Submitted:
05 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
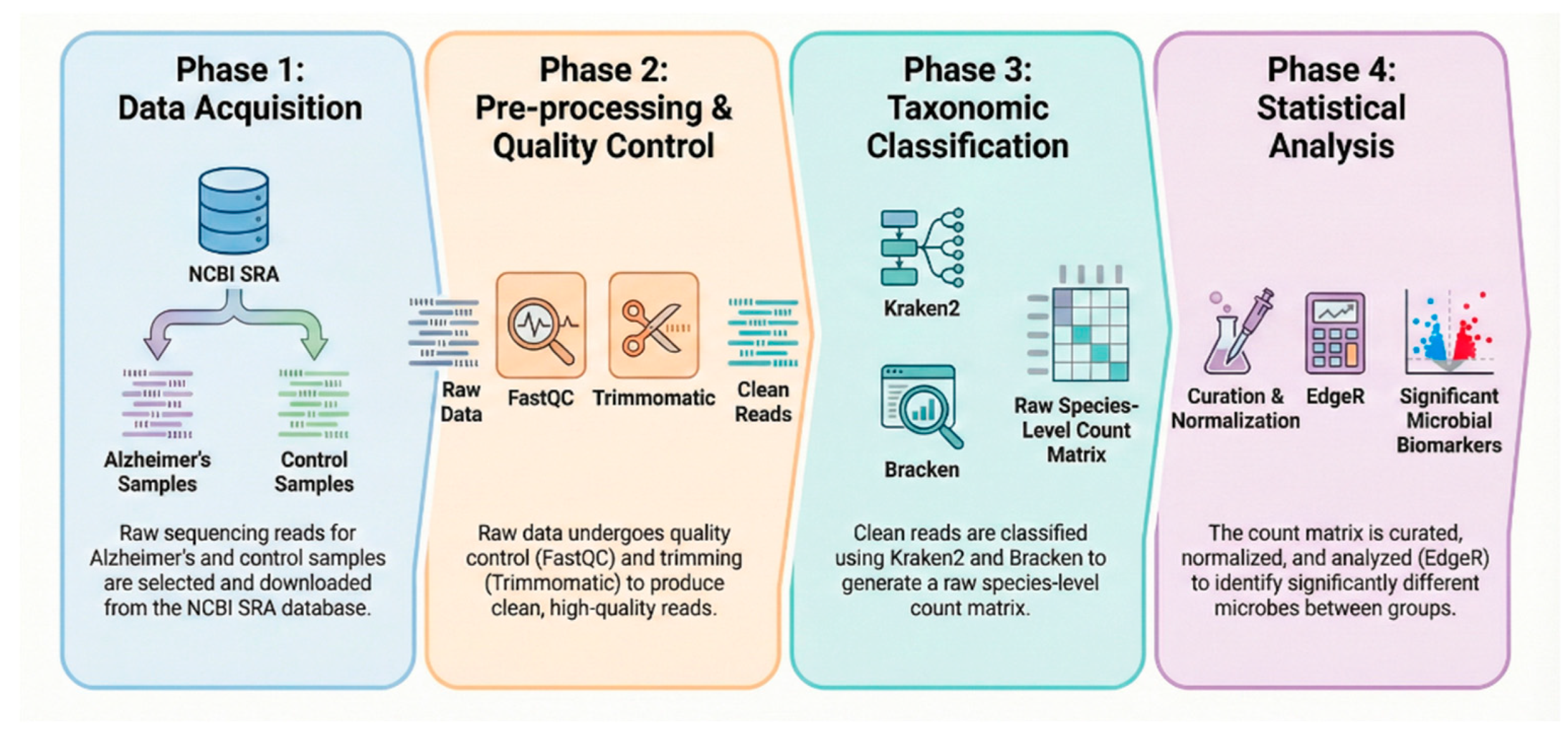
2.2. Bioinformatic Workflow and Data Processing
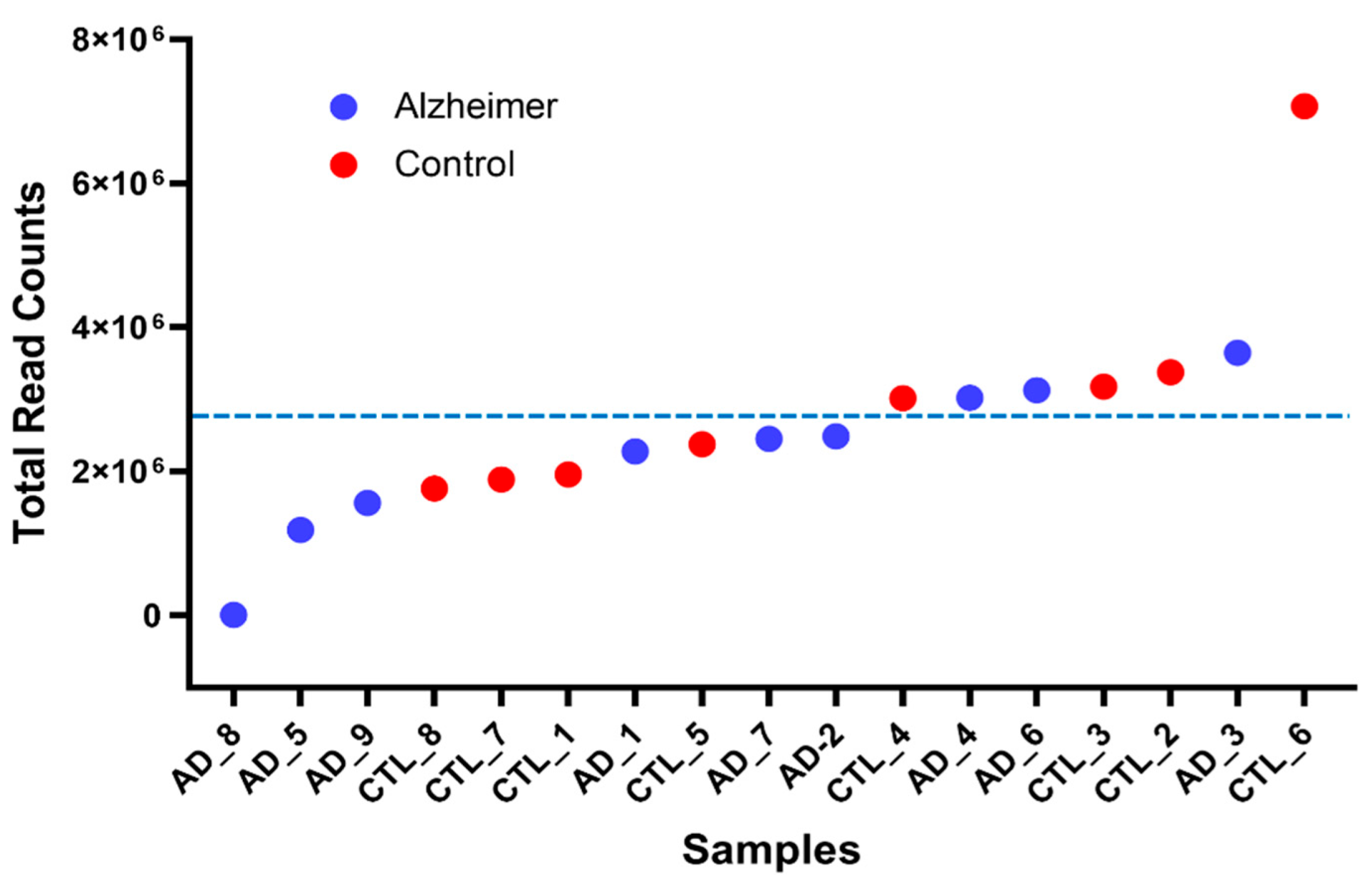
2.3. Sequencing Depth and Library Size Distribution
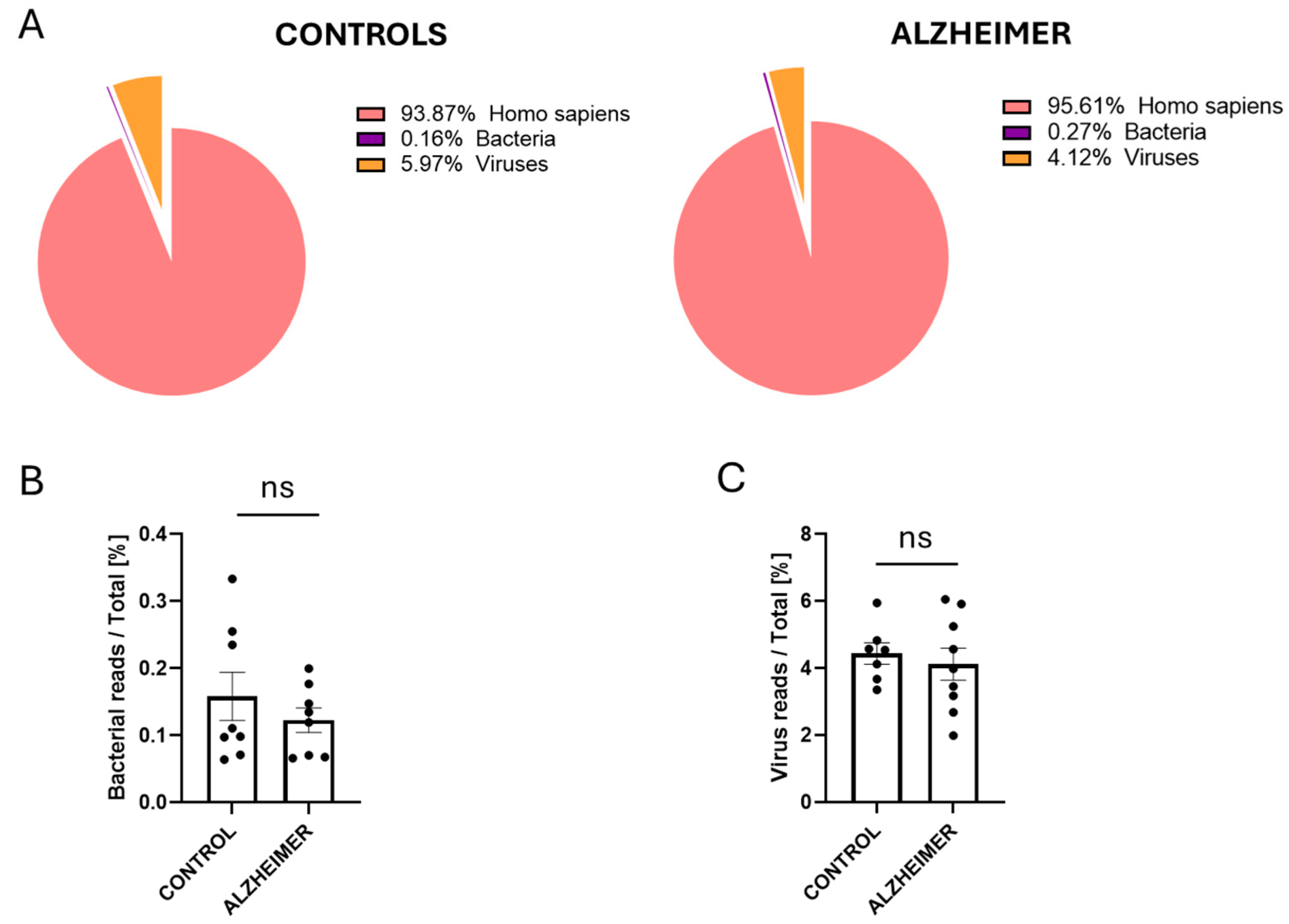
2.4. Global Microbial Composition and Host Contamination
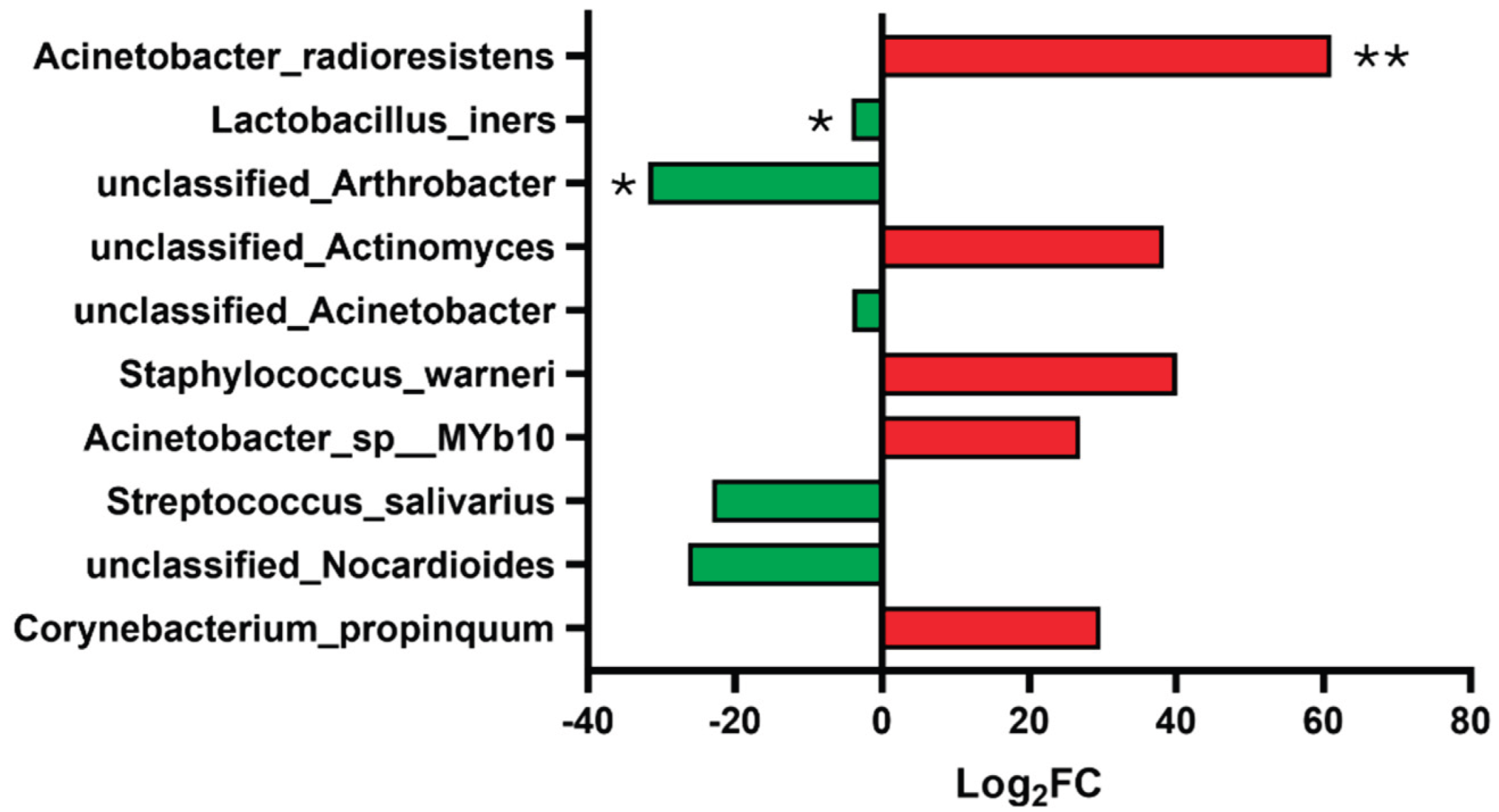
2.5. Differential Abundance Analysis Reveals an Acinetobacter Signature
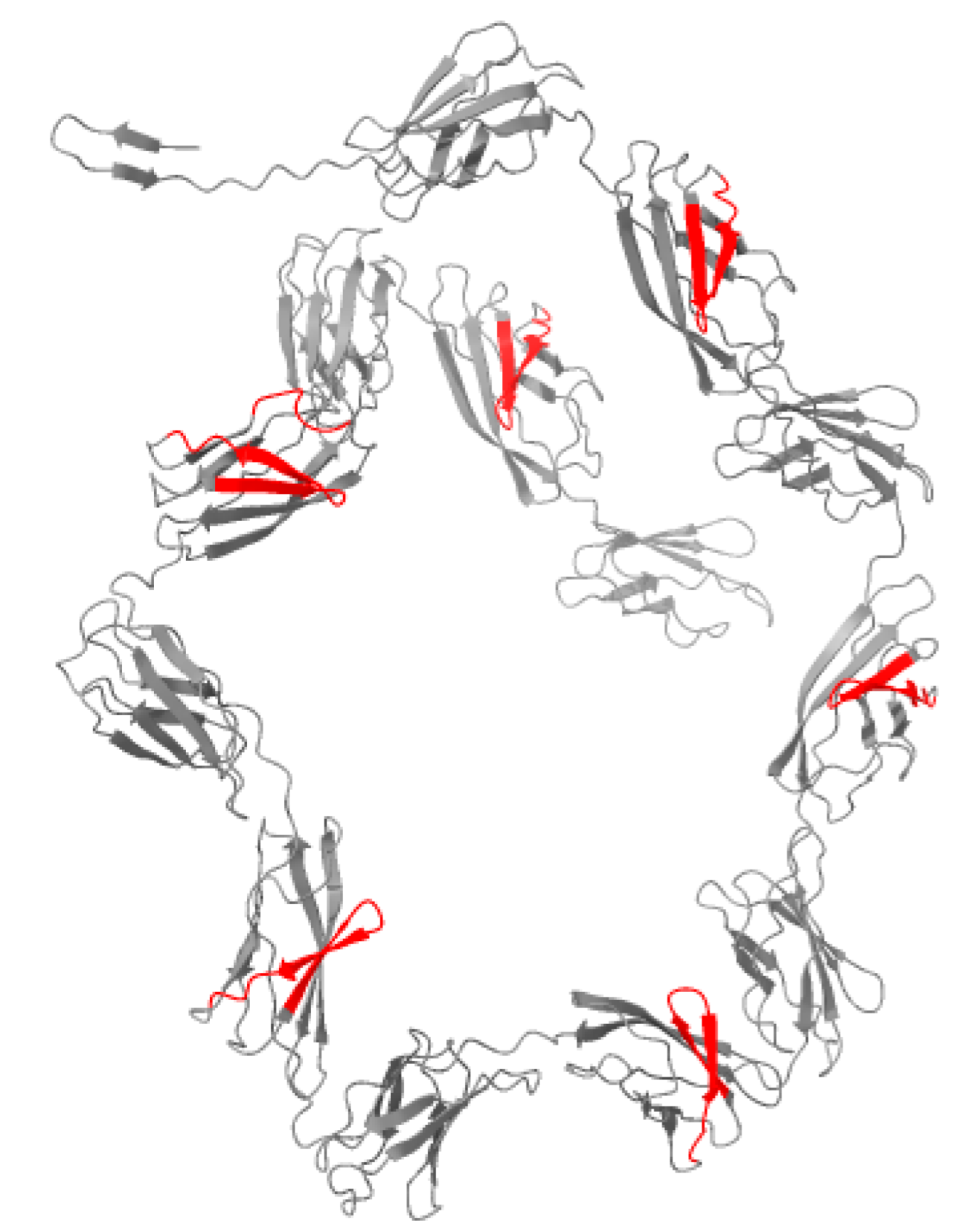
2.6. In Silico Characterization of a Putative Amyloid Cross-Seeding Agent
3. Discussion
4. Materials and Methods
4.1. Data Acquisition and Cohort Selection
4.2. Preprocessing and Quality Control
4.3. Taxonomic Classification
4.4. Data Matrix Generation and Statistical Analysis
4.5. Structural Modeling and Amyloid Prediction
5. Conclusions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| SD | Standard Deviation |
| FC | Fold Change |
| AD | Alzheimer’s disease |
| SEM | Standard Error of the Mean |
References
- Selkoe, D. J.; Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Molecular Medicine 2016, 8(6), 595–608. [Google Scholar]
- van Dyck, C. H.; Swanson, C. J.; Aisen, P.; et al. Lecanemab in early Alzheimer’s disease. New England Journal of Medicine 2023, 388, 9–21. [Google Scholar]
- Sims, JR; Zimmer, JA; Evans, CD; et al. Investigators. Donanemab in Early Symptomatic Alzheimer Disease: The TRAILBLAZER-ALZ 2 Randomized Clinical Trial. JAMA 2023, 330(6), 512–527. [Google Scholar] [CrossRef]
- Kunkle, B. W.; Grenier-Boley, B.; Sims, R.; Bis, J. C.; Damotte, V.; Naj, A. C.; et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nature Genetics 2019, 51, 414–430. [Google Scholar] [CrossRef]
- Piacentini, R; De Chiara, G; Li Puma, DD; et al. HSV-1 and Alzheimer's disease: more than a hypothesis. Front Pharmacol. 2014, 5, 97. [Google Scholar] [CrossRef]
- Readhead, B.; Haure-Mirande, J.-V.; Funk, C. C.; et al. Multiscale analysis of independent Alzheimer’s cohorts finds disruption of molecular, genetic, and clinical networks by human herpesvirus. Neuron 2018, 99(1), 64–82.e7. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, R. Controversial role of herpesviruses in Alzheimer’s disease. PLoS Pathogens 2020, 16(6), e1008575. [Google Scholar] [CrossRef]
- Allnutt, M. A.; Johnson, K.; Bennett, D. A.; Connor, S. M.; Troncoso, J. C.; Pletnikova, O.; Albert, M. S.; De Jager, P. L.; Jacobson, S. Human herpesvirus 6 (HHV-6) detection in Alzheimer’s disease cases and controls across multiple cohorts. Neuron 2020, 105(6), 1027–1035.e2. [Google Scholar] [CrossRef] [PubMed]
- Dominy, SS; Lynch, C; Ermini, F; Benedyk, M; Marczyk, A; Konradi, A; Nguyen, M; Haditsch, U; Raha, D; Griffin, C; Holsinger, LJ; Arastu-Kapur, S; Kaba, S; Lee, A; Ryder, MI; Potempa, B; Mydel, P; Hellvard, A; Adamowicz, K; Hasturk, H; Walker, GD; Reynolds, EC; Faull, RLM; Curtis, MA; Dragunow, M; Potempa, J. Porphyromonas gingivalis in Alzheimer’s disease brains: Evidence for disease causation and treatment with small-molecule inhibitors. Science Advances 2019, 5(1), eaau3333. [Google Scholar] [CrossRef] [PubMed]
- Villar, A; Paladini, S; Cossatis, J. Periodontal disease and Alzheimer’s: Insights from a systematic literature network analysis. Journal of Prevention of Alzheimer’s Disease 2024, 11(4), 1148–1165. [Google Scholar] [CrossRef]
- Sarmiento-Ordóñez, JM; Brito-Samaniego, DR; Vásquez-Palacios, AC; Pacheco-Quito, EM. Association between Porphyromonas gingivalis and Alzheimer’s disease in older adults: A comprehensive review. Infection and Drug Resistance 2025, 18, 2119–2136. [Google Scholar] [CrossRef]
- Soscia, SJ; Kirby, JE; Washicosky, KJ; Tucker, SM; Ingelsson, M; Hyman, B; Burton, MA; Goldstein, LE; Duong, S; Tanzi, RE; Moir, RD. The Alzheimer’s disease-associated amyloid β-protein is an antimicrobial peptide. PLoS ONE 2010, 5(3), e9505. [Google Scholar] [CrossRef]
- Moir, R. D.; Lathe, R.; Tanzi, R. E. The antimicrobial protection hypothesis of Alzheimer’s disease. Alzheimer’s & Dementia 2018, 14(12), 1602–1614. [Google Scholar]
- Eimer, WA; Vijaya Kumar, DK; Navalpur Shanmugam, NK; Rodriguez, AS; Mitchell, T; Washicosky, KJ; György, B; Breakefield, XO; Tanzi, RE; Moir, RD. Alzheimer’s disease-associated β-amyloid is rapidly seeded by Herpesviridae to protect against brain infection. Neuron 2018, 99(1), 56–63.e3. [Google Scholar] [PubMed]
- Friedland, R. P.; Chapman, M. R. The role of microbial amyloid in neurodegeneration. PLoS Pathogens 2017, 13(12), e1006654. [Google Scholar]
- Venegas, C; Kumar, S; Franklin, BS; Dierkes, T; Brinkschulte, R; Tejera, D; Vieira-Saecker, A; Schwartz, S; Santarelli, F; Kummer, MP; Griep, A; Gelpi, E; Beilharz, M; Riedel, D; Golenbock, DT; Geyer, M; Walter, J; Latz, E; Heneka, MT. Microglia-derived ASC specks cross-seed amyloid-β in Alzheimer’s disease. Nature 2017, 552(7685), 355–361. [Google Scholar] [PubMed]
- Taquet, M; Dercon, Q; Todd, JA; Harrison, PJ. The recombinant shingles vaccine is associated with lower risk of dementia. Nature Medicine 2024, 30(10), 2777–2781. [Google Scholar] [CrossRef]
- Eyting, M; Xie, M; Michalik, F; Heß, S; Chung, S; Geldsetzer, P. A natural experiment on the effect of herpes zoster vaccination on dementia. Nature 2025, 641(8062), 438–446. [Google Scholar] [CrossRef] [PubMed]
- Salter, SJ; Cox, MJ; Turek, EM; Calus, ST; Cookson, WO; Moffatt, MF; Turner, P; Parkhill, J; Loman, NJ; Walker, AW. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biology 2014, 12, 87. [Google Scholar] [CrossRef]
- Eisenhofer, R.; Minich, J. J.; Marotz, C.; Cooper, A.; Knight, R.; Weyrich, L. S. Contamination in low microbial biomass microbiome studies: Issues and recommendations. Trends in Microbiology 2019, 27(2), 105–117. [Google Scholar] [CrossRef]
- Fierer, N; Leung, PM; Lappan, R; Eisenhofer, R; Ricci, F; Holland, SI; Dragone, N; Blackall, LL; Dong, X; Dorador, C; Ferrari, BC; Goordial, J; Holmes, SP; Inagaki, F; Korem, T; Li, SS; Makhalanyane, TP; Metcalf, JL; Nagarajan, N; Orsi, WD; Shanahan, ER; Walker, AW; Weyrich, LS; Gilbert, JA; Willis, AD; Callahan, BJ; Shade, A; Parkhill, J; Banfield, JF; Greening, C. Guidelines for preventing and reporting contamination in low-biomass microbiome studies. Nat Microbiol. 2025, 10(7), 1570–1580. [Google Scholar]
- Wang, M; Beckmann, ND; Roussos, P; Wang, E; Zhou, X; Wang, Q; Ming, C; Neff, R; Ma, W; Fullard, JF; Hauberg, ME; Bendl, J; Peters, MA; Logsdon, B; Wang, P; Mahajan, M; Mangravite, LM; Dammer, EB; Duong, DM; Lah, JJ; Seyfried, NT; Levey, AI; Buxbaum, JD; Ehrlich, M; Gandy, S; Katsel, P; Haroutunian, V; Schadt, E; Zhang, B. The Mount Sinai cohort of large-scale genomic, transcriptomic and proteomic data in Alzheimer’s disease. Scientific Data 2018, 5, 180185. [Google Scholar] [PubMed]
- Amabebe, E.; Anumba, D. O. C. The vaginal microenvironment: The physiologic role of Lactobacilli. Frontiers in Medicine 2018, 5, 181. [Google Scholar] [CrossRef] [PubMed]
- Zheng, N.; Guo, R.; Wang, J.; Zhou, W.; Ling, Z. Contribution of Lactobacillus iners to vaginal health and diseases: A systematic review. Frontiers in Cellular and Infection Microbiology 2021, 11, 792787. [Google Scholar] [CrossRef]
- Vaneechoutte, M. Lactobacillus iners, the unusual suspect. Research in Microbiology 2017, 168(9–10), 826–836. [Google Scholar] [CrossRef]
- Suzuki, A; Stern, SA; Bozdagi, O; Huntley, GW; Walker, RH; Magistretti, PJ; Alberini, CM. Astrocyte–neuron lactate transport is required for long-term memory formation. Cell 2011, 144(5), 810–823. [Google Scholar] [CrossRef]
- Yang, J; Ruchti, E; Petit, JM; Jourdain, P; Grenningloh, G; Allaman, I; Magistretti, PJ. Lactate promotes plasticity gene expression by potentiating NMDA signaling in neurons. Proceedings of the National Academy of Sciences USA 2014, 111(33), 12228–12233. [Google Scholar] [CrossRef]
- Barka, E. A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Klenk, H.-P.; Clément, C.; Ouhdouch, Y.; van Wezel, G. P. Taxonomy, physiology, and natural products of Actinobacteria. Microbiology and Molecular Biology Reviews 2015, 80(1), 1–43. [Google Scholar] [CrossRef]
- Ma, Y.; Wang, J.; Liu, Y.; Wang, X.; Zhang, B.; Zhang, W.; Chen, T.; Liu, G.; Xue, L.; Cui, X. Nocardioides: “Specialists” for hard-to-degrade pollutants in the environment. Molecules 2023, 28(21), 7433. [Google Scholar] [CrossRef]
- Hritcu, L.; Stefan, M.; Brandsch, R.; Mihasan, M. 6-hydroxy-L-nicotine from Arthrobacter nicotinovorans sustain spatial memory formation by decreasing brain oxidative stress in rats. Journal of Physiology and Biochemistry 2013, 69(1), 25–34. [Google Scholar] [CrossRef]
- Hritcu, L; Stefan, M; Brandsch, R; Mihasan, M. Enhanced behavioral response by decreasing brain oxidative stress to 6-hydroxy-L-nicotine in Alzheimer’s disease rat model. Neuroscience Letters 2015, 591, 41–47. [Google Scholar] [PubMed]
- Boiangiu, RS; Mihasan, M; Gorgan, DL; Stache, BA; Petre, BA; Hritcu, L. Cotinine and 6-hydroxy-L-nicotine reverse memory deficits and reduce oxidative stress in Aβ25–35-induced rat model of Alzheimer’s disease. Antioxidants 2020, 9(8), 768. [Google Scholar] [CrossRef] [PubMed]
- Montagne, A; Barnes, SR; Sweeney, MD; Halliday, MR; Sagare, AP; Zhao, Z; Toga, AW; Jacobs, RE; Liu, CY; Amezcua, L; Harrington, MG; Chui, HC; Law, M; Zlokovic, BV. Blood–brain barrier breakdown in the aging human hippocampus. Neuron 2015, 85(2), 296–302. [Google Scholar]
- Nation, DA; Sweeney, MD; Montagne, A; Sagare, AP; D'Orazio, LM; Pachicano, M; Sepehrband, F; Nelson, AR; Buennagel, DP; Harrington, MG; Benzinger, TLS; Fagan, AM; Ringman, JM; Schneider, LS; Morris, JC; Chui, HC; Law, M; Toga, AW; Zlokovic, BV. Blood–brain barrier breakdown is an early biomarker of human cognitive dysfunction. Nature Medicine 2019, 25, 270–276. [Google Scholar] [CrossRef]
- De Gregorio, E.; Del Franco, M.; Martinucci, M.; Roscetto, E.; Zarrilli, R.; Di Nocera, P. P. Biofilm-associated proteins: news from Acinetobacter. BMC Genomics 2015, 16, 933. [Google Scholar] [CrossRef] [PubMed]
- Noble, JM; Scarmeas, N; Celenti, RS; Elkind, MS; Wright, CB; Schupf, N; Papapanou, PN; Poole, S.; Singhrao, S. K.; Kesavalu, L.; Curtis, M. A.; Crean, S.; Serum IgG antibody levels to periodontal microbiota are associated with incident Alzheimer disease. Determining the presence of periodontopathic virulence factors in short-term postmortem Alzheimer’s disease brain tissue. PLoS ONE;Journal of Alzheimer’s Disease 2014, 9(12) 36(4), e114959. 50 665–677. [Google Scholar]
- Pereira, AJDSPR; Tavares, AT; Prates, M; Ribeiro, N; Fonseca, LF; Marques, MDR; Proença, F. Brain abscess: A rare clinical case with oral etiology. Case Rep Infect Dis 2022, 2022, 5140259. [Google Scholar]
- Emery, DC; Shoemark, DK; Batstone, TE; Waterfall, CM; Coghill, JA; Cerajewska, TL; Davies, M; West, NX; Allen, SJ. 16S rRNA next generation sequencing analysis shows bacteria in Alzheimer’s post-mortem brain. Frontiers in Aging Neuroscience 2017, 9, 195. [Google Scholar]
- Alonso, R; Pisa, D; Fernández-Fernández, AM; Carrasco, L. Infection of fungi and bacteria in brain tissue from elderly persons and patients with Alzheimer’s disease. Frontiers in Aging Neuroscience 2018, 10, 159. [Google Scholar] [CrossRef]
- Ravaioli, S; De Donno, A; Bottau, G; Campoccia, D; Maso, A; Dolzani, P; Balaji, P; Pegreffi, F; Daglia, M; Arciola, CR. The Opportunistic Pathogen Staphylococcus warneri: Virulence and Antibiotic Resistance, Clinical Features, Association with Orthopedic Implants and Other Medical Devices, and a Glance at Industrial Applications. Antibiotics (Basel) or products referred to in the content. 2024, 13(10), 972. [Google Scholar] [CrossRef]
- Leinonen, R.; Sugawara, H.; Shumway, M. The Sequence Read Archive. Nucleic Acids Research 2011, 39(Database issue), D19–D21. [Google Scholar] [CrossRef]
- Afgan, E; Baker, D; Batut, B; van den Beek, M; Bouvier, D; Cech, M; Chilton, J; Clements, D; Coraor, N; Grüning, BA; Guerler, A; Hillman-Jackson, J; Hiltemann, S; Jalili, V; Rasche, H; Soranzo, N; Goecks, J; Taylor, J; Nekrutenko, A; Blankenberg, D. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Research 2018, 46(W1), W537–W544. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A quality control tool for high throughput sequence data. In Babraham Bioinformatics; 2010. [Google Scholar]
- Bolger, A. M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30(15), 2114–2120. [Google Scholar] [CrossRef]
- Wood, D. E.; Lu, J.; Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biology 2019, 20(1), 257. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Breitwieser, F. P.; Thielen, P.; Salzberg, S. L. Bracken: Estimating species abundance in metagenomics data. PeerJ Computer Science 2017, 3, e104. [Google Scholar] [CrossRef] [PubMed]
- Ondov, B. D.; Bergman, N. H.; Phillippy, A. M. Interactive metagenomic visualization in a Web browser. BMC Bioinformatics 2011, 12, 385. [Google Scholar] [CrossRef]
- Chong, J.; Liu, P.; Zhou, G.; Xia, J. Using MicrobiomeAnalyst for comprehensive statistical, functional, and meta-analysis of microbiome data. Nature Protocols 2020, 15, 799–821. [Google Scholar] [CrossRef]
- Robinson, M. D.; McCarthy, D. J.; Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26(1), 139–140. [Google Scholar] [PubMed]
- Jumper, J; Evans, R; Pritzel, A; Green, T; Figurnov, M; Ronneberger, O; Tunyasuvunakool, K; Bates, R; Žídek, A; Potapenko, A; Bridgland, A; Meyer, C; Kohl, SAA; Ballard, AJ; Cowie, A; Romera-Paredes, B; Nikolov, S; Jain, R; Adler, J; Back, T; Petersen, S; Reiman, D; Clancy, E; Zielinski, M; Steinegger, M; Pacholska, M; Berghammer, T; Bodenstein, S; Silver, D; Vinyals, O; Senior, AW; Kavukcuoglu, K; Kohli, P; Hassabis, D. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596(7873), 583–589. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: making protein folding accessible to all. Nature Methods 2022, 19(6), 679–682. [Google Scholar] [CrossRef]
- Maurer-Stroh, S; Debulpaep, M; Kuemmerer, N; Lopez de la Paz, M; Martins, IC; Reumers, J; Morris, KL; Copland, A; Serpell, L; Serrano, L; Schymkowitz, JW; Rousseau, F. Exploring the sequence determinants of amyloid structure using position-specific scoring matrices. Nature Methods 2010, 7(3), 237–242. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, EF; Goddard, TD; Huang, CC; Meng, EC; Couch, GS; Croll, TI; Morris, JH; Ferrin, TE. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Science 2021, 30(1), 70–82. [Google Scholar] [CrossRef] [PubMed]
| Characteristic |
Alzheimer's Disease (AD) |
Control (CTL) | P-value* |
| Number of Subjects (n) | 9 | 8 | - |
| Age at Death (Mean ± SD) | 84.5 ± 5.2 years | 81.3 ± 4.8 years | > 0.05 (ns) |
| Sex (Female %) | 66.7% (6F / 3M) | 62.5% (5F / 3M) | > 0.05 (ns) |
| Braak Stage (Mean) | 5.4 (Stage V-VI) | 1.4 (Stage I-II) | < 0.001 |
| PMI (Post-Mortem Interval) | ~300 min | ~290 min | > 0.05 (ns) |
| Tissue Region | DLPFC (BM46) | DLPFC (BM46) | - |
| Bacteria | log2FC | Pvalues | FDR |
| Acinetobacter_radioresistens | 61.125 | 5,79E-01 | 0.018 |
| Lactobacillus_iners | -4.161 | 0.0003328 | 0.051 |
| unclassified_Arthrobacter | -31.781 | 0.00075723 | 0.078 |
| unclassified_Actinomyces | 38.251 | 0.0015533 | 0.104 |
| unclassified_Acinetobacter | -4.04 | 0.0016755 | 0.104 |
| Staphylococcus_warneri | 40.077 | 0.0020558 | 0.104 |
| Acinetobacter_sp__MYb10 | 26.921 | 0.002351 | 0.104 |
| Streptococcus_salivarius | -23.078 | 0.0027394 | 0.105 |
| unclassified_Nocardioides | -26.411 | 0.0030489 | 0.105 |
| Corynebacterium_propinquum | 29.678 | 0.0034775 | 0.108 |
| Accession Number | Protein Name | Organism | Sequence Length (aa) |
No. of Amyloidogenic Regions |
Amyloidogenic Region Coordinates |
| XDO94130.1 | Ig-like domain-containing protein | Acinetobacter radioresistens | 3436 | 42 | 135–149; 157–162; 307–321; 329–334; 479–493; 501–506; 651–665; 673–678; 823–837; 845–850; 995–1009; 1017–1022; 1167–1181; 1189–1194; 1339–1353; 1361–1366; 1485–1493; 1571–1578; 1654–1660; 1737–1744; 1839–1847; 1879–1887; 2086–2092; 2169–2175; 2252–2258; 2354–2362; 2394–2402; 2463–2469; 2576–2584; 2603–2609; 2803–2816; 2829–2834; 2837–2845; 2854–2860; 2869–2876; 2950–2958; 2996–3004; 3080–3085; 3113–3118; 3150–3161; 3380–3389; 3431–3436 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




