1. Introduction
Globally, respiratory infections remain a leading cause of mortality, with influenza viruses representing major contributors to seasonal viral respiratory epidemics [
1]. Among the four types of influenza viruses—A, B, C, and D—influenza A viruses are the most frequently detected and are responsible for the majority of influenza epidemics in humans [
2]. Seasonal influenza represents a substantial global disease burden, with approximately one billion infections occurring each year, of which 3 to 5 million are classified as severe, resulting in an estimated 290,000 to 650,000 deaths annually. Reducing the impact of seasonal influenza through improved surveillance, prevention, and control helps countries prepare for future pandemics [
1]. Influenza virus infection causes an acute respiratory illness primarily affecting the airways and is characterized by symptoms including fever, dry cough, headache, chills, and sore throat [
3].
Influenza A viruses (IAVs) are categorized into multiple subtypes based on the presence of two surface glycoproteins, hemagglutinin (HA) and neuraminidase (NA), which are primarily found in wild birds. To date, IAVs have been classified into 18 HA subtypes (H1–H18) and 11 NA subtypes (N1–N11), which can combine in various ways to produce subtypes such as H1N1, H3N2, and others [
2]. Influenza A viruses (IAVs) belong to the family Orthomyxoviridae. Their genome consists of eight segments of negative-sense single-stranded RNA, with a total length of approximately 13,000 nucleotides, encoding at least 12 viral proteins [
4].
Historically, influenza pandemics have occurred every few decades, with their timing and severity influenced by various ecological and epidemiological factors. In 1918, influenza A viruses caused one of the most devastating pandemics in recorded history, known as the Spanish flu, which affected approximately one-third of the world’s population and resulted in an estimated 50–100 million deaths [
5,
6]. In Libya, during the 2009 influenza pandemic, the first cases of influenza A(H1N1) were identified in July 2009 through the surveillance system of the Libyan National Center for Disease Control (NCDC) [
7].
In the Northern Hemisphere, influenza activity typically peaks between January and March [
8]. As Libya, is located in the Northern Hemisphere, it is expected to follow a similar seasonal influenza pattern [
1,
9]. Although several mechanisms have been proposed to explain the seasonality of influenza in humans—including climatic factors, human behavioral patterns, and the effects of environmental changes on the immune system—the seasonal dynamics of respiratory viruses remain incompletely understood [
10].
Influenza A viruses (IAVs) possess a well-established capacity to generate new subtypes through rapid genetic changes, which underlies their potential for global spread. Understanding their genetic evolution is therefore essential for predicting outbreak risks and guiding the development of effective vaccines and antiviral therapies [
11]. According to the World Health Organization, A(H1N1)pdm09 was the most active influenza A subtype in North Africa between September 2023 and January 2024 [
12].
Despite extensive global surveillance efforts, significant challenges persist because influenza A viruses (IAVs) continuously evolve, often reducing vaccine effectiveness when antigenic mismatches occur [
13]. Consequently, a comprehensive understanding of influenza A virus genetic diversity is critical for optimizing vaccine design and strengthening future pandemic preparedness [
14]. Whole-genome sequencing plays a crucial role in detecting novel IAV reassortants and characterizing their molecular properties, including host range and virulence [
15]. Influenza A(H1N1) RNA remains stable for up to 56 days at −80 °C or −20 °C and after up to nine freeze–thaw cycles when extracted, whereas RNA in nasopharyngeal swab specimens remains stable for at least three years at −80 °C or after up to seven freeze–thaw cycles when re-extracted. Consequently, 0.5–1.0 mL of influenza-positive samples can be stored at −80 °C for up to three years without significant loss of viral RNA, allowing accurate viral load assessment in retrospective and clinical studies [
16]. In modern molecular epidemiology, phylogenetic analysis has become an indispensable tool for tracking the evolutionary dynamics and transmission pathways of seasonal influenza viruses. The use of real-time genomic surveillance platforms, such as Nextstrain, facilitates rapid sharing and visualization of analyses for important pathogens, including influenza A(H1N1) and A(H3N2) [
17].
The aim of this study was to detect, subtype, and sequence influenza A viruses (IAVs) in human samples collected through the NCDC sentinel surveillance system in Libya, during recent influenza seasons, in order to address the existing gap in influenza genomic data from the country.
2. Materials and Methods
2.1. Influenza Surveillance in Libya
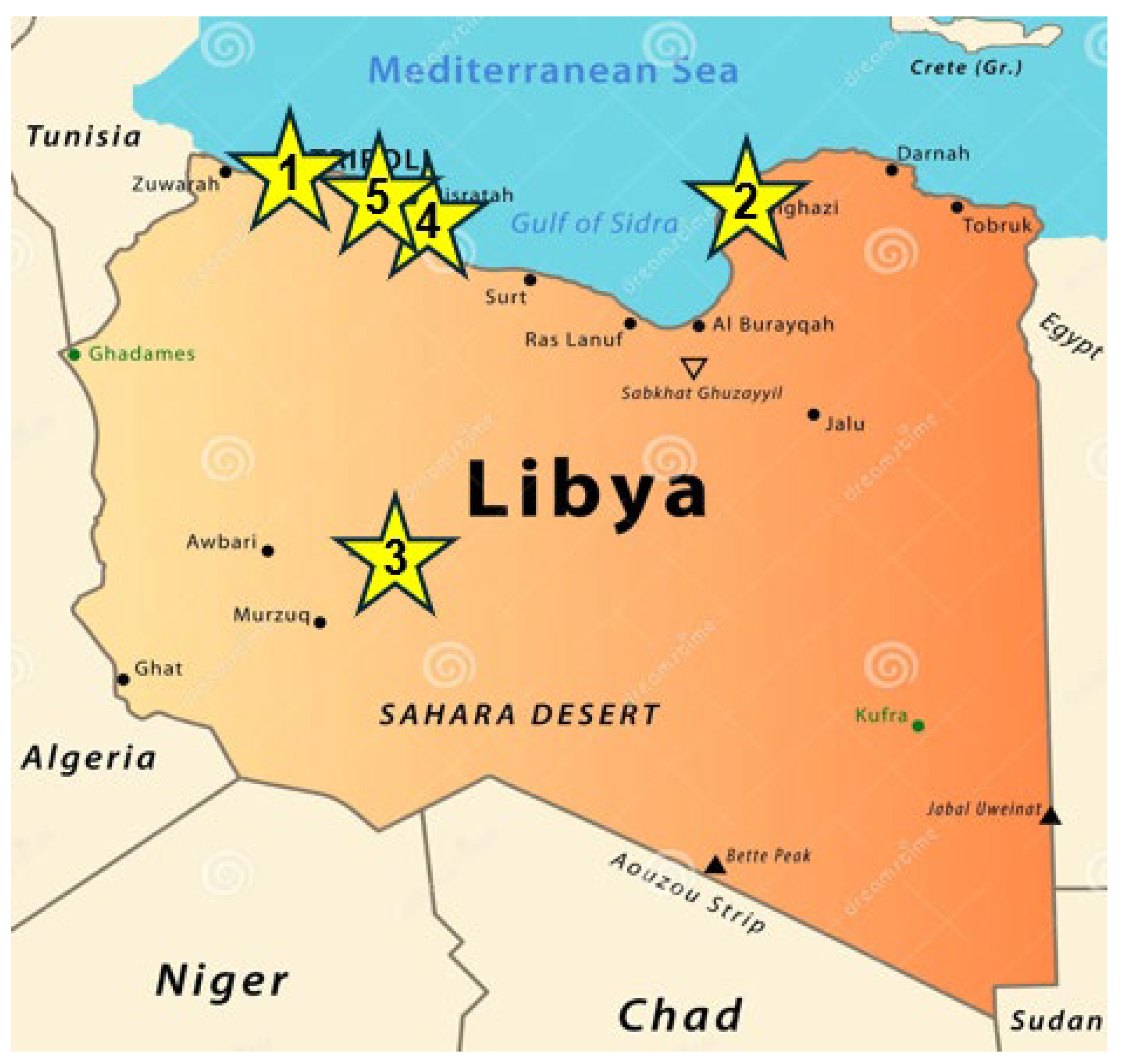
The NCDC established an influenza sentinel surveillance program in 2022, initially comprising a single sentinel site in Tripoli. By the end of 2025, the network had expanded to 15 sentinel sites across five cities in Libya (
Figure 1).
2.2. Collection of Samples
A total of 1,864 clinical samples were obtained from patients presenting with influenza-like symptoms as part of the routine surveillance program conducted by the NCDC across 15 sentinel sites in five Libyan cities. Samples were collected and tested during the influenza seasons from early 2022 to the end of 2024. All influenza-positive nasopharyngeal swabs were stored at −80 °C to preserve RNA integrity.
2.3. Laboratory Analysis
A total of 1864 nasopharyngeal samples were tested using the GeneXpert system in the laboratories of all influenza sentinel sites to detect four viruses (influenza type A, influenza type B, SARS-CoV-2 and RSV). Out of the positive samples by GeneXpert, a total of 29 samples of influenza type A were randomly selected, representing the recent influenza seasons, for further analysis in this study.
One milliliter from each of the 29 selected samples was used to confirm the presence of influenza A virus at the National Research Center for Tropical and Transboundary Diseases (NRCTTD) laboratory. Viral RNA was extracted using the Sterilab NC-15 Plus Automated System with the Alpha & Gene extraction kit. Subsequently, real-time RT-PCR was performed using the Long Gene RT-PCR system to detect IAVs, according to the manufacturer’s instructions.
The remaining portions of the 29 samples were then transferred to the National Influenza Center Tunisia for further confirmation, subtyping, and sequencing. RNA was re-extracted using the QIAGEN EZ1 Connect MDx system (Hilden, Germany) with the EZ1 DSP Virus Kit. RNA quantity and quality were assessed using a Qubit Fluorometer.
Real-time RT-PCR was performed using the Applied Biosystems 7500 Fast RT-PCR System with the CDC rRT-PCR Influenza Panel, strictly following the kit instructions. Influenza A subtyping was subsequently conducted on the same platform using the CDC Influenza Virus RT-PCR Subtyping Panel, in accordance with the recommended protocol add ref.
Finally, RNA library preparation was carried out using Illumina RNA Prep with Enrichment and a respiratory virus panel. Sequencing was performed using the Illumina iSeq 100 System.
2.4. Bioinformatic Analysis
Following completion of the Illumina iSeq 100 sequencing run, raw sequencing data were converted into FASTQ files using Illumina Local Run Manager. Genome Detective was used to perform read quality control, trimming, de novo assembly, and reference-based alignment against influenza A virus reference genomes. Based on these analyses, consensus genome sequences were generated and exported in FASTA format. The resulting consensus FASTA sequences were subsequently submitted to the GISAID EpiFlu™ database. Mutation analysis was performed using FluSurver (GISAID) to identify nucleotide and amino acid substitutions related to reference influenza A strains.
Phylogeographic analyses were conducted based on the HA gene (HA1 and HA3). Closely related reference sequences were selected using the Audacity Instant tool in GISAID. Two datasets were generated: 118 HA1 sequences (
https://doi.org/10.55876/gis8.260101zo) and 58 HA3 sequences (
https://doi.org/10.55876/gis8.260101fs). All data were used according to GISAID terms, and DOIs were generated. These datasets were analyzed for phylogenetic and spatial diffusion analyses using the Nextstrain pipeline.
3. Results
3.1. Detection of Influenza Type A, Influenza Type B, SARS-CoV-2 and RSV
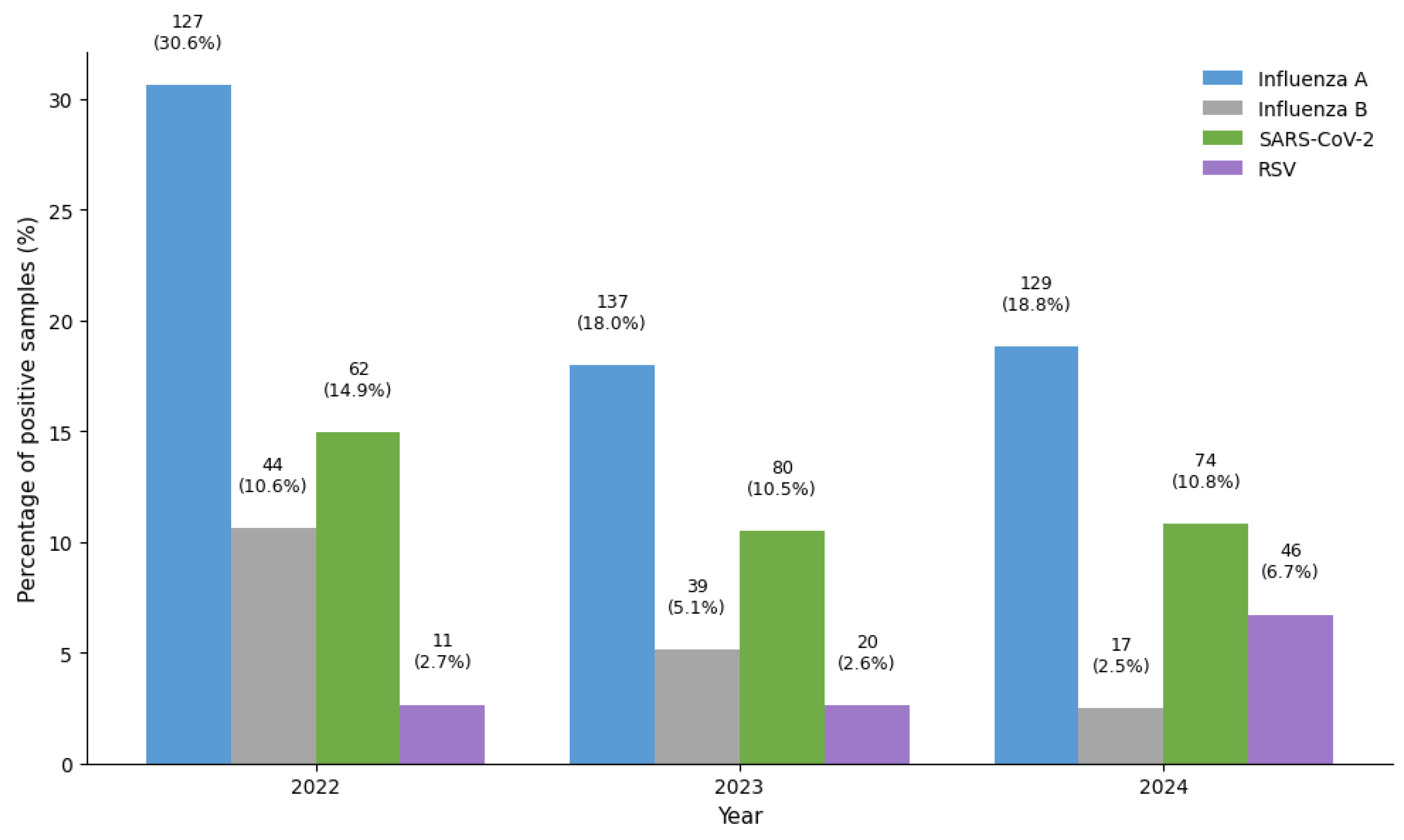
Out of 1864 samples collected by the NCDC sentinel sites in Libya during 2022–2024 they were tested for influenza type A, influenza type B, SARS-CoV-2, and RSV. Influenza virus was detected in 26.4% (493/1864) of samples [influenza type A was detected in 21.1% (393/1864) of samples and influenza B in 5.4% (100/1864)], SARS-CoV-2 was detected in 11.6% (216/1864) of samples, and RSV was detected in 4.1% (77/1864) of samples (
Figure 2).
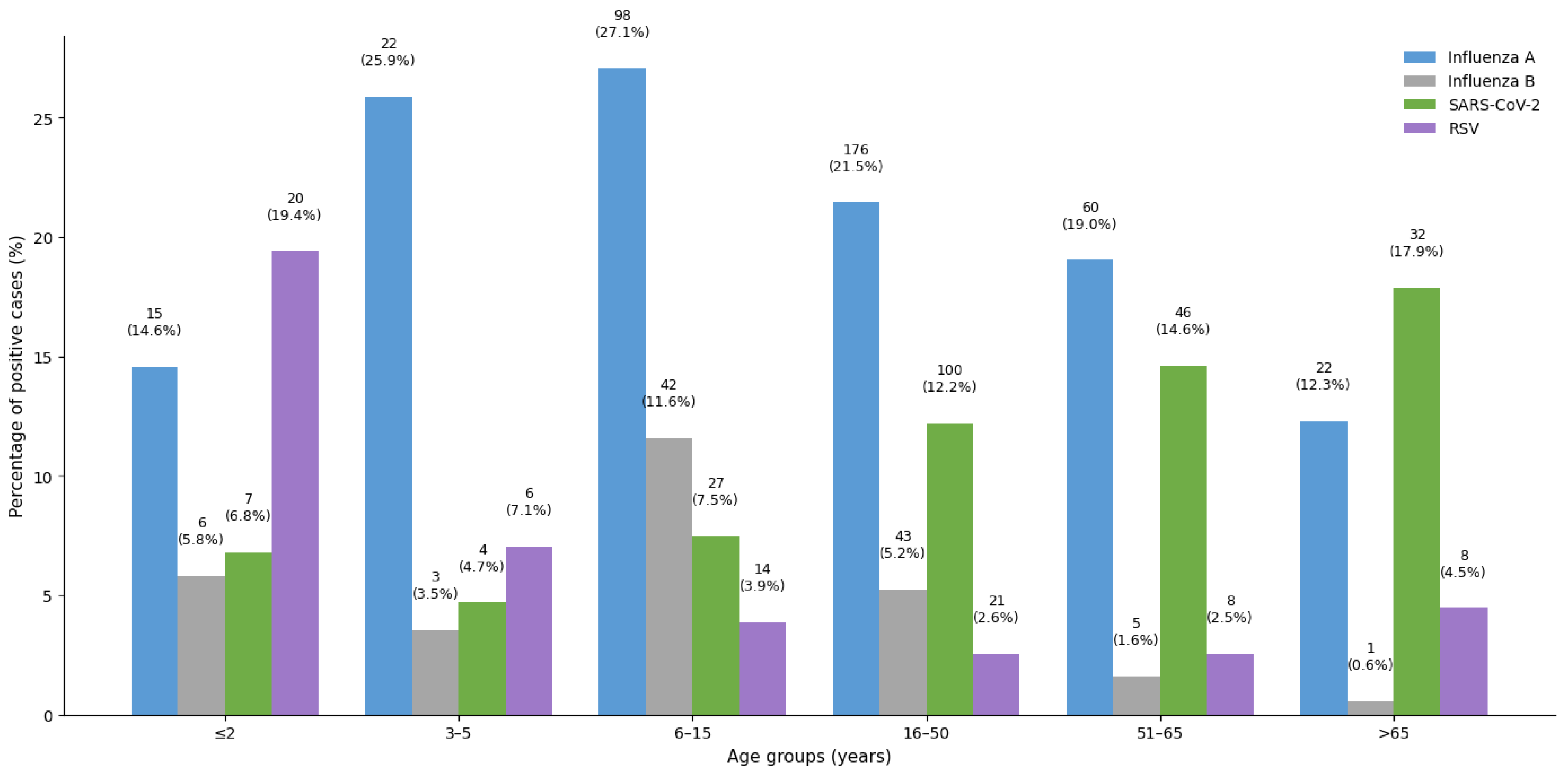
The results of the number and percentage for each age group of positive samples for influenza type A, influenza type B, SARS-CoV-2, and RSV among tested samples (2022-2024) are shown in
Figure 3.
The twenty-nine previously described samples, which were initially positive for influenza A by GeneXpert and stored at −80 °C, were selected for further analyses. Upon retesting these samples at the NRCTTD and National Influenza Center of Tunisia only eighteen of the samples were confirmed positive in both laboratories (
Table 1).
3.2. Subtypes Identified on Confirmed Positive Samples
Of the 18 samples confirmed as positive at the National Influenza Center of Tunisia, the majority were identified as influenza A(H1N1) pdm09 (n = 13), whereas the remaining samples were classified as influenza A(H3N2) (n = 5). Thirteen of these samples were subsequently selected for sequencing.
3.3. Sequencing Output
From the influenza A–positive samples, 13 were selected for whole genome sequencing. Complete genome sequences covering all eight gene segments were successfully obtained for seven H1N1 isolates, while partial genome sequences were obtained for the remaining samples, including high-quality HA sequences from three additional H1N1 and two H3N2 isolates, as well as an NA sequence from one additional H1N1 isolate.
Genetic analysis showed that all A(H1N1) pdm09 strains belonged to clade 6B.1A.5a.2a (5a.2a), while the A(H3N2) strains clustered within clade 3C.2a1b.2a.2a.3a.1 (2a.3a.1) based on the HA mutation profile. No mutations associated with antiviral resistance were detected based on the NA gene analysis.
Table 1.
Ct values of 29 selected Influenza A samples tested and re-confirmed by RT-PCR.
Table 1.
Ct values of 29 selected Influenza A samples tested and re-confirmed by RT-PCR.
| NCDC CODE |
Age |
Gender |
Collection Date |
FLU A1 Ct |
FLU A Ct NRCTTD |
Flu A Ct CN |
Flu subtype |
| 2651 |
48 |
F |
21/11/2023 |
28.9 |
NEG |
NEG |
- |
| 2646 |
7 |
M |
21/11/2023 |
22.1 |
28 |
29 |
H3N2 |
| 2663 |
33 |
F |
29/11/2023 |
18.3 |
29 |
30 |
H1N1 |
| 2681 |
58 |
M |
05/12/2023 |
20.1 |
28 |
35 |
H1N1 |
| 2757 |
21 |
F |
26/12/2023 |
20 |
33 |
32 |
H1N1 |
| 2755 |
50 |
F |
26/12/2023 |
24.1 |
31 |
32 |
H1N1 |
| 2756 |
18 |
M |
26/12/2023 |
15.7 |
30 |
27 |
H1N1 |
| 2771 |
15 |
F |
26/12/2023 |
23.3 |
32 |
33 |
H1N1 |
| 2777 |
45 |
F |
26/12/2023 |
20.2 |
NEG |
NEG |
- |
| 2769 |
9 |
F |
26/12/2023 |
33 |
NEG |
NEG |
- |
| 2765 |
47 |
M |
26/12/2023 |
24.8 |
NEG |
NEG |
- |
| 2786 |
60 |
M |
27/12/2023 |
22.1 |
NEG |
NEG |
- |
| 2849 |
45 |
F |
02/01/2024 |
32 |
NEG |
NEG |
- |
| 2839 |
90 |
F |
02/01/2024 |
14.8 |
29 |
28 |
H1N1 |
| 2827 |
23 |
F |
02/01/2024 |
21.8 |
NEG |
37 |
H3N2 |
| 3620 |
75 |
F |
17/12/2024 |
24.8 |
35 |
35 |
H3N2 |
| 3623 |
45 |
M |
22/12/2024 |
24.3 |
34 |
35 |
H1N1 |
| 3627 |
29 |
F |
25/12/2024 |
19.4 |
32 |
31 |
H1N1 |
| 3646 |
60 |
F |
29/12/2024 |
16.6 |
30 |
31 |
H3N2 |
| 3634 |
58 |
F |
29/12/2024 |
27.5 |
35 |
35 |
H3N2 |
| 3648 |
40 |
M |
31/12/2024 |
17.1 |
33 |
34 |
H3N2 |
| 3654 |
48 |
M |
31/12/2024 |
18.9 |
36 |
NEG |
- |
| 024-2025 |
42 |
F |
05/01/2025 |
23.2 |
NEG |
NEG |
- |
| 034-2025 |
2 |
M |
06/01/2025 |
19.5 |
21 |
34 |
H1N1 |
| 082-2025 |
52 |
F |
07/01/2025 |
35.3 |
NEG |
NEG |
- |
| 108-2025 |
29 |
F |
08/01/2025 |
16.5 |
36 |
33 |
H1N1 |
| 167-2025 |
37 |
M |
16/01/2025 |
23.9 |
37 |
NEG |
- |
| 210-2025 |
65 |
F |
21/01/2025 |
23.6 |
NEG |
NEG |
- |
| 213-2025 |
43 |
F |
22/01/2025 |
32.6 |
NEG |
NEG |
- |
3.4. Molecular Characterization
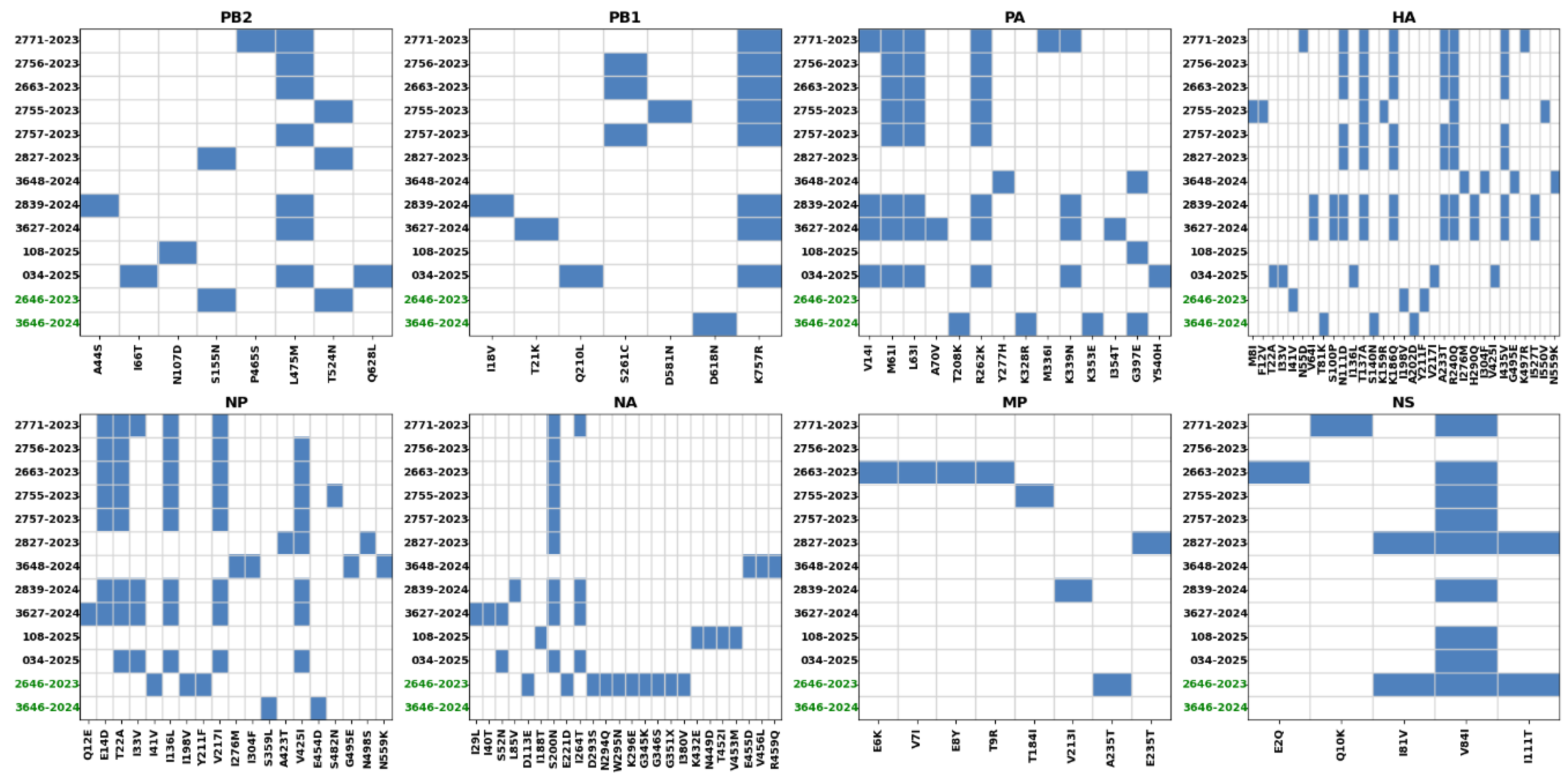
Multiple amino acid substitutions were identified in both HA1 and HA2 regions. In A(H1N1)pdm09 isolates, HA1 substitutions included I25V, E50K, G53N, T65K, N96S, N122D, S124N, I140K, S145N, I192F, I223V, K276E, and N49S, along with signal peptide–associated changes. Additional substitutions were observed in the HA2 region. In contrast, A(H3N2) isolates exhibited a distinct mutation profile, with HA1 substitutions including E50K, G53N, N96S, K121E, N122D, I140K, I192F, F195Y, Q197K, N49S, G150E, and N214K, and HA2 substitutions such as I276M, I304F, G495E, N559K, T81K, S140N, and A202D.
Analysis of the neuraminidase (NA) gene revealed further amino acid variability, including N294Q, W295N, K296E, G345K, I264T, L85V, I29L, I40T, S52N, Q51K, N58L, N59S, and C-terminal changes (K432E, N449D, T452I, V453N). The S200N substitution was recurrent across multiple sequences, often appearing alongside I264T and/or S52N, highlighting common mutation patterns among circulating strains. Variation in NA mutation profiles was observed among different isolates.
Whole-genome sequencing also demonstrated amino acid substitutions across additional viral proteins, including PB2, MP, NS, and PA-X, with some substitutions recurring among isolates. One isolate presented only partial NA sequence data, but overall, these findings reflect genome-wide genetic diversity among the sequenced IAVs.
The detailed amino acid substitutions identified in HA, NA, and internal gene segments are summarized in
Figure 4.
This study represents the first submission of human influenza A virus sequences from Libya to the GISAID database. The deposited sequences were assigned the following accession numbers: EPI_ISL_20311925 (A/Libya/3646/2024), EPI_ISL_20311841 (A/Libya/034/2025), EPI_ISL_20311731 (A/Libya/3627/2024), EPI_ISL_20311675 (A/Libya/2839/2024), EPI_ISL_20311593 (A/Libya/2771/2023), EPI_ISL_20311567 (A/Libya/2757/2023), EPI_ISL_20311527 (A/Libya/2756/2023), EPI_ISL_20311468 (A/Libya/2755/2023), and EPI_ISL_20311368 (A/Libya/2663/2023).
3.5. Phylogenetic Analysis of the Hemagglutinin (HA) Gene Segment
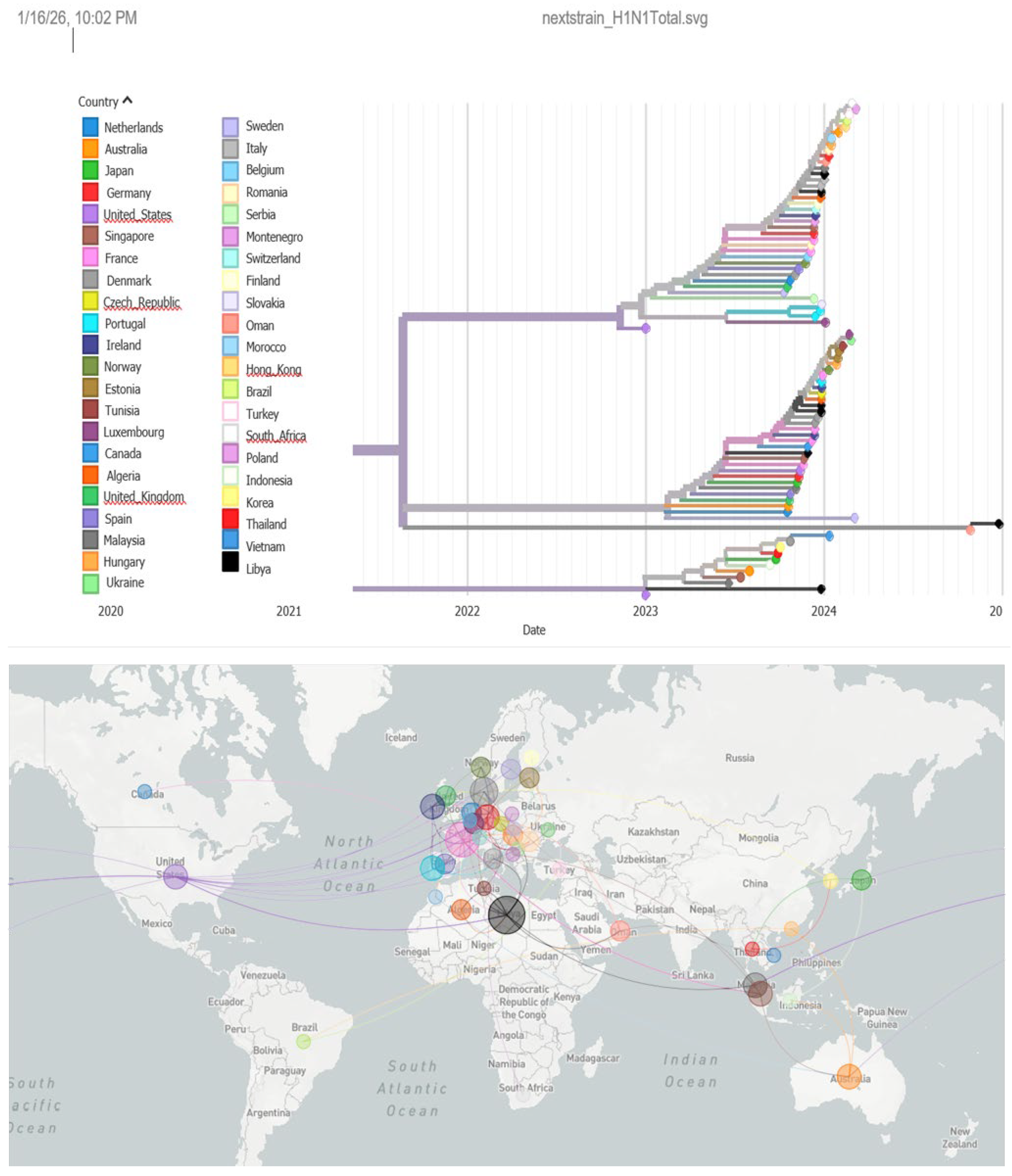
Phylogenetic analysis of the HA gene was performed using two datasets comprising 118 H1N1 sequences (including seven Libyan isolates) sampled during 2021–2024 and 58 H3N2 sequences (including two Libyan isolates) sampled during 2022–2024, representing the global diversity of circulating viruses.
The H1N1 phylogeny showed that Libyan isolates clustered within multiple branches of the global tree and were closely related to European strains, particularly from Sweden, the Netherlands, and Spain, suggesting recent introductions from Europe into Libya (
Figure 5). The limited genetic divergence among the Libyan sequences indicates either a single introduction followed by local transmission or multiple introductions of closely related strains. The interspersion of Libyan sequences with those from other North African countries supports regional connectivity and ongoing cross-border viral movement (
Figure 5).
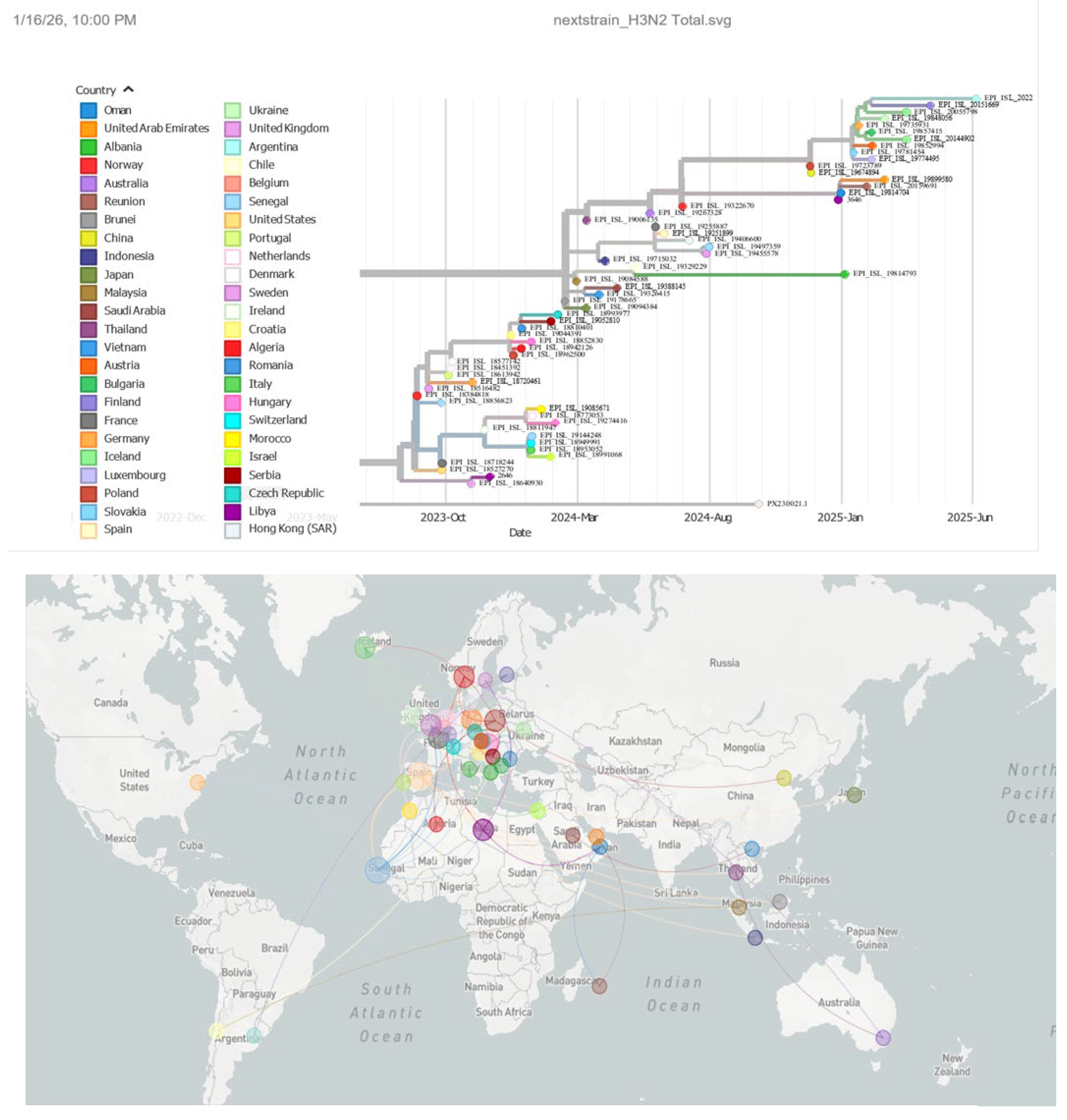
In contrast, the Libyan H3N2 isolates belonged to a lineage that emerged in Europe during the 2021–2022 season and was introduced into Africa primarily via West Africa before reaching Libya (
Figure 6). This lineage showed limited circulation within North Africa and low genetic diversity among Libyan isolates, consistent with a single or few introduction events and restricted local transmission (
Figure 6).
Overall, these findings reveal distinct dispersal patterns of influenza A subtypes in North Africa, with broader regional circulation of H1N1 compared to H3N2, highlighting the importance of continuous molecular surveillance to monitor viral spread at both regional and global scales.
4. Discussion
4.1. Comparison of Influenza A Detection with Other Respiratory Viruses
Previous studies in Libya have reported variable detection rates of influenza A across different epidemiological periods [
18]. For example, only 2.8% of tested samples were positive for influenza A during the 2021–2022 season in Libya. This low detection rate was likely attributable to public health interventions implemented during the COVID-19 pandemic, including widespread mask use, reduced social gatherings, and increased public awareness of respiratory infection prevention. Collectively, these measures contributed to a substantial decline in the circulation of seasonal influenza viruses.
In contrast, Elahmer et al. [
7] reported a markedly higher influenza A positivity rate (35%) during the 2009 H1N1 pandemic, reflecting the enhanced transmission dynamics typically observed during pandemic periods. At the regional level, influenza A has been reported to predominate, accounting for approximately 67% of influenza cases in the Eastern Mediterranean and North African regions [
19]. Similar trends have been reported globally, with influenza A accounting for 91.8% of detected influenza cases in China between December 2022 and January 2023 [
20].
In the present study, surveillance data from the NCDC demonstrated that influenza A was the most frequently detected respiratory virus during the 2022, 2023, and 2024 influenza seasons. An increasing trend in influenza A detection was observed across these seasons, indicating a clear resurgence of seasonal influenza activity following the COVID-19 pandemic. This pattern is consistent with the relaxation of non-pharmaceutical interventions, including reduced mask use and increased population mobility, which likely facilitated the re-emergence and sustained circulation of influenza viruses. The continued predominance of influenza A underscores its public health significance and highlights the need for sustained molecular surveillance, particularly in regions such as Libya, where influenza genomic data remain limited.
In addition to epidemiological factors, pre-analytical conditions likely influenced viral detection efficiency, as sample storage, handling, and transportation are critical determinants of RNA integrity [
21]. In the present study, repeated freeze–thaw cycles and delays in sample processing may have contributed to RNA degradation. Although 29 samples were initially positive, only 18 yielded detectable viral RNA upon re-analysis. Additionally, cycle threshold (Ct) values were notably higher than those obtained during initial testing, indicating reduced viral RNA concentrations, likely due to RNA instability associated with suboptimal storage conditions. These findings highlight the importance of minimizing freeze–thaw cycles and ensuring optimal sample preservation to enhance molecular detection sensitivity.
4.2. Subtyping of Influenza A
In this study, subtyping was performed on 18 influenza A–positive samples, representing the first molecular characterization of circulating human influenza A subtypes in Libya. The A(H1N1)pdm09 subtype predominated, accounting for approximately 72% of cases, while A(H3N2) comprised roughly 27% of cases.
Although previous studies in Libya have reported influenza A detection, most did not include subtype differentiation or were limited to non-human settings, highlighting the novelty of the present findings. The observed subtype distribution is consistent with reports from the Middle East and North Africa (MENA) region [
22], where A(H1N1)pdm09 has predominated in recent seasons, followed by A(H3N2). This pattern is further supported by the World Health Organization, which reported that A(H1N1)pdm09 was the most active influenza A subtype in North Africa between September 2023 and January 2024 [
12]. In contrast, surveillance data from Tunisia during the 2022 influenza season, derived from patients admitted to the medical intensive care unit (MICU), indicated that influenza A(H3N2) was the dominant circulating subtype [
23]. This finding highlights regional variability in influenza subtype circulation and suggests that H3N2 may be more strongly associated with severe disease rather than higher community prevalence, potentially influenced by differences in population immunity, viral evolution, and local epidemiological dynamics.
4.3. Sequencing of H1N1 and H3N2
According to the GISAID platform, multiple genetic clades have circulated globally in recent influenza seasons, including several subclades of 6B.1A for A(H1N1)pdm09 and predominantly 3C.2 and its subclades for A(H3N2). Notably, no human influenza A sequences from Libya were available in GISAID during these periods, highlighting the scarcity of publicly available genomic data and underscoring the importance of the present study. Whole-genome sequencing in this work demonstrated that all Libyan A(H1N1)pdm09 strains belonged to clade 6B.1A.5a.2a, whereas the A(H3N2) strains clustered within clade 3C.2a1b.2a.2a.3a.1, based on hemagglutinin (HA) mutation profiles.
Molecular analysis of the neuraminidase (NA) gene identified multiple amino acid substitutions; however, none corresponded to well-established neuraminidase inhibitor resistance–associated mutations. Recurrent substitutions, including S200N, I264T, and S52N, likely represent natural polymorphisms, and mutations at positions 294–296 differed from the classical resistance-associated N294S substitution [
24]. These findings suggest that the analyzed strains remain susceptible to neuraminidase inhibitors and emphasize the need for continued surveillance.
In addition, amino acid substitutions were detected in several internal proteins, including PB2, MP, and NS. These changes were scattered and recurrent among some isolates, consistent with natural genetic diversity and ongoing viral evolution, and no mutations previously associated with increased virulence or antiviral resistance were identified [
11,
24].
4.4. Phylogenetic Analysis Discussion
The H1N1 phylogeny indicated that Libyan isolates were distributed among multiple clusters within the contemporary global diversity and were closely related to European strains, suggesting repeated introductions from Europe. Phylogeographic analyses further supported viral movement from North America to Europe and subsequently into North Africa, including Libya. The limited genetic divergence among Libyan H1N1 isolates suggests either a single introduction followed by local transmission or multiple introductions of closely related strains, while their interspersion with sequences from neighboring countries highlights active regional connectivity [
12,
15,
22].
In contrast, the H3N2 phylogeny revealed a more restricted pattern, with Libyan isolates clustering within a single lineage that emerged in Europe during the 2021–2022 season and was subsequently introduced into Africa via West Africa before reaching Libya. The low genetic diversity among Libyan H3N2 isolates is consistent with a single or a few introduction events followed by limited local transmission [
12,
15,
22].
Overall, these findings demonstrate heterogeneous dispersal patterns of influenza A subtypes in North Africa, with broader regional circulation of H1N1 compared to the more limited spread of H3N2. This study also highlights the value of genomic surveillance platforms, such as Nextstrain, for reconstructing transmission pathways and contextualizing local strains within global diversity. Although the number of Libyan sequences remains limited, the close genetic relationship between Libyan and European strains suggests that Northern Hemisphere vaccine recommendations are likely applicable to Libya and underscores the need to expand routine genomic surveillance in the region [
12,
17,
22,
23].
5. Conclusions
During the recent influenza seasons, influenza A was the most prevalent among the detected respiratory viruses. Among influenza A–positive samples, the H1N1pdm09 subtype predominated, followed by H3N2, and no other influenza A subtypes were detected. Genetic sequencing of the influenza virus samples revealed no unusual or previously unreported mutations, and the analyzed viral isolates did not exhibit mutations associated with antiviral resistance. Comparative sequence analysis further demonstrated a high degree of genetic similarity between the strains identified in this study and the vaccine strain used in Libya during the study period.
Despite the limited number of analyzed samples, this study represents the first molecular and genetic characterization of human influenza A viruses in Libya and provides baseline data that can serve as a reference for future surveillance and research. The findings underscore the importance of continuous molecular monitoring of influenza viruses in humans and animals to detect emerging variants, track viral evolution, and identify potential antiviral resistance at an early stage.
Author Contributions
Conceptualization, M.A., O.E., and I.E.; methodology, M.A., S.F., O.E., R.O., S.A., A.E., A.K., S.S., H.E., I.B., and I.E.; validation, O.E. and I.E.; investigation, M.A., O.E., and I.E.; resources, A.B., S.S., H.E., and I.E.; data curation, M.A., O.E., A.E., and I.E.; writing—original draft preparation, M.A., S.F., S.A., and I.E.; writing—review and editing, M.A., S.F., O.E., R.O., A.B., S.A., A.E., A.K., S.S., H.E., I.B., and I.E.; supervision, O.E., I.B., and I.E.; project administration, I.E.; funding acquisition, A.B., H.E., and I.E. All authors have read and agreed to the published version of the manuscript.
Funding
Sample collection was performed by sentinel teams and NCDC staff on a voluntary basis. Genetic and sequencing analyses were supported by the WHO Eastern Mediterranean Regional Office (EMRO).
Institutional Review Board Statement
Ethical approval for this study was obtained from the Scientific Research Ethics Committee at the University of Tripoli, Libya (Ref. No. SREC/010/110). Strict confidentiality and anonymity were maintained for all data and samples, and no personal identifiers were associated with the specimens.
Informed Consent Statement
Written informed consent was obtained from all subjects involved in the study, including consent to publish the data presented in this paper.
Data Availability Statement
All data generated or analyzed during this study are included in this manuscript.
Acknowledgments
The authors thank all influenza sentinel surveillance teams across Libyan cities for their dedicated work and assistance in sample collection.
Conflicts of Interest
The authors declare no conflict of interest regarding the publication of this study.
Abbreviations
The following abbreviations are used in this manuscript:
| NCDC |
National Center for Disease Control |
| IAVs |
Influenza A viruses |
| RSV |
respiratory syncytial virus |
| SARS-CoV-2 |
severe acute respiratory syndrome-coronavirus-2 |
| GISAID |
Global Initiative on Sharing All Influenza Data |
| HA |
Hemagglutinin |
| NA |
Neuraminidase |
References
- World Health Organization. Influenza (Seasonal). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal) (accessed on 15 December 2025).
- Centers for Disease Control and Prevention. Types of Influenza Viruses. Available online: https://www.cdc.gov/flu/about/viruses/types.htm (accessed on 25 November 2025).
- Kalil, A.C.; Thomas, P.G. Influenza virus-related critical illness: pathophysiology and epidemiology. Crit. Care 2019, 23, 258. [CrossRef]
- Webster, R.G.; Bean, W.J.; Gorman, O.T.; Chambers, T.M.; Kawaoka, Y. Orthomyxoviridae: The Viruses and Their Replication; Springer: New York, NY, USA, 1992; pp. 1–59.
- Taubenberger, J.K.; Morens, D.M. Influenza: the once and future pandemic. Public Health Rep. 2010, 125 (3_suppl), 15–26. [CrossRef]
- Berche, P. The Spanish flu. La Presse Médicale 2022, 51, 104127.
- Elahmer, O.; Abudher, A.; Bashein, A.; Smeo, M.; Alkhattali, F.; Alkorbaa, A.; Elshweidi, M.; Abubaker, A.; Elkayekh, W.; Zorgani, A.; Ziglam, H. Influenza A/H1N1 in Libya: implementation during pandemic influenza. Abstract presented at the 10th May 2011 Conference, Abstract R2516.
- Dave, K.; Lee, P.C. Global geographical and temporal patterns of seasonal influenza and associated climatic factors. Epidemiol. Rev. 2019, 41, 51–68.
- Tamerius, J.; Shaman, J.; Alonso, W.J.; Bloom-Feshbach, K.; Uejio, C.K.; Comrie, A.; Viboud, C. Environmental predictors of seasonal influenza epidemics across temperate and tropical climates. PLoS Pathog. 2013, 9, e1003194.
- Neumann, G.; Kawaoka, Y. Seasonality of influenza and other respiratory viruses. EMBO Mol. Med. 2022, 14, e15352.
- Fauci, A.S.; Collins, F.S. Benefits and risks of influenza research: lessons learned. Science 2012, 336, 1522–1523.
- World Health Organization. Influenza Update No. 523 [Global respiratory virus activity weekly update]. 23 April 2025. Available online: https://www.who.int/publications/m/item/influenza-update-n--523 (accessed on 15 November 2025).
- Briand, S.; Mounts, A.; Chamberland, M. Challenges of global surveillance during an influenza pandemic. Public Health 2011, 125, 247–256.
- Darvishian, M.; Dijkstra, F.; van Doorn, E.; Bijlsma, M.J.; Donker, G.A.; de Lange, M.M.; Meijer, A. Influenza vaccine effectiveness in the Netherlands from 2003/2004 through 2013/2014: The importance of circulating influenza virus types and subtypes. PLoS ONE 2017, 12, e0169528. [CrossRef]
- Deng, Y.M.; Spirason, N.; Iannello, P.; Jelley, L.; Lau, H.; Barr, I.G. A simplified Sanger sequencing method for full genome sequencing of multiple subtypes of human influenza A viruses. J. Clin. Virol. 2015, 68, 43–48. [CrossRef]
- Granados, A.; Petrich, A.; McGeer, A.; Gubbay, J.B. Measuring influenza RNA quantity after prolonged storage or multiple freeze/thaw cycles. J. Virol. Methods 2017, 247, 45–50. [CrossRef]
- Gardy, J.; Loman, N.J.; Rambaut, A. Real-time digital pathogen surveillance—the time is now. Genome Biol. 2015, 16, 155. [CrossRef]
- Alhudiri, I.M.; Saad, S.R.; Abusrewil, Z.; Amer, A.O.; El Meshri, S.E.; Abdallah, M.N.B.; Elzagheid, A. A preliminary study on the frequency of influenza infections during early 2022 amid the COVID-19 epidemic in Libya. Ibnosina J. Med. Biomed. Sci. 2022, 14, 130–134.
- Soudani, S.; Mafi, A.; Al Mayahi, Z.; Al Balushi, S.; Dbaibo, G.; Al Awaidy, S.; Amiche, A. A systematic review of influenza epidemiology and surveillance in the Eastern Mediterranean and North African region. Infect. Dis. Ther. 2022, 11, 15–52. [CrossRef]
- Xie, Y.; Lin, S.; Zeng, X.; Tang, J.; Cheng, Y.; Huang, W.; Wang, D. Two peaks of seasonal influenza epidemics—China, 2023. China CDC Wkly. 2024, 6, 905.
- Balish, A.L.; Katz, J.M.; Klimov, A.I. Influenza: propagation, quantification, and storage. Curr. Protoc. Microbiol. 2013, 29, 15G.1.
- Caini, S.; El--Guerche Séblain, C.; Ciblak, M.A.; Paget, J. Epidemiology of seasonal influenza in the Middle East and North Africa regions, 2010–2016: circulating influenza A and B viruses and spatial timing of epidemics. Influenza Other Respir. Viruses 2018, 12, 344–352. [CrossRef]
- Boussarsar, M.; Ennouri, E.; Habbachi, N.; Bouguezzi, N.; Meddeb, K.; Gallas, S.; Ben Salah, A. Epidemiology and burden of severe acute respiratory infections (SARI) in the aftermath of the COVID-19 pandemic: a prospective sentinel surveillance study in a Tunisian medical ICU, 2022/2023. PLoS ONE 2023, 18, e0294960. [CrossRef]
- Song, Y.; Zhang, X.; Ji, J.; Li, L.; Zhou, Y.; Ren, G.; Zhu, G. Genetic characteristics analysis of influenza A (H1N1) virus in Jiaxing, China, in the postepidemic era. BMC Infect. Dis. 2025, 25, 905. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |