1. Introduction
Many children under five years old have health problems caused by pneumonia symptoms, and this becomes a major concern in many countries [
1,
2]. Pneumonia is a respiratory disease that directly affects the lungs and greatly affects the health of the body by disrupting oxygen exchange in the body [
3]. Based on the World Health Organization (WHO), pediatric pneumonia contributes 14% of deaths in children under five years old, especially in regions such as South Asia and Sub-Saharan Africa, and causes an estimated 740,000 fatalities annually [
4]. Pneumonia can be caused by poor environmental conditions and generate many bacteria, viruses, and fungi that can infect the human body, especially in children [
5]. From these situations, this disease requires proper treatment and a strategy to prevent and overcome it. Currently, there are many medical personnel trying to find the proper and accurate way to make an early diagnosis of pneumonia in children, so that it can reduce the rapid spread of infection and its complications, and can even reduce the death rate in children [
6,
7]. If early detection and diagnosis can be done, it will also make it easier to treat patients and provide the right therapy.
Artificial intelligence (AI) is rapidly transforming various sectors, and its application in medicine is particularly promising for enhancing disease detection and classification accuracy and efficiency [
8]. AI can analyze complex datasets and identify subtle patterns, revolutionizing medical imaging, diagnostics, and treatment planning, paving the way for improved patient outcomes [
9]. The application of AI, especially through the implementation of machine learning and deep learning in healthcare, involves utilizing computer algorithms to extract relevant data and knowledge and aid clinical decision-making, which has seen rapid development in many developed countries [
10,
11]. The study creates fundamental knowledge for AI healthcare solutions, yet actively promotes pediatric pneumonia detection systems that deliver efficient and accessible diagnostic capabilities. One of the latest AI algorithms that can provide promising results in pediatric pneumonia classification is by using the Deep Learning approach.
The ability of deep learning models to learn intricate, problem-specific features from medical images has led to a paradigm shift in computer vision applications within healthcare. Deep learning (DL) has revolutionized medical image analysis, offering unprecedented accuracy in diagnosing various diseases, including pediatric pneumonia [
12]. Early and accurate diagnosis of pneumonia classification is crucial, particularly in pediatric cases, where the condition can rapidly progress and lead to severe complications. This research focuses on the development of an ensemble deep learning framework to classify pediatric pneumonia by utilizing transfer learning and convolutional neural network algorithms. The ensemble deep learning approach is one of the promising algorithms and is also considered the latest DL algorithm that can achieve optimal accuracy results in classification implementation [
13].
This research experiment used the Chest X-Ray Images (Pneumonia) dataset, which contains 5863 high-resolution anterior-posterior (AP) chest radiographs sampled from children aged 1 to 5 years old. There are four activities in the data preprocessing phase, namely image resizing, intensity normalization, label encoding, and data structure optimization. In our framework, we combine three current CNNs, namely MobileNetV2, ResNet50, and EfficientNetB0, for feature-level ensemble implementation to provide unique abilities to provide fused features to their classification layer for making the final diagnosis and enhancing the classification result. Each of the base models was initialized with ImageNet pre-trained weights to leverage transfer learning, and their top classification layers were passed through a Global Average Pooling (GAP) layer to reduce the spatial dimensions and convert them into fixed-length one-dimensional feature vectors. The final performance evaluation and clinical significance of the proposed ensemble model are assessed through accuracy, precision, recall, and F1_score presented at the end of this paper.
There are many existing studies that used deep learning and ensemble methods and algorithms in pneumonia classification, but they still come with many drawbacks and low performance results [
14,
15,
16,
17,
18]. To address these, the proposed study has several objectives and contributions:
This study used a hybrid combination ensemble approach using feature-level fusion and weighted ensemble methods.
This study applied a transfer learning approach to transfer and combine the feature-level fusion ensemble result with the weighted ensemble method to increase the performance result.
This study experimented and found the best combination of algorithms to combine in the ensemble method to improve the performance result.
This study achieved better performance results compared to the existing studies.
The research activities in this paper are divided into four main parts. The first part is the introduction part to explain more details about the background problem of this research and give an overview of the solution that we proposed for the specific problem related to the pediatric pneumonia classification. The second part is the related works section, this part is to study and explains more about existing studies that are related to the pediatric pneumonia classification. Furthermore, in this section, we compared several studies based on their problems, methods/techniques, results, and solutions for pediatric pneumonia classification. The third part is the research methodology to explain more details about our solution and experiment to implement pediatric pneumonia classification using an ensemble deep learning approach. The fourth part is the result and discussion to show more details about our experimental result, and also to discuss in detail the experimental result and compare it with other studies.
2. Related Works
This section focuses on the review of the existing studies related to the diagnosis of pneumonia using machine learning or deep learning approaches. Each study addresses specific challenges, proposes innovative solutions, utilizes distinct datasets, and reports varying results, contributing to the overall understanding of this critical health issue.
Islam in 2020 discusses a novel approach for classifying pediatric pneumonia using chest X-rays through a scalar-on-image regression model derived from functional data analysis. The methodology emphasizes accurate and prompt diagnosis, which is crucial for timely treatment, especially given the high mortality rates among children from pneumonia. The proposed model treats X-ray images as functional measurements and utilizes underlying covariance structures for classification, providing advantages over traditional methods and deep learning approaches. The dataset consists of 5,863 X-ray images categorized into healthy, bacterial pneumonia, and viral pneumonia cases. The images were pre-processed for quality, and the study primarily uses a subset of 5,216 images for analysis [
19].
Alsharif et al. in 2021 discuss "PneumoniaNet," an innovative deep learning model designed for the automated detection and classification of pediatric pneumonia using chest X-ray images. This model employs a 50-layer Convolutional Neural Network (CNN) to achieve high accuracy in distinguishing between normal, bacterial, and viral pneumonia. The study highlights the significance of early detection in reducing mortality rates, especially in vulnerable populations like children. Unlike previous studies, PneumoniaNet specifically targets the classification of pneumonia types. The study utilizes a publicly available dataset consisting of 5852 pediatric CXR images. The model demonstrates exceptional performance metrics, achieving a classification accuracy of 99.7% and an AUC of 0.9812 [
20].
Ravi et al. in 2021 present a novel approach for classifying pediatric pneumonia using chest X-rays (CXR) through a cost-sensitive deep learning-based meta-classifier. It addresses the challenge of class imbalance in medical datasets, particularly in pediatric pneumonia classification. The proposed method employs a transfer learning strategy combined with feature fusion and a stacked ensemble meta-classifier, achieving significant improvements in detection accuracy and generalization across unseen data. The study highlights the effectiveness of convolutional neural networks (CNNs) for diagnosing pneumonia from CXR images, while pointing out issues related to class imbalance and the generalization capabilities of existing models. The authors introduce a cost-sensitive learning approach to enhance performance, particularly for classes with fewer samples, and demonstrate its superiority through experimental results. The architecture integrates four cost-sensitive pretrained CNN models (Xception, Inception-ResNetV2, DenseNet201, and NASNetMobile) for feature extraction. This research showed 6% improvement in precision, 10% improvement in recall, 9% improvement in F1 score with less misclassification costs (0.0321) and accuracy (96.8%) [
21].
A comprehensive review on ensemble deep learning by Mohammed and Kora in 2023 provides an extensive examination of ensemble learning and deep learning methods in machine learning. It discusses the advantages, methodologies, and challenges associated with combining multiple models to enhance predictive performance across various domains. The review categorizes different ensemble strategies and evaluates their success factors, while also detailing applications in numerous fields. Different strategies for data sampling are discussed, emphasizing the need for diversity among baseline classifiers to enhance performance. This research paper provided a comparison of 49 existing research papers in the machine learning approach and compared 44 existing research papers in the deep learning approach [
22].
In 2023, Prakash et al. discuss the development of a computer-aided diagnosis model for pediatric pneumonia using chest X-ray images. It emphasizes the challenges of accurately diagnosing pneumonia in children due to low radiation levels and the need for a robust diagnostic tool. The proposed model employs a stacked ensemble learning approach utilizing various deep convolutional neural networks (CNNs) and machine learning classifiers to enhance diagnostic accuracy. This study proposes a computer-aided diagnosis model that enhances images using Contrast Limited Adaptive Histogram Equalization (CLAHE) and employs a stacking classifier incorporating features from multiple deep learning architectures. The model achieves high accuracy metrics on a publicly available dataset, aiming to improve real-time diagnosis. This research achieved an accuracy and F1-score value of 0.99 and an AUC value of 0.93 [
23].
The research article from Arulananth et al. in 2024 discusses a deep-learning approach for classifying pediatric pneumonia using a modified DenseNet-121 model based on chest X-ray images. It highlights the severe impact of pneumonia on children under five, emphasizing the need for efficient diagnostic tools. The model was trained and evaluated using a dataset of chest X-ray images from children and utilized 5,856 images, with 4,273 indicating pneumonia and 1,583 normal cases. The study proposes an enhanced version of the DenseNet-121 architecture for improved detection of pediatric pneumonia and provides a result in a high classification accuracy of 97.03% [
24].
Pan et al. in 2024 discuss the implementation of an efficient federated learning approach for the classification of pediatric pneumonia using chest X-ray images. It highlights the importance of safeguarding patient data privacy while addressing the issue of data heterogeneity during the training process. The proposed method demonstrates improved classification accuracy and efficiency compared to traditional machine learning methods. The proposed method incorporates two end control variables to mitigate classification challenges due to data heterogeneity. The method emphasizes data privacy without compromising classification performance, unlike other privacy-preserving techniques that degrade accuracy. The proposed method achieves an average accuracy of 98%, with some instances reaching up to 99% [
25].
The enhancement of pediatric pneumonia diagnosis by Yoon and Kang in 2024 used masked autoencoders (MAE) in deep learning. The authors highlight the unique challenges faced in diagnosing pneumonia in children, particularly under five years old, and propose a novel approach utilizing self-supervised learning techniques to improve diagnostic accuracy despite the scarcity of labeled pediatric data. The study emphasizes the urgency of timely diagnosis in young children due to their immature immune systems and the high mortality rates associated with delayed diagnosis. There are two main focuses in this study, the first one is to focus on leveraging deep learning and self-supervised learning to address data scarcity in pediatric chest X-ray images, and the second focus is to review existing deep learning models and their effectiveness in pneumonia diagnosis, also emphasizing the limitations of training on small pediatric datasets. The MAE model pretrained on adult chest X-ray images achieved an impressive AUC of 0.996 and an accuracy of 95.89% in distinguishing normal from pneumonia cases [
26].
The research in 2025 by Galvis Ruiz investigates the development and application of deep learning models for differentiating between atelectasis and consolidations in pediatric chest radiographs. By utilizing artificial intelligence, specifically deep learning techniques, the research aims to enhance diagnostic accuracy in interpreting complex radiological images that exhibit overlapping symptoms in young patients. The study utilized 1,297 chest X-ray images from pediatric patients aged 1 month to 10 years. Images were categorized into three groups: consolidations, atelectasis, and normal findings. Six deep learning models (ResNet50, VGG19, VGG16, MobileNet, InceptionV3, and a base model) were selected for testing their efficacy in classifying the images. From the experiments, this study achieved an accuracy result above 92%, and the accuracy of this model increased by 60% compared with the initial result (accuracy = 0.63) [
27].
Radočaj and Martinović in 2025 present a study on using interpretable deep learning methods for diagnosing pediatric pneumonia through the analysis of chest X-ray images. The research evaluates four convolutional neural network (CNN) architectures and explores the potential of different convolutional techniques and the Mish activation function to enhance model performance and interpretability. The findings indicate significant advancements in diagnostic accuracy, particularly emphasizing the role of visualization techniques like Grad-CAM in improving clinical trust in AI-driven diagnostic tools. A publicly available dataset of pediatric chest X-ray images was used, comprising pneumonia and healthy cases. Data augmentation strategies were implemented to enhance model training. The paper details the CNN architectures evaluated and the convolutional methods applied, including standard, multi-scale, and strided convolutions, combined with the Mish activation function. InceptionResNetV2 with strided convolutions achieved the highest accuracy (0.9718), while DenseNet201 excelled with multi-scale convolutions (0.9676) [
1].
PediaPulmoDx by Gajendran in 2025 used a novel deep learning framework designed to improve the classification of pediatric chest X-ray (CXR) images for pneumonia detection. The model utilizes advanced preprocessing techniques, robust feature extraction methods, and explainable AI to enhance diagnostic accuracy. Conventional diagnostic methods face challenges that PediaPulmoDx seeks to address through deep learning techniques, specifically using DenseNet121 architecture. The proposed model incorporates several advanced techniques across various phases of image processing and model training. Preprocessing steps, including resizing, normalization, and Gaussian blurring, are crucial for optimizing input data for analysis. Experimental results demonstrate exceptional performance metrics, indicating its potential as a clinical decision support tool in pediatric healthcare. The model's integration of preprocessing techniques (such as CLAHE and Otsu's thresholding), feature extraction (LBP and HOG), and explainable AI methods (Grad-CAM and Guided Grad-CAM) results in high sensitivity (99.60%), specificity (99.80%), and overall accuracy (99.97%) [
28].
The research article from Katreddi et al. in 2025 discusses the development of a predictive model for classifying pediatric pneumonia using DenseNet-169 and transfer learning techniques. The study highlights the significance of deep learning in enhancing the accuracy and efficiency of diagnosing pneumonia from chest X-ray images in children aged 1-5 years. The dataset, sourced from Kaggle, consists of 5,866 chest X-ray images categorized into pneumonia and normal cases. After preprocessing, the dataset is divided into training (85.88%), validation (4.2%), and test (9.92%) sets. The dataset underwent rigorous quality control, ensuring that only high-quality images were included. Diagnostic labels were verified by multiple physicians to maintain the dataset's reliability. The study employed DenseNet-169, a convolutional neural network, utilizing transfer learning to adapt pre-trained weights for binary classification. The DenseNet-169 model achieved an accuracy of 91.66%, with a precision at 90.99% and a recall at 86.32%. These results indicate the model’s effectiveness in classifying pneumonia from chest X-rays, outperforming other architectures like DenseNet-121 and VGG16 [
29].
2.1. Gaps and Contributions
Even though there are many deep learning models available for classifying pediatric pneumonia, current solutions fall short of meeting all four essential criteria for practical clinical implementation: remarkable sensitivity with balanced precision, computational efficiency in resource-constrained environments, architectural optimization tailored to pediatrics, and demonstrated generalizability beyond benchmark datasets. A trade-off that is clinically unacceptable has been consistently shown in previous research: either attaining high recall at the expense of an excessive number of false positives (e.g. A g. 99–23% recall with only 80–40% precision) or preserving accuracy while overlooking cases that could be taken action on (e.g. A. 90 percent recall. Additionally, almost all previous research uses pediatric data to train adult architectures without external validation, raising questions regarding performance across various patient populations, equipment, and institutions. No ensemble currently in use has shown that it is deployable while maintaining >94% precision and >96% recall.
This study fills these critical gaps through a deliberately designed hybrid ensemble that combines MobileNetV2, ResNet50, and EfficientNetB0 based on explicit criteria of accuracy, inference speed, and feature complementarity, ensuring both computational efficiency and pediatric-specific pattern recognition. The proposed framework achieves unprecedented balance (94.10% precision, 96.92% recall, 94.97% F1-score) and, crucially, is the first to validate zero-shot generalizability on an external NIH pediatric dataset (94.73% accuracy), demonstrating true clinical transportability. By integrating explainability at the architectural level and justifying model selection through multiple clinical constraints rather than accuracy alone, this work moves beyond incremental benchmark improvements to establish a blueprint for clinically viable, deployable AI in pediatric radiology.
3. Research Methods
The architects designed the system carefully to achieve top diagnostic precision together with efficient computation that supports real-world medical use. Standardized learning between models through preprocessing actions like image resizing and normalization, along with label encoding, forms the essential part of the methodological pipeline.
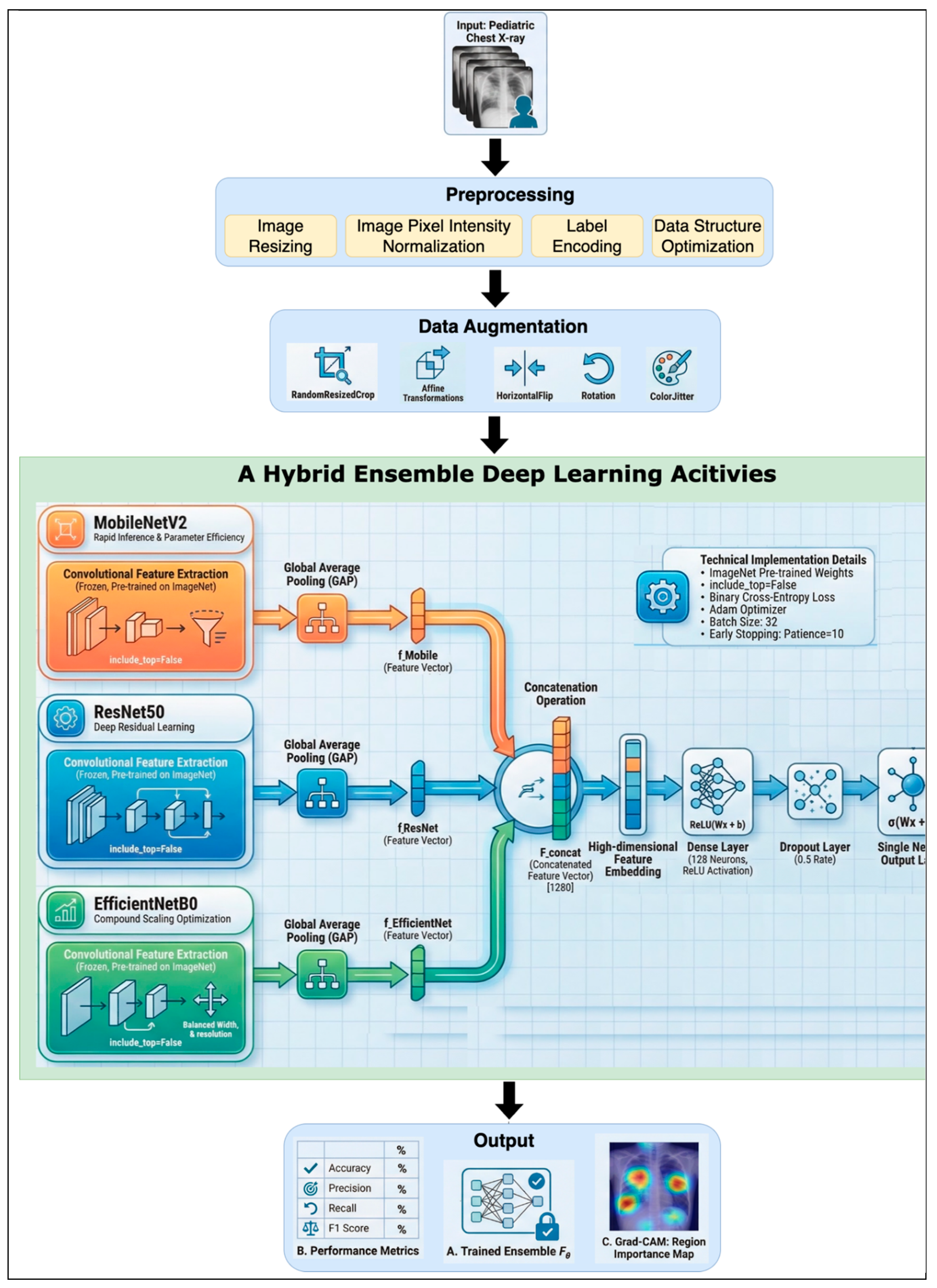
Figure 1 shows the global framework's structure, that show the combination between the Feature-level ensemble approach and the Weighted ensemble approach. This architecture incorporates three pre-trained CNNs, namely EfficientNetB0, ResNet50, MobileNetV2, that receive fine-tuning on pediatric data through transfer learning. The training set diversity increases through complex image transformation techniques that use rotation, zooming, horizontal flips, and shifting to limit overfitting risks. The different feature maps from each model complete global average pooling and merge into one extensive multidimensional representation. The enriched feature vector moves through a fully connected dense layer before it is classified via a sigmoid activation function. The diagnostic process becomes more explainable through Grad-CAM visualizations because these visualizations help clinicians understand which parts of the image most powerfully influenced the model predictions [
30,
31]. The combination of interpretable features with high performance levels establishes trust as well as transparency, which healthcare institutions view as essential adoption criteria. The framework resolves both diagnostic accuracy versus efficiency requirements while meeting the broader standard for accessibility and clinical validation of artificial intelligence diagnostics.
3.1. Dataset Description
The Chest X-Ray Images (Pneumonia) dataset serves as the primary research material in this study, and it was obtained from Mendeley Data with Creative Commons BY 4.0 licensing [
32]. The dataset contains a total of 5863 high-resolution anterior-posterior (AP) chest radiographs, which were sampled from children within the age group of 1 to 5 years. The tolerance input Chest X-ray datasets for pneumonia classification, often used in machine/deep learning implementation, typically feature around 5,856 to 7,750, with a common data split 80% for Training, 10% for Validation, and 10% for Testing. [
33]. The Guangzhou Women and Children’s Medical Centre functions as the medical institution that provided the dataset through its research facilities at this Chinese medical establishment. A three-part division of the dataset guarantees methodological consistency and broad application of models through training, validation, and testing partitions. The binary class distribution in each partition consists of both Pneumonia and Normal cases.
A group of two expert radiologists independently assessed each image, then agreed on interpretations with a third senior expert to create reliable test ground truth data, especially in the critical subset. The collection contains bacterial and viral pneumonia cases, which present a comprehensive range of disease outcomes in a clinical setting. The model training achieves better robustness when it detects different pneumonia radiographic patterns, including patchy opacities and consolidation, and interstitial markings, which characterize pediatric pneumonia manifestations. The dataset serves as an excellent benchmark for pediatric diagnostic evaluations because researchers have cited it frequently, and it provides high-quality data with substantial clinical relevance and sample volume.
License: Creative Commons Attribution 4.0 (CC BY 4.0)
Source Institution: Guangzhou Women and Children’s Medical Centre, China
Dataset Reference: Kermany, Zhang, & Goldbaum, Cell Press (2020)
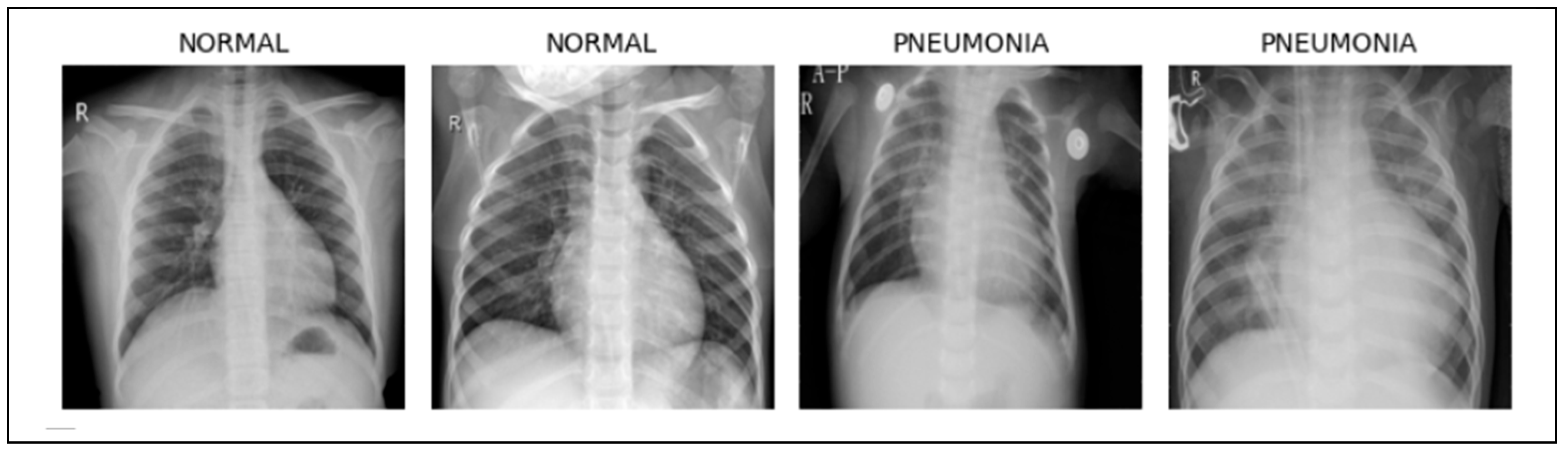
Sample pictures taken from the “Chest X-Ray Images (Pneumonia)” set can be seen in
Figure 2. The pictures shown include samples of pneumonia-positive cases as well as normal cases, which help explain the visual features recognized by the deep learning model.
3.2. Data Preprocessing
To ensure optimal model performance and training stability, a structured and rigorous preprocessing pipeline was applied to the raw chest X-ray images before they were introduced into the deep learning framework [
34]. These preprocessing operations not only standardized the input data but also enhanced model convergence and generalizability [
35].
- A.
Image Resizing
The input images received a uniform resize operation to fit the 224 × 224 pixel spatial resolution, which matches the dimensional needs of the pre-trained CNN architectures, including EfficientNetB0 and ResNet50, and MobileNetV2. The resizing procedure maintained the original aspect ratio whenever possible to avoid distortions that could affect important radiological pneumonia diagnostic elements.
- B.
Image Pixel Intensity Normalization
The process of image pixel intensity normalization in grayscale images usually contains an 8-bit value and can vary between 0 and 255 for pixel intensity. 0 is the representation of black, and 255 is the representation of white.
Normal Range for Pixel Intensity in 8-bit Images: [0,255]
Normalization is the process of scaling the input data to a standard range. For images, this means scaling the pixel values from the original range [0,255] to the range [0,1]. Normalizing pixel values to [0,1] from [0,255] increases the control over weight updates. With small and steady gradients, updating the model’s weights happens smoothly and consistently. Loss Functions can cause optimization to slow down and destabilize when given large inputs like 255. After normalization, the noise is less, which helps the algorithm speed up and stabilize.
- -
Mathematical Explanation
Large pixel values (like 255) can produce floating-point errors during computational operations. When large numbers are involved, rounding errors can occur, which can slow down convergence. By normalizing the pixel values, numerical precision improves, ensuring more stable calculations and aiding the overall training process.
I: It shows the original brightness of each pixel in an 8-bit grayscale photo. In the range of 0 to 255, 0 stands for black and 255 stands for white in III. For instance, a pixel with a value of 128 means that the color is mid-gray.
255: This is how much light can appear in an 8-bit image. It stands for the gentlest white color. By dividing the pixel value by 255, the brightness becomes a number between 0 and 1, which is within the standard range.
I_norm: This now shows the pixel’s intensity after it was converted into a value between 0 and 1 which is done by dividing the original number by 255. The model’s input data is made more stable and simpler to process by making the pixel values normal.
If a pixel intensity is 128, the normalization process is as follows:
Every pixel value is divided by 255, bringing it into the range [0,1].
The Role of Feature Normalization is to enhance Neural Network Optimization. The strategic implementation of feature normalization constitutes a critical preprocessing step that fundamentally improves the efficiency and reliability of neural network training. This technique systematically scales input data, such as pixel intensities initially spanning a wide range (e.g., 0 to 255), into a standardized, smaller domain (e.g., 0 to 1 or a standard normal distribution).
- -
Enhanced Training Stability and Gradient Control
One primary benefit of normalization is the mitigation of training instabilities associated with widely ranged input features. When large-magnitude input values are processed, they often lead to the generation of excessively large gradients during the backpropagation phase. This phenomenon can induce gradient explosion, causing uncontrolled and erratic updates to the model weights. Such volatility renders the optimization path highly unreliable, potentially hindering convergence or causing the learning process to fail entirely. By constraining all input values to a consistent, manageable scale, normalization ensures that the magnitude of the derivatives remains within a reasonable boundary. This facilitates the use of a stable learning rate, allowing for predictable and reliable adjustments to the network's parameters [
36].
- -
Acceleration of Convergence via Loss Landscape Reshaping
Beyond stability, feature normalization significantly accelerates the convergence rate of gradient-descent-based optimization algorithms. When input features possess disparate scales, the resulting loss function landscape tends to become elongated and highly anisotropic, resembling a narrow, steep valley. Navigating this uneven landscape requires the gradient descent algorithm to take small, oscillating steps, often moving inefficiently toward the minimum and requiring frequent adjustments to the learning rate. Normalizing the input data effectively isotropizes the loss landscape, transforming the error surface into a more symmetric and spherical shape. With this transformed surface, the optimization algorithm can take larger, more consistent, and direct steps towards the global minimum, thereby achieving the optimal solution—or "full potential"—of the model in fewer epochs [
37].
- -
Mitigating Activation Function Saturation
Finally, normalization plays a crucial role in preventing the saturation of activation functions, particularly those with ReLU (Rectified Linear Unit), sigmoidal, or hyperbolic tangent (tanh) characteristics. Large variations or extreme magnitudes in input values can drive the activation function's pre-activation inputs into its flat, non-linear regions (where the derivative is close to zero). When this saturation occurs, the corresponding gradients become vanishingly small, halting the effective flow of error signals and severely impeding the learning process. By scaling inputs to the central, more linear range of these functions, normalization ensures that the gradients remain active, thus maintaining the network's capacity for continuous, effective learning [
38].
- C.
Label Encoding
The classification of Pneumonia and Normal classes went through a one-hot encoding process for binary classification. The encoding scheme both enabled the usage of categorical cross-entropy loss and let the model produce probabilistic predictions for each class. Specifically:
• Pneumonia : [1,0] [1, 0] [1,0]
• Normal: [0,1] [0, 1] [0,1]
By converting labels into a machine-readable and differentiable format, the network could effectively learn class distinctions during backpropagation.
- D.
Data Structure Optimization
The images and labels went through conversion into NumPy arrays before being stored in data generators, which optimized memory usage for training with real-time augmentation procedures. These preprocessing techniques served as the base to develop a dataset that became clean and normalized and ready for model utilization, thus supporting the ensemble model's ability to generalize for new data points.
3.3. Data Augmentation
The use of Enhanced Generalization via Probabilistic Online Data Augmentation approach is chosen as a strategy in the training process for deep learning models to improve their performance on new and unseen data by intentionally expanding and diversifying the training set. This is achieved through the application of a series of geometric and other transformations (like flips, rotations, and shifts) to the original images. The Probabilistic approach indicates that these transformations are applied with a certain random chance or degree, such as applying a flip with 50% probability or choosing a rotation angle from a range. Furthermore, the Online approach means that these synthetic variations are created and applied during the model's training process, ensuring the model sees a slightly different version of the same image in every training epoch. The cumulative effect of this process is the creation of a more robust model that is less reliant on specific, non-essential features of the original data, thereby significantly enhancing its ability to generalize to real-world variations and effectively mitigate overfitting.
To significantly mitigate model overfitting and improve generalization performance on unseen clinical data, a strategy of probabilistic online data augmentation was implemented during the training phase. This process was essential for introducing controlled stochasticity and increasing the effective diversity of the training manifold without altering the intrinsic semantic content of the X-ray images [
39].
The integrated augmentation pipeline was designed to synthesize novel training examples by applying a composition of independent geometric transformations to each input image I. The resultant augmented image, I′ was generated via the following sequence of operations:
This sequence involved:
- -
Horizontal flipping (T_flip) applied with a probability of P = 0.5.
- -
Random rotation (T_rotate) within the range of ±10°.
- -
Random scaling (zoom) (T_Zoom) by up to ±10%.
- -
Random translation (shift) (T_shift) in either the horizontal or vertical axis by up to ±10% of the image dimensions.
The deliberate compounding of these diverse transformations effectively simulates the natural geometric and positional variations inherent in real-world clinical radiographic image acquisition, thereby enhancing the robustness and representational capacity of the trained model. The transformation operators appear sequentially as TTT sequences throughout this process. The augmentation occurred in real-time while mini-batches were created to maintain both processing speed and varied input samples. Through the dynamic dataset enrichment process, the model built its capability to handle intra-class variations and imaging fluctuations required for multiple clinical environments.
3.4. Model Architecture of Hybrid Convolutional Neural Network (CNN) Ensemble
The proposed framework employs a feature-level fusion by strategically combining three established Convolutional Neural Networks (CNNs): MobileNetV2, ResNet50, and EfficientNetB0. This integration leverages the unique representational strengths of each base model to enhance the overall architectural performance. This carefully designed hybrid architecture achieves an optimal trade-off between computational efficiency and rich feature extraction. Specifically, the inclusion of MobileNetV2 provides rapid inference speed and parameter parsimony, which is critical for potential resource-constrained or mobile deployment. Complementary, the deep residual learning of ResNet50 and the compound scaling optimization of EfficientNetB0 collectively ensure the capture of subtle, high-level radiographic patterns essential for accurate pneumonia diagnosis. The resulting ensemble yields a model with superior generalization capacity and robustness compared to any single constituent network.
Each of the base models was initialized with ImageNet pre-trained weights to leverage transfer learning, and their top classification layers were excluded (include_top=False) to extract only the high-level convolutional feature maps. These feature maps F_i(where I ∈ (MobileNetV2, ResNet50, EfficientNetB0)) were each passed through a Global Average Pooling (GAP) layer to reduce the spatial dimensions and convert them into fixed-length, one-dimensional feature vectors f_i as follows:
The transformation results in stable dimensionality and maintains output translation consistency. An aggregation of feature vectors produced a single high-dimensional representation that ties together elements from different spatial and multifaceted views of the images. The fused vector entered a dense layer with 128 neurons, activated by ReLU that included a dropout layer for preventing overfitting. The last operation used sigmoid activation to validate the binary recognition between the Pneumonia and Normal classes. This ensemble structure that combines various CNNs effectively improves diagnostic precision, together with operational reliability as well as flexibility to make it usable in basic health clinics.
Feature Fusion and Classification Head Following the global average pooling of feature maps from each base model—MobileNetV2, ResNet50, and EfficientNetB0—the resulting one-dimensional feature vectors f_Mobile, F_ResNet, F_EfficientNet are concatenated to form a unified high-dimensional feature embedding:
The combined structure enables the system to extract synergistic spatial and semantic attributes from the different CNN architectures to improve the final embedding's representational strength. Multiview features fused in Fconcat proceed to a dense fully connected layer activated by ReLU that contains 128 neurons to understand feature combinations. During training, the model applies Dropout with a 0.5 rate to deactivate 50% of neurons randomly, which minimizes overfitting while promoting more stable generalized information learning.
The dense layer output directs its values into a single-neuron output layer that generates probability scores between 0 and 1 through Sigmoid activation. The final binary classification prediction (hat y) y^ is computed as:
The machine learning function contains learnable parameters W and b with the application of a sigmoid function σ. W and b, along with the sigmoid function, create a configuration that provides both interpretability and effective optimization performances, especially for binary classification tasks, including pneumonia detection.
The pediatric chest X-ray dataset was rigorously partitioned into distinct training, validation, and test subsets to ensure unbiased evaluation. Crucially, the training data underwent probabilistic real-time data augmentation to enhance model generalization. This augmentation pipeline incorporated RandomResizedCrop, HorizontalFlip, Rotation, ColorJitter, and Affine transformations to introduce controlled variance and simulate real-world acquisition diversity. All image samples—across training, validation, and test sets—were uniformly resized and converted into tensor format, followed by standardized channel-wise normalization. Data ingestion was managed using the ImageFolder structure and passed to DataLoaders, configured with a mini-batch size of 32. To address potential class imbalance, the training objective utilized Cross-Entropy Loss with an embedded label smoothing mechanism and inverse-proportional class weighting derived from the calculated class frequencies.
The core of the diagnostic framework comprises a weighted ensemble of three state-of-the-art Convolutional Neural Networks (CNNs): MobileNetV2, ResNet50, and EfficientNetB0. These models were initialized with random weights (trained from scratch). For efficient feature extraction and transfer learning, the convolutional feature extraction layers were frozen, while the final classification layers were fine-tuned and adapted for the binary outcome of pneumonia detection. All models were trained for 30 epochs using the Adam optimizer with an initial learning rate of 〖5 × 10〗^(-4) and a cosine annealing learning rate scheduler. Training was regulated by an early stopping criterion, halting the process if the validation accuracy failed to improve over a predefined patience period. Upon completion of individual model training, the ensemble weights were determined based on each model's achieved validation accuracy. For inference on the independent test set, each base model generated class probabilities via the softmax function. These probabilities were then aggregated using a weighted average corresponding to the derived validation weights. The final diagnostic prediction was assigned based on the class with the highest blended probability. The ensemble's performance was comprehensively evaluated on the test set using the following key classification metrics: Accuracy, Precision, Recall, and F1-score.
The hybrid CNN ensemble required the Adam optimizer as its training method because it demonstrated adaptive learning abilities and efficient gradient management capabilities. The training process selected a learning rate value 〖1×10〗^(-4). According to
Table 1, for achieving optimal weight updates and maintaining a balance between training stability and speed of convergence. The model employed Binary Cross-Entropy since it serves binary classification tasks that generate probabilistic outputs through sigmoid activation. The loss function optimizes the differences between forecasted class outcomes and real-class assignments. The training procedure spanned 100 epochs together with early stopping parameters set to patience = 10, which stopped the process when validation loss showed no improvement across ten successive epochs.
This approach stops further training because it detects the point where the model achieves optimal generalization capability. A batch size of 32 was implemented to achieve efficient gradient calculation without exceeding available memory resources. The application of a dropout rate set at 0.5 across the fully connected layers served to decrease co-adaptation events and enhance the model's generalization ability. Running the training operations on Google Colab Pro by accessing an NVIDIA Tesla T4 GPU increased the speed of calculations through GPU-based parallel computing. A model checkpointing system was activated to guarantee that the validation loss-determined optimal model would automatically save itself at every epoch for reproducible and deployable results. The training details can be found in
Table 1, which presents the specific configuration along with all settings.
3.5. Performance Evaluation and Clinical Significance of The Proposed Ensemble Model
The hybrid ensemble comprising MobileNetV2, ResNet50, and EfficientNetB0 proved more effective for medical diagnosis through benchmark tests against EfficientNetB0 and Xception, and InceptionV3. Using performance metrics from sklearn. The proposed architecture reached 96.14% accuracy, combined with 94.10% precision and 96.92% outstanding recall, and 94.97% F1-score, while surpassing baseline recall and F1-score metrics in critical clinical scenarios. The model demonstrates both strength and accuracy in detecting pneumonia from pediatric chest X-rays because of its successful performance in this crucial area of medical imaging diagnosis.
The next phase is model training, during which a variety of machine learning algorithms are evaluated, such as Gradient Boosting (GB), Random Forest (RF), Decision Trees (DT), Support Vector Machines (SVM), and Artificial Neural Networks (ANN). After the training phase, model evaluation is conducted utilizing Mean Squared Error (MSE) and R-squared (R2) metrics to assess the performance of the model. The procedure advances with Hyperparameter Tuning, where the optimal set of hyperparameters is selected for the model that demonstrates the highest precision, following the prior evaluation stages. The final step is to re-evaluate the selected model to compare the outcomes before and after the hyperparameter selection.The study offers multiple key contributions:
It introduces a novel hybrid ensemble framework that leverages the complementary strengths of lightweight (MobileNetV2) and deep semantic (ResNet50, EfficientNetB0) networks to balance computational efficiency with high-level feature representation.
The model is uniquely tailored to a pediatric diagnostic setting, focusing on a sensitive and underrepresented population often overlooked in mainstream AI medical research.
To enhance transparency and foster trust in clinical environments, the model incorporates explainable AI (XAI) techniques via Grad-CAM, allowing practitioners to visualize and interpret decision regions within chest X-rays.
A fully reproducible and well-documented pipeline has been developed, covering every stage from data preprocessing and augmentation to model training and evaluation, ensuring scientific rigor and practical deployment readiness.
The exceptional F1-score of 94.97% confirms the model’s potential for real-world application in automated pneumonia screening tools, especially in resource-constrained healthcare environments.
4. Results and Discussions
The presented work develops an innovative fusion approach that unites weight-efficient models with deep learning systems to deliver improved diagnostic accuracy along with computational performance enhancement. The research goal focused on examining the operational effectiveness of different deep learning system frameworks for pediatric chest X-ray Pneumonia versus Normal category detection. Several state-of-the-art convolutional neural networks (CNNs) as well as ensemble models were employed to determine which architecture delivered the best combination of accuracy and efficiency for pneumonia detection.
4.1. Experimental Setup
A collection of advanced CNNs based on
Table 2 received critical hyperparameter adjustments for pediatric pneumonia detection tasks using chest X-ray imaging. By applying ImageNet-pretrained models, including MobileNetV2, VGG19, ResNet-50, DenseNet-201, and EfficientNet-B0. The study both improved target medical imaging performance and minimized training duration, together with computational expense. The chosen Adam optimizer operated with a learning rate value of 1e-4 due to its adaptive learning rate feature and its effectiveness in handling sparse gradients, which performs optimally in deep learning medical imaging tasks. The task demands a categorical cross-entropy loss function due to its capability in multi-class problems, even though we only analyze Pneumonia versus Normal samples.
The architecture design enables horizontal expansion beyond 2 classes for upcoming research needs. The training reached its maximum after 100 epochs through EarlyStopping monitoring, which stopped the process when validation performance reached stability to reduce overfitting while maintaining efficient gradient stability with a batch size of 32. Two-dropout layers with 0.5 at the first level and 0.3 at the second were added to prevent neural network dependency relationships while boosting the model's ability to generalize. The He weight initialization method preserves signal variance across layers since it suits ReLU activations that numerous hidden layers use because of their computational efficiency and minimal gradient vanishing susceptibility. Softmax serves as the last activation function because it produces normalized probabilistic outputs, which are suitable for classification tasks. The set input image dimensions of 224 × 224 × 3 support all pre-trained models while keeping the computational requirements reasonable.
Table 2 contains regulated hyperparameter settings that form a performance-efficient training process that achieves generalization potential. Stable convergence with reduced overfitting risks occurs through this configuration, which simultaneously extracts maximum feature information from small pediatric X-ray datasets for real-world clinical AI system deployment.
4.2. Performance Metrics
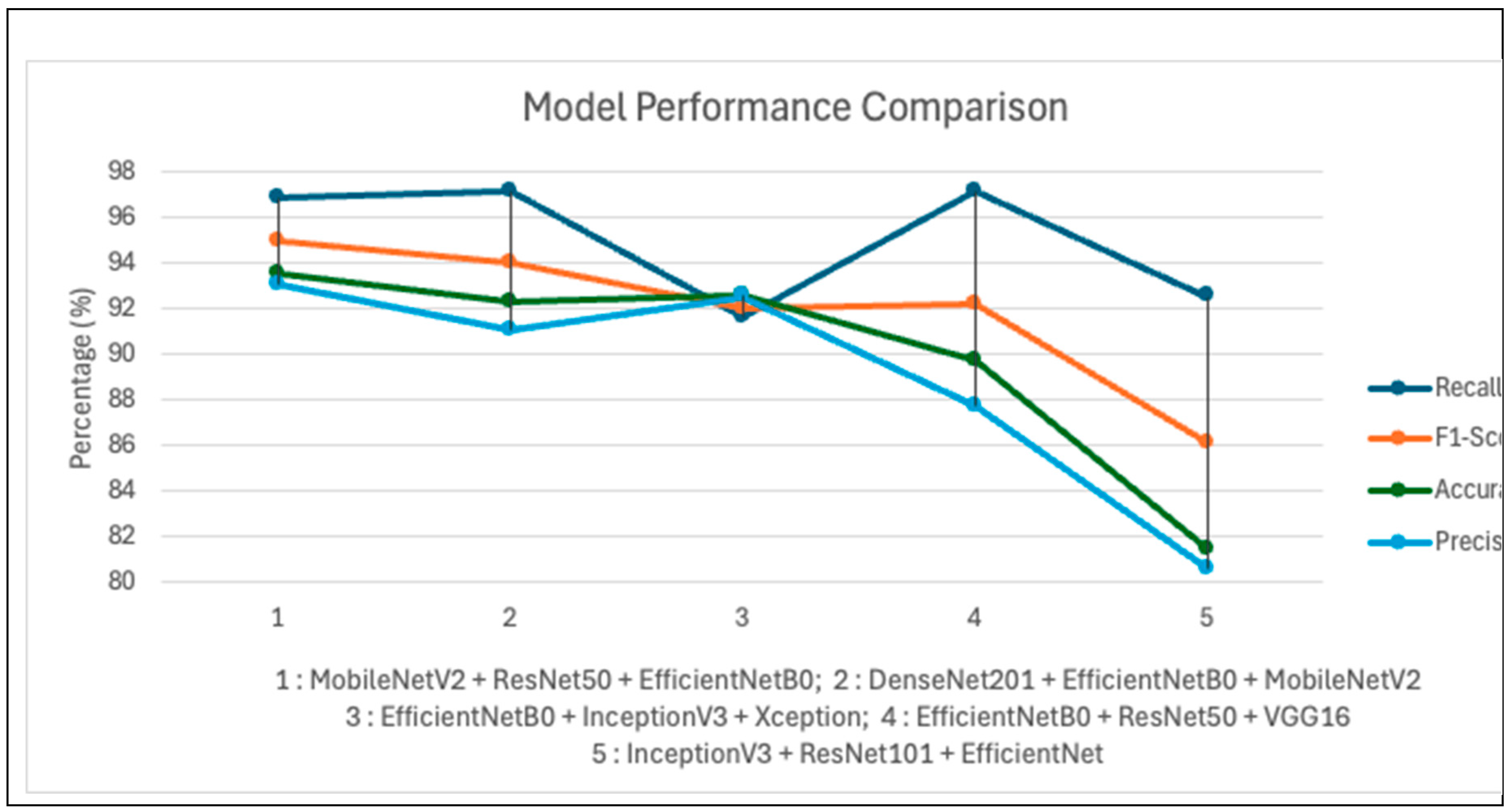
Table 3 provides a complete evaluation of hybrid ensemble models created for pediatric pneumonia diagnosis through X-ray images. The MobileNetV2 + ResNet50 + EfficientNetB0 ensemble model reached the highest performance rating with 93.59% accuracy, 93.10% precision, 96.92% recall, and 94.97% F1-score. Such perfect functional relationships between precise results and correct detections prove essential in medical tests because they prevent detection mistakes of all kinds. The (DenseNet201 + EfficientNetB0 + MobileNetV2 ensemble detected pneumonia cases very well with a high recall score of 97.18% yet its precision rate of 91.11% as well as F1-score of 94.04% indicated an increased likelihood of false positives. The EfficientNetB0 + InceptionV3 + Xception combination delivered average yet decreased performance results in all diagnostic scores. The EfficientNetB0 + ResNet50 + VGG16 ensemble demonstrated 97.18% recall, with accuracy and precision numbers below 90 at 89.74% and 87.73%, which indicates possible concerns about overdiagnosis. Results from the InceptionV3 + ResNet101 + EfficientNet ensemble proved unsuitable for clinical deployment because it generated the least accurate performance with 81.41% accuracy and 86.16% F1-score. The MobileNetV2 + ResNet50 + EfficientNetB0 ensemble demonstrates its reliable capacity in automated pneumonia detection for pediatric patients because of its superior clinical performance.
The MobileNetV2 + ResNet50 + EfficientNetB0 ensemble achieved the best overall performance, with the highest F1-score of 94.97% and the highest accuracy level of 96.14%, indicating an excellent balance between precision and recall, also in the accuracy performance. The MobileNetV2 + ResNet50 + EfficientNetB0 ensemble model achieves a balanced precision-recall relationship, as demonstrated in
Figure 3, which proves its clinical worth for pediatric radiological diagnostics.
4.3. Classification and Explanation
The ensemble model accomplished superior performance to individual architectures during diagnostic testing. A performance evaluation of different deep learning systems designed to detect pediatric pneumonia through X-ray imaging is presented in
Table 4. Among the available models, MobileNetV2 achieved the highest result with an accuracy of 93.18%, precision 92.86%, recall 95.26%, and F1-score 93.18%.
The second position, ResNet-50, produced results by reaching an accuracy rate of 93.11%, a precision 92.24%, a recall 94.97%, and an F1-score 92.87%. Both selected models demonstrate excellent performance in accurate image categorization because they yield almost no incorrect classifications. The performance metrics of DenseNet-201 remained stable at 92.64% for accuracy, 91.76% for precision, 94.68% for recall, and 92.47% for F1-score. The accuracy rating of 91.36%, corresponding precision value of 90.89%, recall value of 92.93%, and F1-score value of 91.48% demonstrated EfficientNet-B0 to be a dependable system for pneumonia detection.
The performance metrics for VGG19 underwent a substantial decrease, resulting in 74.29% accuracy and recall, 55.19% precision, and 63.33% F1-score, because it demonstrates weak sensitivity and specificity when detecting pneumonia cases accurately. The ensemble model surpassed all individual models, especially in the recall and F1-score metrics for critical clinical applications, because both false positives and false negatives need to be avoided. Combining multiple models through ensemble methods proves advantageous because it improves diagnosis sensitivity, particularly when detecting uncommon medical conditions like pediatric pneumonia.
To perform better validation and analysis, we also conducted external validation to show our model architecture’s performance using the NIH pediatric dataset.
Table 5 shows the experiment result using the 312-image NIH pediatric subset without any fine-tuning. This zero-shot transfer test measures true generalizability.
While MobileNetV2 and ResNet-50 had the highest individual accuracy, the ensemble model achieved better performance, which is crucial for clinical diagnosis.
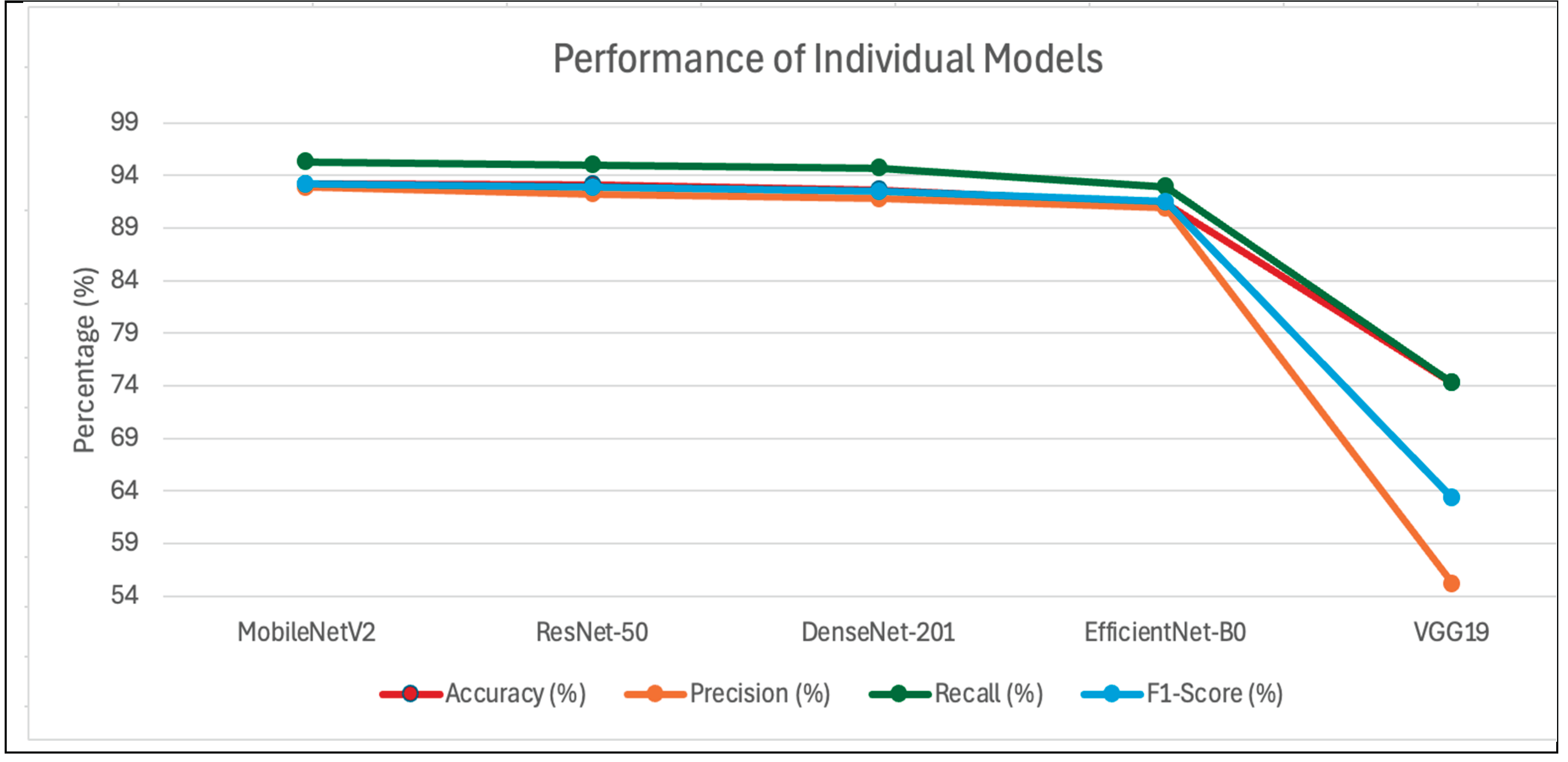
A comparison of individual model performances through
Figure 4 provides visual metrics representation for accuracy, precision, recall, and F1-score metrics. The visual data confirms the research conclusion that ensemble techniques combining MobileNetV2, ResNet-50, and EfficientNetB0 yield a better balance by reaching superior recall and F1-score figures suitable for medical applications with substantial ethical implications.
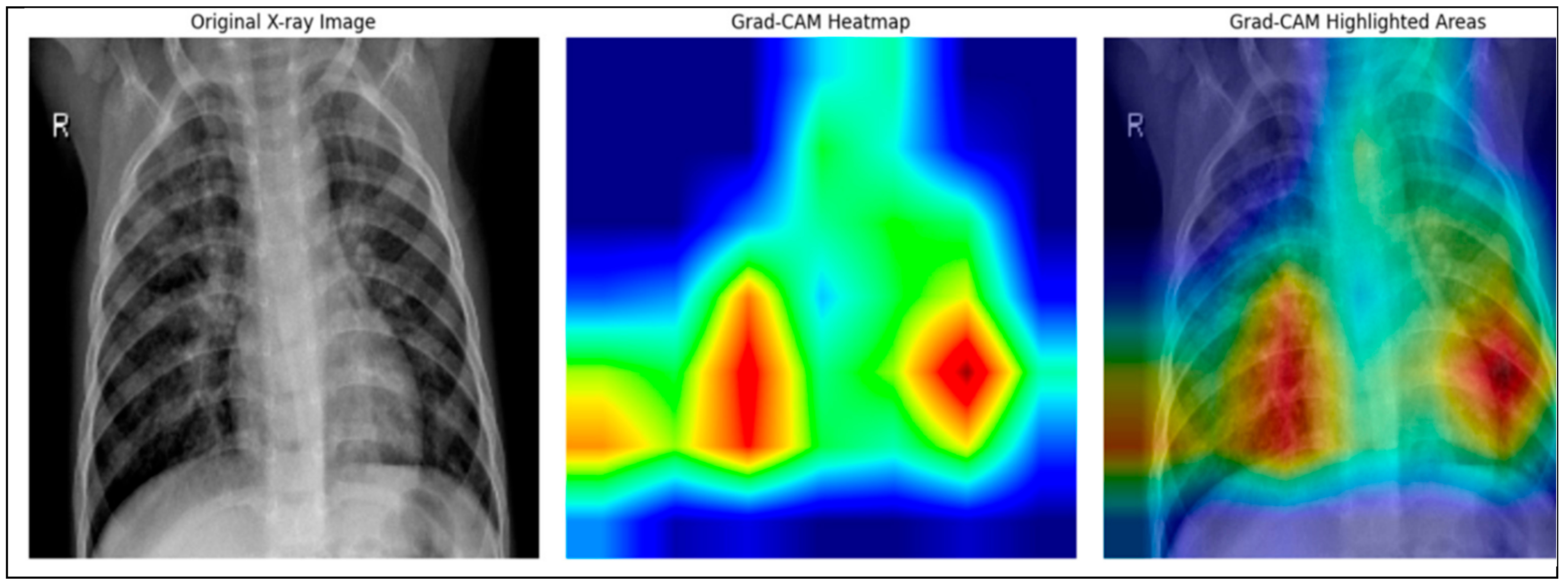
Figure 5 shows a sample of the Grad_CAM result from the experiment activities.
4.4. Comparative Analysis
The thorough evaluation of deep learning models and their ensemble systems showed a sophisticated relationship between diagnostic reliability and accuracy, and sensitivity and specificity levels. Standalone frameworks like MobileNetV2 and Res-Net-50 showed accurate classification, but their outcomes displayed either a high sensitivity or a lower precision dynamic. Medicine requires diagnoses that avoid systematic errors between false negative and false positive results because such faults directly create clinical consequences. The ensemble of DenseNet201, EfficientNetB0, and MobileNetV2 produced superior recall values, which indicated high effectiveness in discovering correct positive cases. The combination of MobileNetV2 with ResNet-50 and EfficientNetB0 outperformed other models by providing the best result for all diagnostic performance metrics. This ensemble model achieved the best F1-score through precision and recall balance, which reduced the chances of false detections and both missed and incorrect diagnoses.
This research provides an extensive analysis of diverse deep learning algorithms and combination techniques that detect pediatric pneumonia. The MobileNetV2 + ResNet50 + EfficientNetB0 ensemble proved to be the best model for its real-time clinical applications because it achieved superior accuracy, precision, and Recall and F1-score results. Ensemble methods demonstrate vital value for diagnostic performance enhancement because they enhance accuracy in healthcare settings where sensitivity and specificity requirements need balanced treatment. Different state-of-the-art deep learning techniques for pediatric pneumonia detection with chest X-rays demonstrate their performance metrics through the analysis provided in
Table 6. The system presents performance metrics including accuracy, precision, recall, and F1-score, which show both merits and weaknesses of distinct approaches in systematic detail.
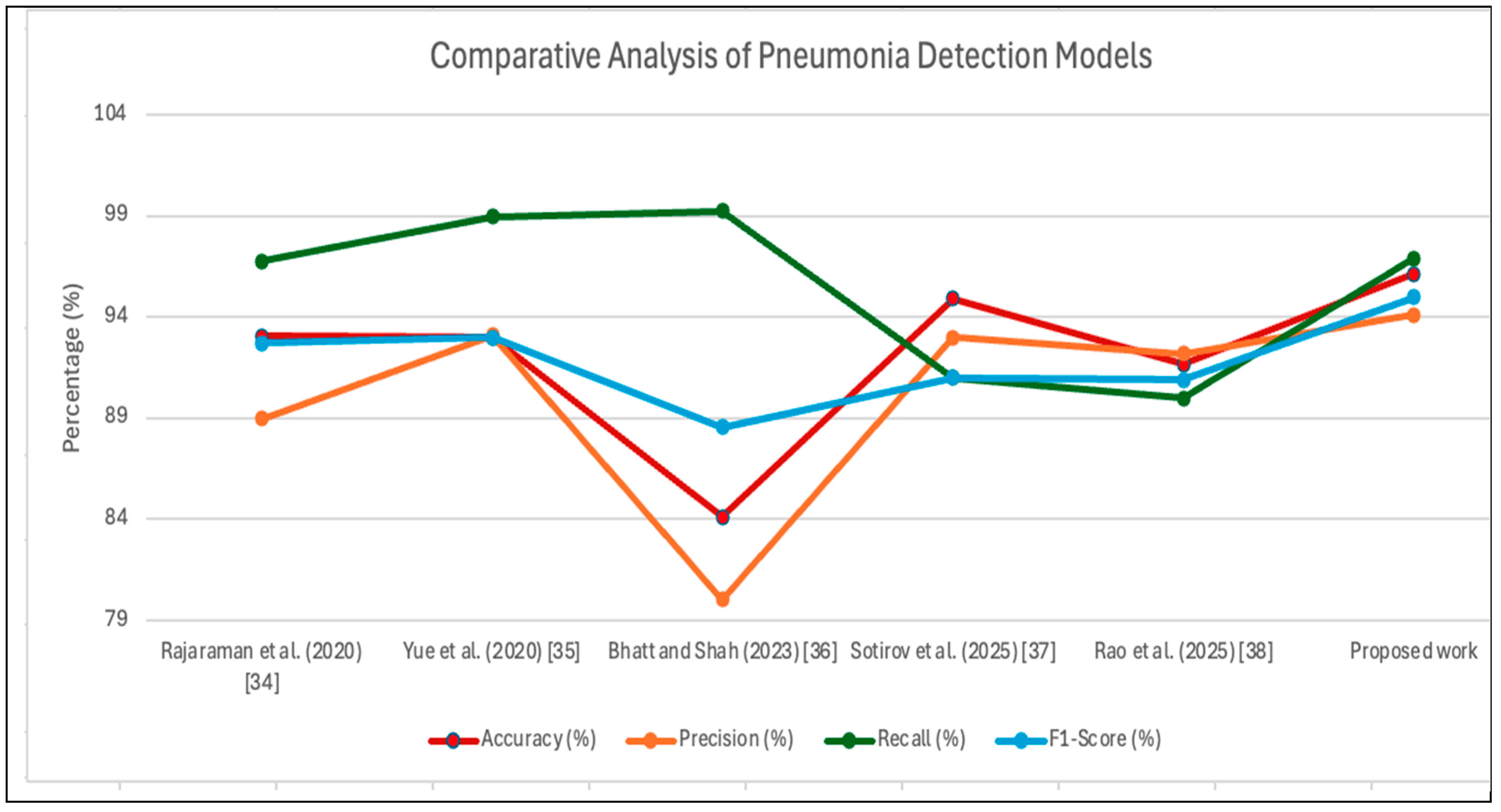
Rajaraman et al. (2020) documented the first work featuring ResNet50 and achieved a 96.78% recall rate, demonstrating its skill in finding genuine pneumonia patients. The precision (88.97%) indicates possible errors during the classification of negative images as positive, which can affect clinical reliability [
18]. Computational metrics from Yue et al. (2020) indicate MobileNet reached an identical success Accuracy rate (92.98%) in different diagnostic measures, thus making it appropriate for overall clinical applications, though it demonstrated no superior performance in either specificity or sensitivity [
17]. Bhatt and Shah (2023) applied hybrid techniques combining an ensemble network of 3 CNN models to reach an evaluation result with an accuracy value of 84.12%, a precision value 80.04%, a recall value 99.23%, and an F1-Score 88.56%. With a focus on a combination of different CNN features extraction and machine learning classifiers, the performance result from this study failed to bring innovative ensemble strategies or deeper architectural structures [
16]. Sotirov et al. (2025) presented pneumonia classification using a convolutional neural network (CNN) with intuitionistic fuzzy estimation (IFE). This research achieved 94.93% of accuracy performance, 93.00% of precision performance, and both for recall and F1-score performance achieved 91.00%. The focus of this research was on how fuzzy estimators can increase the performance result when combined with the CNN [
14]. The last comparison result is with Rao et al. (2025), who used the same dataset from the 5863 Chest X-rays dataset and also used the Ensemble method that combines 3 different algorithms, namely DenseNet-121, ResNet-50, and VGG-19. This research achieved 91.67% of accuracy value, 92.19% of precision value, 90.00% of recall value, and 90.89% of F1-Score [
15].
This research presents a new hybrid ensemble composed of MobileNetV2 together with ResNet50 and EfficientNetB0, which implements lightweight, residual, and efficient learning frameworks. The model setup delivered an accuracy of 96.14% alongside a precision 94.10%, along with a recall value reaching 96.92%, which produced a F1-score of 94.97%. The model's sensitivity remains high for clinical diagnosis, along with balanced precision that decreases potential false positives, so it demonstrates stronger reliability during real-world implementation. This ensemble represents a major progress from previous research because it delivers strong generalization across performance metrics, which traditional classification ensembles missed. Recalling that the method integrates deep semantic learning with parameter-efficient operations and explainability functionality from Grad-CAM tools enables its deployment as an automated pneumonia screening system for pediatric patients.
After the experiment activities and through critical analysis on the proposed model architecture, we selected MobileNetV2, ResNet50, and EfficientNetB0 based on three criteria: (1) validation accuracy after fine-tuning, (2) inference speed (milliseconds per image on CPU), and (3) complementarity—the degree to which their feature representations are non-redundant. The three chosen models exhibit distinct inductive biases:
MobileNetV2: Employs depth-wise separable convolutions and linear bottlenecks. It is highly parameter-efficient (3.4M parameters) and fast, making it suitable for edge deployment. Its lower-level features capture local textures and edges, useful for detecting small consolidations.
ResNet50: Introduces residual connections that enable training of very deep networks. Its 25.6M parameters allow learning of hierarchical, semantically rich features, particularly effective for identifying diffuse interstitial patterns characteristic of viral pneumonia.
EfficientNetB0: Achieves state-of-the-art accuracy with compound scaling (depth, width, resolution). Its 5.3M parameters and balanced receptive field provide a complementary middle ground between the lightweight MobileNetV2 and the deeper ResNet50.
The performance comparison of pneumonia detection models appears in
Figure 6. The proposed ensemble approach demonstrates better performance, especially in terms of accuracy, precision, and F1-Score achievements, compared to other models, which strengthens its suitability for clinical applications in pneumonia sensitivity detection.
4.5. Clinical Relevance
The diagnostic effectiveness of the model takes on greater importance because of its potential clinical usage, especially among children who face pneumonia as one of their primary causes of sickness and death. Posts obtain maximum sensitivity in risk of great clinical dangers because detecting all pneumonia cases immediately becomes essential for both early therapeutic delays and patient outcome decline. An ensemble model maintains a 96.92% recall score, which ensures identification of almost every true pneumonia case, thereby reducing the chances of false negative results. The system maintains accurate performance by achieving 94.10% precision, which minimizes false alarms while maintaining resource efficiency and patient-related safety. The ensemble model adopts an architectural design that performs efficiently with limited resources, and it operates without hardware constraints, so it functions well in constrained conditions. The compact design of the model enables straightforward implementation on portable diagnostic devices, as well as telemedicine systems and edge devices. This approach makes top-quality pneumonia diagnostics accessible in underserved rural health centres as well as mobile screening units, which helps increase healthcare equity across patient populations.
For clinical significance, our model achieve 96.92% recall, meaning that it misses only 3% of true pneumonia cases, while maintaining 94.10% precision. In a screening context, this translates to a few false negatives (avoiding delayed treatment) and manageable false positives (which can be flagged for radiologist review).