1. Introduction
Ionizing radiation is widely utilized in cancer radiotherapy, nuclear medicine, and industrial applications, yet radiation-induced toxicity remains a significant dose-limiting factor compromising treatment efficacy and patient quality of life[
1]. The gastrointestinal tract represents one of the most radiosensitive organs, with acute radiation-induced intestinal injury developing within days of exposure, clinically manifesting as severe diarrhea, abdominal pain, and hematochezia[
2,
3]. Beyond therapeutic contexts, accidental radiation exposure scenarios—such as nuclear accidents or radiological terrorism—necessitate effective medical countermeasures against intestinal acute radiation syndrome (IARS), for which no FDA-approved prophylaxis currently exists[
4,
5]. This critical unmet need underscores the importance of identifying molecular targets that govern intestinal epithelial resilience to radiation toxicity.
The pathophysiology of radiation-induced intestinal toxicity involves a complex cascade initiated by direct DNA damage to crypt stem cells, subsequent impairment of epithelial renewal, and loss of mucosal barrier integrity[
6]. Once the epithelial barrier is breached, luminal bacterial translocation triggers robust inflammatory responses that amplify tissue damage, creating a vicious cycle of injury. Intestinal epithelial cells, characterized by lifelong rapid self-renewal, not only mediate nutrient absorption but also constitute the critical barrier preventing microbial invasion and orchestrating mucosal immunity[
7,
8]. Understanding the molecular determinants that preserve epithelial integrity under radiation stress is therefore essential for developing targeted toxicity mitigation strategies.
PIKfyve (phosphoinositide kinase, FYVE-type zinc finger containing) is the sole known kinase catalyzing the phosphorylation of phosphatidylinositol-3-phosphate (PI3P) to phosphatidylinositol-3,5-bisphosphate [PI(3,5)P2], a lipid second messenger critical for endolysosomal homeostasis, vesicular trafficking, and autophagy regulation[
9,
10,
11,
12]. Pharmacological inhibition of PIKfyve has emerged as a promising therapeutic strategy in cancer treatment, with inhibitors such as apilimod demonstrating efficacy in B-cell lymphoma and solid tumors by disrupting lysosomal function and autophagy[
13,
14,
15]. However, the potential impact of PIKfyve modulation on normal tissue radiation toxicity remains completely unexplored—a critical knowledge gap given the concurrent use of radiotherapy and emerging PIKfyve-targeted therapies.
Given PIKfyve’s essential role in maintaining intestinal epithelial homeostasis and its emerging significance in cancer therapeutics, we hypothesized that PIKfyve deficiency would exacerbate radiation-induced intestinal toxicity. To test this hypothesis, we generated an inducible intestinal epithelial-specific PIKfyve-knockout mouse model and subjected these mice to clinically relevant abdominal irradiation. Our findings reveal that PIKfyve serves as a previously unrecognized protective factor against radiation-induced intestinal toxicity, with important implications for radiation countermeasure development and clinical safety assessment of PIKfyve inhibitors in radiotherapy patients.
2. Materials and Methods
Experimental Animals
The PIKfyve intestinal epithelial gene knockout mice were obtained from The Jackson Laboratory16 and subsequently propagated through independent breeding. After breeding and cultivation, mice with the desired genotype were obtained. The induction and irradiation experiments were conducted when the male mice reached an appropriate weight(22–26 g) and were 6-8 weeks old. Animals were housed in SPF barrier facilities (22 ± 2 °C, 50–60% humidity, 12 h light/dark cycle) with ad libitum access to standard chow and water.
Experimental Groups
Mice were randomly assigned to the following groups:
| FIGURE NUMBER |
EXPERIMENT |
GROUP DESIGN |
MICE PER GROUP |
GROUPS |
MICE |
| Figure 1A,B&C |
Histology / BrdU (3.5d) |
WT;cKO |
3 |
2 |
6 |
|
Figure 2A
|
Histology / BrdU (3.5d) |
IR WT;IR cKO |
3 |
2 |
6 |
|
Figure 2D
|
Olfm4 IHC staining |
WT;cKO;IR WT;IR cKO |
3 |
4 |
12 |
|
Figure 2F
|
Survival(13 Gy) |
IR WT;IR cKO |
10 |
2 |
20 |
|
Figure 3A
|
FITC-Dextran permeability(10 Gy) |
WT;cKO;IR WT;IR cKO |
6 |
4 |
24 |
|
Figure 3B
|
RT-PCR |
WT;cKO |
3 |
2 |
6 |
|
Figure 3C
|
RT-PCR |
IR WT;IR cKO |
3 |
2 |
6 |
|
Figure 3D
|
Caspase-3 IHC |
WT;cKO;IR WT;IR cKO(6 h) |
3 |
4 |
12 |
|
Figure 3E
|
TUNEL |
WT;cKO;IR WT;IR cKO(6 h) |
3 |
4 |
12 |
|
Figure 4A
|
Intestine organoid |
WT;cKO
Intestinal crypts were isolated from mouse small intestine for organoid culture and passaging. Subsequent experiments involved direct irradiation of the organoids without the use of mice. |
3 |
2 |
6 |
| TOTAL |
|
|
|
|
110 |
Mouse Irradiation and Treatment
The RS-2000 biological irradiator was used for local irradiation. The settings were as follows: voltage 220 kV, current 25 mA, dose rate 1.325 Gy/min, and irradiation dose 10 Gy. Prior to irradiation, mice were anesthetized with Tribromoethanol injected into the abdominal cavity. Subsequently, mice were immobilized in a custom-made mouse irradiation box, and the chest and lower abdominal regions were shielded with lead plates, exposing only the entire abdominal area. Mice received tamoxifen administration for 5 days (100mg/kg) prior to irradiation, and tamoxifen was also given to each group of mice.
Western Blot Analysis
On the 7th day post-induction, tissues from the small intestine, colon, spleen, and lung of each group of mice were collected for protein extraction. After electrophoresis, proteins were transferred to a nitrocellulose (NC) membrane. The NC membrane was blocked with 5% skimmed milk-TBST blocking solution for 2 hours, followed by overnight incubation with primary antibodies at 4 ℃. After washing off excess primary antibodies with TBST, the NC membrane was incubated with secondary antibodies for 1 hour. After another round of TBST washing to remove excess secondary antibodies, the membrane was subjected to visualization and imaging.
Tissue HE Staining
At day 7 post-induction, mouse intestinal segments were collected, and “Swiss rolls” were prepared. Tissues were fixed in 4% paraformaldehyde for 24 hours. Subsequently, the intestinal tissues were dehydrated, embedded in paraffin, and sectioned along the longitudinal and transverse sections of the intestinal ring and “Swiss rolls” at a thickness of 5 μm. After deparaffinization, HE staining was performed, and the slides were observed under a microscope.
Tissue Immunohistochemical Staining
Tissue sections of mouse intestines were processed similarly to section 1.4. Tissue sections were placed in citrate sodium repair solution, underwent high-temperature repair for 15 minutes, and naturally cooled to room temperature. After blocking endogenous peroxidase activity for 30 minutes at room temperature, tissues were incubated in serum to block tissues for 1 hour at room temperature. Primary antibodies were diluted according to the recommended concentration, incubated with tissues, and placed in a humid chamber at 4 ℃ overnight. After rewarming, excess primary antibodies were washed off with PBS, and tissues were incubated with enzyme-labeled goat anti-mouse/rabbit IgG at room temperature for 1 hour. After washing with PBS, DAB chromogenic solution was applied, and after counterstaining, dehydration, sealing, and drying, slides were observed under a microscope.
FD4 Content Measurement
Mice were fasted and deprived of water for 3 hours on day 3 post-irradiation. Each mouse was orally administered 150 μl of 4 kDa FITC-dextran (referred to as FD4) at a concentration of 80 mg/mL. After 3 hours of administration, 100 μl of blood was taken from the tail vein and incubated at 37 ℃ for 1 hour before transferring to 4 ℃ for 2 hours. After centrifugation at room temperature for 10 minutes at 3000 ×g, 20 μl of the upper serum was taken and diluted with 80 μl of water. FD4 solution and diluted serum were added to a 96-well plate, and absorbance was measured. FD4 content in the serum was calculated to reflect intestinal permeability.
Real-time Quantitative PCR
RNA samples were extracted using Trizol, and after measuring RNA concentration, cDNA was obtained by reverse transcription using the HiScript III All-in-one RT SuperMix Perfect for qPCR kit. Primers were designed using Primer Premier 5.0 software, and their sequences are provided below. RT-QPCR was performed using SYBR Green Pro Taq HS pre-mixed qPCR reagent and the Roche LightCycler® 96 Instrument. The reaction conditions were as follows: pre-incubation at 95 ℃ for 30 s, two-step amplification at 95 ℃ for 10 s, 60 ℃ for 30 s, and 45 cycles; dissolution at 95 ℃ for 10 s, 65 ℃ for 60 s, and 97 ℃ for 1 s. RT-QPCR results were analyzed using the 2-ΔΔCt method.
Table 1.
The sequences of primers for RT-PCR.
Table 1.
The sequences of primers for RT-PCR.
| Gene |
Primer |
Sequence (5′→3′) |
| TNF-α |
F |
CAGGCGGTGCCTATGTCTC |
| R |
CGATCACCCCGAAGTTCAGTAG |
| TGF-β |
F |
GGCCAGATCCTGTCCAAGC |
| R |
GTGGGTTTCCACCATTAGCAC |
| IL-6 |
F |
CTGCAAGAGACTTCCATCCAG |
| R |
AGTGGTATAGACAGGTCTGTTGG |
| CXCL5 |
F |
GTTCCATCTCGCCATTCATGC |
| R |
GCGGCTATGACTGAGGAAGG |
| CCL2 |
F |
TAAAAACCTGGATCGGAACCAAA |
| R |
GCATTAGCTTCAGATTTACGGGT |
IL-1β
|
F |
ATGATGGCTTATTACAGTGGCAA |
| R |
GTCGGAGATTCGTAGCTGGA |
| GAPDH |
F |
ATGGTGAAGGTCGGTGTGAA |
| R |
TGGAAGATGGTGATGGGCTT |
Mouse Small-Intestinal Organoid Experiment
Following the induction protocol, the entire small intestine was harvested from mice. The tissue was opened longitudinally, villi were removed, and the mucosa was washed fifteen times with sterile PBS. Crypts were isolated with EDTA, resuspended in Matrigel, and seeded at 50 µL per well in 24-well plates. Upon Matrigel solidification, mouse small-intestinal organoid medium was added. Cultures were monitored daily, passaged as required, and then irradiated. Post-irradiation growth was recorded, and RNA was extracted for sequencing.
Inclusion / exclusion criteria:
Mice were included only if they met the following pre-established criteria: (i) body weight within 22–26 g, (ii) normal baseline stool consistency, and (iii) completion of the full radiation protocol. Animals showing wound infection, anesthesia-related mortality within 24 h, or technical failure of irradiation were pre-specified for exclusion. No animals or data points were excluded from the final analyses.
Randomization and blinding:
Group allocation was randomized by a computer-generated random sequence stratified by body weight. Cage position on the rack and order of procedures were rotated daily to minimize environmental confounders. Cage location and order of procedures were randomized across groups to eliminate spatial and temporal biases. Investigators performing histological scoring, organoid imaging, and survival checks were blinded to treatment allocation; blinding was broken only after final data lock.
RNA sequencing
Total RNA was extracted from freshly isolated Organoid (C57BL/6 mice) using TRIzol reagent (Invitrogen, USA) according to the manufacturer’s instructions. RNA integrity was assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies, USA), and samples with RNA Integrity Number (RIN) ≥ 7.0 were selected for subsequent analysis.
After RNA quality control, mRNA was enriched from total RNA using HieffNGS™ mRNA Isolation Master Kit (Yeasen, Cat#12603) with oligo(dT)-conjugated magnetic beads. The enriched mRNA was then fragmented into short pieces using fragmentation buffer. First-strand cDNA was synthesized using random hexamer primers, followed by second-strand synthesis with dNTPs, RNase H, and DNA Polymerase I. The resulting double-stranded cDNA was subjected to end repair, A-tailing, and ligation of sequencing adapters. After adapter ligation, the cDNA libraries were amplified by PCR and purified. The final libraries were quantified using a Qubit Fluorometer (Thermo Fisher Scientific) and assessed for fragment size distribution using an Agilent 2100 Bioanalyzer (Agilent Technologies). Libraries that passed quality control were used for sequencing. Paired-end sequencing (150 bp) was performed on the DNBSEQ-T7 platform (BGI, China). On average, approximately 10.27 Gb of clean data were generated per sample.
Statistical Analysis
GraphPad Prism 8.0 software and SPSS Statistics 26.0 software were used for data plotting and statistical analysis, respectively. Data results are expressed as mean ± standard deviation. Statistical comparisons were performed using t-tests, and a significance level of P < 0.05 was considered statistically significant.For multiple group comparisons (e.g.,
Figure 2E and 3A), one-way ANOVA was applied followed by Tukey’s honestly significant difference (HSD) post hoc test for pairwise comparisons. Survival curves were analyzed using the log-rank test.A P value of less than 0.05 was considered statistically significant.
3. Results
3.1. Knockout of PIKfyve in Intestinal Epithelial Cells Had No Significant Impact on the Structural Integrity of the Mouse Intestines
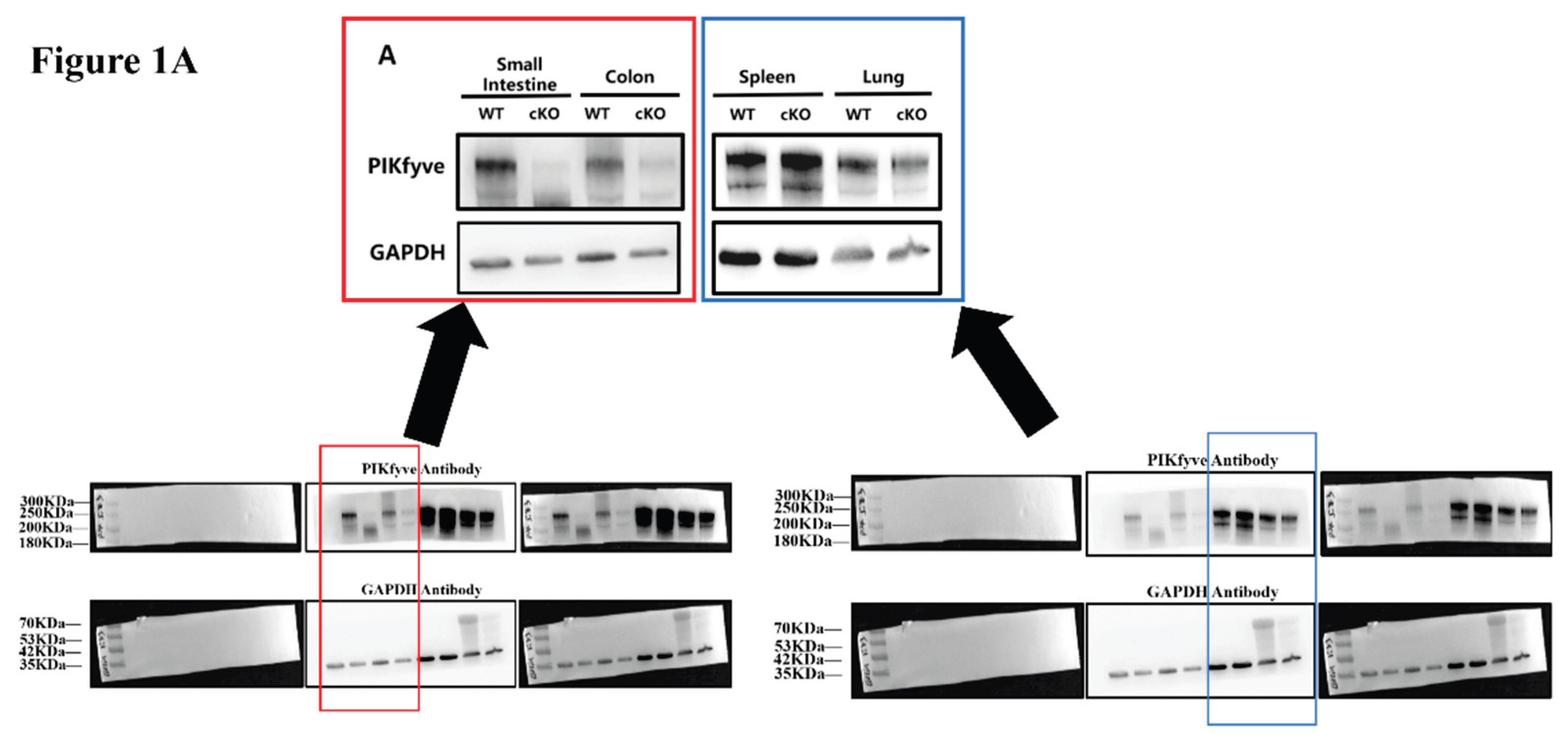
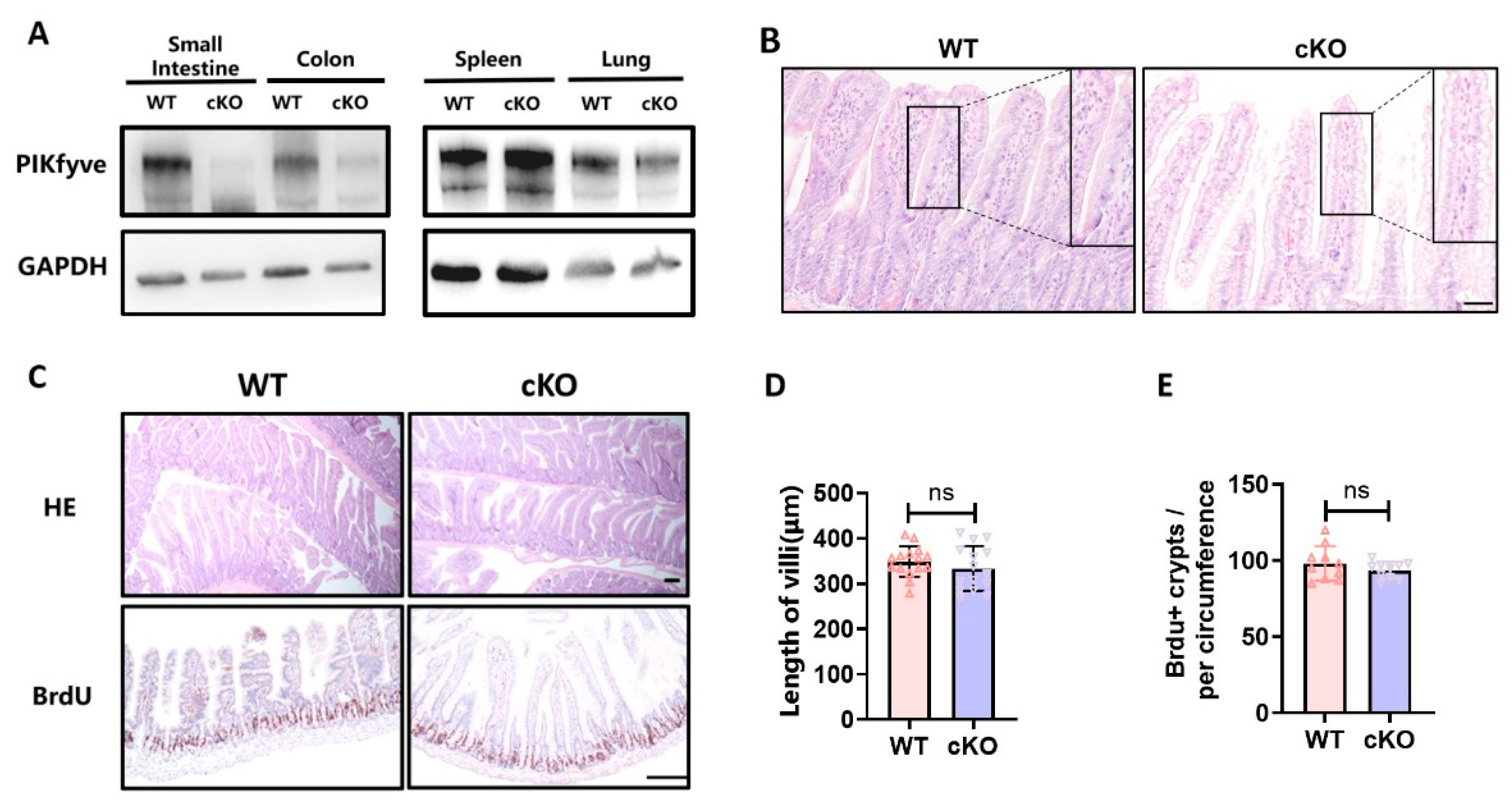
By assessing the levels of PIKfyve protein in tissues, we confirmed the specific deletion of PIKfyve only in the intestine (
Figure 1A). And the deletion of PIKfyve results in the accumulation of vesicles in intestinal epithelium, indicative of significant vacuolization pathology (
Figure 1B). Subsequently, we evaluated the impact of PIKfyve deletion on mouse intestinal villi and crypts. In PIKfyve cKO mice, the arrangement of intestinal villi was compact, and crypts were intact and in a normal proliferative state, showing no significant alterations compared to the control group, except for vacuolization (
Figure 1C-E). These suggest that PIKfyve deletion in intestinal epithelium has no discernible impact on the structure and function of the intestine.
3.2. Deletion of PIKfyve in Intestinal Epithelial Cells Exacerbates Post-Irradiation Intestinal Injury and Impedes the Recovery of Intestinal Stem Cells
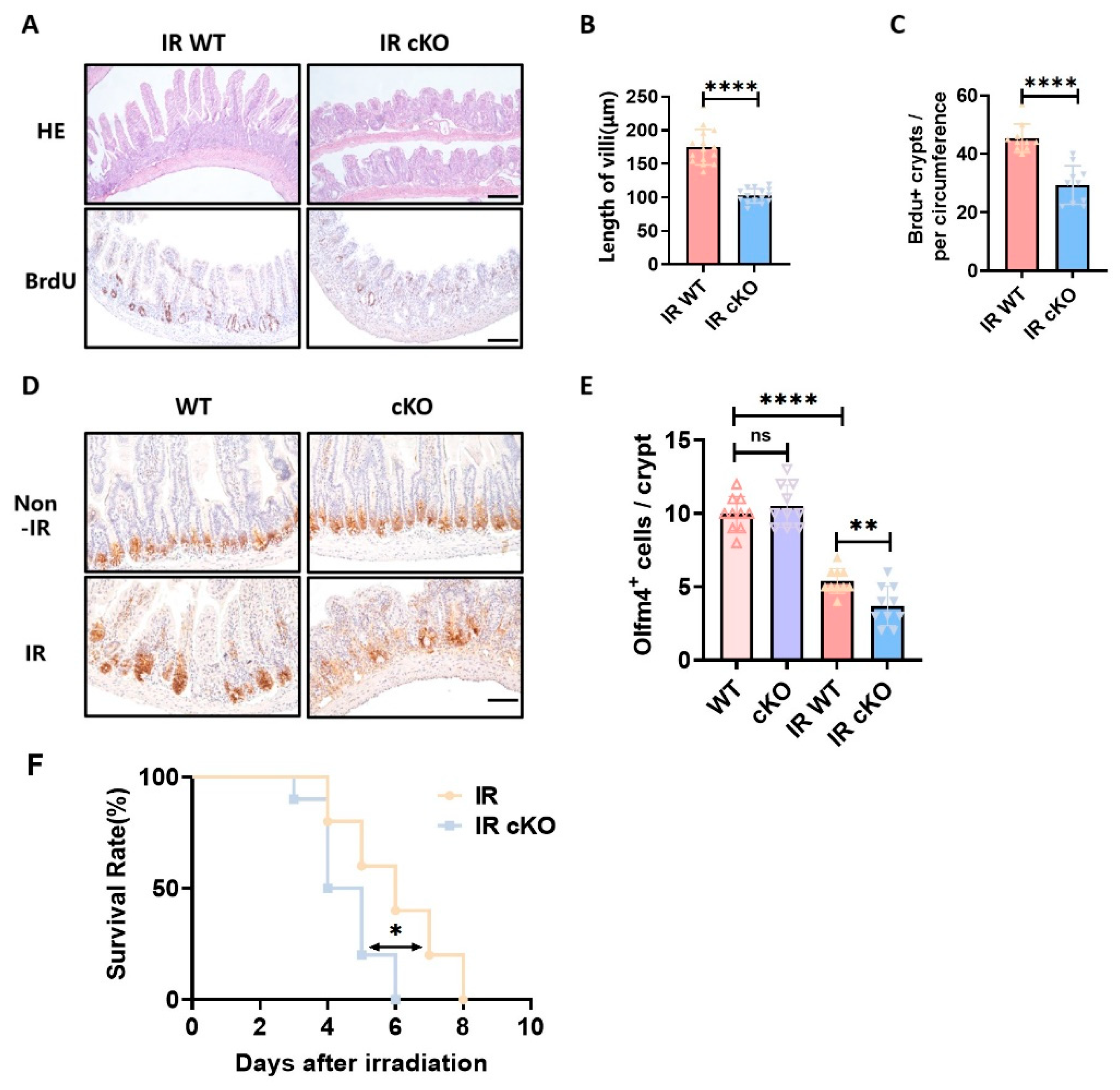
However, following localized abdominal irradiation at 10 Gy, PIKfyve ckO mice exhibited significant villus sloughing and pronounced contraction, along with severe disruption of intestinal epithelial structure compared to the control group (
Figure 2A, B). Notably, in PIKfyve knockout mice, irradiation resulted in compromised crypt morphology and reduced proliferative crypt numbers (
Figure 2A, C), potentially disrupting the normal renewal of intestinal epithelial cells. Staining of intestinal stem cells revealed a significant reduction in stem cell markers within the crypts of PIKfyve cKO mice, which could severely impair normal crypt function(
Figure 2D, E). These results indicate that PIKfyve deletion leads to structural damage in the intestines of irradiated mice, with impaired timely repair processes. PIKfyve-cKO mice exhibit heightened post-irradiation mortality due to a compromised epithelial barrier and severe mucosal disintegration that facilitate bacterial invasion and nutrient loss. (
Figure 2 F) .
3.3. Intestinal Epithelial PIKfyve Deletion Compromises Post-Irradiation Barrier Function in Murine Gut
After radiation-induced damage to the intestine, injury to epithelial and endothelial cells leads to barrier dysfunction[
13], subsequently triggering a severe inflammatory response in the intestine. Intestinal immune response and inflammation are generally considered crucial mechanisms in the development of radiation-induced intestinal damage.
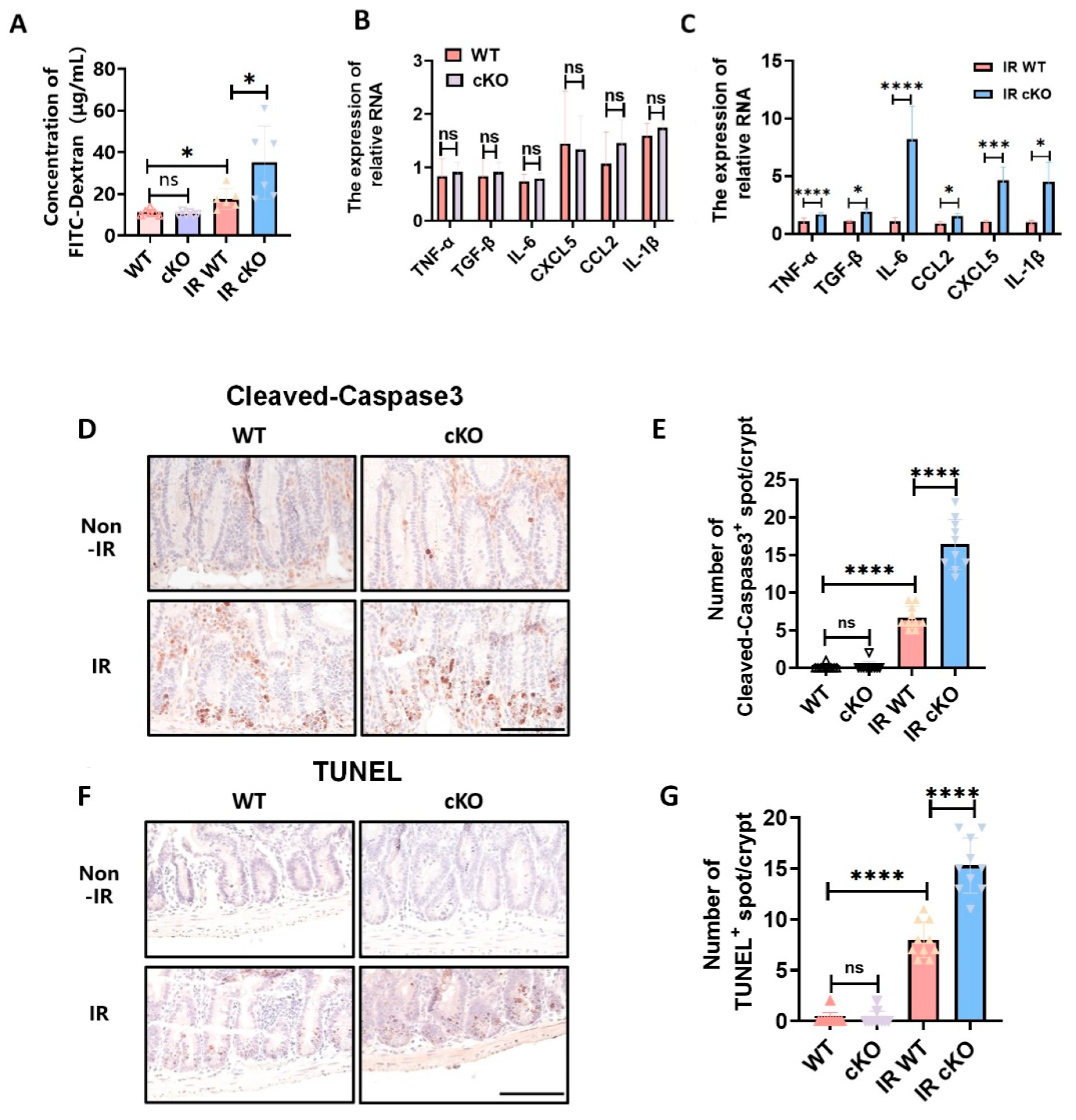
Through the measurement of FITC-dextran levels in peripheral blood of mice, we observed that under steady-state conditions, PIKfyve deletion does not affect intestinal permeability, enabling resistance against bacterial invasion and preventing the onset of severe inflammatory reactions (
Figure 3A, B). After irradiation, PIKfyve-cKO mice exhibit profound mucosal-barrier disruption and elevated intestinal permeability. Consequently, the intestine fails to establish an immune barrier, resulting in severe immune inflammatory responses (
Figure 3 C). These findings confirm that deletion of PIKfyve in intestinal epithelial cells increases radiation sensitivity in mice.
Radiation exposure increases the apoptosis rate of intestinal epithelial cells and crypt cells, and significant crypt apoptosis typically leads to impaired intestinal epithelial regeneration[
14], resulting in compromised function. By assessing changes in apoptosis-related markers at 6 hours post-irradiation, we found that under steady-state conditions, there was no significant impact on mouse intestinal crypt cells after PIKfyve deletion compared to the control group. However, following 10Gy irradiation, there was a significant increase in Cleaved-Caspase3 and TUNEL levels in PIKfyve-/- intestinal crypt cells (
Figure 3 D-G), indicating that PIKfyve deletion increases apoptosis of intestinal crypt cells post-irradiation. This, to some extent, explains the phenomenon of epithelial dysfunction observed after PIKfyve deletion.
3.4. PIKfyve cKO Exacerbates Apoptosis and Inflammatory Responses in Post-Irradiation Murine Enteroids
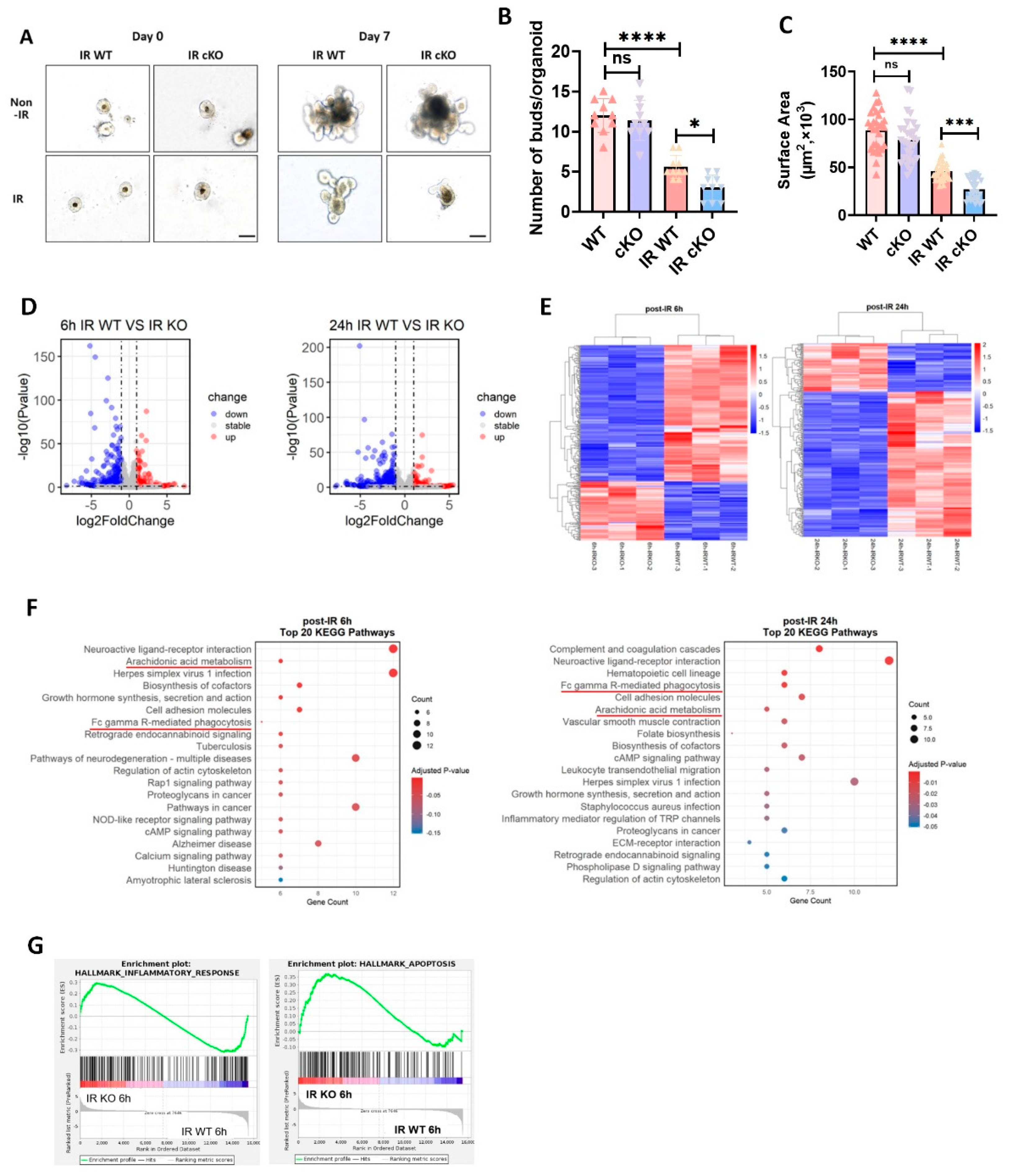
Following the induction protocol, PIKfyvefl/fl;Villin-CreERT2 mice received tamoxifen to ablate PIKfyve specifically in the intestinal epithelium. Intestinal crypts were then isolated and expanded as organoids. Passaged organoids were irradiated with 3 Gy on day 1 and growth was monitored. In unirradiated cultures, both genotypes proliferated normally and generated buds (
Figure 4 A). After irradiation, PIKfyve-deficient organoids displayed reduced expansion and markedly fewer buds, indicating pronounced growth inhibition(
Figure 4 B,C). To define the role of PIKfyve in murine intestinal radiation injury, we performed RNA sequencing on small-intestinal organoids from PIKfyve-knockout and control mice at 6 and 24 h after irradiation. Differential expression was depicted by volcano and heat-map plots (
Figure 4 D,E). At 6 h, 347 genes (e.g., Reg1, Reg3b, Reg3g, Cox6b2) were down-regulated and 154 genes (e.g., Parp3, Alox12b, Alox5ap, Mcoln3, Ace) were up-regulated. By 24 h, 401 genes (e.g., Dmbt1, Ctse, Cdh3) were down-regulated and 152 genes (e.g., Parp3, Rac2, Bmx, Defa2, Defa33) were up-regulated. KEGG enrichment placed Arachidonic acid metabolism and Fc-γ receptor-mediated phagocytosis among the top pathways at 6 h (
Figure 4 F), aligning with the pronounced DNA-damage and inflammatory signatures observed in PIKfyve-deficient organoids[
15,
16]. GSEA enrichment analysis revealed that genes related to inflammatory responses and apoptosis were highly enriched in the irradiated knockout group(
Figure 4 G).These pathways remained within the top 20 at 24 h, indicating that PIKfyve loss intensifies and prolongs inflammation, thereby delaying epithelial repair and amplifying radiation-induced gut damage.In summary, intestinal epithelial PIKfyve deletion amplifies radiation enteropathy by sustaining Alox12b- and Rac2-driven inflammation while repressing Reg1-mediated repair, thereby intensifying crypt apoptosis, compromising barrier integrity, and increasing murine radiosensitivity.
3.5. Figures, Tables and Schemes
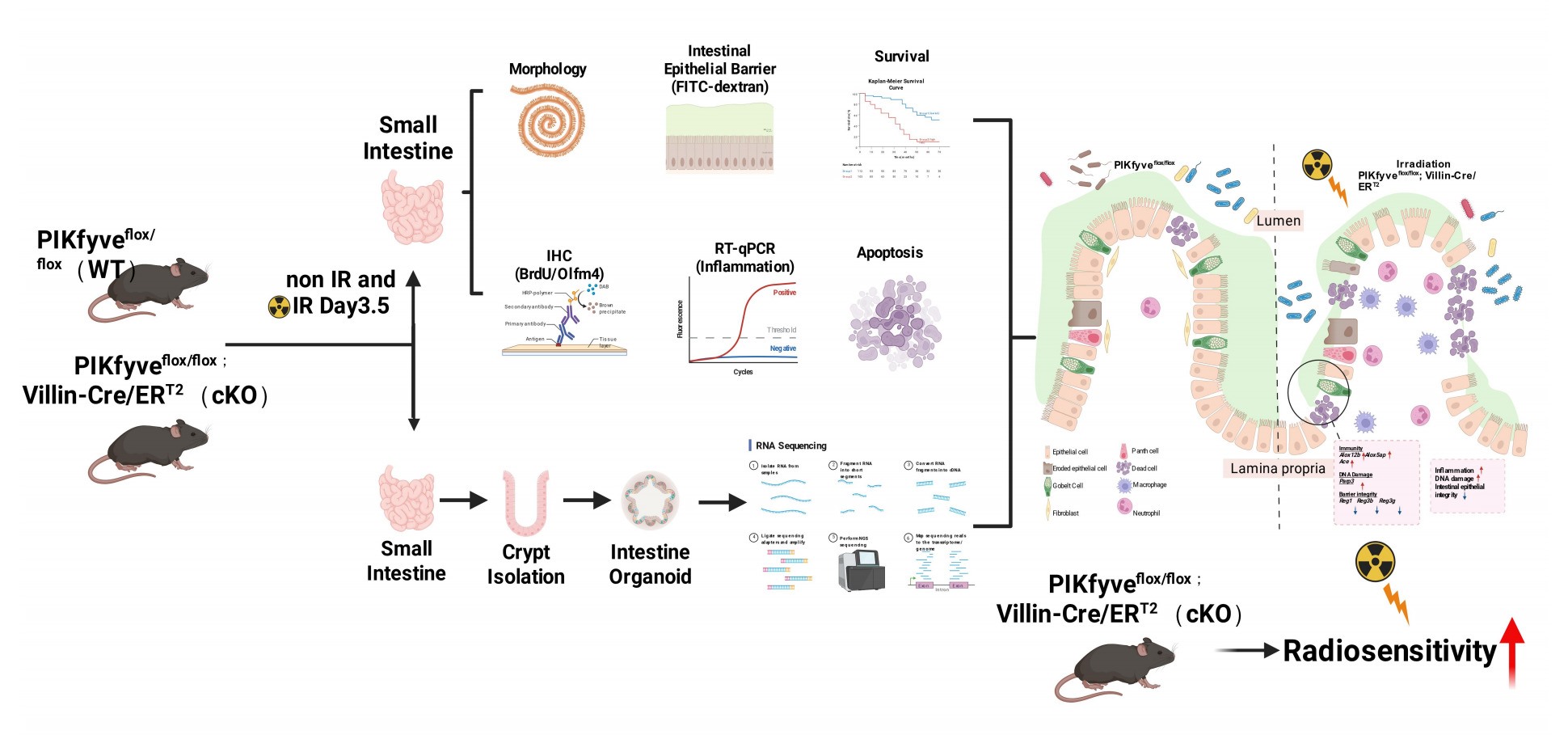
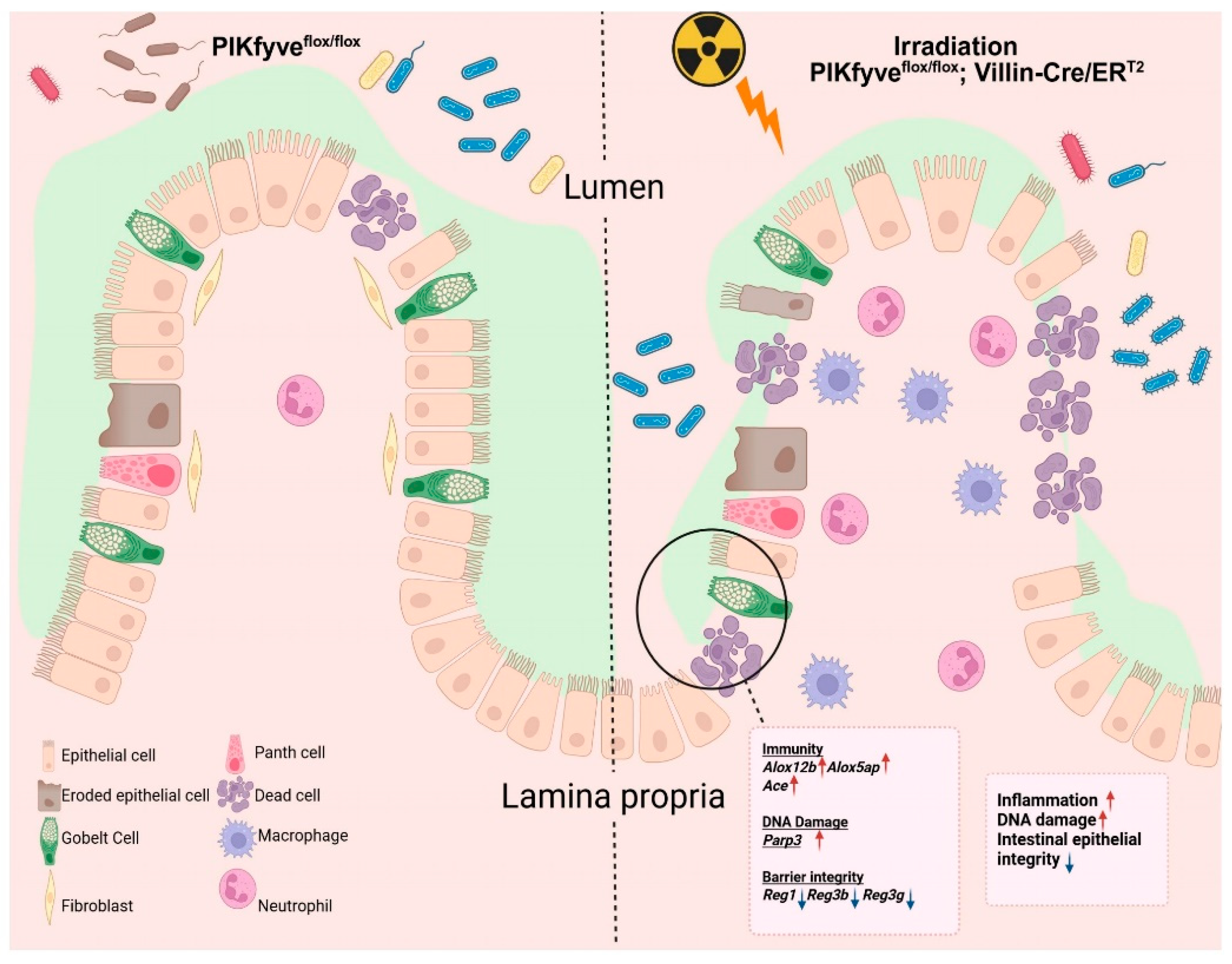
Figure 5.
Model elucidating the impact of intestinal epithelial PIKfyve deletion on the radiosensitivity of the murine small intestine.Under homeostatic conditions, intestinal epithelial lineages coordinate to preserve barrier integrity. Ionizing radiation disrupts this coordination, impairing cellular function and increasing permeability. PIKfyve deletion further compromises DNA repair and epithelial restitution, amplifying structural defects. Luminal microbes and microbial products consequently translocate across the mucosa, inciting a pronounced inflammatory response that augments intestinal radiosensitivity. The principal genes involved are illustrated in the figure.
Figure 5.
Model elucidating the impact of intestinal epithelial PIKfyve deletion on the radiosensitivity of the murine small intestine.Under homeostatic conditions, intestinal epithelial lineages coordinate to preserve barrier integrity. Ionizing radiation disrupts this coordination, impairing cellular function and increasing permeability. PIKfyve deletion further compromises DNA repair and epithelial restitution, amplifying structural defects. Luminal microbes and microbial products consequently translocate across the mucosa, inciting a pronounced inflammatory response that augments intestinal radiosensitivity. The principal genes involved are illustrated in the figure.
4. Discussion
In this study, we identify PIKfyve as a critical guardian of intestinal epithelial integrity against radiation-induced toxicity. Our findings demonstrate that inducible ablation of PIKfyve in intestinal epithelial cells precipitates severe post-irradiation damage characterized by crypt stem cell depletion, exacerbated apoptosis, barrier dysfunction, and lethal inflammation. These results establish PIKfyve-mediated endolysosomal homeostasis as an essential component of the intestinal DNA damage response and position this kinase as a potential target for radiation toxicity mitigation.
Previous studies by Takasuga et al. have elucidated essential roles for PIKfyve in mammalian development and adult intestinal homeostasis using germline and constitutive intestine-specific knockout strategies[
16]. In their landmark study, systemic PIKfyve deficiency resulted in pre-implantation or early embryonic lethality (by E8.5) due to failure of visceral endoderm nutrient transport, whereas constitutive intestinal epithelial deletion (VilCre-driven) caused spontaneous severe malnutrition, bloody stool, diarrhea, and inflammation resembling Crohn’s disease, with 80% mortality by 100 days of age. While these models elegantly demonstrated PIKfyve’s necessity for embryonic development and basal intestinal function, the profound developmental defects and spontaneous pathology preclude their use as platforms to specifically evaluate radiation sensitivity in adult intestines.
To circumvent these developmental confounders and specifically interrogate PIKfyve’s role in radiation response, we employed a tamoxifen-inducible Cre-LoxP system (Villin-Cre^ERT2^) to achieve PIKfyve deletion in adult intestinal epithelium prior to irradiation. This strategy allowed us to isolate PIKfyve’s specific function in DNA damage repair and regenerative responses without the compounding effects of developmental abnormalities or pre-existing intestinal inflammation. Notably, under steady-state conditions, our inducible PIKfyve cKO mice exhibited normal intestinal architecture and barrier function despite vacuolization, indicating that acute loss of PIKfyve in mature intestines is tolerated until challenged by radiation stress. This distinction underscores the utility of our inducible model for dissecting radiation-specific vulnerabilities while avoiding the lethal developmental phenotypes observed in constitutive knockouts.
The intestinal epithelium’s remarkable regenerative capacity depends on Lgr5⁺ crypt base columnar stem cells, which drive continuous epithelial renewal. Olfm4 (Olfactomedin-4) has emerged as a robust and specific marker for these intestinal stem cells, serving as a critical readout for crypt regenerative capacity following genotoxic injury[
14,
17]. In our study, radiation exposure in PIKfyve-deficient intestines resulted in a dramatic reduction of Olfm4⁺ stem cells compared to controls, indicating that PIKfyve is indispensable for maintaining the stem cell pool under radiation stress. This depletion of Olfm4-positive cells likely represents the primary driver of the observed crypt hypoplasia and impaired epithelial restitution, as the loss of these regenerative progenitors compromises the intestine’s ability to replace damaged epithelium, ultimately leading to barrier collapse and lethal sepsis.
Our findings indicate that post-irradiation PIKfyve deletion exacerbates inflammation, perpetuating intestinal damage and exacerbating post-irradiation mortality in mice. These effects may be attributed to increased apoptosis of intestinal crypt cells in PIKfyve knockout mice following irradiation. To dissect the role of PIKfyve in radiation-induced intestinal injury, we derived small-intestinal organoids from PIKfyve-cKO mice. Irradiation markedly impaired their growth. Transcriptomic profiling of organoids harvested at sequential intervals after exposure revealed persistent up-regulation of the DNA-repair gene Parp3, indicative of ongoing DNA damage (Supplement
Figure S2) [
18]. Concurrently, Alox12b and Alox5ap—encoding enzymes that generate leukotrienes and lipoxins—were elevated, signifying an acute inflammatory state[
19]. Enhanced angiotensin-converting enzyme (Ace) expression further promoted vasoconstriction and increased mucosal permeability, amplifying the inflammatory cascade and, consequently, radiosensitivity[
20,
21,
22]. Notably, the regenerative protein Reg1 was down-regulated in PIKfyve-deficient organoids and was almost abolished 24 h after irradiation. Reg1, induced by IL-6/IL-22 via STAT3, stimulates crypt-cell proliferation and fortifies the epithelial barrier by up-regulating claudin-3/4, thereby limiting bacterial translocation[
23,
24,
25]. Loss of PIKfyve blunted this reparative axis, resulting in defective epithelial renewal, heightened permeability, and sustained post-irradiation inflammation that exacerbated intestinal damage.
The translational significance of our findings is underscored by the emerging clinical development of PIKfyve inhibitors as cancer therapeutics. Apilimod, a first-in-class PIKfyve kinase inhibitor, has demonstrated promising efficacy in Phase I/Ⅱ clinical trials for B-cell non-Hodgkin lymphoma, Crohn’s disease and amyotrophic lateral sclerosis[
26,
27,
28]. Additionally, preclinical studies have revealed that PIKfyve inhibition can enhance antitumor immunity by upregulating MHC-I expression and potentiating immune checkpoint blockade in prostate cancer models[
29,
30]. Other PIKfyve-targeting agents currently under investigation include next-generation inhibitors with improved pharmacokinetic properties.
Given the concurrent use of radiotherapy in the management of these malignancies, our results raise critical safety concerns regarding the potential for PIKfyve inhibitors to exacerbate radiation-induced gastrointestinal toxicity. The heightened radiosensitivity observed in our inducible knockout model suggests that patients receiving PIKfyve inhibitors may experience enhanced intestinal toxicity, including severe diarrhea, mucositis, and potentially life-threatening enteritis, when undergoing abdominal or pelvic radiotherapy. This interplay necessitates careful optimization of dosing schedules to temporally separate PIKfyve inhibition from radiation exposure, or the implementation of prophylactic intestinal radioprotective strategies in combined modality protocols. Conversely, our findings also suggest that transient, localized PIKfyve inhibition might sensitize intestinal tumors to radiation while sparing normal tissue, though this therapeutic window requires careful preclinical validation.
5. Conclusions
In summary, our study establishes PIKfyve as a previously unrecognized protective factor against radiation-induced intestinal toxicity. By utilizing an inducible knockout strategy that avoids the developmental lethality of germline deletion, we demonstrate that PIKfyve maintains crypt stem cell viability, limits apoptotic cell death, and preserves barrier integrity following radiation exposure. These findings not only advance our understanding of intestinal radiobiology but also provide crucial guidance for the clinical development of PIKfyve-targeted therapies, emphasizing the need for rigorous assessment of radiation toxicity risks in patients undergoing combined treatment regimens.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org, Figure S1: title; Table S1: title; Video S1: title.
Author Contributions
Aoqiang Ji and Xing Shen conceived and designed the study and performed the primary analysis. Chunan Zhao, Aoqiang Ji, Xing Shen, Zhaopeng Weng, and Xuewen Zhang conducted the experiments. Chunan Zhao and Aoqiang Ji wrote and edited the manuscript. Zuyin Yu, Shuang Xing, and Kai-kai Yu provided professional guidance on experimental details, revised the manuscript, and offered additional professional expertise and support. All authors contributed to the manuscript and approved the final submitted version.
Funding
This research received no external funding.
Ethics Approval
This study was conducted with the approval of the relevant ethics committee. All animal experiments were approved by the Institutional Animal Care and Use Committee (Academy of Military Science). The approved project is titled “PIKfyve Gene Intestinal Epithelium-Specific Knockout Affects Radiation Sensitivity in Mouse Intestine” (IACUC Approval No. IACUC-DWZX-2025-646), issued on 2025.06.19. All protocols adhered with ARRIVE guidelines 2.0.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request. The raw RNA-seq data reported in this study have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject accession number PRJNA1428555.
Acknowledgments
We acknowledge the contributions of our colleagues in the laboratory to this study. Generative artificial intelligence (GenAI) has not been used in this paper.
Conflicts of Interest
The authors declare no competing interests.
Abbreviations
The following abbreviations are used in this manuscript:
| Ace |
angiotensin-converting enzyme |
| Alox12b |
arachidonate lipoxygenase 12B |
| Alox5ap |
arachidonate 5-lipoxygenase-activating protein |
| BrdU |
bromodeoxyuridine |
| CCL2 |
C-C motif chemokine ligand 2 |
| cKO |
conditional knockout |
| CreERT2 |
Cre-estrogen receptor T2 |
| CXCL5 |
C-X-C motif chemokine ligand 5 |
| DAB |
3,3’-diaminobenzidine |
| EDTA |
ethylenediaminetetraacetic acid |
| FD4 |
fluorescein isothiocyanate-dextran (4 kDa) |
| Fig |
figure |
| GAPDH |
glyceraldehyde-3-phosphate dehydrogenase |
| GSEA |
Gene Set Enrichment Analysis |
| Gy |
Gray |
| HE |
hematoxylin and eosin |
| IARS |
intestinal acute radiation syndrome |
| IL |
interleukin |
| KEGG |
Kyoto Encyclopedia of Genes and Genomes |
| Lgr5 |
leucine-rich repeat-containing G-protein coupled receptor 5 |
| MHC-I |
major histocompatibility complex class I |
| NC |
nitrocellulose |
| ns |
not significant |
| Olfm4 |
olfactomedin-4 |
| PARP3 |
poly(ADP-ribose) polymerase 3 |
| PBS |
phosphate-buffered saline |
| PI(3,5)P2 |
phosphatidylinositol-3,5-bisphosphate |
| PI3P |
phosphatidylinositol-3-phosphate |
| PIKfyve |
phosphoinositide kinase, FYVE-type zinc finger containing |
| qPCR |
quantitative polymerase chain reaction |
| Reg |
regenerating gene |
| RIN |
RNA integrity number |
| RT-qPCR |
reverse transcription quantitative PCR |
| SPF |
specific pathogen-free |
| STAT3 |
signal transducer and activator of transcription 3 |
| TBST |
Tris-buffered saline with Tween-20 |
| TGF-β |
transforming growth factor-beta |
| TNF-α |
tumor necrosis factor-alpha |
| TUNEL |
terminal deoxynucleotidyl transferase dUTP nick end labeling |
| WT |
wild type |
| μL |
microliter |
| μm |
micrometer |
References
- Winters, T.A.; Marzella, L.; Molinar-Inglis, O.; Price, P.W.; Han, N.C.; Cohen, J.E.; Wang, S.J.; Fotenos, A.F.; Sullivan, J.M.; Esker, J.I.; et al. Gastrointestinal Acute Radiation Syndrome: Mechanisms, Models, Markers, and Medical Countermeasures. Radiation research 2024, 201, 628–646. [Google Scholar] [CrossRef]
- Gong, W.; Guo, M.; Han, Z.; Wang, Y.; Yang, P.; Xu, C.; Wang, Q.; Du, L.; Li, Q.; Zhao, H.; et al. Mesenchymal stem cells stimulate intestinal stem cells to repair radiation-induced intestinal injury. Cell death & disease 2016, 7, e2387–e2387. [Google Scholar] [CrossRef]
- Singh, V.K.; Seed, T.M. A review of radiation countermeasures focusing on injury-specific medicinals and regulatory approval status: part I. Radiation sub-syndromes, animal models and FDA-approved countermeasures. International journal of radiation biology 2017, 93, 851–869. [Google Scholar] [CrossRef]
- Kenchegowda, D.; Bolduc, D.L.; Kurada, L.; Blakely, W.F. Severity scoring systems for radiation-induced GI injury – prioritization for use of GI-ARS medical countermeasures. International journal of radiation biology 2023, 99, 1037–1045. [Google Scholar] [CrossRef] [PubMed]
- Obrador, E.; Salvador-Palmer, R.; Villaescusa, J.I.; Gallego, E.; Pellicer, B.; Estrela, J.M.; Montoro, A. Nuclear and Radiological Emergencies: Biological Effects, Countermeasures and Biodosimetry. Antioxidants 2022, 11, 1098. [Google Scholar] [CrossRef]
- Hua, G.; Thin, T.H.; Feldman, R.; Haimovitz–Friedman, A.; Clevers, H.; Fuks, Z.; Kolesnick, R. Crypt Base Columnar Stem Cells in Small Intestines of Mice Are Radioresistant. Gastroenterology 2012, 143, 1266–1276. [Google Scholar] [CrossRef]
- Jian, Y.; Zhang, D.; Liu, M.; Wang, Y.; Xu, Z.-X. The Impact of Gut Microbiota on Radiation-Induced Enteritis. 2021, 11–2021. [Google Scholar] [CrossRef]
- Haber, A.L.; Biton, M.; Rogel, N.; Herbst, R.H.; Shekhar, K.; Smillie, C.; Burgin, G.; Delorey, T.M.; Howitt, M.R.; Katz, Y.; et al. A single-cell survey of the small intestinal epithelium. Nature 2017, 551, 333–339. [Google Scholar] [CrossRef]
- Duex, J.E.; Tang, F.; Weisman, L.S. The Vac14p–Fig4p complex acts independently of Vac7p and couples PI3,5P2 synthesis and turnover. Journal of Cell Biology 2006, 172, 693–704. [Google Scholar] [CrossRef]
- Rusten, T.E.; Rodahl, L.M.W.; Pattni, K.; Englund, C.; Samakovlis, C.; Dove, S.; Brech, A.; Stenmark, H. Fab1 Phosphatidylinositol 3-Phosphate 5-Kinase Controls Trafficking but Not Silencing of Endocytosed Receptors. Molecular Biology of the Cell 2006, 17, 3989–4001. [Google Scholar] [CrossRef] [PubMed]
- Kutchukian, C.; Casas, M.; Dixon, R.E.; Dickson, E.J. Disruption of the PIKfyve complex unveils an adaptive mechanism to promote lysosomal repair and mitochondrial homeostasis. Nature communications 2025, 16, 10761. [Google Scholar] [CrossRef] [PubMed]
- Sbrissa, D.; Ikonomov, O.C.; Shisheva, A. PIKfyve, a mammalian ortholog of yeast Fab1p lipid kinase, synthesizes 5-phosphoinositides. Effect of insulin. The Journal of biological chemistry 1999, 274, 21589–21597. [Google Scholar] [CrossRef]
- Gayle, S.; Landrette, S.; Beeharry, N.; Conrad, C.; Hernandez, M.; Beckett, P.; Ferguson, S.M.; Mandelkern, T.; Zheng, M.; Xu, T.; et al. Identification of apilimod as a first-in-class PIKfyve kinase inhibitor for treatment of B-cell non-Hodgkin lymphoma. Blood 2017, 129, 1768–1778. [Google Scholar] [CrossRef]
- Wible, D.J.; Parikh, Z.; Cho, E.J.; Chen, M.-D.; Jeter, C.R.; Mukhopadhyay, S.; Dalby, K.N.; Varadarajan, S.; Bratton, S.B. Unexpected inhibition of the lipid kinase PIKfyve reveals an epistatic role for p38 MAPKs in endolysosomal fission and volume control. Cell death & disease 2024, 15, 80. [Google Scholar] [CrossRef]
- Ikonomov, O.C.; Sbrissa, D.; Shisheva, A. Small molecule PIKfyve inhibitors as cancer therapeutics: Translational promises and limitations. Toxicology and applied pharmacology 2019, 383, 114771. [Google Scholar] [CrossRef] [PubMed]
- Takasuga, S.; Horie, Y.; Sasaki, J.; Sun-Wada, G.H.; Kawamura, N.; Iizuka, R.; Mizuno, K.; Eguchi, S.; Kofuji, S.; Kimura, H.; et al. Critical roles of type III phosphatidylinositol phosphate kinase in murine embryonic visceral endoderm and adult intestine. Proceedings of the National Academy of Sciences of the United States of America 2013, 110, 1726–1731. [Google Scholar] [CrossRef] [PubMed]
- van der Flier, L.G.; Haegebarth, A.; Stange, D.E.; van de Wetering, M.; Clevers, H. OLFM4 is a robust marker for stem cells in human intestine and marks a subset of colorectal cancer cells. Gastroenterology 2009, 137, 15–17. [Google Scholar] [CrossRef] [PubMed]
- Fenton, A.L.; Shirodkar, P.; Macrae, C.J.; Meng, L.; Koch, C.A. The PARP3- and ATM-dependent phosphorylation of APLF facilitates DNA double-strand break repair. Nucleic acids research 2013, 41, 4080–4092. [Google Scholar] [CrossRef]
- Amoah, A.-S.; Pestov, N.B.; Korneenko, T.V.; Prokhorenko, I.A.; Kurakin, G.F.; Barlev, N.A. Lipoxygenases at the Intersection of Infection and Carcinogenesis. 2024, 25, 3961. [Google Scholar] [CrossRef]
- Garg, M.; Royce, S.G.; Tikellis, C.; Shallue, C.; Batu, D.; Velkoska, E.; Burrell, L.M.; Patel, S.K.; Beswick, L.; Jackson, A.; et al. Imbalance of the renin-angiotensin system may contribute to inflammation and fibrosis in IBD: a novel therapeutic target? Gut 2020, 69, 841–851. [Google Scholar] [CrossRef]
- He, L.; Du, J.; Chen, Y.; Liu, C.; Zhou, M.; Adhikari, S.; Rubin, D.T.; Pekow, J.; Li, Y.C. Renin-angiotensin system promotes colonic inflammation by inducing T(H)17 activation via JAK2/STAT pathway. American journal of physiology. Gastrointestinal and liver physiology 2019, 316, G774–g784. [Google Scholar] [CrossRef]
- Fan, J.-F.; Wang, Y.-K.; Liu, M.; Liu, G.-S.; Min, T.-J.; Chen, R.-Y.; He, Y. Effect of angiotensin II on irradiation exacerbated decompression sickness. Scientific reports 2023, 13, 11659. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, C.; Sakuramoto-Tsuchida, S.; Taked, M.; Itaya-Hironaka, A.; Yamauchi, A.; Misu, M.; Shobatake, R.; Uchiyama, T.; Makino, M.; Pujol-Autonell, I.; et al. Expression of REG family genes in human inflammatory bowel diseases and its regulation. Biochemistry and biophysics reports 2017, 12, 198–205. [Google Scholar] [CrossRef]
- Kitayama, Y.; Fukui, H.; Hara, K.; Eda, H.; Kodani, M.; Yang, M.; Sun, C.; Yamagishi, H.; Tomita, T.; Oshima, T.; et al. Role of regenerating gene I in claudin expression and barrier function in the small intestine. Translational research: the journal of laboratory and clinical medicine 2016, 173, 92–100. [Google Scholar] [CrossRef]
- Mao, H.; Jia, J.; Sheng, J.; Zhang, S.; Huang, K.; Li, H.; He, F. Protective and anti-inflammatory role of REG1A in inflammatory bowel disease induced by JAK/STAT3 signaling axis. International immunopharmacology 2021, 92, 107304. [Google Scholar] [CrossRef] [PubMed]
- Harb, W.A.; Diefenbach, C.S.; Lakhani, N.; Rutherford, S.C.; Schreeder, M.T.; Ansell, S.M.; Sher, T.; Aboulafia, D.M.; Cohen, J.B.; Nix, D.; et al. Phase 1 Clinical Safety, Pharmacokinetics (PK), and Activity of Apilimod Dimesylate (LAM-002A), a First-in-Class Inhibitor of Phosphatidylinositol-3-Phosphate 5-Kinase (PIKfyve), in Patients with Relapsed or Refractory B-Cell Malignancies. Blood 2017, 130, 4119–4119. [Google Scholar] [CrossRef]
- Sands, B.E.; Jacobson, E.W.; Sylwestrowicz, T.; Younes, Z.; Dryden, G.; Fedorak, R.; Greenbloom, S. Randomized, double-blind, placebo-controlled trial of the oral interleukin-12/23 inhibitor apilimod mesylate for treatment of active Crohn’s disease. Inflammatory bowel diseases 2010, 16, 1209–1218. [Google Scholar] [CrossRef]
- Babu, S.; Nicholson, K.A.; Rothstein, J.D.; Swenson, A.; Sampognaro, P.J.; Pant, P.; Macklin, E.A.; Spruill, S.; Paganoni, S.; Gendron, T.F.; et al. Apilimod dimesylate in C9orf72 amyotrophic lateral sclerosis: a randomized phase 2a clinical trial. Brain: a journal of neurology 2024, 147, 2998–3008. [Google Scholar] [CrossRef]
- Bao, Y.; Qiao, Y.; Choi, J.E.; Zhang, Y.; Mannan, R.; Cheng, C.; He, T.; Zheng, Y.; Yu, J.; Gondal, M.; et al. Targeting the lipid kinase PIKfyve upregulates surface expression of MHC class I to augment cancer immunotherapy. Proceedings of the National Academy of Sciences of the United States of America 2023, 120, e2314416120. [Google Scholar] [CrossRef]
- Qiao, Y.; Choi, J.E.; Tien, J.C.; Simko, S.A.; Rajendiran, T.; Vo, J.N.; Delekta, A.D.; Wang, L.; Xiao, L.; Hodge, N.B.; et al. Autophagy Inhibition by Targeting PIKfyve Potentiates Response to Immune Checkpoint Blockade in Prostate Cancer. Nature cancer 2021, 2, 978–993. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).