological and biochemical responses in transgenic and non-transgenic genotypes. Plant Physiol. Rep. 2024, 29, 948–74. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Zhao, X.; Yang, S.; Huang, L.; Du, F.; Li, Z.; Zhao, X.; Fu, B.; Wang, W. Overexpression of the transcription factor gene OsSTAP1 increases salt tolerance in rice. Rice 2020, 13, 1–2. [Google Scholar] [CrossRef]
- Cao, W.; Zhang, H.; Zhou, Y.; Zhao, J.; Lu, S.; Wang, X.; Chen, X.; Yuan, L.; Guan, H.; Wang, G.; Shen, W. Suppressing chlorophyll degradation by silencing OsNYC3 improves rice resistance to Rhizoctonia solani, the causal agent of sheath blight. Plant Biotechnol J. 2022, 20(2), 335–49. [Google Scholar] [CrossRef]
- Mukherjee, S.; Mukherjee, A.; Das, P.; Bandyopadhyay, S.; Chattopadhyay, D.; Chatterjee, J.; Majumder, A.L. A salt-tolerant chloroplastic FBPase from Oryza coarctata confers improved photosynthesis with higher yield and multi-stress tolerance to indica rice. PCTOC. 2021, 145, 561–78. [Google Scholar] [CrossRef]
- Nounjan, N.; Mahakham, W.; Siangliw, J.L.; Toojinda, T.; Theerakulpisut, P. Chlorophyll retention and high photosynthetic performance contribute to salinity tolerance in rice carrying drought tolerance quantitative trait loci (QTLs). Agriculture. 2020, 10, 620. [Google Scholar] [CrossRef]
- Liu, C.; Hu, Z.H.; Yu, L.F.; Chen, S.T.; Liu. XM Responses of photosynthetic characteristics and growth in rice and winter wheat to different elevated CO2 concentrations. Photosynthetica 2020, 58, 5. [Google Scholar] [CrossRef]
- Yildiz, M.; Poyraz, İ.; Çavdar, A.; Özgen, Y.; Beyaz, R. Plant responses to salt stress. IntechOpen 2020. [Google Scholar] [CrossRef]
- Basu, S.; Kumari, S.; Kumar, P.; Kumar, G.; Rajwanshi, R. Redox imbalance impedes photosynthetic activity in rice by disrupting cellular membrane integrity and induces programmed cell death under submergence. Physiologia Plantarum 2021, 172(3), 1764–78. [Google Scholar] [CrossRef] [PubMed]
- Kaewneramit, T.; Buaboocha, T.; Sangchai, P.; Wutipraditkul, N. OsCaM1-1 overexpression in the transgenic rice mitigated salt-induced oxidative damage. Biol Plant. 2019, 63, 335. [Google Scholar] [CrossRef]
- Ali, M.; Kamran, M.; Abbasi, G.H.; Saleem, M.H.; Ahmad, S.; Parveen, A.; Malik, Z.; Afzal, S.; Ahmar, S.; Dawar, K.M.; Ali, S. Melatonin-induced salinity tolerance by ameliorating osmotic and oxidative stress in the seedlings of two tomato (Solanum lycopersicum L.) cultivars. J. Plant Growth Regul. 2021, 40, 2236–48. [Google Scholar] [CrossRef]
- Ejaz, S.; Fahad, S.; Anjum, M.A.; Nawaz, A.; Naz, S.; Hussain, S.; Ahmad, S. Role of osmolytes in the mechanisms of antioxidant defense of plants. SAR. 2020, 39, 95–117. [Google Scholar] [CrossRef]
- Gao, Y.; Long, R.; Kang, J.; Wang, Z.; Zhang, T.; Sun, H.; Li, X.; Yang, Q. Comparative proteomic analysis reveals that antioxidant system and soluble sugar metabolism contribute to salt tolerance in alfalfa (Medicago sativa L.) leaves. J. Proteome Res. 2018, 18, 191–203. [Google Scholar] [CrossRef]
- Bai, X.; Zeng, X.; Huang, S.; Liang, J.; Dong, L.; Wei, Y.; Li, Y.; Qu, J.; Wang, Z. Marginal impact of cropping BADH transgenic maize BZ-136 on chemical property, enzyme activity, and bacterial community diversity of rhizosphere soil. Plant Soil. 2019, 436, 527–541. [Google Scholar] [CrossRef]
- Yang, Z.; Shen, L.; Hu, L.; Cai, Y.; Zheng, Q.; Wu, Y. The PEPCK and FBP genes regulate gluconeogenesis metabolism in grape berries in response to light intensity. Horticulturae 2024, 10(12), 1270. [Google Scholar] [CrossRef]
- Anjum, N.; Maiti, M.K. OsNAC121 regulates root development, tillering, panicle morphology, and grain filling in rice plant. Plant Mol. Biol. 2024, 114(4), 1–20. [Google Scholar] [CrossRef]
- Swain, D.M.; Sahoo, R.K.; Srivastava, V.K.; et al. Function of heterotrimeric G-protein γ subunit RGG1 in providing salinity stress tolerance in rice by elevating detoxification of ROS. Planta 2017, 245, 367–383. [Google Scholar] [CrossRef]
- Swain, D.M.; Sahoo, R.K.; Chandan, R.K.; et al. Concurrent overexpression of rice G-protein β and γ subunits provide enhanced tolerance to sheath blight disease and abiotic stress in rice. Planta 2019, 250, 1505–1520. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




