Submitted:
04 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
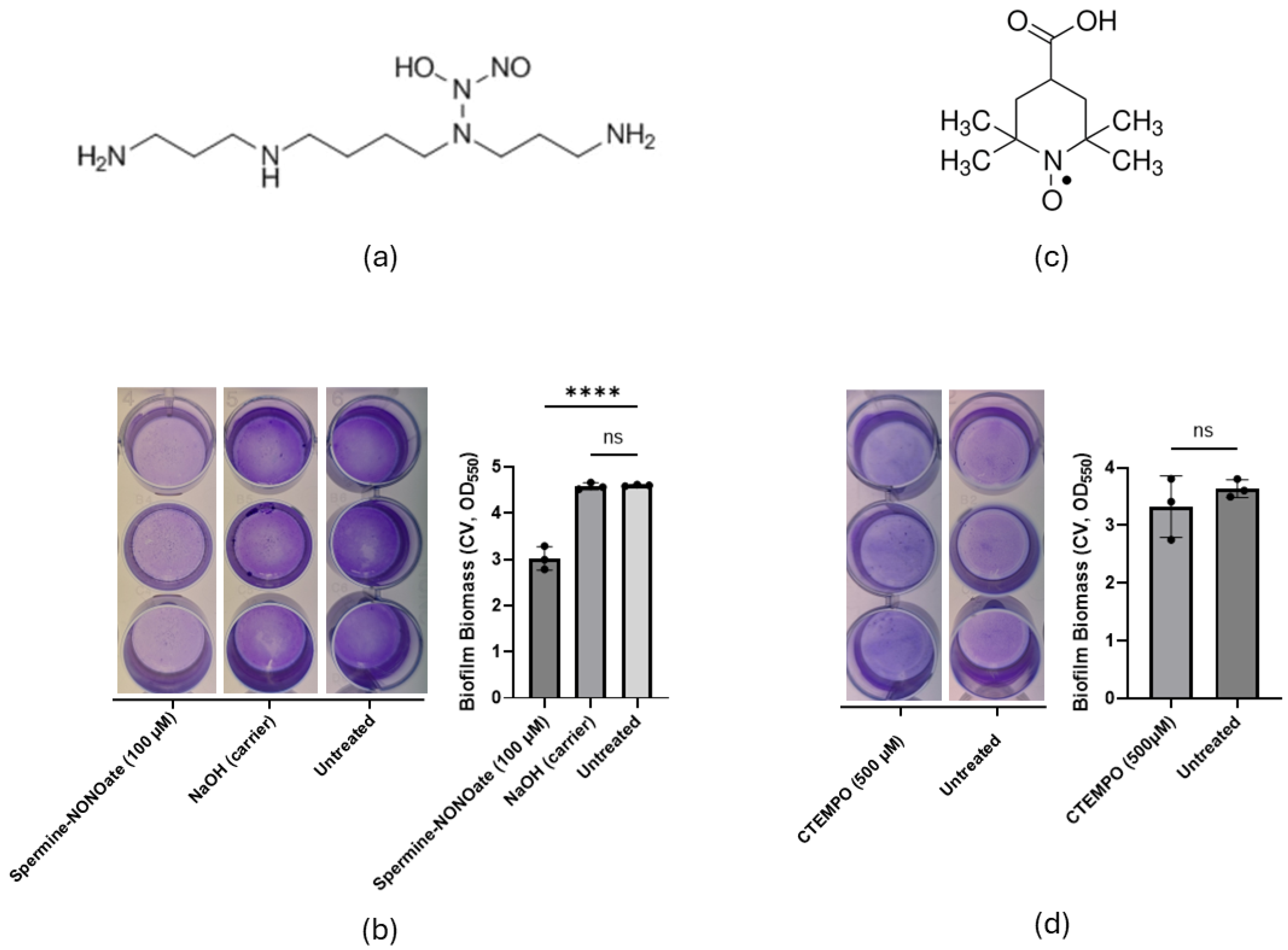
2.1. The NO Donor SP-NONO, but Not the Nitroxide C-TEMPO, Rapidly Induced Biofilm Biomass Reduction of P. aeruginosa Biofilms in Closed Systems
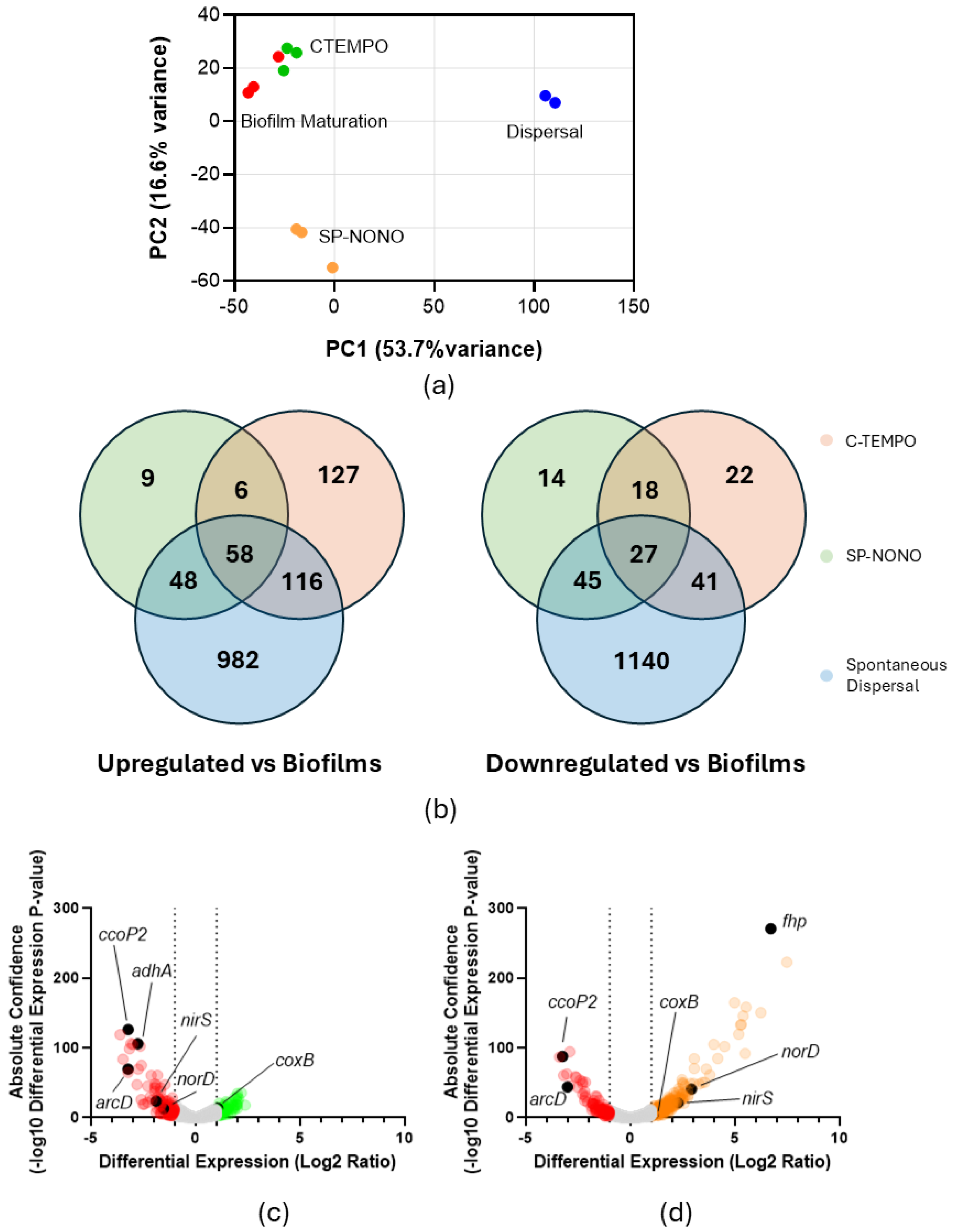
2.2. SP-NONO and C-TEMPO Disrupt ANR-Regulated Energy Production Pathways
2.3. SP-NONO Upregulates Metabolic Pathways of Spontaneous Dispersal
2.4. SP-NONO Treatment of PAO1 Biofilms Upregulates Biomarkers of Spontaneous Dispersal
3. Discussion
4. Materials and Methods
4.1. Strains, Media and Culture Conditions
4.2. Biofilm Dispersal Assays
4.3. RNA Sequencing and Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| NO | Nitric Oxide |
| SP-NONO | Spermine-NONOate |
| C-TEMPO | 4-carboxy-TEMPO |
| RPKM | Reads per kilobase million |
References
- Flemming, H. C.; Wingender, J. ‘The biofilm matrix’. In Nat. Rev. Microbiol.; SUBJMETA, Sep 2010; vol. 8, no. 9, pp. 623–633. [Google Scholar] [CrossRef]
- Ciofu, O.; Tolker-Nielsen, T. Antibiotic tolerance and resistance in biofilms’, in Biofilm Infections; Bjarnsholt, T., Jensen, P. Ø., Moser, C., Høiby, N., Eds.; Springer New York: New York, 2011; Volume ch. 13, pp. 215–229. [Google Scholar] [CrossRef]
- An, S.; Wu, J.; Zhang, L. H. ‘Modulation of pseudomonas aeruginosa biofilm dispersal by a cyclic-di-gmp phosphodiesterase with a putative hypoxia-sensing domain’. Appl. Environ. Microbiol. 2010, vol. 76(no. 24), 8160–8173. [Google Scholar] [CrossRef]
- Sauer, K.; Cullen, M. C.; Rickard, A. H.; Zeef, L. A. H.; Davies, D. G.; Gilbert, P. ‘Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm’. J. Bacteriol. 2004, vol. 186(no. 21), 7312–7326. [Google Scholar] [CrossRef] [PubMed]
- Barraud, N.; Kjelleberg, S.; Rice, S. A. ‘Dispersal from Microbial Biofilms’. Microbiol. Spectr. 2015, vol. 3(no. 6). [Google Scholar] [CrossRef] [PubMed]
- Chambers, J. R.; Cherny, K. E.; Sauer, K. ‘Susceptibility of Pseudomonas aeruginosa Dispersed Cells to Antimicrobial Agents Is Dependent on the Dispersion Cue and Class of the Antimicrobial Agent Used.’. Antimicrob. Agents Chemother. 2017, vol. 61(no. 12), e00846–17. [Google Scholar] [CrossRef]
- Barraud, N.; Storey, M. V.; Moore, Z. P.; Webb, J. S.; Rice, S. A.; Kjelleberg, S. ‘Nitric oxide-mediated dispersal in single- and multi-species biofilms of clinically and industrially relevant microorganisms’. Microb. Biotechnol. 2009, vol. 2(no. 3), 370–378. [Google Scholar] [CrossRef]
- Howlin, R. P. ‘Low-Dose Nitric Oxide as Targeted Anti-biofilm Adjunctive Therapy to Treat Chronic Pseudomonas aeruginosa Infection in Cystic Fibrosis’. Molecular Therapy 2017, vol. 25(no. 9), 2104–2116. [Google Scholar] [CrossRef]
- Zhu, X.; Rice, S. A.; Barraud, N. ‘Nitric Oxide and Iron Signaling Cues Have Opposing Effects on Biofilm Development in Pseudomonas aeruginosa’. Appl. Environ. Microbiol. 2019, vol. 85(no. 3). [Google Scholar] [CrossRef]
- Zhu, X.; Oh, H. S.; Ng, Y. C. B.; Tang, P. Y. P.; Barraud, N.; Rice, S. A. Nitric Oxide-Mediated Induction of Dispersal in Pseudomonas aeruginosa Biofilms Is Inhibited by Flavohemoglobin Production and Is Enhanced by Imidazole’, Antimicrob. Agents Chemother. Accessed. Mar 2018, vol. 62.
- Barnes, R. J. ‘Optimal dosing regimen of nitric oxide donor compounds for the reduction of Pseudomonas aeruginosa biofilm and isolates from wastewater membranes’. Biofouling 2013, vol. 29(no. 2), 203–212. [Google Scholar] [CrossRef]
- Cai, Y. ming; Webb, J. S. ‘Optimization of nitric oxide donors for investigating biofilm dispersal response in Pseudomonas aeruginosa clinical isolates’. Appl. Microbiol. Biotechnol. 2020, vol. 104(no. 20), 8859–8869. [Google Scholar] [CrossRef]
- Bertran i Forga, X.; Hong, Y.; Fairfull-Smith, K. E.; Qin, J.; Totsika, M. Nitric oxide donor sodium nitroprusside serves as a source of iron supporting Pseudomonas aeruginosa growth and biofilm formation’. Microbiol. Spectr. 2025. [Google Scholar] [CrossRef] [PubMed]
- Arai, H.; Hayashi, M.; Kuroi, A.; Ishii, M.; Igarashi, Y. ‘Transcriptional regulation of the flavohemoglobin gene for aerobic nitric oxide detoxification by the second nitric oxide-responsive regulator of Pseudomonas aeruginosa’. J. Bacteriol. 2005, vol. 187(no. 12), 3960–3968. [Google Scholar] [CrossRef]
- Tucker, N. P.; D’Autréaux, B.; Spiro, S.; Dixon, R. ‘Mechanism of transcriptional regulation by the Escherichia coli nitric oxide sensor NorR’. Biochem. Soc. Trans. 2006, vol. 34 no. Pt 1, 191–194. [Google Scholar] [CrossRef]
- Koskenkorva, T.; Aro-Kärkkäinen, N.; Bachmann, D.; Arai, H.; Frey, A. D.; Kallio, P. T. ‘Transcriptional activity of Pseudomonas aeruginosa fhp promoter is dependent on two regulators in addition to FhpR’. Arch. Microbiol. 2008, vol. 189(no. 4), 385–396. [Google Scholar] [CrossRef]
- Kuroki, M.; Igarashi, Y.; Ishii, M.; Arai, H. ‘Fine-tuned regulation of the dissimilatory nitrite reductase gene by oxygen and nitric oxide in Pseudomonas aeruginosa’. Environ. Microbiol. Rep. 2014, vol. 6(no. 6), 792–801. [Google Scholar] [CrossRef] [PubMed]
- Stamler, J. S.; Singel, D. J.; Loscalzo, J. Biochemistry of Nitric Oxide and Its Redox-Activated Forms’, Science (1979).; JOURNAL:JOURNAL:SCIENCE: WGROUP; STRING:PUBLICATION, Dec 1992; vol. 258, no. 5090, pp. 1898–1902. [Google Scholar] [CrossRef]
- Volodarsky, L. B.; Reznikov, V. A.; Ovcharenko, V. I. Synthetic chemistry of stable nitroxides; CRC Press; doi: 10.1201/9780203710159/SYNTHETIC-CHEMISTRY-STABLE-NITROXIDES-VOLODARSKY-REZNIKOV-OVCHARENKO/RIGHTS-AND-PERMISSIONS, 2017. [Google Scholar]
- Volodarsky, L. B.; Reznikov, V. A.; Ovcharenko, V. I. Synthetic chemistry of stable nitroxides, 1st ed.; CRC Press; doi: 10.1201/9780203710159/SYNTHETIC-CHEMISTRY-STABLE-NITROXIDES-VOLODARSKY-REZNIKOV-OVCHARENKO/RIGHTS-AND-PERMISSIONS, 1994. [Google Scholar]
- De La Fuente-Núñez, C.; Reffuveille, F.; Fairfull-Smith, K. E.; Hancock, R. E. W. ‘Effect of nitroxides on swarming motility and biofilm formation, multicellular behaviors in Pseudomonas aeruginosa’. Antimicrob. Agents Chemother. 2013, vol. 57(no. 10), 4877–4881. [Google Scholar] [CrossRef]
- Reffuveille, F.; de la Fuente-Núñez, C.; Fairfull-Smith, K. E.; Hancock, R. E. W. ‘Potentiation of ciprofloxacin action against Gram-negative bacterial biofilms by a nitroxide’. Pathog. Dis. 2015, vol. 73(no. 5), 16. [Google Scholar] [CrossRef]
- Bertran i Forga, X.; Fairfull-Smith, K. E.; Qin, J.; Totsika, M. ‘Transcriptional profiling of Pseudomonas aeruginosa biofilm life cycle stages reveals dispersal-specific biomarkers’. bioRxiv 2025, p. 2025.12.18.695191. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. ‘KEGG: kyoto encyclopedia of genes and genomes’. Nucleic Acids Res. 2000, vol. 28(no. 1), 27–30. [Google Scholar] [CrossRef] [PubMed]
- Belanger, C. R. ‘Surviving the host: Microbial metabolic genes required for growth of Pseudomonas aeruginosa in physiologically-relevant conditions’. Front. Microbiol. 2022, vol. 13, 1055512. [Google Scholar] [CrossRef] [PubMed]
- Revelles, O.; Espinosa-Urgel, M.; Molin, S.; Ramos, J. L. ‘The davDT operon of Pseudomonas putida, involved in lysine catabolism, is induced in response to the pathway intermediate delta-aminovaleric acid’. J. Bacteriol. 2004, vol. 186(no. 11), 3439–3446. [Google Scholar] [CrossRef]
- BURNS, G.; BROWN, T.; HATTER, K.; SOKATCH, J. R. ‘Comparison of the amino acid sequences of the transacylase components of branched chain oxoacid dehydrogenase of Pseudomonas putida, and the pyruvate and 2-oxoglutarate dehydrogenases of Escherichia coli’. Eur. J. Biochem. 1988, vol. 176(no. 1), 165–169. [Google Scholar] [CrossRef] [PubMed]
- Mattevi, A.; Obmolova, G.; Sokatch, J. R.; Betzel, C.; Hol, W. G. J. ‘The refined crystal structure of Pseudomonas putida lipoamide dehydrogenase complexed with NAD+ at 2.45 A resolution’. Proteins 1992, vol. 13(no. 4), 336–351. [Google Scholar] [CrossRef] [PubMed]
- Gliese, N.; Khodaverdi, V.; Görisch, H. ‘The PQQ biosynthetic operons and their transcriptional regulation in Pseudomonas aeruginosa’. Arch. Microbiol. 2010, vol. 192(no. 1), 1–14. [Google Scholar] [CrossRef] [PubMed]
- Orillard, E.; Watts, K. J. ‘Deciphering the Che2 chemosensory pathway and the roles of individual Che2 proteins from Pseudomonas aeruginosa’. Mol. Microbiol. 2021, vol. 115(no. 2), 222–237. [Google Scholar] [CrossRef]
- Del Pozo, J. L.; Patel, R. ‘The challenge of treating biofilm-associated bacterial infections’. Clin. Pharmacol. Ther. 2007, vol. 82(no. 2), 204–209. [Google Scholar] [CrossRef]
- Lewis, K. ‘Riddle of biofilm resistance’. Antimicrob. Agents Chemother. 2001, vol. 45(no. 4), 999–1007. [Google Scholar] [CrossRef]
- Reynolds, D.; Kollef, M. ‘The Epidemiology and Pathogenesis and Treatment of Pseudomonas aeruginosa Infections: An Update’. In Adis; 01 Dec 2021. [Google Scholar] [CrossRef]
- Vestby, L. K.; Grønseth, T.; Simm, R.; Nesse, L. L. ‘Bacterial biofilm and its role in the pathogenesis of disease’; 2020. [Google Scholar] [CrossRef]
- Southey-Pillig, C. J.; Davies, D. G.; Sauer, K. ‘Characterization of temporal protein production in Pseudomonas aeruginosa biofilms’. J. Bacteriol. 2005, vol. 187(no. 23), 8114–8126. [Google Scholar] [CrossRef] [PubMed]
- Gamper, M.; Zimmermann, A.; Haas, D. ‘Anaerobic regulation of transcription initiation in the arcDABC operon of Pseudomonas aeruginosa’. J. Bacteriol. 1991, vol. 173(no. 15), 4742–4750. [Google Scholar] [CrossRef]
- Kawakami, T.; Kuroki, M.; Ishii, M.; Igarashi, Y.; Arai, H. ‘Differential expression of multiple terminal oxidases for aerobic respiration in Pseudomonas aeruginosa’. In Environ. Microbiol.; WEBSITE:WEBSITE:SFAMJOURNALS; STRING:PUBLICATION: WGROUP, Jun 2010; vol. 12, no. 6, pp. 1399–1412. [Google Scholar] [CrossRef]
- Crocker, A.W. ‘Pseudomonas aeruginosa Ethanol Oxidation by AdhA in Low-Oxygen Environments’. J. Bacteriol. 2019, vol. 201(no. 23). [Google Scholar] [CrossRef]
- Barraud, N. ‘Nitric Oxide Signaling in Pseudomonas aeruginosa Biofilms Mediates Phosphodiesterase Activity, Decreased Cyclic Di-GMP Levels, and Enhanced Dispersal’. J. Bacteriol. 2009, vol. 191(no. 23), 7333–7342. [Google Scholar] [CrossRef]
- Feng, Y.; Adams~, E. ‘Glutarate Semialdehyde Dehydrogenase of Pseudomonas’; 1977. [Google Scholar]
- Chou, H. T.; Li, J. Y.; Lu, C. D. ‘Functional Characterization of the agtABCD and agtSR Operons for 4-Aminobutyrate and 5-Aminovalerate Uptake and Regulation in Pseudomonas aeruginosa PAO1’. Current Microbiology 2013, 2013 68:1, vol. 68(no. 1), 59–63. [Google Scholar] [CrossRef]
- Valentini, M.; Storelli, N.; Lapouge, K. ‘Identification of C 4-dicarboxylate transport systems in Pseudomonas aeruginosa PAO1’. J. Bacteriol. 2011, vol. 193(no. 17), 4307–4316. [Google Scholar] [CrossRef] [PubMed]
- Underhill, S. A. M.; Cabeen, M. T. ‘Redundancy in Citrate and cis-Aconitate Transport in Pseudomonas aeruginosa’. J. Bacteriol. 2022, vol. 204(no. 12). [Google Scholar] [CrossRef] [PubMed]
- Zhu, X. ‘Mechanisms of nitric oxide-mediated biofilm dispersal in Pseudomonas aeruginosa’; Nanyang Technological University: Singapore, 2018. [Google Scholar]
- Hong, C. S. ‘Chemotaxis proteins and transducers for aerotaxis in Pseudomonas aeruginosa’. FEMS Microbiol. Lett. 2004, vol. 231(no. 2), 247–252. [Google Scholar] [CrossRef] [PubMed]
- Garvis, S. ‘Caenorhabditis elegans Semi-Automated Liquid Screen Reveals a Specialized Role for the Chemotaxis Gene cheB2 in Pseudomonas aeruginosa Virulence’. PLoS Pathog. 2009, vol. 5(no. 8), e1000540. [Google Scholar] [CrossRef]
| Gene | Fold Change - C-TEMPOa | Fold Change - SP-NONOa | Gene product |
|---|---|---|---|
| Main anaerobiosis regulators | |||
| anr | -1.40 | -1.60 | transcriptional regulator Anr |
| dnr | -2.16 | -1.64 | transcriptional regulator Dnr |
| Oxidative phosphorylation | |||
| ccoP1 | 1.33 | 0.98 | cytochrome C oxidase cbb3-type subunit CcoP |
| ccoQ1 | 1.22 | 0.93 | cytochrome C oxidase cbb3-type subunit CcoQ |
| ccoO1 | 1.16 | 0.94 | cbb3-type cytochrome C oxidase subunit II |
| ccoN1 | 1.15 | 0.82 | cbb3-type cytochrome C oxidase subunit I |
| ccoP2 | -9.30 | -9.48 | cytochrome C oxidase cbb3-type subunit CcoP |
| ccoQ2 | -3.83 | -4.51 | cytochrome C oxidase cbb3-type subunit CcoQ |
| ccoO2 | -8.92 | -10.45 | cbb3-type cytochrome C oxidase subunit II |
| ccoN2 | -6.72 | -8.18 | cbb3-type cytochrome C oxidase subunit Id |
| coxA | 2.08 | 2.12 | cytochrome C oxidase subunit II |
| coxB | 3.90 | 2.95 | cytochrome C oxidase subunit I |
| cioA | 1.34 | 2.15 | cyanide insensitive terminal oxidase |
| cioB | 1.30 | 2.01 | cyanide insensitive terminal oxidase |
| cyoA | 1.09 | 1.80 | cytochrome o ubiquinol oxidase subunit II |
| cyoB | 1.06 | 1.27 | cytochrome o ubiquinol oxidase subunit I |
| cyoC | 1.26 | 1.01 | cytochrome o ubiquinol oxidase subunit III |
| cyoD | 1.31 | 0.87 | cytochrome o ubiquinol oxidase subunit IV |
| cyoE | 1.31 | 0.83 | protoheme IX farnesyltransferase |
| Denitrification | |||
| nirN | -1.25 | 2.70 | cytochrome C |
| PA0510 | -1.62 | 3.55 | uroporphyrin-III C-methyltransferase |
| nirJ | -1.48 | 3.67 | heme d1 biosynthesis protein NirJ |
| nirH | -1.10 | 3.30 | heme d1 biosynthesis protein NirH |
| nirG | -1.80 | 3.86 | heme d1 biosynthesis protein NirG |
| nirL | -2.63 | 2.37 | heme d1 biosynthesis protein NirL |
| nirD | -3.62 | 1.76 | heme d1 biosynthesis protein NirD |
| nirF | -4.88 | 1.54 | heme d1 biosynthesis protein NirF |
| nirC | -5.70 | 2.10 | cytochrome c55X |
| nirM | -5.44 | 3.13 | cytochrome C-551 |
| nirS | -3.70 | 4.80 | nitrite reductase |
| nirQ | -0.76 | 2.86 | denitrification regulatory protein NirQ |
| PA0521 | -0.95 | 6.67 | cytochrome C oxidase subunit |
| PA0522 | -1.20 | 6.08 | hypothetical protein |
| norC | -1.13 | 45.90 | nitric oxide reductase subunit C |
| norB | -1.05 | 31.34 | nitric oxide reductase subunit B |
| norD | -2.88 | 7.54 | denitrification protein NorD |
| nosR | 1.05 | 8.26 | regulatory protein NosR |
| nosZ | 1.02 | 7.28 | nitrous-oxide reductase |
| nosD | 0.95 | 3.93 | copper-binding periplasmic protein |
| nosF | 1.16 | 4.09 | copper ABC transporter ATP-binding protein |
| nosY | 0.88 | 3.47 | membrane protein NosY |
| nosL | 0.99 | 1.54 | acessory protein NosL |
| Arginine deiminase pathway | |||
| arcD | -9.37 | -8.02 | arginine/ornithine antiporter |
| arcA | -12.14 | -9.16 | arginine deiminase |
| arcB | -11.07 | -7.48 | ornithine carbamoyltransferase |
| arcC | -3.08 | -3.58 | carbamate kinase |
| Alcohol oxidation pathway | |||
| adhA | -6.79 | -5.97 | alcohol dehydrogenase |
| exaA | -1.27 | 1.34 | quinoprotein ethanol dehydrogenase |
| exaB | -1.39 | 1.84 | cytochrome C550 |
| exaC | -2.23 | 1.48 | NAD+ dependent aldehyde dehydrogenase ExaC |
| Gene | Locus Tag | Fold Change - SP-NONO a | Fold Change - Spontaneous dispersal a | Gene product |
|---|---|---|---|---|
| Quorumsensing | ||||
| agtA | PA0604 | 4.63 | 3.20 | ABC transporter |
| agtB | PA0605 | 3.61 | 2.27 | ABC transporter permease |
| agtC | PA0606 | 2.36 | 2.10 | ABC transporter permease |
| PA1617 | 2.04 | 4.32 | AMP-binding protein | |
| mexG | PA4205 | 2.43 | 14.12 | hypothetical protein |
| mexH | PA4206 | 2.48 | 15.14 | resistance-nodulation-cell division (RND) efflux membrane fusion protein |
| mexI | PA4207 | 2.07 | 8.51 | resistance-nodulation-cell division (RND) efflux transporter |
| Sulphur metabolism | ||||
| cysW | PA0281 | -2.33 | -3.01 | sulfate transporter CysW |
| cysT | PA0282 | -2.73 | -2.81 | sulfate transporter CysT |
| cysP | PA1493 | -2.39 | -2.04 | sulfate ABC transporter substrate-binding protein |
| PA3449 | -3.23 | -3.05 | hypothetical protein | |
| PA3936 | -2.16 | -2.95 | taurine ABC transporter permease | |
| PA3937 | -2.45 | -2.85 | taurine ABC transporter ATP-binding protein | |
| PA3938 | -2.57 | -2.11 | taurine-binding protein | |
| cysN | PA4442 | -2.50 | -5.10 | bifunctional sulfate adenylyltransferase subunit1/adenylylsulfate kinase |
| cysD | PA4443 | -2.08 | -3.43 | sulfate adenylyltransferase subunit 2 |
| ABC transporters | ||||
| lhpK | PA1255 | 2.60 | 4.38 | trans-3-hydroxy-L-proline dehydratase |
| lhpO | PA1256 | 2.48 | 3.63 | amino acid ABC transporter ATP binding protein |
| lhpM | PA1258 | 2.41 | 2.25 | ABC transporter permease |
| nosF | PA3394 | 4.08 | 3.89 | copper ABC transporter ATP-binding protein |
| opuCA | PA3891 | 3.56 | 7.06 | ABC transporter ATP-binding protein |
| PA4193 | -2.25 | -8.63 | ABC transporter permease | |
| PA4194 | -2.13 | -4.14 | ABC transporter permease | |
| PA4195 | -2.08 | -2.55 | ABC transporter | |
| PA5095 | 2.10 | 2.55 | ABC transporter permease | |
| Two-component systems | ||||
| PA0034 | -2.00 | -8.11 | two-component response regulator | |
| PA0752 | 2.28 | 4.89 | hypothetical protein | |
| PA0753 | 2.57 | 4.35 | hypothetical protein | |
| PA0754 | 2.81 | 6.96 | hypothetical protein | |
| dctA | PA1183 | 44.63 | 7.57 | C4-dicarboxylate transport protein |
| ansB | PA1337 | 2.33 | 3.36 | glutaminase-asparaginase |
| PA3356 | 3.66 | 2.57 | hypothetical protein | |
| pprB | PA4296 | 2.14 | 8.57 | two-component response regulator PprB |
| tadB | PA4301 | 2.00 | 23.10 | type II secretion system protein TadB |
| rcpA | PA4304 | 2.11 | 28.25 | type II/III secretion system protein |
| rcpC | PA4305 | 2.19 | 30.48 | hypothetical protein |
| PA4648 | 2.36 | 21.86 | hypothetical protein | |
| PA4649 | 2.10 | 8.75 | hypothetical protein | |
| PA4908 | 3.14 | 3.25 | ornithine cyclodeaminase | |
| PA4909 | 2.68 | 3.27 | ABC transporter ATP-binding protein | |
| PA4910 | 3.14 | 3.03 | ABC transporter ATP-binding protein | |
| PA4911 | 3.97 | 3.07 | branched-chain amino acid ABC transporter permease | |
| PA4912 | 2.77 | 2.30 | branched-chain amino acid ABC transporter | |
| PA4913 | 2.01 | 2.14 | ABC transporter | |
| Alanine, aspartate and glutamate metabolism | ||||
| davD | PA0265 | 5.03 | 2.08 | glutarate-semialdehyde dehydrogenase DavD |
| davT | PA0266 | 4.06 | 2.66 | 5-aminovalerate aminotransferase DavT |
| ansB | PA1337 | 2.33 | 3.36 | glutaminase-asparaginase |
| PA3356 | 3.66 | 2.57 | hypothetical protein | |
| PA3758 | 3.39 | 2.13 | N-acetylglucosamine-6-phosphate deacetylase | |
| PA3759 | 3.36 | 2.73 | aminotransferase | |
| PA3760 | 3.41 | 2.45 | N-acetyl-D-glucosamine phosphotransferase system transporter | |
| PA5522 | 4.23 | 3.25 | glutamine synthetase | |
| PA5523 | 5.50 | 2.91 | aminotransferase | |
| Valine, leucine and isoleucine metabolism | ||||
| braC | PA1074 | 2.13 | 2.91 | branched-chain amino acid ABC transporter substrate-binding protein BraC |
| bkdB | PA2249 | 2.10 | 3.56 | branched-chain alpha-keto acid dehydrogenase complex lipoamide acyltransferase |
| lpdV | PA2250 | 2.69 | 2.93 | branched-chain alpha-keto acid dehydrogenase complex dihydrolipoyl dehydrogenase |
| leuD | PA3120 | -3.36 | -3.18 | isopropylmalate isomerase small subunit |
| leuC | PA3121 | -2.53 | -4.11 | 3-isopropylmalate dehydratase large subunit |
| PA3417 | 2.66 | 13.09 | pyruvate dehydrogenase E1 component subunit alpha | |
| ldh | PA3418 | 2.19 | 17.88 | leucine dehydrogenase |
| ilvI | PA4696 | -2.07 | -11.16 | acetolactate synthase 3 catalytic subunit |
| Arginine and proline metabolism | ||||
| PA4908 | 3.14 | 3.25 | ornithine cyclodeaminase | |
| PA4909 | 2.68 | 3.27 | ABC transporter ATP-binding protein | |
| PA4910 | 3.14 | 3.03 | ABC transporter ATP-binding protein | |
| PA4911 | 3.97 | 3.07 | branched-chain amino acid ABC transporter permease | |
| PA4912 | 2.77 | 2.30 | branched-chain amino acid ABC transporter | |
| PA4913 | 2.01 | 2.14 | ABC transporter | |
| Bacterial chemotaxis | ||||
| PA0173 | 3.12 | 4.17 | chemotaxis response regulator protein-glutamate methylesterase | |
| PA0174 | 3.46 | 2.95 | hypothetical protein | |
| cheR2 | PA0175 | 3.41 | 5.06 | chemotaxis protein methyltransferase |
| aer2 | PA0176 | 2.30 | 5.98 | aerotaxis transducer Aer2 |
| PA0177 | 2.01 | 4.56 | purine-binding chemotaxis protein | |
| PA0179 | 2.00 | 9.99 | two-component response regulator | |
| Porphyrin metabolism | ||||
| cobP | PA1278 | -2.30 | -5.70 | bifunctional adenosylcobinamide kinase/adenosylcobinamide-phosphate guanylyltransferase |
| cobU | PA1279 | -2.38 | -5.03 | nicotinate-nucleotide--dimethylbenzimidazole phosphoribosyltransferase |
| PA1280 | -2.33 | -7.16 | hypothetical protein | |
| cobV | PA1281 | -2.35 | -6.06 | adenosylcobinamide-GDP ribazoletransferase |
| PA4088 | 2.27 | 2.64 | aminotransferase | |
| PA5523 | 5.50 | 2.91 | aminotransferase | |
| Nicotinate and nicotinamide metabolism | ||||
| pntAB | PA0195.1 | -2.23 | -2.17 | NAD(P) transhydrogenase subunit alpha |
| pntB | PA0196 | -2.48 | -2.35 | pyridine nucleotide transhydrogenase subunit beta |
| nadE | PA4920 | -2.51 | -3.05 | NAD synthetase |
| Pyrroloquinoline quinone biosynthesis | ||||
| pqqA | PA1985 | 2.51 | 14.03 | coenzyme PQQ synthesis protein A |
| pqqD | PA1988 | 3.18 | 2.14 | coenzyme PQQ synthesis protein D |
| pqqE | PA1989 | 3.73 | 2.85 | coenzyme PQQ synthesis protein E |
| pqqH | PA1990 | 3.36 | 3.76 | peptidase |
| Locus Tag | Fold Change – C-TEMPO a |
Fold Change – SP-NONO a | Gene product |
|---|---|---|---|
| PA0111 | 5.15 | 6.62 | hypothetical protein |
| cheR2 | 1.44 | 3.41 | chemotaxis protein methyltransferase |
| PA0743 | 1.32 | 1.83 | NAD-dependent L-serine dehydrogenase |
| PA1353 | 1.70 | 1.47 | hypothetical protein |
| pqqA | 1.21 | 2.52 | coenzyme PQQ synthesis protein A |
| cdpR | 1.39 | -1.01 | transcriptional regulator |
| amrZ | 1.16 | 1.40 | alginate and motility regulator Z |
| tadA | 2.01 | 2.21 | ATPase TadA |
| rcpA | 1.58 | 2.12 | type II/III secretion system protein |
| rcpC | 1.50 | 2.18 | hypothetical protein |
| flp | 2.20 | 2.27 | type IVb pilin Flp |
| PA4523 | -1.13 | 1.22 | hypothetical protein |
| cupE1 | 1.79 | 2.36 | fimbrial subunit CupE1 |
| cupE2 | 1.46 | 2.09 | fimbrial subunit CupE2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





